Abstract
Our aim was to evaluate whether automation for the preanalytical phase improves data quality. Blood from 100 volunteers was collected into two vacuum tubes. One sample from each volunteer was respectively assigned to (G1) traditional processing, starting with centrifugation at 1200g for 10 min, and (G2) the MODULAR PRE-ANALYTICALS EVO-MPA system. The routine clinical chemistry tests were performed in duplicate on the same instrument Cobas 6000 <c501> module. G1 samples were uncapped manually and immediately placed into the instrument. G2 samples were directly fed from the MPA system to the instrument without further staff intervention. At the end, (1) the G1 samples were stored for 6 h at 4 °C as prescribed in our accredited laboratory and (2) the G2 samples were stored for 6 h in the MPA output buffer. Results from G1 and G2, before and after storage, were compared. Significant increases were observed in G1 compared with G2 samples as follows: (1) before storage for alkaline phosphatase (ALP), lactate dehydrogenase (LDH), phosphate (P), magnesium (MG), iron (FE), and hemolysis index and (2) after storage for total cholesterol (COL), triglycerides (TG), total protein (TP), albumin (ALB), blood urea nitrogen (BUN), creatinine (CRE), uric acid (UA), ALP, pancreatic amylase, aspartate aminotransferase (AST), alanine aminotransferase (ALT), g-glutamyltransferase (GGT), LDH, creatine kinase (CK), calcium (CA), FE, sodium (NA), potassium (K), and hemolysis index. Moreover, significant increases were observed in (3) G1-after versus G1-before storage samples for COL, high-density lipoprotein cholesterol, TG, TP, ALB, BUN, CRE, UA, AST, ALT, GGT, LDH, P, CA, MG, FE, NA, K, and hemolysis index and (4) G2-after versus G2-before storage only for BUN, AST, LDH, P, and CA. In conclusion, our results show that the MPA system improves the quality of laboratory testing.
Most technological devices are increasingly devised to perform repetitive tasks so far pertaining to human operators, with advantages in terms of accuracy, speed, convenience, and cost. At the beginning of the 21st century, robotic automation is poised to revolutionize laboratory practices. 1 Laboratory testing is an integral part of the decision-making process, wherein results of laboratory testing often influence diagnosis and treatment of a variety of human disorders. 2 The final goal of automation and consolidation in clinical laboratories is to ensure reliable information to assist physicians in patient management and improving clinical outcomes.3–8 There is a long history of quality requirements in laboratory medicine, yet the concern has mainly regarded the analytical phase of this process. 9 However, greater appreciation of the prevalence of errors in the pre- and postanalytical phases and their potential for patient harm has led to increasing requirements for laboratories to take greater responsibility for activities outside their immediate control to guarantee patient safety. 10 Owing to substantial advances in technology, laboratory automation, and analytical quality, there is mounting evidence that further quality improvements should be targeted to extra-analytical activities of the total testing process.11–17 Several studies have shown that automation in laboratory medicine is effective to reduce the turnaround time.4,18,19 The aim of this study was to assess whether laboratory automation for the preanalytical phase improves quality and is also effective for improving patient safety.
Materials and Methods
Study Design
We selected a group of 100 adult ambulatory patients of both genders, 54 female and 46 male, with an average age of 49 years (range, 46–69 years) who were enrolled as volunteers in this study. The investigation was submitted to the Internal Review Board and approved by the local Human Research Ethics Committee. All volunteers signed an informed consent. All laboratory equipment (both for support and analyses) were calibrated, and the metrological traceability was guaranteed as recommended by the ISO 15189 document. 20
Collection of Diagnostic Blood Specimens
The collection of all blood samples was performed by a single, expert phlebotomist, according to the recommendations of the Clinical Laboratory Standard Institute (CLSI) 21 and producer. All volunteers, after 12 h of fasting, were maintained seated for 15 min prior to phlebotomy in order to prevent possible interferences of blood distribution due to posture. 22 After this time interval, a vein was located on forearm by a subcutaneous tissue transilluminator device, without tourniquet placement (Venoscópio IV plus, Duan do Brasil, Brazil) to prevent interference from venous stasis,23–25 and 7 mL of whole blood was collected by venipuncture with a 20 G straight needle (Terumo Europe NV, Leuven, Belgium) directly into two identical plasma vacuum tubes with 52.5 USP lithium heparin and gel separator Venosafe 3.5 mL (lot 1107001, Terumo Europe NV). To eliminate any potential interference due to either the contact phase or tissue factor, ~2 mL of blood was preliminarily collected in a discard tube without additive (Vacuette lot A101004D, Greiner Bio-One GmbH, Kremsmünster, Austria). The entire blood collection process was performed using needles and vacuum tubes of the same lot. The tubes, one from each volunteer, were separately processed as follows.
