Abstract
Study design
Retrospective cohort study.
Objectives
Develop and validate a model combining clinical data, deep learning radiomics (DLR), and radiomic features from lumbar CT and multisequence MRI to predict high-risk patients for adjacent segment degeneration (ASDeg) post-lumbar fusion.
Methods
This study included 305 patients undergoing preoperative CT and MRI for lumbar fusion surgery, divided into training (n = 192), internal validation (n = 83), and external test (n = 30) cohorts. Vision Transformer 3D-based deep learning model was developed. LASSO regression was used for feature selection to establish a logistic regression model. ASDeg was defined as adjacent segment degeneration during radiological follow-up 6 months post-surgery. Fourteen machine learning algorithms were evaluated using ROC curves, and a combined model integrating clinical variables was developed.
Results
After feature selection, 21 radiomics, 12 DLR, and 3 clinical features were selected. The linear support vector machine algorithm performed best for the radiomic model, and AdaBoost was optimal for the DLR model. A combined model using these and clinical features was developed, with the multi-layer perceptron as the most effective algorithm. The areas under the curve for training, internal validation, and external test cohorts were 0.993, 0.936, and 0.835, respectively. The combined model outperformed the combined predictions of 2 surgeons.
Conclusions
This study developed and validated a combined model integrating clinical, DLR and radiomic features, demonstrating high predictive performance for identifying high-risk ASDeg patients post-lumbar fusion based on clinical data, CT, and MRI. The model could potentially reduce ASDeg-related revision surgeries, thereby reducing the burden on the public healthcare.
Keywords
Introduction
Lumbar degenerative disease is a major cause of lower back pain and reduced mobility. Clinical symptoms typically include progressively worsening lower back pain and neurological impairment in lower limbs, often significantly affecting the daily lives of patients. When conservative treatments fail, lumbar fusion surgery is a standard treatment option. Adjacent segment degeneration (ASDeg) refers to radiographic degeneration in segments adjacent to a previous spinal fusion surgery. 1 Studies suggest lumbar fusion may accelerate ASDeg,2-4 which is often asymptomatic in its early stages but may require revision surgery or additional intervention as it progresses. This significantly increases the burden on the healthcare system and impacts patient labor force participation and productivity. Therefore, accurately identifying high-risk patients for ASDeg preoperatively is crucial to reducing surgical risks, optimizing treatment strategies, and improving prognosis.
