Abstract
Elucidation of the heterogeneity of cells is a challenging task due to the lack of efficient analytical tools to make measurements with single-cell resolution. Microfluidics has emerged as a powerful platform for single-cell analysis with the ability to manipulate small volume and integrate multiple sample preparation steps into one device. In this review, we discuss the differentiating advantages of microfluidic platforms that have been demonstrated for single-cell protein analysis.
Introduction
Proteins regulate almost all cellular functions. Conventional protein analysis techniques require a large population of cells. While these studies provide rich information on cell population, population-averaged results do not allow measuring cell-to-cell variation, which can provide important biological insights.1–4 Cells under seemingly identical extrinsic environmental conditions often display heterogeneous behaviors. An example is nuclear factor (NF)–κB trafficking: cells challenged with tumor necrosis factor–α (TNF-α) display quantitative differences from cell to cell in NF-κB nuclear localization.1,2 Similar examples have also been reported in tumor progression, stem cell differentiation, and host-pathogen interactions.3,5 However, protein analysis at a single cell level is challenging. The hurdles lie in the low abundance of proteins in single cells, 6 the large dynamic range of many protein constituents within one cell, 7 and the transitory existence of cellular proteins responding to stimulation.8,9 To overcome these hurdles, several analytical instrumentation and techniques such as flow cytometry,10,11 mass cytometry,12,13 affinity arrays, 14 and mass spectrometry15,16 have been developed over the years. For example, the invention of fluorescence-activated cell sorting (FACS) greatly improved single-cell analysis with high-throughput and high-content information.10,11 One limitation of FACS, though, is that because sample preparation is done manually, a large number of cells are often required, which makes it difficult to analyze rare samples such as primary cells and biopsy samples.
Microfluidics has emerged as a powerful platform for single-cell analysis with the capability to manipulate small volume and integrate multiple sample preparation steps into one device. Microfluidic chips have characteristic dimensions at the same scale as the dimension of a cell, which allows very rapid exchange of buffers and reagents, thereby minimizing gradients of stimulants experienced by cells. The small volume contained in microfluidic devices allows experiments requiring a lower amount of less expensive reagents/rare cells and enables a higher number of experiments from a given amount of sample. Other advantages include precise control of the cellular microenvironment and the ability to encapsulate a single cell in droplets. Because of these distinct capabilities, many state-of-the-art microfluidic platforms have been developed for single-cell protein analysis in the past few years.17–22