Traditional Processing of Diagnostic Blood Specimens (Gold Standard from CLSI)
One hundred tubes (one from each volunteer) were left in the upright position for 10 min at room temperature (20 °C) to allow complete in vitro anticoagulation before centrifugation 26 at 1200g for 10 min at room temperature (according to the instructions from Terumo) on the same centrifuge Rotanta 460R (Andreas Hettich GmbH & Co. KG, Tuttlingen, Germany). All primary tubes processed by traditional method were uncapped manually and immediately placed into the Cobas 6000 <c501> module (Roche Diagnostics GmbH, Mannheim, Germany). No signs of visual hemolysis were detected in any sample, and no specimen was discarded because of unsatisfactory attempts, problems in locating venous access, missing vein, or manifest lipemia.
Preanalytical Phase’s Laboratory Automation (Automated Processing)
Diagnostic Blood Specimen Processing
The other 100 tubes, one from each volunteer, were left in the upright position for 10 min at room temperature (20 °C) to allow complete in vitro anticoagulation. The tubes were then placed in racks suited for the following process into one MODULAR PRE-ANALYTICALS EVO-MPA system (Roche Diagnostics GmbH, Mannheim, Germany).
System Layout and Data Flow
The tubes followed the automated analytical process presently functioning in the laboratory, which includes the Laboratory Information System (LIS), the Process Systems Manager (PSM; formerly Preanalytic Systems Manager), Total System Manager (TSM), the MODULAR PRE-ANALYTICALS EVO-MPA modules, and the online MODULAR ANALYTICS EVO-MA systems. The TSM controls the operation of the MPA modules and their connection to the MA systems. The TSM computer has network interface cards for communicating with the LIS/PSM and the MPA over Ethernet connections. Communications between (1) TSM and the MPA and (2) the TSM and the LIS/PSM are strictly separated from each other. The MPA system is an automated, software-controlled system for processing patient samples before analysis. 27 The MPA system includes modules that centrifuge, aliquot patient samples, and automatically perform removal and insertion of container stoppers and barcode labeling. The automatic centrifuge unit (ACU) is a module inside MPA used to centrifuge samples based on parameters (such as rotation rate and duration) defined in the TSM. The centrifuge is self-balancing, and a single ACU can accommodate up to 40 samples (8 racks) at a time. A MODULAR PRE-ANALYTICS EVO system can be configured with either one or two ACU modules. For this purposes of this study, only an ACU was used for centrifugation at 1200g for 10 min at room temperature (according to the instructions from Terumo). Then only one aliquot from each primary vacuum tube was performed by MPA system. Moreover, this software-controlled system guarantees the traceability of all aliquots generated.
Laboratory Testing
The routine clinical biochemistry tests were performed in duplicate to minimize the analytical coefficient of variability (CVa), so the CVa from analyses in duplicate is smaller than the CVa from internal quality control. All samples were assayed on the same instrument, Cobas 6000 <c501> module (Roche Diagnostics GmbH), according to the manufacturer’s specifications and using proprietary reagents. The panel of tests included the following: total cholesterol (COL), high-density lipoprotein cholesterol (HDL), triglycerides (TG), total protein (TP), albumin (ALB), blood urea nitrogen (BUN), creatinine (CRE), C-reactive protein (CRP), uric acid (UA), alkaline phosphatase (ALP), amylase (AMYL), pancreatic amylase (AMY-P), aspartate aminotransferase (AST), alanine aminotransferase (ALT), g-glutamyltransferase (GGT), lactate dehydrogenase (LDH), lipase (LIP), creatine kinase (CK), total bilirubin (BT), direct bilirubin (BD), phosphate (P), calcium (CA), magnesium (MG), iron (FE), sodium (NA), potassium (K), and hemolysis index (HI). The instrument was calibrated against appropriate proprietary reference standard material and verified with the use of proprietary quality controls. Our evaluation of the within-run precision by internal quality control on Cobas 6000 <c501> module yielded very modest coefficients of variation ( Table 1 ). Samples from the MPA system were fed directly into the same Cobas 6000 <c501> module, without further staff intervention. A completely automated sample flow, starting from primary tube through to analysis and disposal, was thereby realized.