Computed tomography (CT) and magnetic resonance imaging (MRI) are both non-invasive, high-resolution imaging techniques. MRI is recognized for its superior tissue contrast and multiplanar imaging, while CT excels at imaging bone structures. Both are essential for evaluating lumbar degenerative disease. However, identifying high-risk patients for ASDeg from medical imaging is still largely dependent on the subjective judgment of clinicians, leading to variability in results. Enhancing the predictive ability of medical images for ASDeg is a crucial clinical need.
Radiomics helps evaluate microstructural changes in the lumbar spine and surrounding tissues. 5 Deep learning, particularly three-dimensional convolutional neural networks (3D CNNs), has been widely used in spinal imaging analysis due to its effectiveness in processing 3D data. In this study, we used radiomics techniques to develop prediction models using lumbar CT and MR images from local medical centers. We compared the performance of these models, which incorporate medical imaging and clinical data, in predicting ASDeg after lumbar fusion surgery. Combined models were constructed using 3 types of features to evaluate their effectiveness in identifying patients at high risk for ASDeg.
Methods
Patients
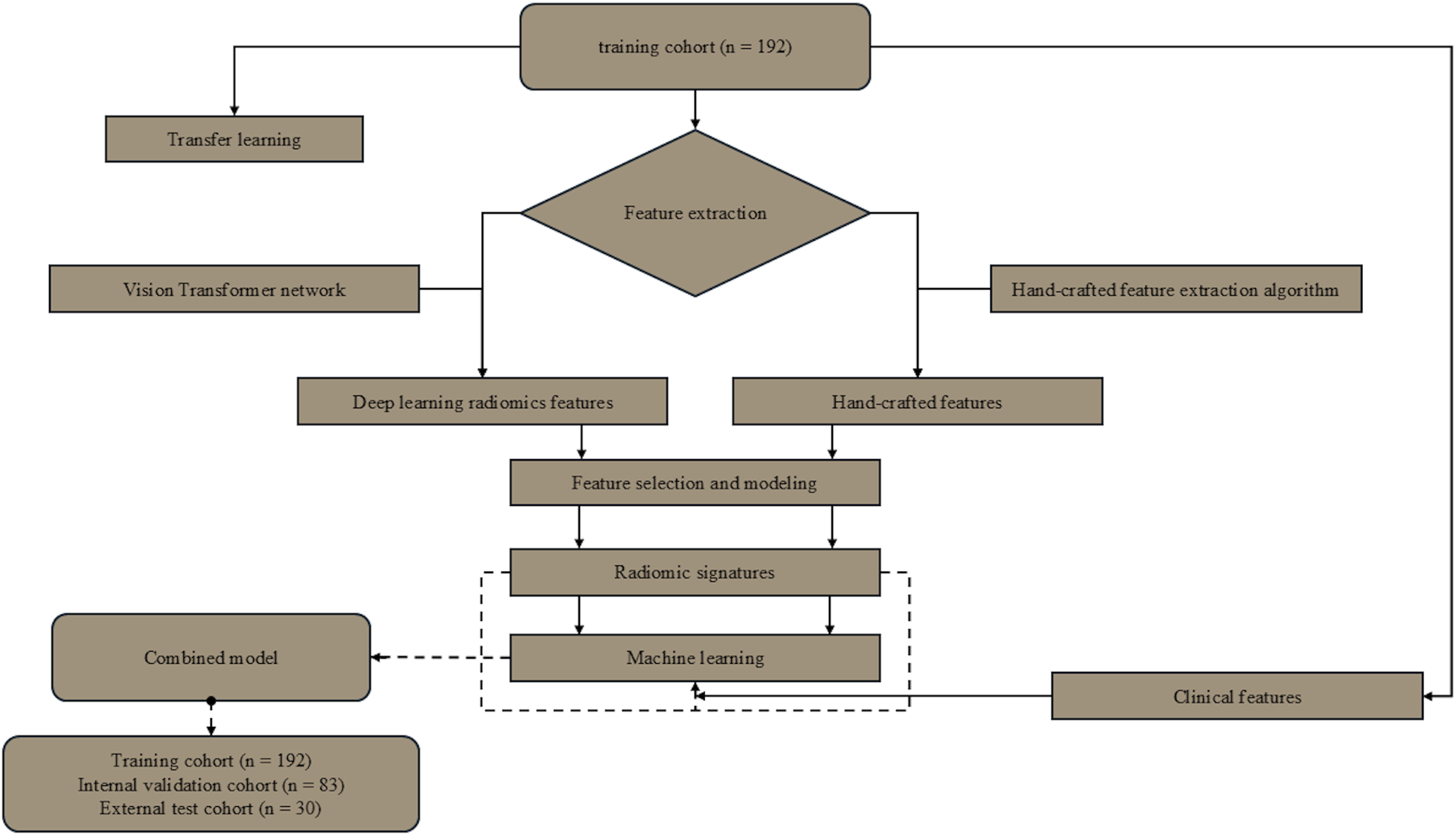
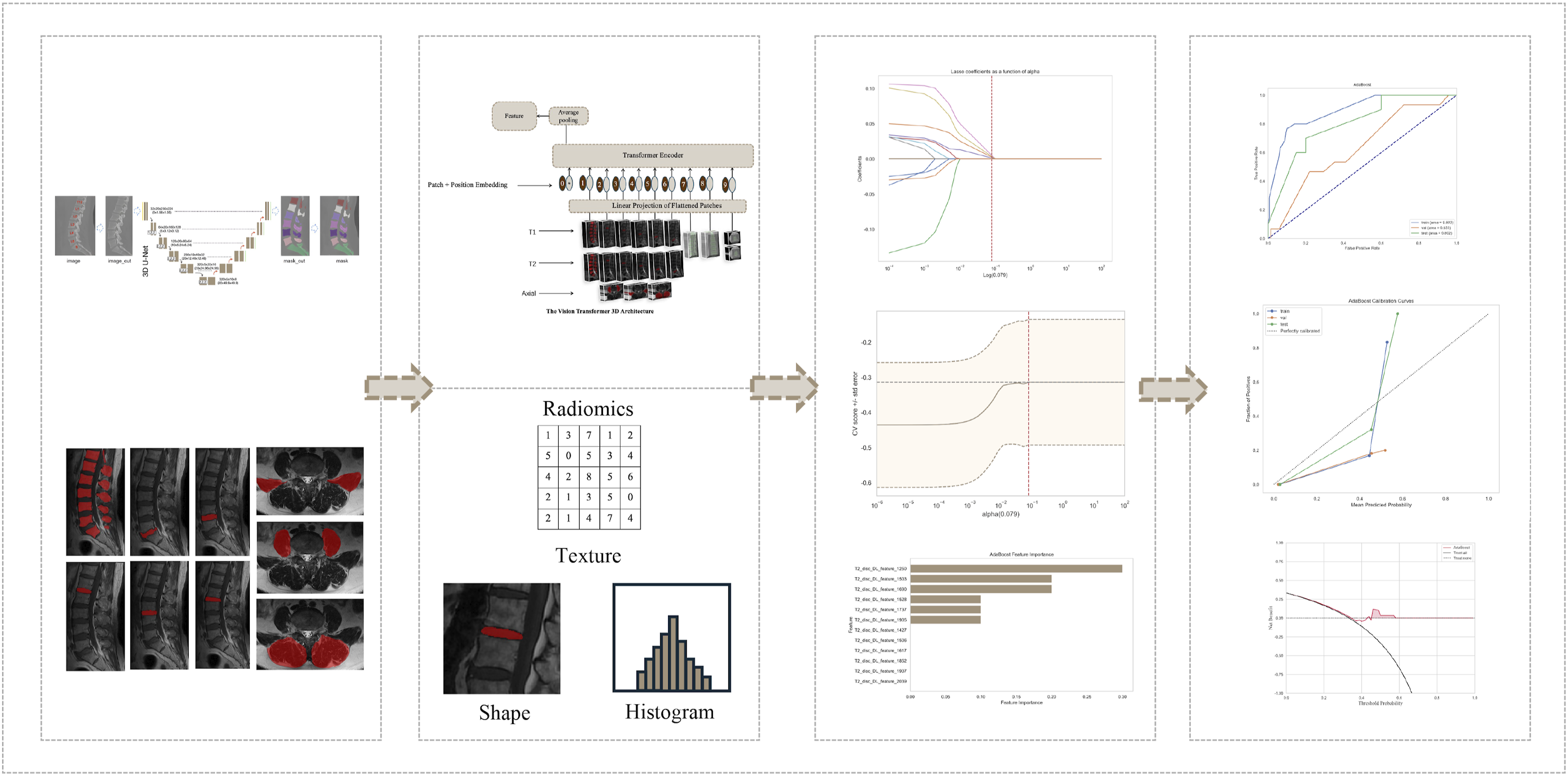
This study used medical imaging data from 3 hospitals in China. We collected baseline clinical and imaging data from patients with lumbar degenerative disease who underwent posterior lumbar interbody fusion (PLIF) between January 2016 and June 2023, establishing internal training and validation cohorts and an external testing cohort. Inclusion criteria were as follows: (1) patients who underwent 1-3 segment PLIF at our hospital and 2 other research centers; (2) patients with at least 6 months of complete medical and postoperative follow-up records; (3) patients with complete clinical data; (4) patients without history of spinal surgery. Exclusion criteria included the following: (1) patients with lumbar diseases related to infection or tumors; (2) Patients with poor image quality or artifacts (as defined in Supplemental Explanation S2) were excluded; (3) patients with incomplete follow-up or medical records. Figure 1 illustrates the detailed flowchart of case selection, with training and internal validation cohorts randomly assigned in a 7:3 ratio. Figure 2 and 3 outline the study design and