Differentiating Capabilities of Microfluidics
Many biochemical analyses have benefited from the unique advantages offered by microfluidics.23,24 Laminar flows in microfluidic channels enable well-defined and stable liquid-liquid interfaces down to cellular dimensions. One of the early examples using this distinct capability is the selective treatment of cellular microdomains, resulting electively in the delivery of membrane-permeable molecules to specific microdomains of single cells ( Fig. 1A ).25,26 Similarly, lateral propagation of epidermal growth factor (EGF) signaling in single live cells was revealed by laminar flows, providing evidence that ligand-independent propagation of EGF signaling depends on the receptor density of the plasma membrane. 27
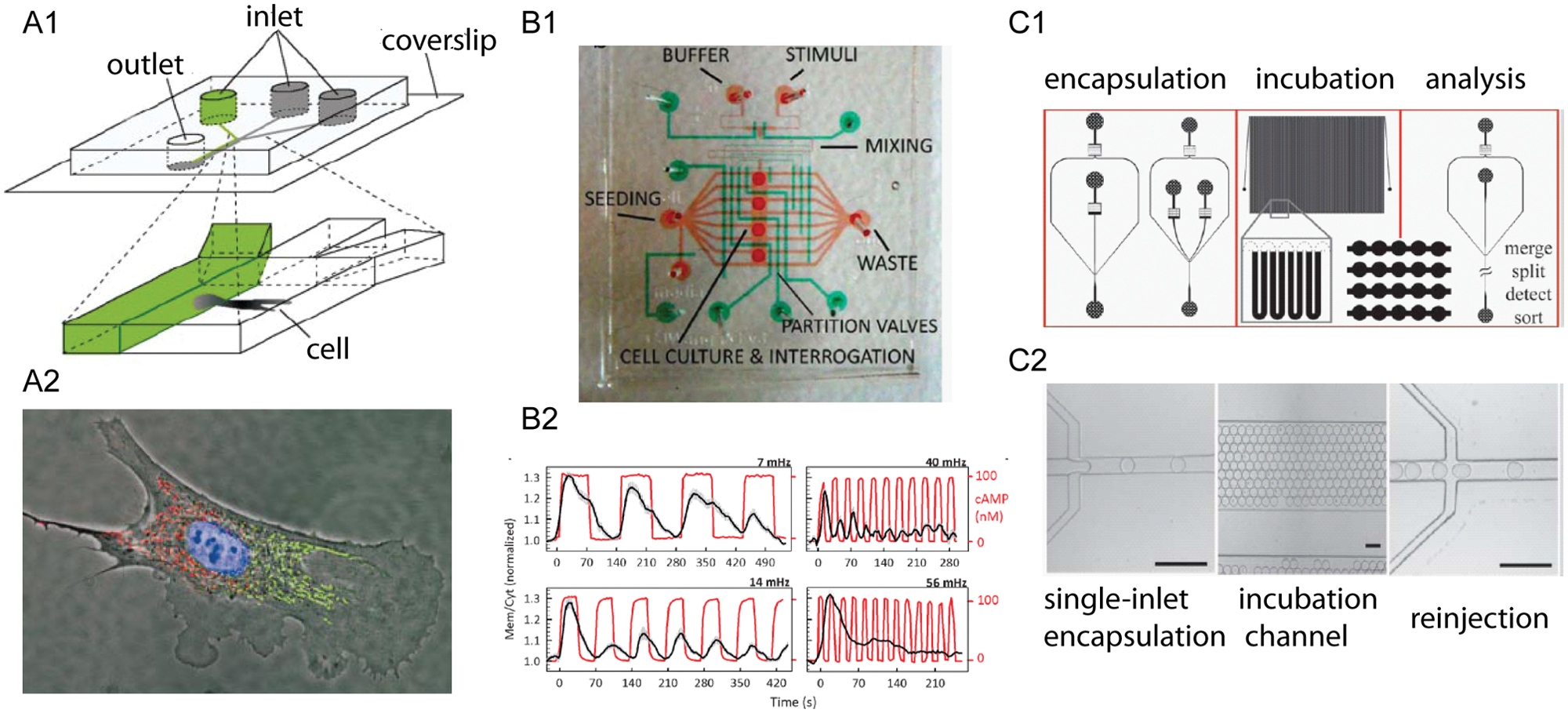
Examples of biological applications benefiting from differentiating capabilities of microfluidics. (
Another unique capability of microfluidics is the generation of spatial gradients of fluidics through progressive serial dilution of multiple input streams into a series of streams spanning several orders of magnitude in concentration. 28 For example, Thompson et al. 29 generated multiple concentrations of stimulus in an upstream microfluidic network to examine the activation of transcription factor NF-κB in HeLa cells.
The advancement in zero dead-volume valves and control software has enabled the precise control of fluidics in space, time, and magnitude. It allows interesting exposure profiles to single cells, including pulsatile, ramp, square-wave, and oscillatory signals,20,30–33 whereas only step flow is feasible in a conventional culture dish. Precise control of fluidic profiles over cells has been used to measure protein levels in fundamental cellular behaviors, including inflammatory responses and cellular signaling pathways. For instance, Mettetal et al. 30 used a flow-cell setup to produce periodic stimuli to measure the frequency dependence of signal processing in the osmo-adaptation pathway. Hersen et al. 32 designed a microfluidic device to reliably manipulate a single cellular environment over a range of pulsatile frequencies and measured the signal transduction by the HOG MAP kinase pathway. More recently, Wang et al. 31 demonstrated the use of a polydimethylsiloxane (PDMS)–based microfluidic function generator to characterize the signal response upon pulses of chemoattractant cyclic adenosine monophosphate (cAMP) ( Fig. 1B ). The precise control of the cellular microenvironment in microfluidic platforms has proven to be indispensable to study dynamic changes of proteins at cellular and subcellular resolution.
