Abstract
Human pluripotent stem cells (hPSCs), including embryonic and induced pluripotent stem cells (hESCs and hiPSCs, respectively), continue to attract researchers and clinicians as well as patients worldwide because of their applicability in regenerative medicine, cell-based therapies, and drug discovery. However, conventional biomedical approaches for stem cell research provide only limited insights regarding how stem cells are regulated in our body and how we could control them. To accelerate the progress of stem cell research and their applications, interdisciplinary research needs to be carried out to enable access of or mimic in vivo physiological conditions and to investigate their functions. In the past few decades, progress in microfabrication engineering has had a tremendous impact in both academia and industry (electronics, mechanics, chemicals, and environments), and many people have realized that microfabricated tools hold great potential for applications in stem cell research because they allow precise manipulation of a wide range of biological materials, including small molecules, DNA/RNA, proteins, cells, and tissues. In this review, I describe advanced microfabricated tools for studying the mechanisms of in vivo stem cell regulation and precise manipulation of stem cells, notably hPSCs, and concerns of using such tools for stem cell research.
Keywords
Introduction
Human pluripotent stem cells (hPSCs), including embryonic and induced pluripotent stem cells (hESCs 1 and hiPSCs,2–5 respectively), hold great potential for applications in basic and clinical studies, since hPSCs have two distinct and very unique characteristics—namely, unlimited self-renewal and differentiation to almost any type of cell in the human body ( Fig. 1 ). Recently, some cell-based therapies using differentiated hESCs went into clinical trials. In addition, Japanese researchers led by Dr. Takahashi proposed the use of hiPSCs for patients with age-related macular degeneration.6,7 Clearly, there is increasing interest and need for applications of hPSCs.
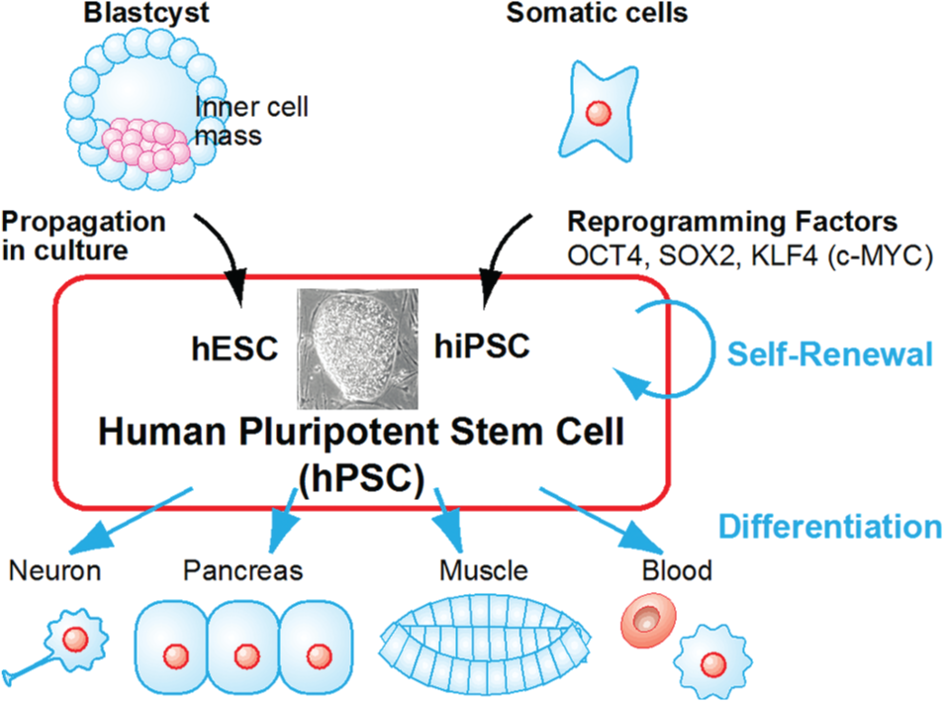
Human pluripotent stem cells (hPSCs). Human embryonic stem cells (hESCs) were harvested from the inner cell mass of a blastocyst and propagated in cell culture. Human-induced pluripotent stem cells (hiPSCs) were generated by introducing reprogramming factors (OCT4, SOX2, KLF4, and c-MYC) into somatic cells. These cells have two distinct characteristics: (1) unlimited self-renewal under proper conditions and (2) the capability to differentiate into almost any type of cell in the human body.
For hPSC applications, several issues need to be addressed. For example, for cell-based therapies, regenerative medicine, and drug screening, 8 large amounts of quality-controlled hPSCs are required because both the efficiency of differentiation to a specific lineage and the survival rate after transplantation are low. Thus, there is a clear need for technological advancement to overcome these issues and facilitate the progress of hPSC applications.
Microfabrication tools for biomedical applications, including hPSCs, hold great potential for facilitating research in the field of cell biology because of their ability to manipulate a wide range of sizes of extracellular environments as well as subcellular components, including proteins and nucleic acids. In this review, I discuss microfabrication approaches for studying hPSCs.
Large Expansion of PSCs
Current Technologies of hPSC Culture
To apply hPSCs for cell-based therapies and regenerative medicine, we require a large number of high-quality cells. In general, cell-based therapies require 109 cells and then need to be induced to differentiate to obtain target cells. The efficiency of obtaining target cells is still insufficient, and the survival rate after cell transplantation is still low. For example, if we use 10-cm dishes to expand hPSCs, it would require 1000 dishes to obtain a sufficient number of cells, and this would be both labor-intensive and time-consuming. Thus, researchers are attempting to establish new culture systems for the expansion of large numbers of high-quality hPSCs. Although there are some reports of the establishment of such methods (spinner flask and suspension culture9,10), each method has its own issues and is not commonly used. There is tremendous scope for improvement in the large expansion system. The common approach to tackle this issue is to establish a 3D culture system. The establishment of a 3D culture system allows efficient use of medium to culture hPSCs with a high cell density. I will introduce and discuss some examples, particularly those that involve the use of a microengineering approach ( Fig. 2 ).