deep learning radiomics (DLR) workflow, covering case collection and grouping, image preprocessing, feature extraction, analysis, and model construction. This study complied with the Declaration of Helsinki and received Ethics Committee approval from the 3 hospitals involved (Approval No.2024-KE-346). Due to the retrospective study design, the Ethics Committee waived the need for informed consent. Flowchart summarizing patient selection and allocation to the training cohort, validation cohort and test cohort. Flowchart of this study. Workflow of deep learning radiomics for ASDeg prediction. The process includes 4 primary stages: (1). Annotation and segmentation: Marking and outlining specific regions of interest (ROIs) on medical imagery. (2). Feature extraction: Employing a deep learning framework based on Vision Transformer (ViT) to derive radiomic characteristics such as texture, shape, and histogram attributes from processed images. (3). Feature selection and model development: Utilizing statistical techniques and machine learning algorithms to identify key predictive features and construct the model. (4). Model performance evaluation: Validating the model’s predictive accuracy and clinical utility through calibration curves, ROC analysis, and decision curves.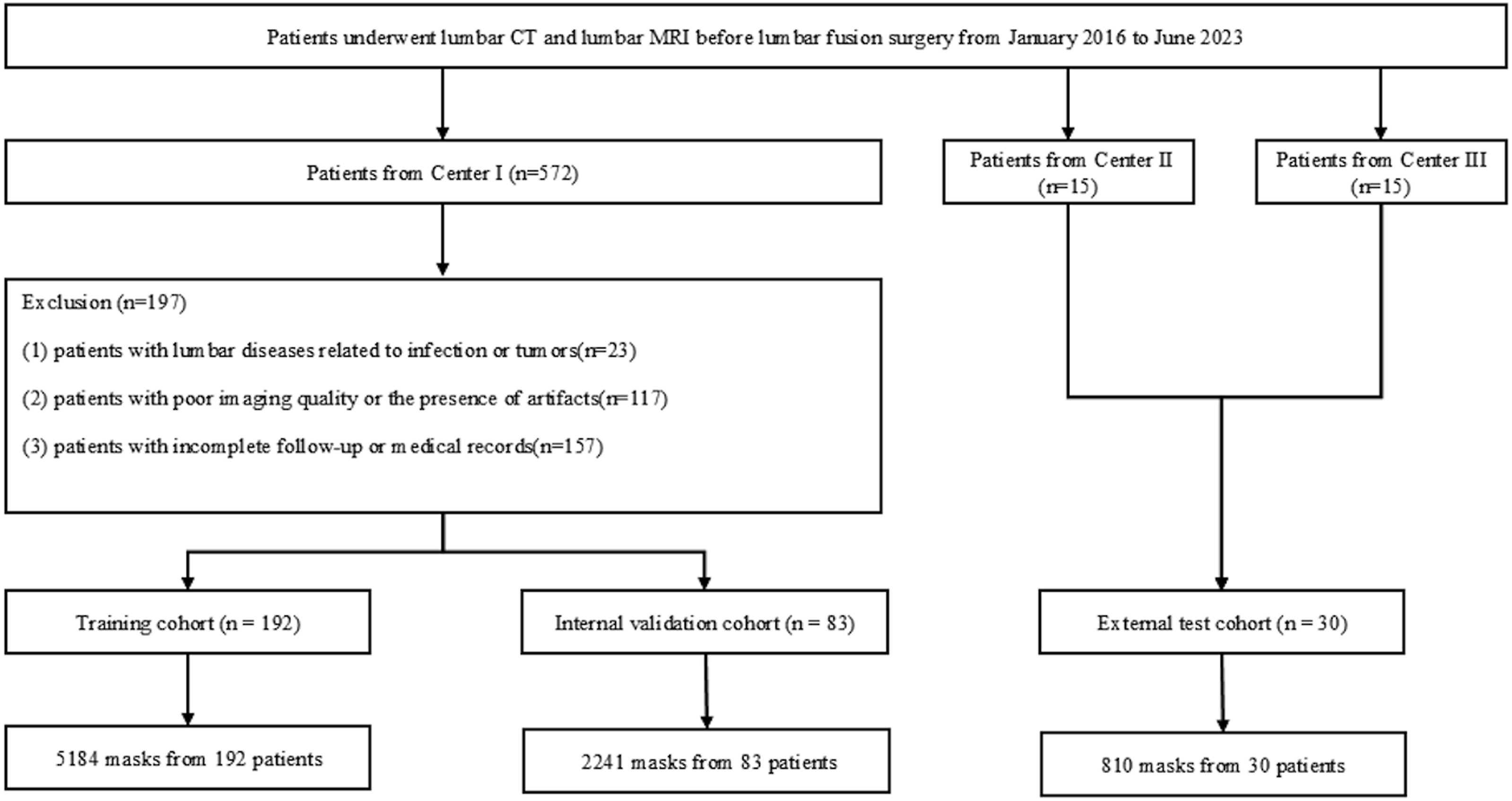
Clinical Characteristics and Medical Imaging Acquisition
Baseline data for all patients, including sex, age, number of surgical segments, body mass index (BMI), osteoporosis status, and diagnosis, were extracted from the clinical records. Details of the CT and MRI equipment and imaging parameters are provided in Explanation S1.
ASDeg in our study was radiographically defined by at least one of the following observed ≥6 months postoperatively: (1) spondylolisthesis >4 mm, angular motion >10°, or disc height loss >10%; (2) UCLA grade ≥2; or (3) Pfirrmann grade IV/V in adjacent segments. The evaluations were conducted independently by 2 spine surgeons each possessing over 5 years of experience. They were unaware of the surgical specifics, and a senior surgeon subsequently verified the final diagnoses.
Image Analysis and Predictions
Two spinal surgeons (A and B), with 6 and 11 years of experience, respectively, independently predicted the likelihood of ASDeg development after surgery based on the same baseline data and clinical images used by the algorithm. In cases where the predictions of the 2 surgeons differed, an expert spinal surgeon (C) with over 20 years of experience served as an adjudicator. Cohen’s Kappa coefficient was calculated to assess the predictive agreement between A and B (Table S1).