Another feature of microfluidics is the ability to compartmentalize individual cells in a nanoliter or smaller volume for biochemical analysis. This has led to analysis of single cells with minimal sample dilution, shorter reaction times, and higher sensitivity.21,34–36 One group encapsulated a desired cell into a droplet and then photolyzed the cell rapidly and conducted fluorescent microscopy–based single-cell assays. 37 Coupled with precise generation and the repeatability of droplet generation, this system has the potential to become a high-throughput platform for cellular proteomics. 38 Another fluorescence-activated droplet sorting platform processed cells at rates of about 2000 droplets per second, allowing researchers to accurately describe the heterogeneity of the cell population. 39 The droplets also serve as a “container” for secreted biomolecules, minimizing sample dilution and boosting detection sensitivity. One can rapidly analyze secreted proteins (e.g., antibodies or cytokines) or released metabolites (e.g., biomarkers or vitamins) from thousands of encapsulated single cells.40–42 Huebner et al. 42 developed a quantitative enzyme assay in microdroplets. Koester et al. 40 also have demonstrated the detection of secreted antibodies in single hydridoma cells in picodrops ( Fig. 1C ). In addition, Joensson et al. 43 described the detection of cell-surface protein biomarkers CCR5 and CD19 on individual human cells (U937) using enzymatic amplification in microfluidic droplets with high sensitivity and high throughput.
Fabrication of Microfluidic Devices
Historically, silicon and glass have been the preferred substrates for the fabrication of microfluidic chips. 44 They remain attractive for many biological applications because of the following properties: large optical transition range (180–2500 nm); high resistance to mechanical stress, heat, and chemicals; and high electric isolation and high biocompatibility through its well-studied surface chemistry. Of the three major glass etching techniques—mechanical, dry, and wet—the most practiced method is isotropic wet etching. This method uses a photolithography mask to define features on the surface of the wafer. Timed exposure to chemical etchants such as HF dissolves the Si-O-Si bonds in the glass isotropically, generating a D-shaped channel with a smooth surface and a constant depth.
Today, the materials that used to fabricate microfluidic devices have been expanded from glass and silicon to polymers, paper, tape, fabric, and thread.45–48 Among those, the most used material is PDMS.49,50 PDMS is optically transparent (240–1100nm), permeable to atmospheric gases, chemically inert, mechanically flexible, and electrically insulating. 51 The transparency allows the use of transmitted light and fluorescent signals for imaging cell activity. Permeability to oxygen and carbon dioxide is beneficial for culturing mammalian cells within devices over extended periods. The flexibility allows easy integration of fluidic tubings and valves. PDMS-based microfluidic devices are attractive for rapid prototyping because the turnaround of PDMS devices can be as fast as a few days. The critical drawbacks of PDMS in biological experiments include the adsorption of small hydrophobic molecules and proteins from the solution to the surface of PDMS.52,53
Microfluidic Platforms for Single-Cell Protein Analysis
There are three essential elements for carrying out single-cell protein analysis: single-cell isolation, cell manipulation, and detection of intracellular proteins. The strategies of isolating individual cells fall into two categories: biochemical and physical. Physical trapping methods include single-cell trapping array, 4 microvalves,54,55 droplet encapsulation, 40 dielectrophoresis, 56 electrophoresis,57,58 and optical trapping.59,60 Biochemical trapping methods include trapping the cells using antibodies 61 and/or cellular-matrix protein-assisted adhesion.62,63 Proteins in isolated single cells can be analyzed in intact (or whole) cells or analyzed after lysis. Sensitive detection of intracellular proteins is desired for quantitative measurements from single cells on a microfluidic device. Main detection technologies that have been coupled with microfluidics for single-cell analysis include fluorescent microscopy,14,33,64 laser-induced fluorescence (LIF),55,65,66 flow cytometry,67,68 and mass spectrometry (MS).15,69 Fluorescence-based technologies provide high-throughput and high-information content analysis but often are limited by the available genetic and chemical probes; MS can be highly sensitive and requires neither prelabeling nor purification of proteins. However, the lack of an effective interface between sample preparation and the detection is an obstacle for MS to be widely applied in single-cell protein analysis via microfluidics. 70 Nevertheless, integrated microfluidic platforms enable single-cell protein analysis by minimizing sample dilution and by offering efficient separation capabilities, which will be discussed in the following sections.