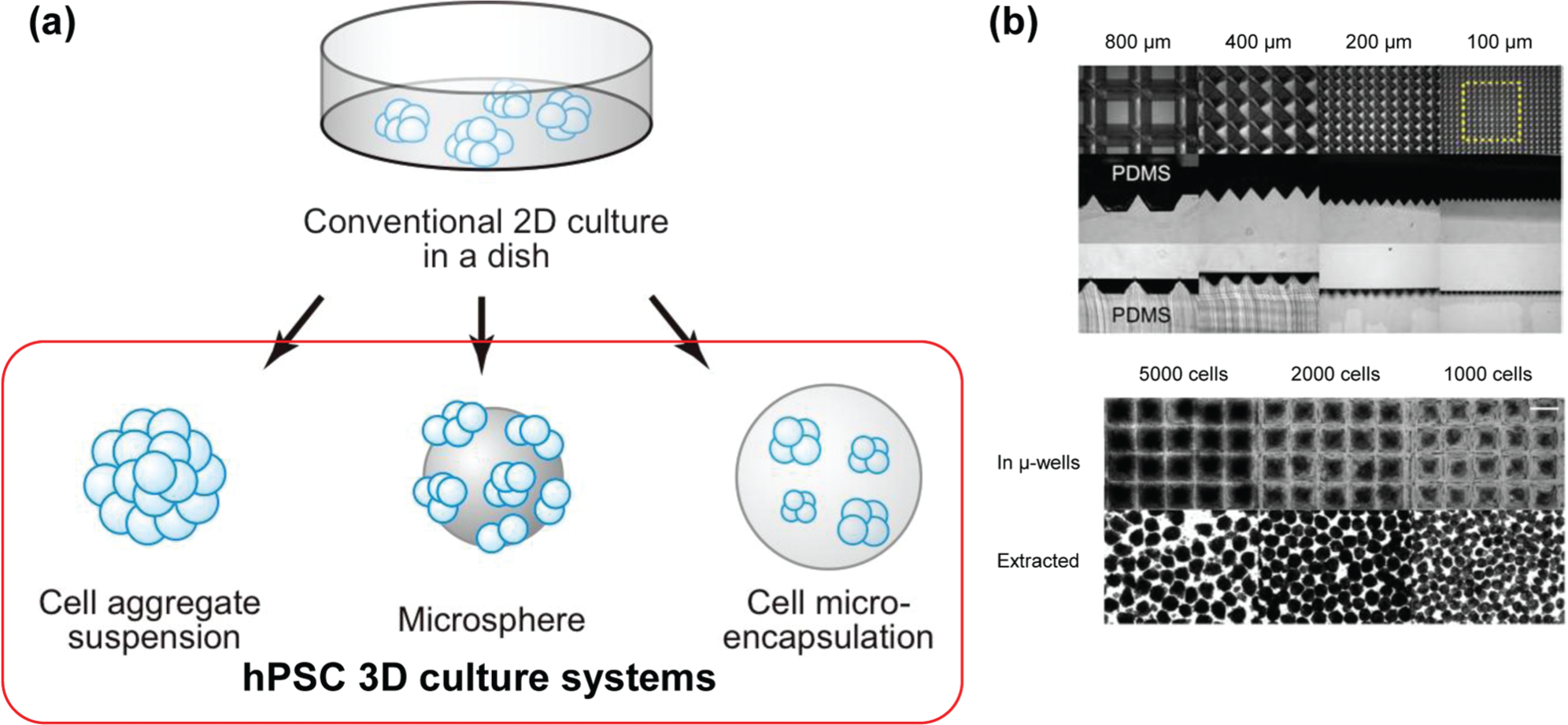
Microsphere Culture
Microspheres can be used to establish a 3D culture system.11–13 Microspheres coated with extracellular matrix (ECM) proteins allow adherence and growth of hPSCs on their surface. By using microspheres, we can increase the ratio of volume to surface area and obtain a larger number of cells. This method is also scalable from small to large culture systems. Compared with other 3D culture methods, we obtain a relatively homogeneous size of cell aggregates using this method, thus demonstrating the reproducibility of this method. However, this method also requires stirring during culture, which causes cell stress and damage. Microspheres also incur additional costs for the culturing systems. For passaging, we need to have additional steps to separate the cells and microspheres. Microspheres also have their own volumes, and therefore, it would be difficult to obtain considerably higher cell numbers after culture.
Cell Microencapsulation
Instead of culturing hPSCs on microspheres, researchers are attempting to encapsulate hPSCs in microspheres, called microencapsulation. For example, Siti-Ismail et al. 14 aimed to introduce hESCs into 1.1% (w/v) calcium alginate hydrogels to maintain them in feeder- and serum-free culture conditions for a long-term period (up to 260 days). This method has similar advantages to microsphere methods such as a high volume-to-surface ratio, reproducibility, and scalability, and in addition to microsphere methods, cell microencapsulation methods protect cells by hydrogel, which causes less cell stress and damage. The most important advantage to the use of cell encapsulation methods is the creation of artificial cellular microenvironments, which provides a more suitable system than microsphere methods owing to the capability of forming 3D scaffolds for the capsulated cells; therefore, we will be able to systematically analyze both physical and chemical microenvironmental cues.14,15 However, some additional issues need to be considered. For example, it would be tedious to encapsulate hPSCs in microspheres, control cell numbers and medium/gas access in spheres, and harvest cells from spheres.
To increase the throughput to obtain optimal hydrogel conditions for cell encapsulation, Kumachev et al. 16 established a microfluidic platform to generate hydrogel beads with different elasticities. Two streams, with either a high or low concentration of agarose mixed with mouse embryonic stem cells (mESCs), were generated with various agarose elasticities in a microfluidic-based droplet generator. They successfully encapsulated mESCs in hydrogel microbeads; these would then be applicable in investigating the role of cellular microenvironmental elasticity on cellular functions.