Image Segmentation
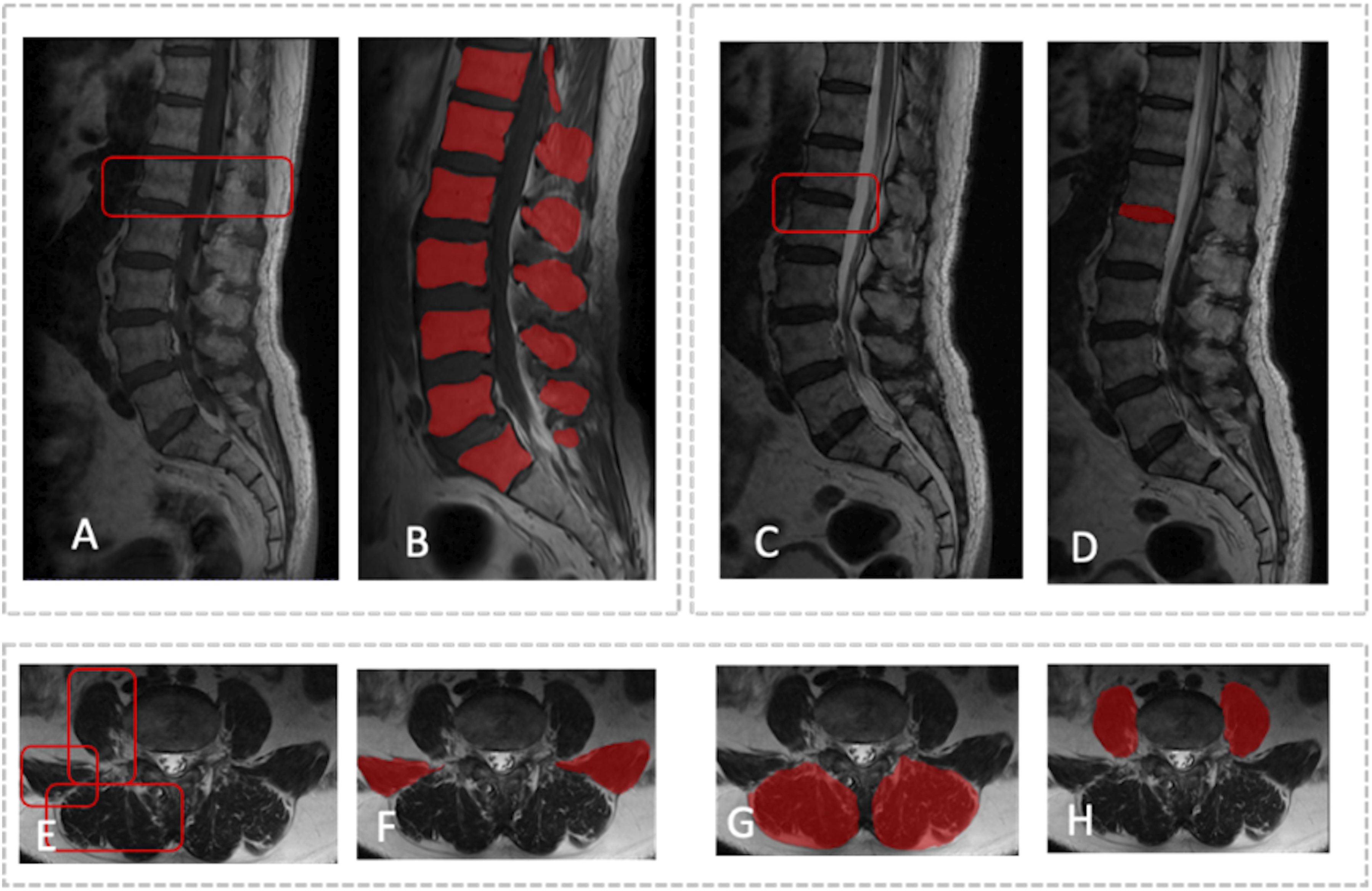
Accurate tissue segmentation is essential for subsequent image analysis. Vertebral body segmentation was performed in 2 steps. For CT images, after preprocessing, a 3D U-Net model was used to segment the vertebral bodies, with results saved as NIfTI mask files (Figure 4). MRI segmentation was manually and independently performed by 2 spinal surgeons. MR images were first imported into ITK-SNAP software (version 3.8.0, https://www.itksnap.org). In sagittal T1 and T2 sequences, vertebral bodies and intervertebral discs were manually annotated. In the axial sequences, the borders of the psoas major, quadratus lumborum, and paraspinal muscles were manually outlined and filled to include spine and surrounding soft tissue data. The MRI segmentation results were then saved as mask files in NIfTI format (Figure 5). One month later, 30 patients from the training cohort were randomly selected, and surgeons A and B independently repeated the segmentation. The intraclass correlation coefficient was calculated to assess inter-observer consistency in vertebral segmentation. Segmentation images based on lumbar CT. Segmentation images based on multi-sequence MRI. Segmentation of both the intervertebral discs and vertebral bodies was performed separately on T1 and T2 MRI sequences to ensure precise delineation of these structures. The process for segmenting the paraspinal muscles, including the quadratus lumborum and psoas major, was also carried out for the corresponding anatomical regions. To preserve the integrity and distinctness of each segmented structure, all segmented images were saved in the NIfTI format (.nii.gz) as individual files. (A and B) Vertebral body segmentation on sagittal MRI images. (C and D) Intervertebral disc segmentation on sagittal MRI images. (E to H) Muscle ROI on axial MRI images. ROI, region of interest.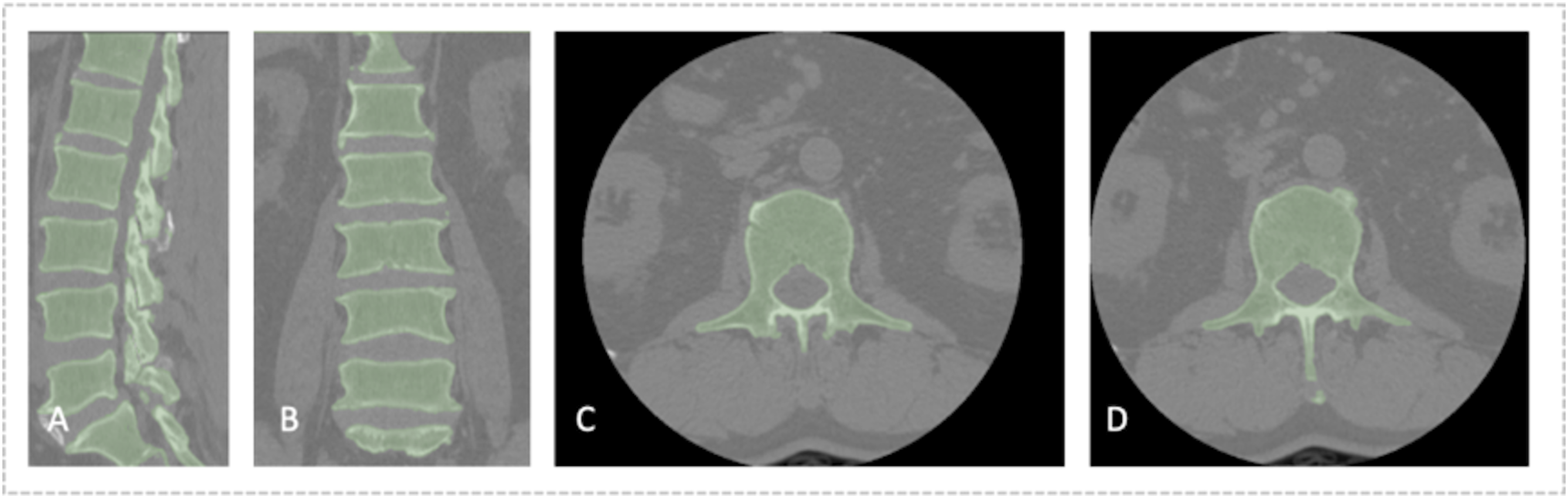
Radiomics Feature Extraction
To prevent data leakage, feature extraction was performed on vertebral bodies, intervertebral discs, and paraspinal muscles across all datasets, while feature selection was conducted only on the training cohort. Before feature extraction, Z-score normalization was applied to all images. The feature extraction algorithms were optimized following the guidelines of the Image Biomarker Standardization Initiative. Radiomic features were extracted using the open-source package Pyradiomics (https://pypi.org/project/pyradiomics/) based on Python (version 3.8). These features included first-order statistics, shape, gray-level co-occurrence matrix, size zone matrix, run-length matrix, neighboring gray-tone difference matrix, and gray-level dependence matrix (GLDM) features. A detailed description of these radiomics features can be found in the Pyradiomics documentation (https://pyradiomics.readthedocs.io). After extraction, robust normalization was applied to standardize all features by calculating the median and quartiles for each feature. This normalization involved subtracting the median from each feature and dividing by the interquartile range to reduce data discrepancies between different centers.
Deep Transfer Learning (DTL) Feature Extraction
The input images were resized to dimensions of 128 × 128 × 128 using linear interpolation, and pixel intensities were normalized to a mean of 0 ± 1. We employed a DTL method in the PyTorch library based on Python (version 3.8), similar to previous studies. 