Intact Cell Analysis
Intact cell analysis refers to detection of proteins without lysing the cells and is typically done by immunostaining or arrays (for secreted cells). The advantage of this approach is that the proteins are maintained at their intracellular concentrations that are orders of magnitude higher than those upon cell lysis. In a typical setting, cells are cultured on chip before external stimulation. Intracellular proteins are then probed with fluorescent antibodies for downstream measurement. Groisman et al.
71
introduced a microfabricated elastomer chip wherein chemostatic conditions for bacterial and yeast are well maintained. Tay et al.
33
demonstrated the use of a versatile microfluidic cell culture system
72
to study how single cells respond to TNF-α. They found that the process of cell activation is digital at the single-cell level but graded at the population level (
Fig. 2A
). To link single-cell genomics with proteomics, Taniguchi et al.
64
reported a microfluidic device–based approach to quantifying the
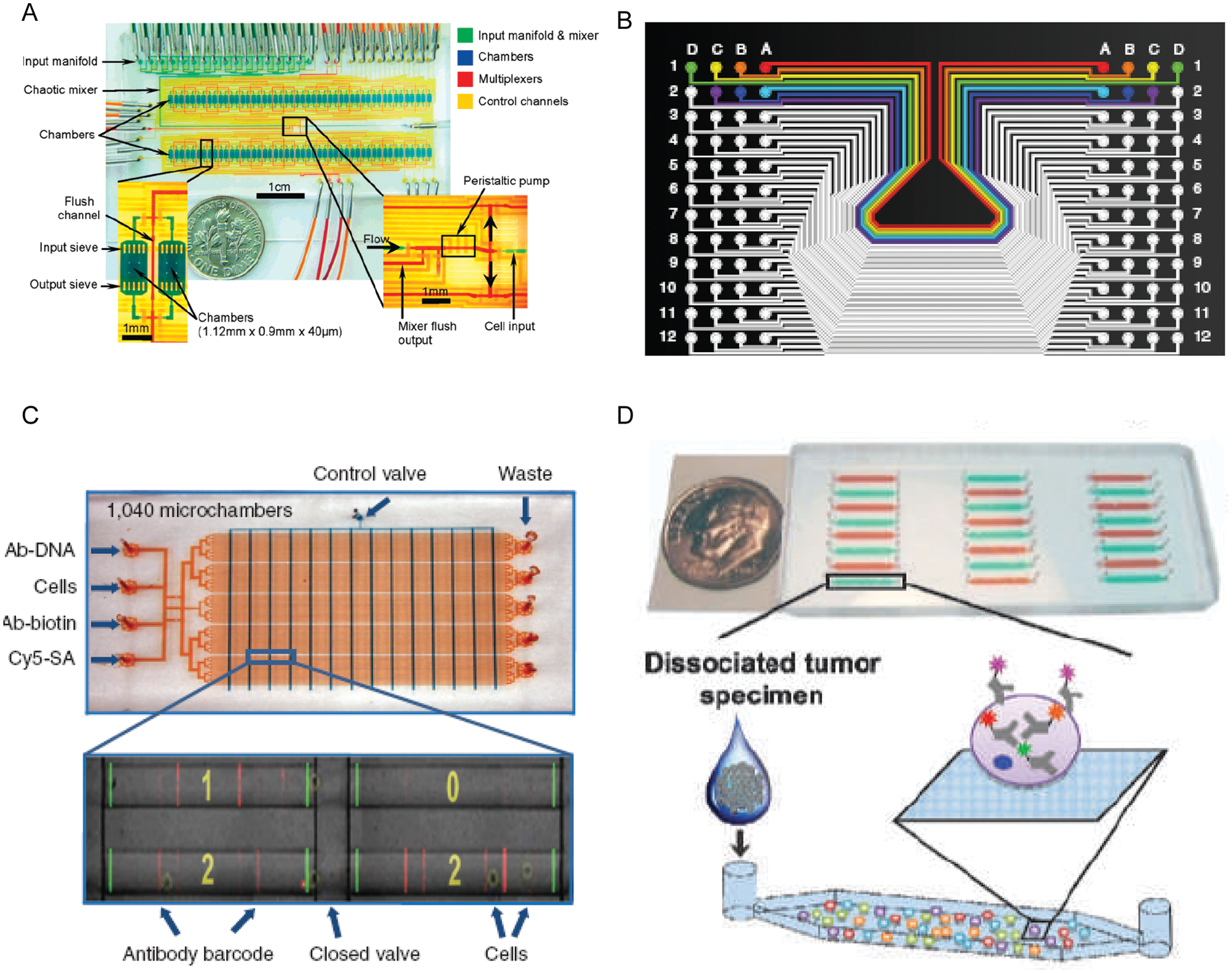
Microfluidic platforms for intact cell analysis. (
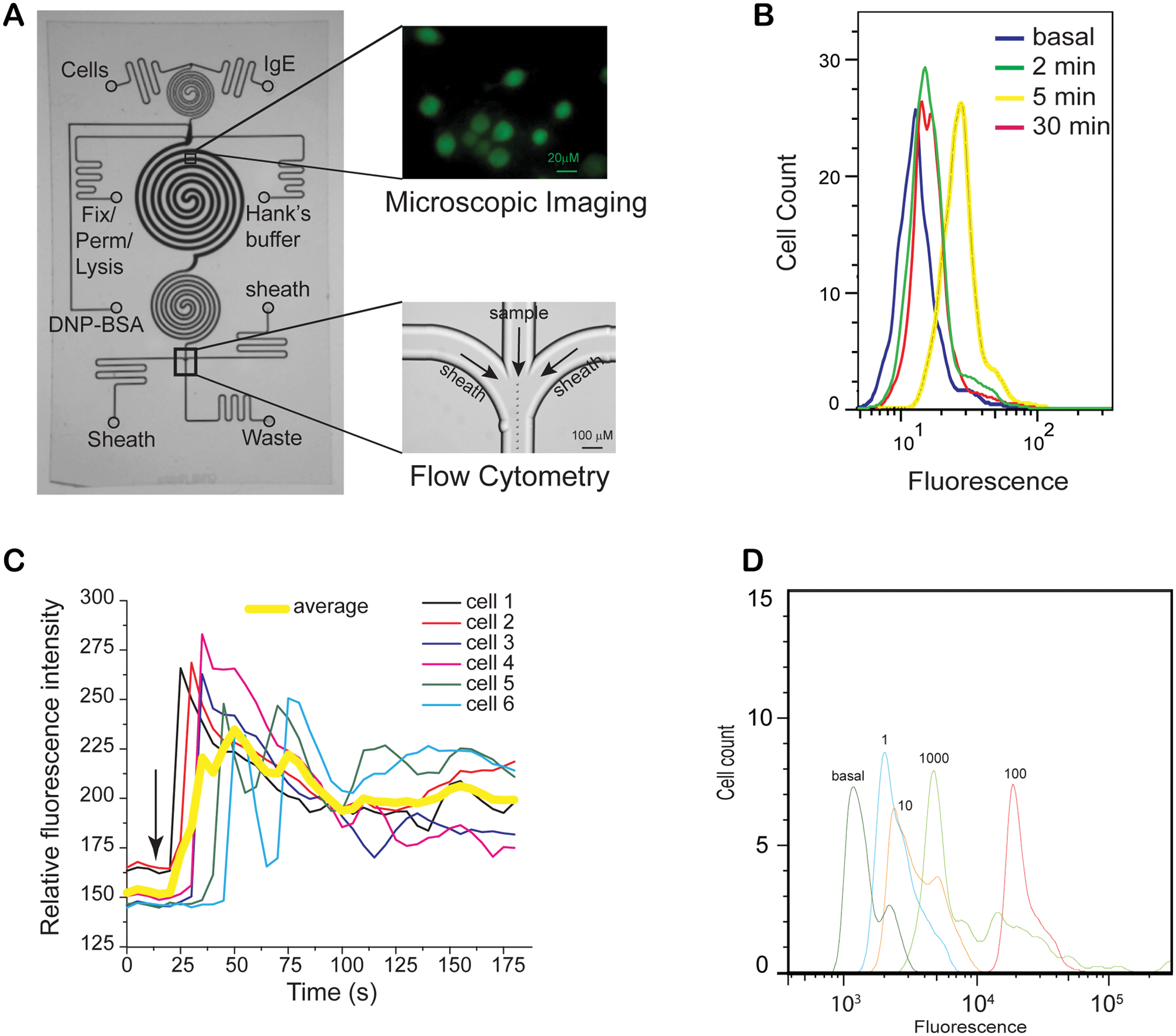
An integrated microfluidic platform for single-cell measurement of allergic pathway. (
Cell Lysate Analysis
This approach is especially useful when the components are impossible to label in intact cells. To achieve efficient lysis, chemical lysis is widely used to disrupt the interaction of cell membrane and the substrate.76,77 Cells can also be lysed by electroporation under a sufficiently