Cell Aggregation for Suspension Culture
Recently, it was found that hPSCs could be maintained in suspension culture conditions even without a cellular scaffold.9,10 One of the difficulties faced with using suspension culture systems is in obtaining high-quality hPSCs in a robust fashion because of the lack of controllability of cellular parameters such as the size of cell aggregates and unexpected cell death. To tackle these issues, Ungrin et al. 17 developed unique microfabricated substrates consisting of an array of truncated pyramid structures for the formation of cell aggregates with a uniform size forced by centrifugation ( Fig. 2b ). Because the experiments could be started with homogeneous cell aggregates, experimental variability was dramatically reduced compared with other suspension culture systems. Therefore, this method is also applicable for the development of the bioprocess and monitoring of differentiating cells.17–20
Miniaturized PSC Culture
Microfabricated hPSC Culture/Assay Platforms
Microfabrication techniques such as micropatterning and microfluidics offer unique experimental settings for cell biology.21–24 They enable the creation of cellular microenvironments in vitro. Cellular microenvironments are very important in the regulation of in vivo stem cell functions such as self-renewal, differentiation, migration, and apoptosis. While the conventional biological experimental setting has limited access to cellular microenvironments, microfabrication techniques allow the creation of artificial regulatory cellular microenvironments to investigate the underlying mechanisms.
Peerani et al. 25 and Discher et al. 26 successfully used micropatterning technology to precisely control the size of hESC colonies and found that SMAD1 was associated with hESC self-renewal and differentiation depending on the size of the hESC colonies. A larger colony size (d = 800 µm) increased the expression of GDF3 and LEFTY1 as well as the numbers of OCT4-positive cells. In contrast, a smaller hPSC colony size (d = 200 µm) had higher levels of phosphorylated SMAD1 in hESCs derived in the extra-embryonic endoderm. Furthermore, these authors found that spatial control of mESC colonies strongly influences Jak-Stat signaling pathways as well as pluripotency-associated genes. 27 They established the modeling system of microenvironmental cues and investigated how endogenous Stat3 was activated by extrinsic cues as well as by the size of the ECM.
To develop a new synthetic hydrogel to maintain hPSCs for a long-term period under defined conditions and eliminate enzymatic passaging procedures, Zhang et al. 28 developed a micropatterned library of thermoresponsive hydrogels that would allow both culturing and harvesting of hPSCs with only a change in temperature. They found that the optimal hydrogel was based on 2-(dimethylamino)ethyl acrylate as a chemically defined thermoresponsive cellular substrate, and it was capable of maintaining hPSCs over 2 to 6 months.
Microfluidic Cell Culture
Microfluidic technologies24,29 allow for the control of soluble factors as well as 3D geometry. Several groups, including ours, have developed microfluidic platforms for culturing and analyzing PSCs.21,30–33 For example, we developed a microfluidic device to miniaturize the culture conditions of hPSCs co-cultured with mouse embryonic fibroblasts (MEFs) and to perform semi-automated culturing and immunocytochemistry ( Fig. 3a ). Furthermore, we established a simple and user-friendly microfluidic device for general biologists to culture hPSCs under serum- and feeder-free culture conditions. 22 The hPSCs cultured in microfluidic channels under such conditions grow with a well-maintained expression of pluripotency-related proteins such as OCT4 and NANOG. Using flow control of microfluidic technologies, Villa-Diaz et al. 30 applied two different phases (culture medium and trypsin, which is used to harvest cells) to a single hESC colony by using laminar flow ( Fig. 3b ). That way, they were able to selectively remove half of the hESC colony but leave the other half intact. The remaining hESCs maintained OCT4 and NANOG gene expression. Titmarsh et al. 34 identified the optimal flow rate for cultivating hPSCs under continuous-flow conditions. They found that low flow rates of 3.6 µL/h were not sufficient to exchange the medium and remove waste and dead cells. The flow rate of the medium was optimal for hPSC growth at approximately 20 to 30 µL/h, whereas higher flow rates reduced the number of cells and colonies grown.