Processing of diagnostic blood specimen’s variability as mean % differences in a routine clinical chemistry laboratory test.
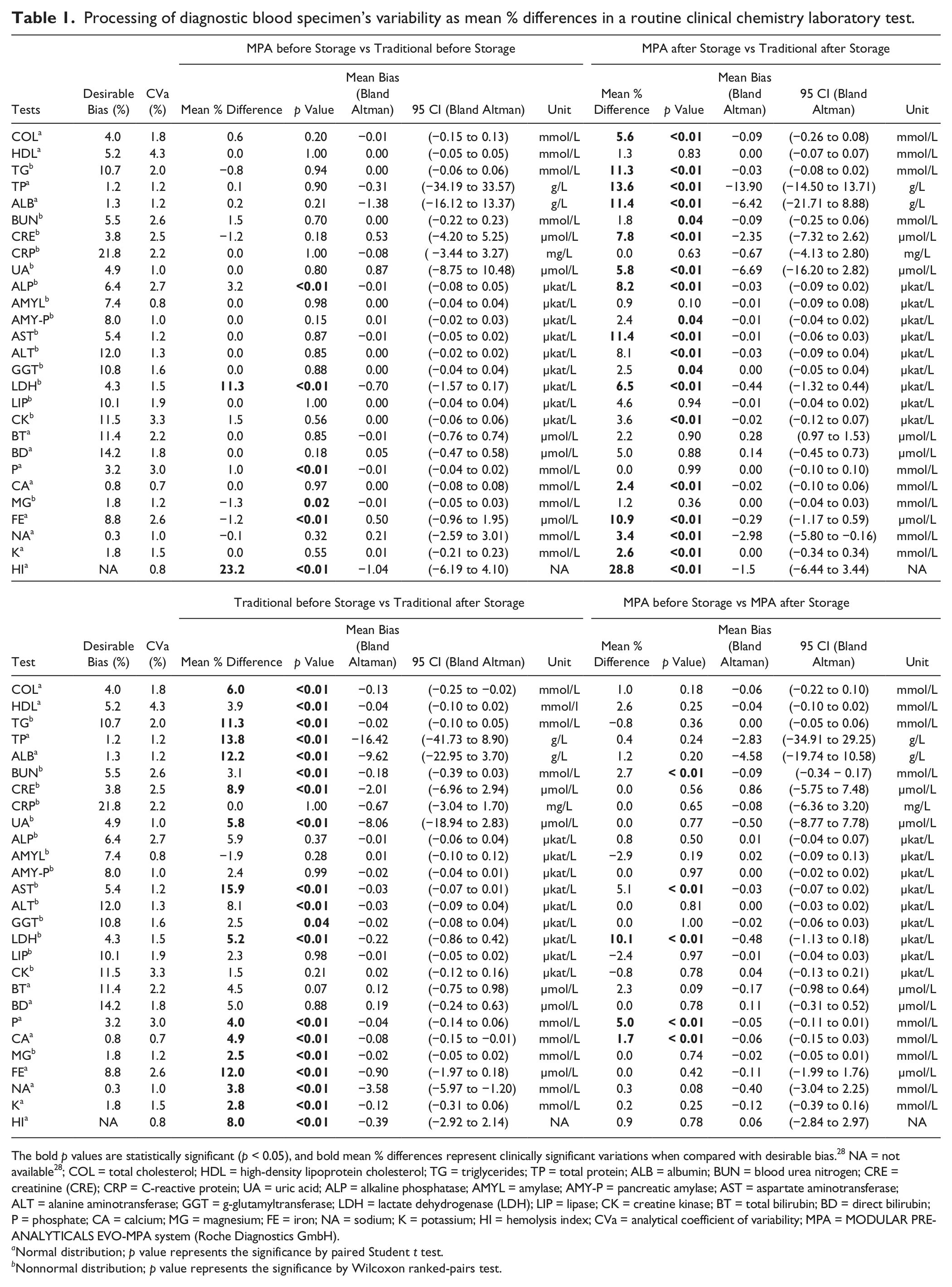
The bold p values are statistically significant (p < 0.05), and bold mean % differences represent clinically significant variations when compared with desirable bias. 28 NA = not available 28 ; COL = total cholesterol; HDL = high-density lipoprotein cholesterol; TG = triglycerides; TP = total protein; ALB = albumin; BUN = blood urea nitrogen; CRE = creatinine (CRE); CRP = C-reactive protein; UA = uric acid; ALP = alkaline phosphatase; AMYL = amylase; AMY-P = pancreatic amylase; AST = aspartate aminotransferase; ALT = alanine aminotransferase; GGT = g-glutamyltransferase; LDH = lactate dehydrogenase (LDH); LIP = lipase; CK = creatine kinase; BT = total bilirubin; BD = direct bilirubin; P = phosphate; CA = calcium; MG = magnesium; FE = iron; NA = sodium; K = potassium; HI = hemolysis index; CVa = analytical coefficient of variability; MPA = MODULAR PRE-ANALYTICALS EVO-MPA system (Roche Diagnostics GmbH).