high electric field.78,79 Laser photolysis can rapidly lyse cells so that cells are frozen at the moment of photolysis.37,80 For cellular proteins that cannot be amplified, efficient separation and sensitive detection are desired for identification and analysis. The separation modalities such as chromatography and electrophoresis within microchannels can be incorporated into the same chip to perform high-resolution analysis after efficient sample transfer. Several approaches have been established to integrate sample preparation, lysate separation, and subsequent identification on-chip. Microchip capillary electrophoresis (CE) has shown great promise for separating lysate. Wu and coworkers 55 demonstrated the use of chemical cytometry to analyze the chemical content of a single cell. The same group then improved the multistate valve to detect low-copy-number proteins (less than 1000 molecules/cell) in individual cells ( Fig. 4A ). 66 The development of PDMS-glass hybrid chips and the detection of amino acids through chemiluminescence also improved the usability of microfluidic CE.81,82
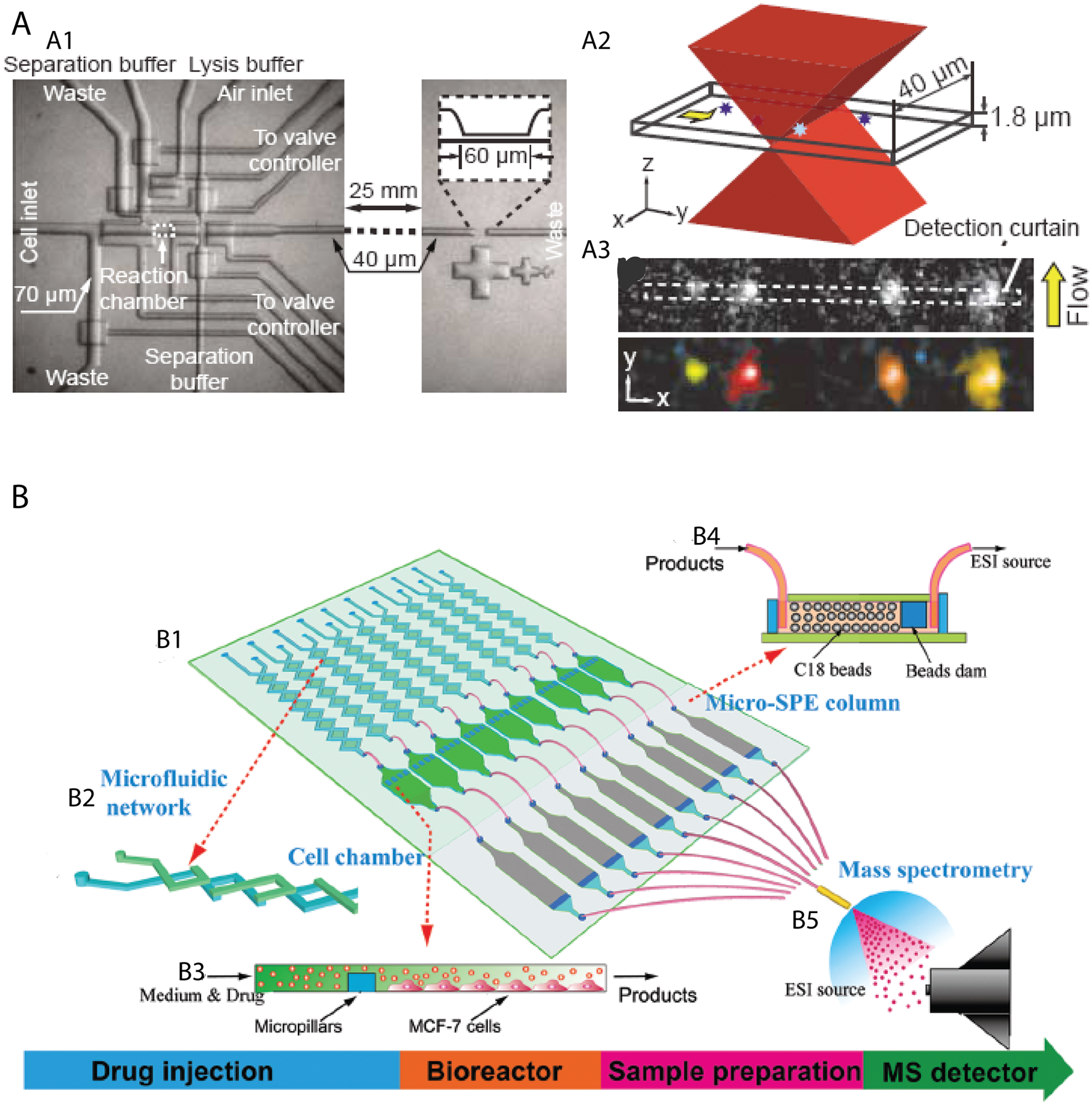
Microfluidic platforms for cell lysate analysis. (
To address the lack of an effective interface between miniaturized sample preparation and the mass spectrometer, 70 Mellors et al. 15 reported continuous on-chip lysis of erythrocytes with real-time electrophoretic separation and electrospray ionization MS. More recently, Chen et al. 69 demonstrated a isotope labeling–assisted microfluidic chip electrospray ionization MS platform for quantitative and qualitative analysis of multiplex metabolites in MCF-7 cells ( Fig. 4B ).