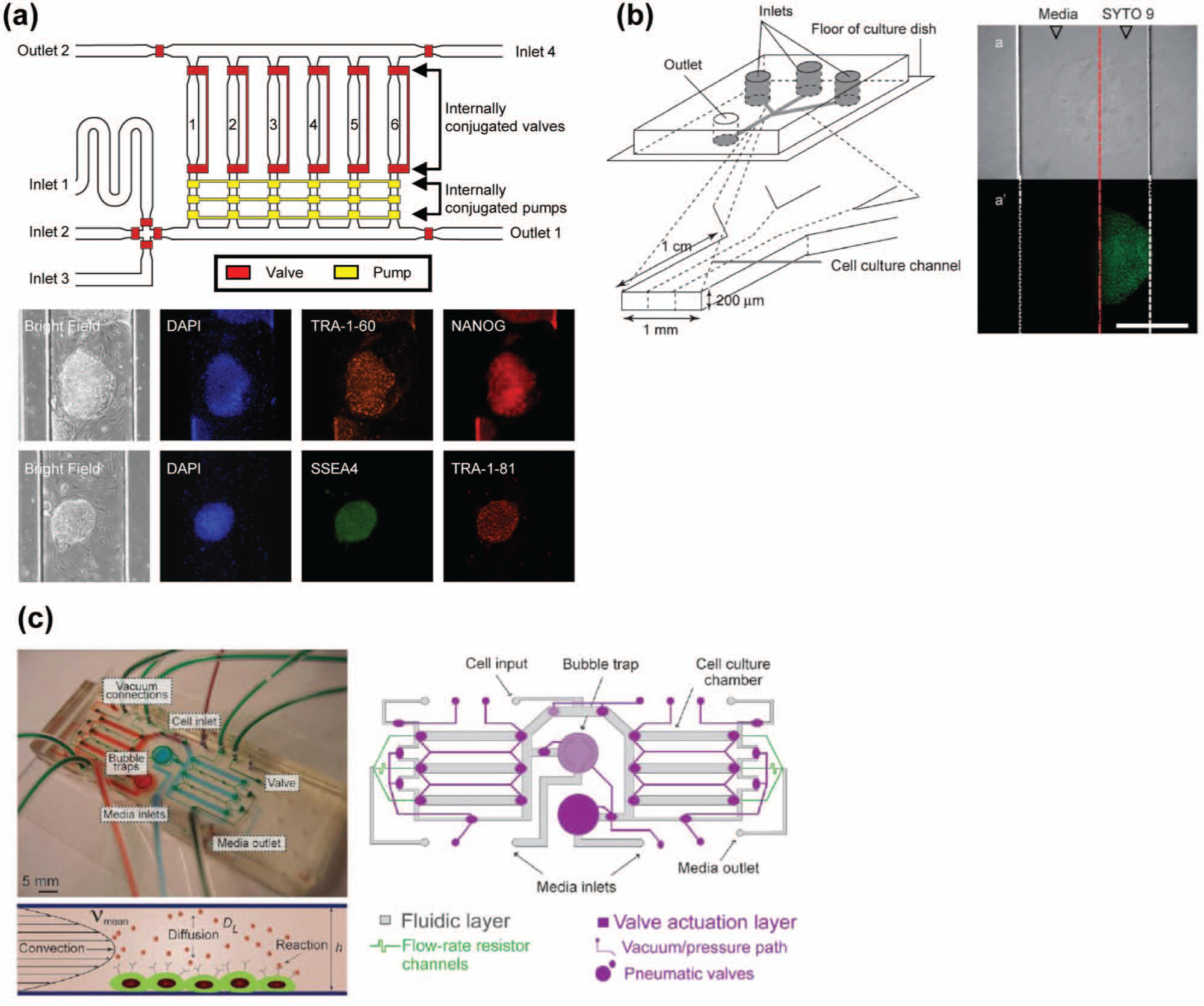
Microfluidic platforms to investigate the role of extrinsic cues on self-renewal of pluripotent stem cells (PSCs). (
To investigate the function of autocrine and paracrine signaling on mESC self-renewal and differentiation, researchers have used a microfluidic device ( Fig. 3c ).35–38 In general, it was well known that LIF (leukemia inhibitory factor) and BMP4 (bone morphogenetic protein 4) maintain mESCs, but it was unclear whether these were sufficient. Przybyla and Voldman35,36 revealed that, under continuous perfusion conditions for mESCs to perturb cell-secreted soluble factors, ECM proteins secreted from mESCs were removed and mESCs transitioned into a primed state. In static conditions, the underlying mechanisms of self-renewal of mESCs regulated by extrinsic cues could not be determined, and a microfluidic device played an important role in this context. Furthermore, in combination with an in silico approach, Moledina et al. 39 showed that local endogenous autocrine and paracrine signaling heterogeneously activated STAT3 signaling in individual mESCs. They simulated flow rate as well as cell density to investigate how soluble factors secreted from mESCs were able to effectively stimulate cells and thus validated their results by using a microfluidic experimental approach. The precise controllability of liquid flow by microfluidic platforms provided agreed well with the computational theoretical approach. 40 Unfortunately, these approaches have been not carried out for hPSCs. Because the mechanisms of self-renewal of hPSCs regulated by extrinsic cues are quite different from those of mESCs, it would be crucial to investigate their roles for hPSCs.
Comparison of Large-Scale Cell Culture and Microfluidic Culture Systems
Microfluidic-based culture platforms would be applicable for the establishment of a high-throughput screening system for cell culture optimization. In addition, by using hydrogel, microfluidics allows for the creation of 3D culture environments, which mimic in vivo conditions and investigate their roles on stem cell functions and fate decisions. As described above, microfluidics also permits studying the effects of chemical (e.g., autocrine and paracrine signaling) and physical (e.g., shear stress) factors. However, scalability is a limitation owing to their size. However, large-scale cell culture systems obtain high cell numbers but are not suitable for high-throughput systems to identify optimal conditions for maintaining or differentiating hPSCs. As each system has both pros and cons for hPSC research, we need to carefully choose the most suitable system for the research objectives.
PSC Differentiation and Tissue Engineering
In general, to obtain cells of interest from PSCs, there are several ways to induce differentiation, such as embryoid body (EB) formation and direct differentiation from PSCs. Even though there have been tremendous efforts in establishing protocols for inducing differentiation into specific lineages from hPSCs, only a small number of successful cases have been reported thus far. Most cases have very low efficiencies of obtaining target cells and are long multistep procedures. To tackle these issues, researchers have developed many microfluidic devices using these requirements.
Uniform EB Formation
An EB is a spherically formed cell aggregate of PSCs in suspension culture and contains a variety of differentiated cells with all three germ layers. Thus, EB formation is commonly used as a first step of a differentiation protocol. Because EBs contain various differentiated cells and the ratio of these cells is varied with time and size of the EBs, it is necessary to have a method to obtain a homogeneous EB size to increase the efficiency of inducing differentiation to a specific lineage. To tackle this issue, researchers have established the hanging drop method,41,42 but this method requires a labor-intensive procedure with less robustness, and thus, there is a clear need to establish a more robust and easy method for EB formation. For example, Valamehr et al. 43 successfully enhanced EB formation by using hydrophobic surfaces, such as alkanethiols on an Au surface or polydimethylsiloxane (PDMS). Torisawa et al. 44 developed a microfluidic device with compartmentalized microchambers for EB formation with uniform sizes, and others have also attempted to use microfluidic45–47 or microwell array48,49 devices for EB formation ( Fig. 4a ). Furthermore, some researchers have treated EBs derived from hPSC-induced differentiation with early stage fate specification as well as mesodermal lineage-committed cells with concentration gradients of Wnt3a, activin A, BMP4, and their inhibitors by using microfluidic bioreactors.50,51 In addition, as described above, researchers have developed substrates with an array of microscaled and truncated pyramid structures for forming uniformly sized EBs forced by centrifugation.18–20 Sorting technology is also beneficial to obtain a uniform size of EBs. 52 Although microfluidic technology is often used for cell sorting, it is also used to sort much larger cell aggregates such as EBs. EBs can be separated on the basis of size, and uniform EBs have been obtained without any damage or influence in their differentiation capability.