6 The Vision Transformer (ViT) 3D was selected as the base model, with the learning rate carefully adjusted to enhance transfer performance. The fundamental concept of the ViT is to partition the image into small patches and input them into a neural network for processing. Each patch is processed independently, and the outputs of these patches are combined, enabling the network to learn the global structure and features of the image. In summary, the ViT decomposes clinical images into patches, processes each patch individually, and concatenates the results to generate a representation of the entire image. After processing the raw input data, we used the ViT outputs to predict the occurrence of ASDeg through a binary classifier. This model learns complex features associated with ASDeg, enabling accurate predictions for new images. Transfer features were extracted from the penultimate layer of the model (for example, the average pooling layer, which provides a global image representation by averaging patch features). Model parameters were divided into 2 parts: the backbone and the task-specific components. Backbone parameters were initialized using the RadImageNet pretrained model, 7 while task-specific parameters were randomly initialized.
Feature Selection
Early fusion, or feature-level fusion, was applied to radiomics features extracted from CT images of vertebral bodies and from vertebral bodies, intervertebral discs, and paraspinal muscles on multi-sequence MR images. Specifically, the radiomics features from these anatomical structures were concatenated into a single radiomics feature vector. To select radiomics features with high repeatability and low redundancy, we conducted an independent-sample F-test on the co-occurrence matrix histogram of regional features, removing features with P > .05. Pearson’s correlation coefficients were then calculated between features, retaining only one feature from pairs with correlation coefficients >0.9. To maximize representational capability, we adopted a greedy recursive deletion strategy, removing the most correlated feature at each step until the feature set redundancy was acceptable. We then used the least absolute shrinkage and selection operator (LASSO) algorithm to incorporate stable radiomic features by constructing a penalty function λ that shrinks regression coefficients to zero. A 10-fold cross-validation was performed to maximize the mean cross-validation score, determining the optimal λ value. Based on this optimal λ, radiomics parameters with non-zero coefficients and their weights were selected, ultimately identifying independent and stable radiomics features.
Feature-level fusion of the DLR features from the vertebral bodies, intervertebral discs, and paraspinal muscles was achieved by concatenating them into a single feature vector. Z-score normalization was applied to the DLR features before principal component analysis (PCA) dimensionality reduction, converting each feature to a standard normal distribution by subtracting the mean and dividing it by the standard deviation. PCA was then used to extract principal components, enhancing model generalization ability and reducing the risk of overfitting. Finally, LASSO regression was employed for feature selection, retaining significant features with non-zero coefficients.
Models Construction and Validation
After feature selection, we used the scikit-learn machine learning library to construct classification models, including logistic regression (LR), Naive Bayes, linear support vector machines (linear SVM); polynomial, sigmoid, and radial basis function kernels; decision trees; random forests; Extremely Randomized Trees; eXtreme Gradient Boosting; AdaBoost; Light Gradient Boosting Machine; multi-layer perceptrons (MLP); Gradient Boosting Machine. We utilized grid search algorithms on the training set to optimize the hyperparameters of all models, allowing for adjustments to commonly used parameters in each model. A 10-fold cross-validation was applied to evaluate the performance of different classification models and select the optimal hyperparameters. Receiver operating characteristic (ROC) curves were plotted, and the area under the curve (AUC), accuracy, sensitivity, F1 score, and specificity were calculated to evaluate the predictive model performance. We established separate handcrafted radiomics models and DLR models, defining the output probabilities of the optimal classifiers as the radiomics signature (Rad-sign) and deep learning signature (DL-sign).