Conclusions and Future Outlook
In recent years, the microfluidic community has shifted its focus from technology development to biological applications, realizing the urgent need of bringing the technology from promise to practice. As a result, we have seen many noteworthy applications such as elucidating the heterogeneities of cell populations, monitoring intracellular protein dynamics by precisely controlling the cellular microenvironment, and detecting a low copy number of proteins in single cells.
However, nontrivial challenges need to be addressed before microfluidic devices for single-cell analysis gain wide acceptance from the biological research community. First, we need to develop assays with higher sensitivity. The most interesting proteins in cells tend to be present at low concentrations and cannot be measured by antibody-based approaches. Advances in detection technologies and improved affinity reagents can also play a major role. Second, it is difficult to measure proteins in living cells. Development of genetically engineered fluorescent proteins has been revolutionary in interrogating proteins in living cells, but the technology is limited to analysis of cell lines. Another limitation has been measurement not only of the proteins but also of DNA and RNA so that one can correlate them to get a systems-level picture. Finally, seamless integration of materials, fluidics handling, and hardware control and detection systems is desired. Microfluidics has the potential to address some critical issues of single-cell proteomics, but it cannot be a system with much complexity and poor stability. It is imperative to translate the proof-of-concept experiments into robust and easy-to-use techniques that biologists will adopt.
Despite all these challenges, the future of microfluidics in single-cell protein analysis is promising. Microfluidics offers many differentiating capabilities that could not be possible without microfluidics. The deep collaboration between chemists, biologists, and engineers would bring microfluidics to a new level toward elucidation of a single-cell proteome.
Footnotes
Acknowledgements
We thank Dr. Mais J. Jebrail for critical reading of this manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Financial support for preparation and some of the work included was provided by the following grants: R01 DE020891, funded by the National Institute of Dental and Craniofacial Research; P50GM085273 (the New Mexico Spatiotemporal Modeling Center), funded by the National Institute of General Medical Sciences; and ENIGMA, a Lawrence Berkeley National Laboratory Scientific Focus Area Program supported by the U.S. Department of Energy, Office of Science, Office of Biological and Environmental Research. Sandia National Laboratories is a multiprogram laboratory managed and operated by Sandia Corporation, a wholly owned subsidiary of Lockheed Martin Corporation, for the U.S. Department of Energy’s National Nuclear Security Administration under contract DE-AC04-94AL85000.