Normal distribution; p value represents the significance by paired Student t test.
Nonnormal distribution; p value represents the significance by Wilcoxon ranked-pairs test.
Storage Evaluation
When all the laboratory testing was finished, (1) the samples from the traditional processing were stored at 4 °C, as common practice in laboratory diagnostics and as prescribed in our accredited laboratory, and (2) the primary tubes from the laboratory automation for the preanalytical phase were stored inside the output buffer (OPB) in the same MPA system. 27 The OPB stores racks containing the primary samples intended for offline processing at room controlled temperature. All samples were stored during 6 h, and the same panel of tests was then performed on the same Cobas 6000 <c501> module.
Statistical Analysis
The significance of the differences between samples was assessed by paired Student t test after checking for normality (with D’Agostino-Pearson’s omnibus test). As nonnormal distribution was found for TG, BUN, CRE, CRP, UA, AMYL, AMY-P, AST, ALT, GGT, LDH, LIP, CK, and MG, results were assessed by Wilcoxon ranked-pairs test. The level of statistical significance was set at p < 0.05. Finally, the biases from blood specimens subjected to (1) traditional processing before and after storage and (2) automated processing before and after storage were compared with the current desirable quality specifications for bias (B), derived from biological variation according to the formula B < 0.25 (CVw2 + CVg2)1/2, where CVw and CVg are within- and between-subject CVs, 28 as well as using Bland and Altman plot analysis.
Results
The main results of this study are shown in Table 2 . Overall, significant differences were observed from comparison between (1) traditional processing versus automated processing both before storage for ALP, LDH, P, MG, FE, and HI; (2) traditional processing versus automated processing both after storage for COL, TG, TP, ALB, BUN, CRE, UA, ALP, AMY-P, AST, ALT, GGT, LDH, CK, CA, FE, NA, K, and HI; (3) traditional processing before storage versus traditional processing after storage for COL, HDL, TG, TP, ALB, BUN, CRE, UA, AST, ALT, GGT, LDH, P, CA, MG, FE, NA, K, and HI; and (4) automated processing before storage versus automated processing after storage for BUN, AST, LDH, P, and CA. Clinically significant variations as compared with the current desirable quality specifications 28 and bias calculated with Bland and Altman plot are shown in Table 1 . The biases were higher than as specified 28 for (1) LDH when traditional processing versus automated processing both before storage were compared; (2) COL, TG, TP, ALB, CRE, UA, ALP, AST, LDH, CA, FE, NA, and K when traditional processing versus automated processing both after storage were compared; (3) COL, TG, TP, ALB, CRE, UA, AST, LDH, P, CA, MG, FE, NA, and K when traditional processing before storage versus traditional processing after storage were compared; and (4) LDH, P, and CA when automated processing before storage versus automated processing after storage were compared.
Comprehensive results of routine clinical chemistry laboratory tests from two different processing methods of diagnostic blood specimens.