Binary LR analysis was performed to evaluate baseline clinical variables, including surgical segments, sex, age, diagnosis, BMI, and osteoporosis. Statistically significant clinical variables were combined with Rad-sign and DL-sign to construct a combined model using the machine learning algorithms described above. This was done to further assess the effectiveness of the model in identifying high-risk patients for ASDeg after lumbar fusion surgery. The modeling was performed using the PixelMed AI platform (https://github.com/410312774).
Statistical Analysis
All statistical analyses were performed using Python (version 3.8.2; https://www.python.org). Continuous variables with a normal distribution are expressed as means ± standard deviation, and comparisons between groups were made using independent-sample t-tests. Non-normally distributed continuous variables are expressed as median (interquartile range), and comparisons were made using the Mann–Whitney U test. Levene’s test was used to assess the homogeneity of variances before conducting t-tests. Categorical variables are presented as counts (n) and percentages (%), and comparisons between groups were made using the chi-square or Fisher’s exact test. All statistical tests were two-sided, with statistical significance set at P < .05. The performance of classification models was evaluated using ROC curves and AUC. Decision curve analysis (DCA) was conducted by quantifying net benefit at different threshold probabilities to assess the clinical value of the models. The DeLong test was used to compare AUCs between models.
Results
Clinical Baseline Characteristics
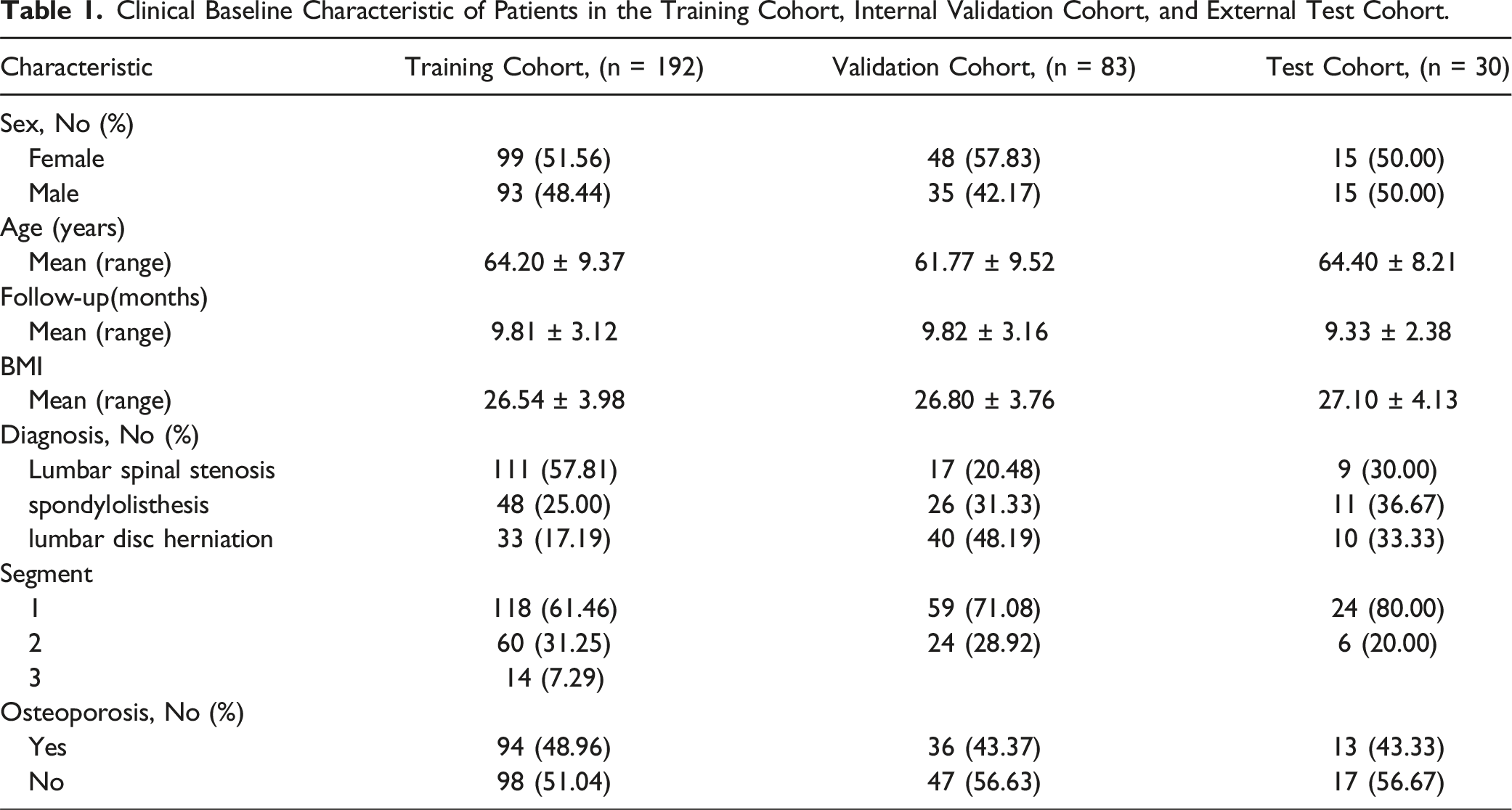
Clinical Baseline Characteristic of Patients in the Training Cohort, Internal Validation Cohort, and External Test Cohort.