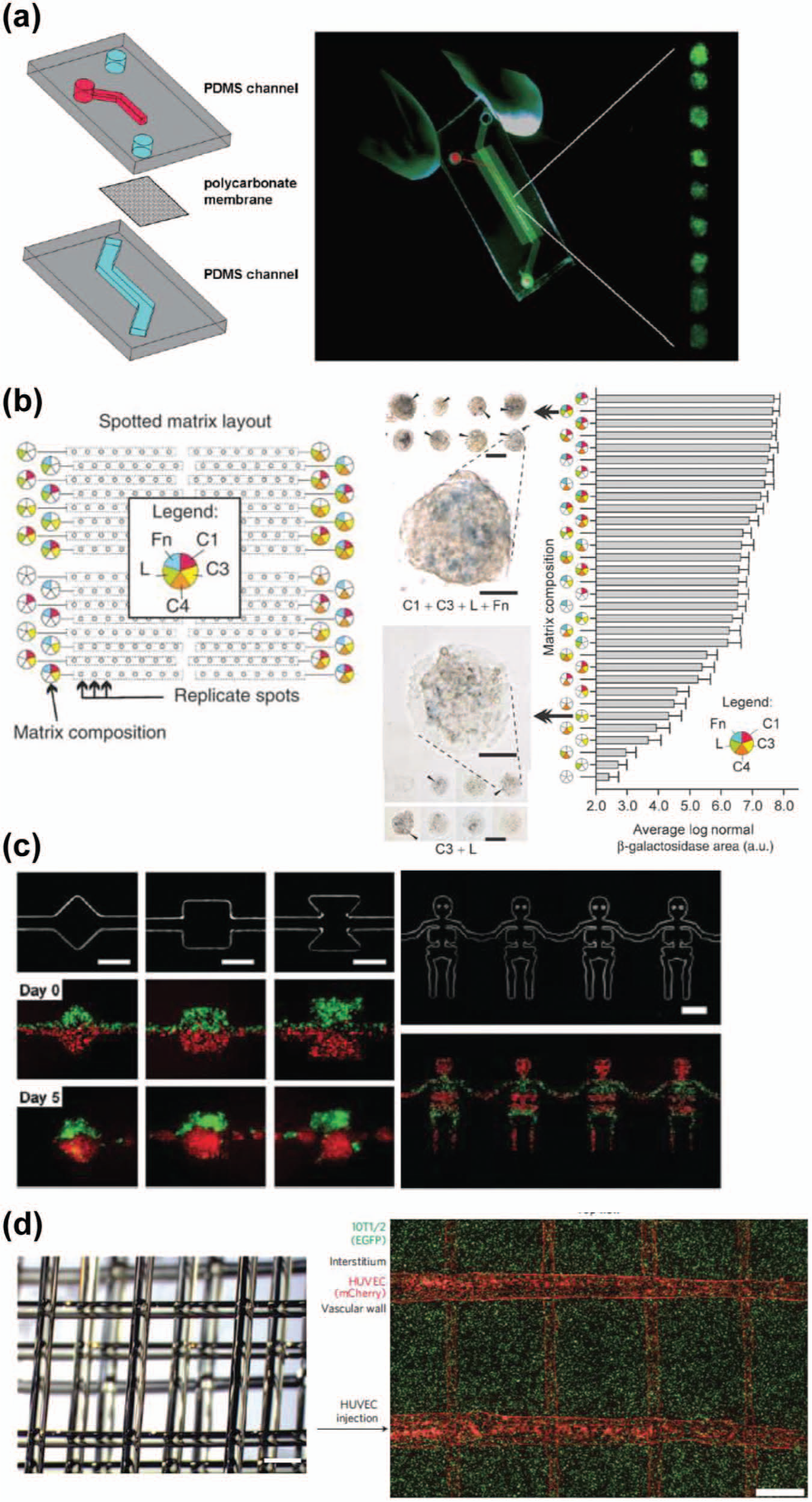
Microfabricated platforms for inducing differentiation of pluripotent stem cells (PSCs). (
Direct Differentiation on Micro/Nanofabricated Substrates
To identify good extracellular environments for inducing differentiation, researchers have used microfabrication technologies such as microfluidics 53 and micropatterning,54–56 which provide an advanced platform to systematically analyze the effects of such environments on cell behavior and phenotype. Micropatterning technology in conjunction with microarray technology has been used to screen a combinatorial library of ECM proteins ( Fig. 4b ). 54 mESCs were introduced on spotted patterns of a combination of five ECM proteins (fibronectin, laminin, and collagen I, III, and IV), and the optimal combination was identified to induce their differentiation into hepatocytes. Recently, Gobaa et al. 55 developed a high-throughput microarray consisting of hydrogel with variations in chemical and physical parameters in combination with proteins to form an artificial niche (cellular microenvironment) at the single-cell level and to identify a better niche for adipogenic or osteogenic differentiation from human mesenchymal stem cells. Solanki et al. 56 also revealed that structures of micropatterned ECMs strongly influence the lineage specification of neural differentiation. They introduced neural stem cells (NSCs) on the substrate with grid, square, and stripe structures and found that striped ECM induced the differentiation of NSCs into astrocytes but not neurons. Conversely, a grid ECM shape resulted in more neural differentiation than other substrates.