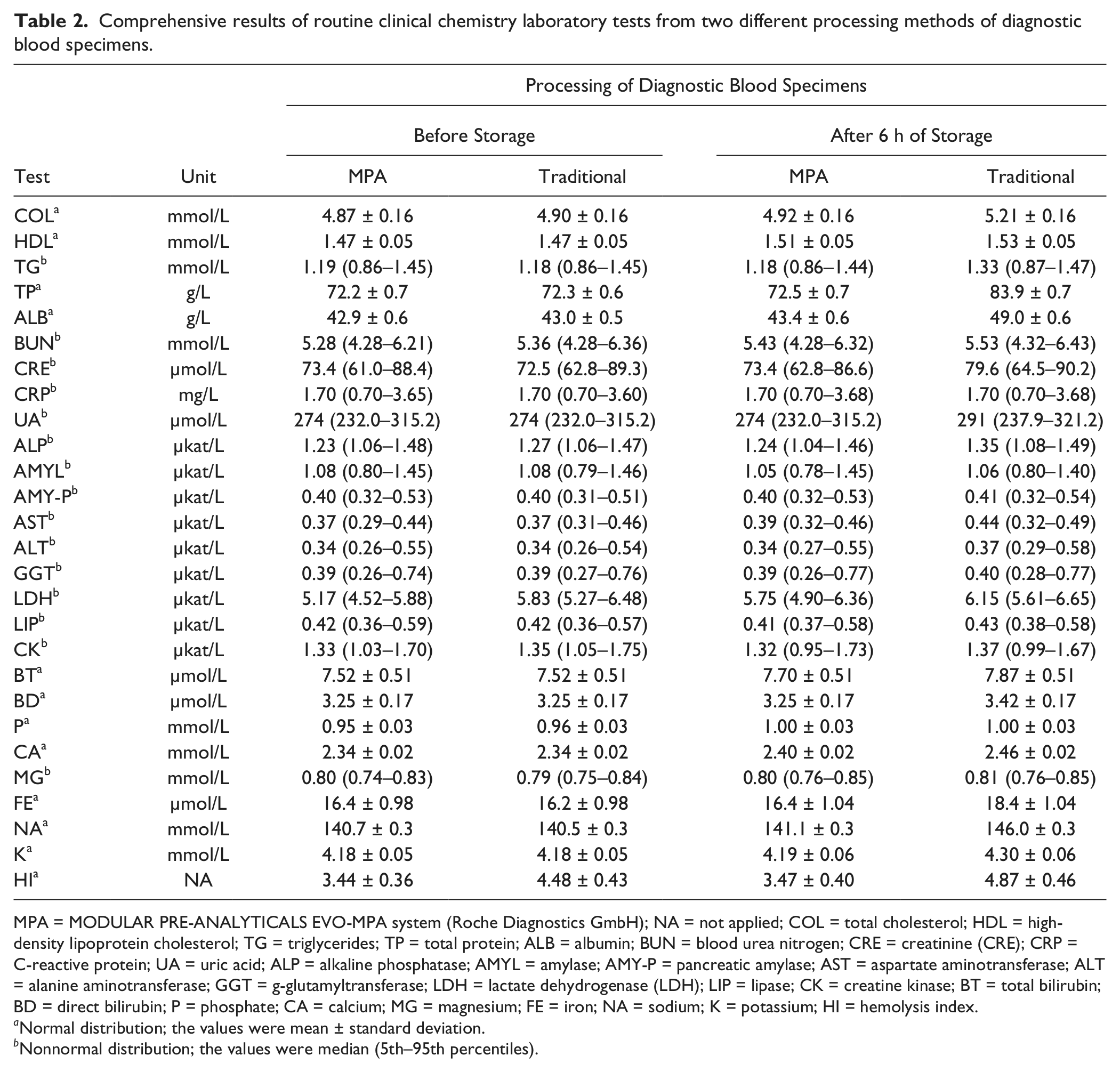
MPA = MODULAR PRE-ANALYTICALS EVO-MPA system (Roche Diagnostics GmbH); NA = not applied; COL = total cholesterol; HDL = high-density lipoprotein cholesterol; TG = triglycerides; TP = total protein; ALB = albumin; BUN = blood urea nitrogen; CRE = creatinine (CRE); CRP = C-reactive protein; UA = uric acid; ALP = alkaline phosphatase; AMYL = amylase; AMY-P = pancreatic amylase; AST = aspartate aminotransferase; ALT = alanine aminotransferase; GGT = g-glutamyltransferase; LDH = lactate dehydrogenase (LDH); LIP = lipase; CK = creatine kinase; BT = total bilirubin; BD = direct bilirubin; P = phosphate; CA = calcium; MG = magnesium; FE = iron; NA = sodium; K = potassium; HI = hemolysis index.
Normal distribution; the values were mean ± standard deviation.
Nonnormal distribution; the values were median (5th–95th percentiles).