Radiomics Feature Selection and Construction of Prediction Model (Radiology-Based)
Following feature fusion, we applied the LASSO regression model for feature selection and dimensionality reduction, using a penalty coefficient of λ = 0.012 (Figures 6A and 6B). This process retained 21 radiomics features (Figure 6C). LASSO regression-based selection of radiomics features. The optimal λ value of 0.012 was selected. (A) Feature coefficients corresponding to the value of parameter λ. Each line represents the change trajectory of each independent variable. (B) The most valuable features were screened out by tuning λ using LASSO regression. The dotted vertical line represents the optimal log(λ) value determined through cross-validation. (C) Feature importance ranking based on the LASSO-selected radiomic features using linear SVM. The y-axis indicates the selected radiomic features, and the x-axis represents their relative importance.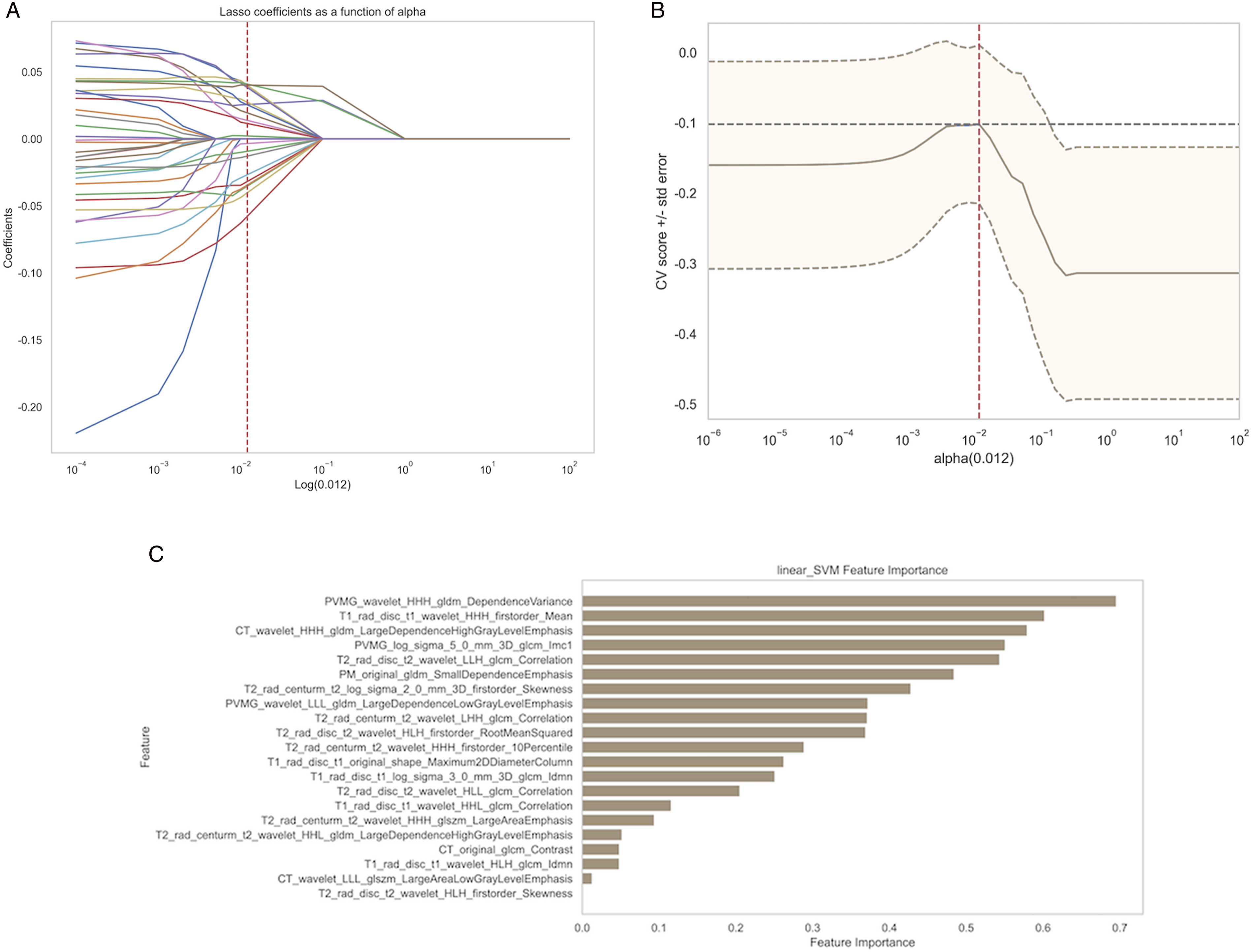
Based on these fused features and their regression coefficients, we constructed a model using preoperative clinical imaging handcrafted radiomics features to predict ASDeg after lumbar fusion surgery. Among all models, the linear SVM performed best, achieving