Microwell Array
To carry out reproducible EB formation leading to robust differentiation efficiency into target cells, researchers should use microwell arrays instead of 2D microcontact printing technology due to the capability to provide precisely controlled 3D geometry for EBs.48,49,57–60 Azarin et al. 57 found that the defined physical environments determined by microwells modulated the intercellular interaction of hESCs via the WNT/β-catenin signaling pathway, and EBs obtained from a microwell increased the genes associated with cardiogenesis as compared with EBs derived from hESCs cultured on 2D substrates. These results indicate that microwell arrays provide a platform not only for generating EBs with a uniform size but also for modulating intercellular interactions.
Microtissue Engineering
In general, cells alone in a culture dish cannot have their own function. In our body, cells are well organized into a 3D structure and have their functions within a tissue. To obtain functional tissues, we need to create well-organized 3D-structured cell aggregates using multiple types of cells. To address this issue, Torisawa et al. 29 applied hydrodynamic focusing for multiple types of cells into a single-cell culture chamber in a microfluidic device and successfully demonstrated formation of 3D heterogeneous cell patterning with mESCs and HepG2 cells ( Fig. 4c ). For inducing neural progenitor cells from hESCs, Park et al. 61 also successfully applied protein concentration gradients generated by a microfluidic device to mimic in vivo early developmental processes, such as forming both dorso-vental and anterior-posterior axes in a 3D manner. In addition, using microfabricated wells combined with electrospun polymer nanofibers as cellular scaffolds, cells (i.e., hepatocytes, fibroblasts, pancreatic cells, ESCs, and cardiomyocytes) formed 3D clusters in a microwell. 62 To create functional tissues in vitro, a bio-inspired approach can also be used and reconstructed in vitro. One example is to mimic the retina by using microfabrication technology. 63 To re-create retinal tissues derived from hESCs, an array of microchannels that mimic the rod structure of a retinal tissue was fabricated, and dissociated cells differentiated from hESCs were introduced into the array. After co-culturing with retinal pigment epithelium, photoreceptor cells derived from hESCs showed similar characteristics of infiltration, morphology, and expression of both rod and Müller glia differentiation markers. Even though there are some reports of the formation of 3D microtissues by using microfabrication techniques as described above, few studies have demonstrated “in vivo–like” tissue functions, and these tissues have not been fully functional or matured.64,65 It would be necessary to organize multiple cell types in a 3D fashion to have real in vivo functions of the targeted tissues, and further development for the next generation of 3D cell arrangement technology is needed to create 3D functionalized tissues.
Vascular Network
To form functional 3D tissues for future applications in tissue engineering, there is a need for the formation of a vascular network to supply growth factors and supplements for 3D tissues. Without a vascular network, developed 3D tissues would become necrotic and lack their actual functions. Even though there have not been reports of the use of blood vessels derived from hPSCs, the methods for in vitro vascular formation have been developed for applications in studying angiogenesis and thrombosis as well as for 3D tissue engineering.66–68 Miller et al. 68 developed a method of enabling formation of a vascular network in a living 3D tissue ( Fig. 4d ). Here, they printed a rigid 3D filament network of carbohydrate glass, introduced endothelial cells into the network to form a vascular wall, and then added the medium. To establish an artificial liver using their method, they confirmed that their vascular networks supplied enough supplements for liver cells as well as gain of liver functions.
Functional Analyses of hPSC-Derived Differentiated Cells
To use differentiated cells from hPSCs for various applications, we need to evaluate their proper functionality for the targeted tissues or cells.69,70 For example, Shin et al. 69 used a microfluidic device for compartmentally cultured ESC-derived neurons to evaluate their capability for axon propagation. In this device, they isolated somatic cell bodies on one side of a microfluidic channel and observed axon propagation to the other side of a microchannel via narrow microchannels. In addition, Myers et al. 71 developed label-free methods to distinguish the desired functional cells, in this case cardiomyocytes, from a mixture of differentiated cells. They noticed that electrically excitable cells (i.e., cardiomyocytes, neurons, and smooth muscle cells) had unique electrophysiological properties. Because the other types of somatic cells and stem cells do not have such electrical activity, they found that this property could be used as a sorting parameter as an alternative to cell surface markers, which may influence cell phenotypes. They developed electrophysiology-activated cell cytometry with a microfluidic device in combination with multiple electrodes to stimulate cells and measure the extracellular field potential. They were able to specifically sort cardiomyocytes derived from induced PSCs (iPSCs). This approach was unique because the targeted cells were sorted by measuring their functional activity, and this method will be significant in the field of stem cell research.
Reprogramming Somatic Cells
Since Takahashi and Yamanaka 72 developed reprogramming techniques for generating iPSCs by introducing a simple combination of several transcription factors into somatic cells, these techniques have attracted considerable attention from many researchers. There are several methods to reprogram cells, such as nuclear transfer, cell fusion, and introduction of egg extract.
Cell Fusion
Cell fusion is an important process for the maturation of cells in some tissues such as muscle and bone. In addition, it is also used to generate hybridomas, which are fusions of antibody-producing B cells and myeloma cells, for the production of monoclonal antibodies in culture. 73 Alternatively, forced cell fusion is also used for cell reprogramming. 74 Although many methods for cell fusion have been established, there are some issues in performing cell fusion in vitro, such as low efficiency because of random cell-cell pairing. To overcome this issue, researchers have established single cell-cell pairing and fusion, but this requires labor-intensive procedures and lower throughput. In an attempt to resolve this issue, Skelley et al. 75 developed a microfluidic device, which allowed the capture and pairing of a large number of cells, and integrated electrodes into the device for performing electrofusion ( Fig. 5 ). They successfully performed cell fusion with both electrofusion and chemical fusion using polyethylene glycol in their microfluidic device and found that electrofusion gave a better cell fusion efficiency than the chemical method. Using their device, the efficiency of cell fusion was increased to 50%, and reprogramming was observed with the fused cells by using both mESCs and MEFs. Subsequently, a number of studies describing cell pairing and fusion of somatic and stem cells for reprogramming were published, 76 but considerable progress needs to be made to generate hiPSCs in a robust and highly efficient fashion, since the generation of hiPSCs by cell fusion has not been achieved owing to the dramatically lower reprogramming efficiency of human cells than that of mouse cells.