Discussion
Rapid advances in computer technology and robotic automation, from sample identification to collection, processing, analysis, and reporting of test results, are disclosing a promising scenario for drastic improvements that will ultimately lead to miniaturization and cost containment. 29 The MODULAR PRE-ANALYTICALS EVO-MPA system (Roche Diagnostics GmbH) effectively reduces the labor associated with specimen processing and decreases the number of laboratory errors that occur with specimen sorting, labeling, and aliquoting. This is a predictable advantage, if one considers that the instrument does not require routine human manipulation. The more interesting finding is that the MODULAR PRE-ANALYTICALS EVO-MPA system (Roche Diagnostics GmbH) preserves the integrity of sample handling throughout the activities of specimen processing. Some suggestions can be advanced to explain our findings: the primary tubes are recapped and stored for further analyses (when needed) more rapidly in the MPA system than in traditional processing. When the process takes place starting after the MPA system, all tests are directly performed without a break on aliquoted-barcoded cups on the Cobas 6000 <c501> module, whereas the primary tube remain open inside the Cobas 6000 <c501> module (as in all brands of clinical chemistry analyzers) during the measuring process using traditional (manual) processing. Then, blood tubes are manually recapped and stored at the end of the process, when the rack tray in the analyzer is full. The evidence of clinically significant increases in COL, TG, TP, ALB, CRE, UA, ALP, AST, LDH, CA, FE, NA, and K when traditional processing was compared with automated processing after storage can be explained by the fact that either high- or low-molecular-weight analytes are affected by water evaporation in open tubes. This is confirmed by previous evidence that tubes remaining opened for a long time display significant variation in the concentration of macromolecules, enzymes, and ions.30,31 This hypothesis also supports the clinically significant increases observed between traditional processing before storage and traditional processing after storage for COL, TG, TP, ALB, CRE, UA, AST, LDH, P, CA, MG, FE, NA, and K. Although it might be provocatively concluded that the results of this study ( Table 2 ) may be considered clinically irrelevant, this would not consider the current quality specifications for bias, derived from biological variation. 28 Obviously, the quality specifications derived from biological variation are considered of pivotal importance, and their usefulness is acknowledged in daily practice by quality managers of medical laboratories.32–35 As such, the caring physicians, who are unaware of the real patient situation, might abstain from appropriate treatments as a consequence of biased results from traditional processing of diagnostic blood specimens. It is also noteworthy that the relative (percentage) increases of the various analytes was rather broad and heterogeneous (i.e., ranging from −1.9% to +15.8%) and thereby unpredictable a priori. For a rational explanation of the consequences of uncontrolled evaporation, it should be considered that in serum/plasma where the pH is about 7.4, the interfacial water molecules between air (which is over the interface) and liquid bulk (which is under the interface) are polarized with stronger hydrogen bonding in the surface than in the bulk. This creates water’s solvation and ionization properties that change at the interface with a depth of about 250 nm, with ions and hydrophilic solutes such as proteins generally becoming less soluble. Moreover, plasma proteins that are mainly in the form of anions at pH 7.4 are progressively repelled by the evaporating surface, thus concentrating in the liquid bulk. During the process, the proteins more proximal to the interface can partially dehydrate and shrink, with conformational changes possibly involving chemical reactivity. Complex molecules, such as lipoproteins, are expected to behave according to the lipid/apoprotein ratio.36,37 The significant increase in HI for all comparisons, except for automated processing before storage versus automated processing after storage, is surprising and somehow unexpected. The HIs that can interfere with test results, as reported by Roche, 38 are ≥10 for BD; ≥15 for LDH; ≥20 for AST; ≥50 for BT; ≥100 for K; ≥200 for ALP, AMY-P, GGT, CK, and FE; ≥300 for P; ≥500 for AMYL; ≥700 for COL, TG, and ALT; and ≥1000 for GLU, TP, ALB, BUN, CRE, CRP, UA, LIP, CA, MG, NA, and HDL. Our results showed a mean HI ~3.5 for automated processing both before and after storage as well as ~4.5 before and ~4.9 after storage for traditional processing. Obviously, these HIs must be considered still negligible and do not justify the changes observed in analyte concentrations but still mirror an enhanced effect on erythrocytes using traditional sample processing. In conclusion, our results showed that the MODULAR PRE-ANALYTICALS EVO-MPA system (Roche Diagnostics GmbH) can really improve the quality in the preanalytical phase of laboratory testing. We suggest that laboratory quality managers perform similar evaluation in other automated preanalytical laboratory processing.
Footnotes
Acknowledgements
The authors acknowledge the quality manager Ms. Sandra Meneghelli and all clinical biochemistry staff from Laboratory of Clinical Biochemistry, University of Verona, for their skillful technical support. Special thanks for Mr. Claudio Brentegani and Ms. Marika Pantani. The authors also thank Ms. Samdra Sampaio from Roche Diagnostics Brazil for providing the supplementary file.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