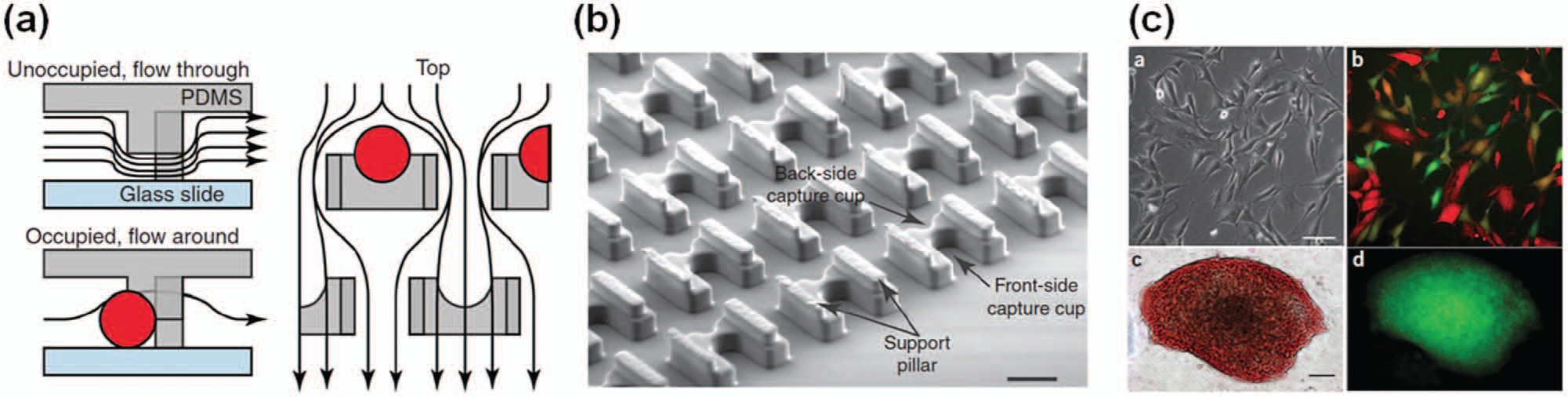
Cell fusion platform for individual cell-cell pairing for reprogramming cells and generating hybridomas. 75 PDMS, polydimethylsiloxane. Reproduced with permission from The Nature Publishing Group.
Drug Testing and Screening
Because hPSCs have the ability of unlimited expansion and differentiation into almost all types of cells in a body, hPSCs offer a source of an in vitro cell-based assay for drug screening as well as toxicological tests.77,78 In addition, a number of disease models using hPSCs have been established,77–79 and it is becoming more important to investigate disease mechanisms to discover novel methods for curing disease. Indeed, microfabricated devices with hPSC-derived cells have been used for developing in vitro functional cell assays for pharmacological as well as toxicological analyses.65,80 Although there are a limited number of reports on the use of microfluidic devices for hPSC-based drug screening, this would be a promising approach to reduce reagent/sample consumption as well as establish robust cell-based assay platforms. For the establishment of high-throughput screening, a microfluidic device was designed to fit with a commonly used robotic liquid dispenser with an SBS/ANSI standard for automated cell/sample loading. 81 This microfluidic device has a passive pumping system for liquid loading and does not require any tubing to connect or control the solution in the device.
Analysis of Cellular Heterogeneity
Cells naturally form heterogeneous populations in culture as well as in tissues and in the body. Particularly, during induction of differentiation from stem cells to a specific lineage as well as during cell reprogramming, we often obtain not only the target type of cells but also undesired random cells. To increase the efficiency to induce a specific differentiation/reprogramming process, it is very important to trace information of individual cells.82–84 Thus, cellular heterogeneity is now becoming a key issue to investigate developmental processes from stem cells and control stem cell functions as desired.
Cell Sorting and Capturing
Cell sorting, such as fluorescence-activated cell sorting (FACS) and magnet-activated cell sorting (MACS), is commonly used to measure cellular heterogeneity by detecting the expression of markers as well as phosphorylated proteins. Many studies in the literature describe that the use of microfluidic devices enable FACS or MACS. Dielectrophoresis also has been used to capture cells by their dielectricity by subjecting them to a nonuniform electric field. One issue with FACS and MACS is the detection of rare cells within a cell population, for which cell capturing would be suitable. At present, there is a growing interest to develop a microfluidic device with an array of microposts conjugated with EpCAM antibodies to capture circulating tumor cells, which are hardly detectable in the blood of cancer patients, to diagnose their metastatic status.85,86 Furthermore, for application of cell capturing techniques, Lawrenz et al. 87 systematically investigated the effects of the geometry of microfluidic chambers and facilitated the capturing efficiencies for various types of cells, including hESCs. For cell transplantation using differentiated hPSCs, contamination of undifferentiated hPSCs would carry a risk of tumorigenicity, and thus, it is very important to investigate whether injected cells contain any undifferentiated hPSCs before cell transplantation.
Single-Cell Analyses
Conventional -omics technologies require large numbers of cells to obtain reproducible data, but at the same time, these data average out individual cells and miss their information. However, several issues need to be considered. For example, because each cell has a small amount of targetable molecules such as DNA, RNA, and proteins, the experimental setup for single-cell analysis should have a sensitive detection system. Moreover, these systems require large numbers of cells to understand the degree of cell heterogeneity; hence, the experimental procedures to observe and analyze individual cells are labor-intensive. To establish a new method for single-cell analysis, microfluidic devices are advantageous because of their unique capabilities of precise liquid handling as well as automation. Thorsen et al. 88 developed largely integrated and automated microfluidic devices with mechanical valves and then established a “digital polymerase chain reaction (PCR) array” for enabling large numbers of single-cell reverse transcription (RT)–PCRs in a single device.89,90 White et al. 91 also developed microfluidic-based high-throughput single-cell RT-PCR to analyze the gene expression of 3300 individual cells in a single device. Microfluidic-based RT-PCR systems are maturing in terms of sensitivity and selectivity, and they can be applied in various research and clinical settings.92,93 However, although single-cell RT-PCR is commonly used by many researchers, single-cell genomic sequencing is still challenging because each cell has only femtograms of genomic DNA below the detection levels of the system. Furthermore, methodological development of other single-cell -omics analyses (i.e., proteomics, epigenetics, and metabolomics) is now even more challenging. For development in stem cell research, these methodological developments are necessary, and I believe microfluidic technology would be beneficial for this development.
Nonviral Gene Transfection
To investigate the role of cellular signaling networks, gene transfection (plasmids or siRNA) is generally used to perturb signaling nodes by overexpressing or silencing target genes of interest, but the efficiency of gene transfection into hPSCs is much lower than that of other cell types. Although a number of studies have described gene transfection using microfluidic platforms in conjunction with electroporation for facilitating gene transfection efficiency,94–96 there are few examples for mESCs, 97 and it is still challenging to apply these techniques to hPSCs. Recently, gene-editing technologies have been developed,98–100 and it is crucial to enhance gene transfection efficiency into hPSCs.
Cautions for the Use of Microfabricated Tools in hPSC Research
As I described in this review, microfabrication technologies are promising for facilitating hPSC research and its applications. However, for further development of these technologies, some issues need to be solved.
Complexity of Microfabricated Tools
One of the issues is that these technologies often require special instruments and software to control the microfabricated tools, as well as require complicated fabrication processes sometimes. Some microfabricated tools are very attractive because of their sophisticated designs, but their functioning is affected because of their complexity. Researchers in stem cell biology would prefer a simple design of microfabricated tools for daily use. Some researchers have tackled these issues by developing very simple devices.22,81 These tools do not require any special instruments and work with a general pipette as well as a robotic liquid dispenser. Of course, sophisticated designs of microfabricated tools provide more functions and precision for experiments, but it does not make sense if they are not used. Therefore, it is important to strike a balance between simplicity and complexity.
Microfabrication Materials
To begin experiments for stem cell research using microfabricated platforms, we also need to know the effects of the materials used for these tools because some materials may influence experimental conditions and cause artifacts in the obtained data. Several groups, including ours, are working on investigating the effects of these materials on cells.101–104 For example, PDMS is commonly used for fabricating microfluidic devices, but it is also well known that PDMS absorbs hydrophobic small molecules in solution.101,102 For cellular experiments, because cell culture medium contains hydrophobic molecules, the absorption of these molecules would affect cellular viability in a PDMS-based microfluidic device. In addition, some drug candidates often possess hydrophobic parts that might reduce drug efficacy on cells in a microfluidic device. We also investigated the effects of microfabrication materials, including PDMS and negative photoresists, on cellular phenotypes as well as gene expression, which is related to hPSC self-renewal and differentiation. 104 We found that these materials did not change the expression of pluripotent markers (i.e., alkaline phosphatase and SSEA4) but altered the expression of differentiation-related genes as confirmed by a multiplex quantitative RT-PCR array. Those materials preferentially induced MYF5 expression, which is related to muscle differentiation, and suppressed expression of LEFTY2, a self-renewal–related gene. These results suggest that the material itself may cause induction of hPSC differentiation, and we need to consider our obtained results from experiments by using microfabricated tools and find the optimal materials for experimental purposes.
In summary, cutting-edge microfabricated tools have already been developed for hPSC research. After the establishment of hiPSCs by Takahashi and his colleagues, 2 the big wave of research on developing microfabricated tools progressed to applications for hiPSC research. As shown in this review, some tools were successfully used by many researchers and were useful for solving many questions and problems. However, the others were on a “proof-of-concept” stage and did not move to the next level. For example, as mentioned above, microfabricated tools are promising for the establishment of fully functional and mature tissues, but this has not been achieved because of the lack of a precise control of multiple cells and blood vessels within a microtissue. In addition, although a number of types of microfluidic platforms have been developed, few stem cell biologists have used them for solving problems owing to the lack of simplicity of use. Additionally, they were often out of the needs or interests of stem cell research. This is because the interaction between stem cell biologists and microengineers has not been sufficient for solving these problems. hPSC-based products and therapies will be available on the market in the near future, and for its realization, we need to have strong interactions and stimulations among researchers in microengineering, stem cell biology, and clinical settings, as well as in academia, industry, and hospitals. Even though there have been a number of discussions over the years to find “killer applications” of microfluidics and related technologies, especially for stem cell research,105–107 I believe that stem cell biology and its applications in the real world will be “killer applications” for microfabricated tools and that these tools will contribute to the acceleration of progress.
Footnotes
Acknowledgements
The author thanks all laboratory members and colleagues for helpful discussion of this review.
Declaration of Conflicting Interests
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by the Japan Society for the Promotion of Science (JSPS; Grants-in-Aid for Young Scientists [A] 23681028 and Challenging Exploratory Research [24656502]) as well as the New Energy and Industrial Technology Development Organization (NEDO) in Japan (11100210). The WPI-iCeMS is supported by the World Premier International Research Center Initiative (WPI), MEXT, Japan.