AUC scores of 0.917, 0.766, and 0.870 on the internal training, internal validation, and external test sets, respectively (Figure 7). Performance of the machine learning model based on the linear SVM algorithm. (A) ROC curve. (B) Calibration curve. (C) Decision curve analysis.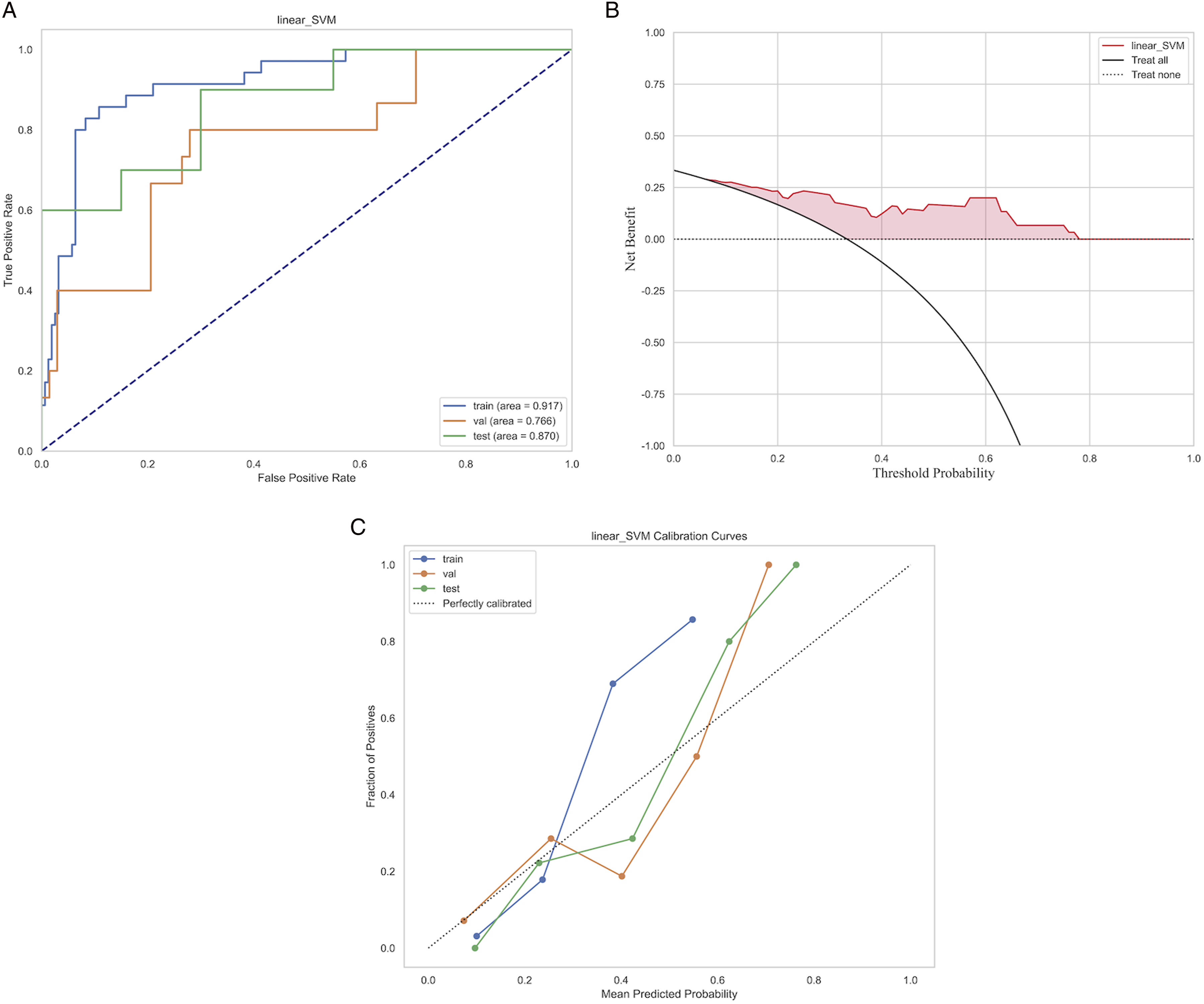
Feature Selection and Construction of Prediction Model (Based on DL)
After combining the features, we applied the LASSO regression model for feature selection and dimensionality reduction, using a penalty coefficient of λ = 0.079 (Figures 8A and 8B). This process retained 12 DLR features (Figure 8C). LASSO regression-based selection of deep learning radiomics features. The optimal λ value of 0.079 was selected.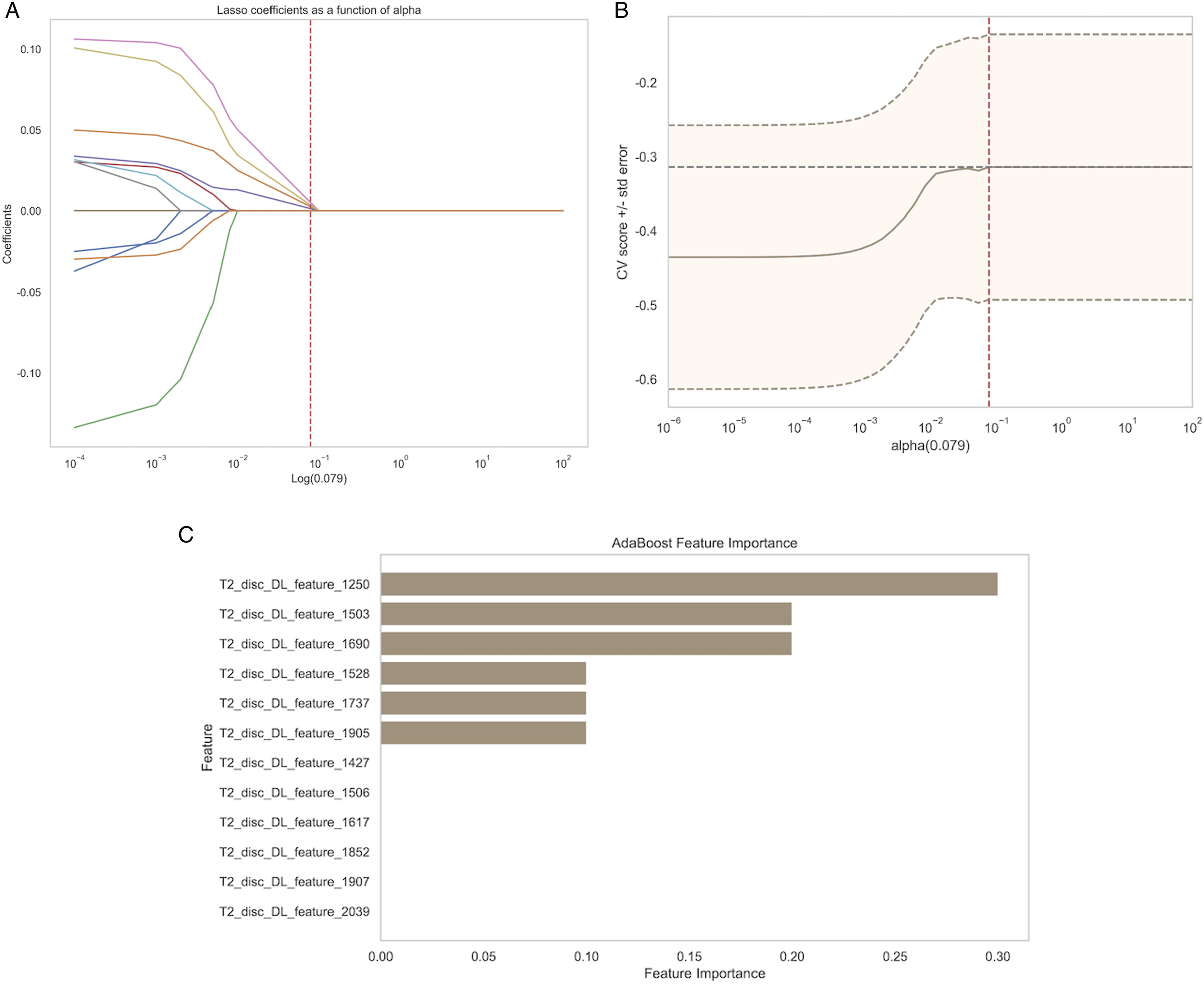
Based on these fused features and their regression coefficients, we constructed a model using preoperative clinical imaging DLR features to predict ASDeg after lumbar fusion surgery. Among all models, AdaBoost performed best (Figure 9) with AUC values of 0.893, 0.633, and 0.802 for the internal training, internal validation, and external test sets, respectively. Performance of the machine learning model based on the AdaBoost algorithm. (A) ROC curve. (B) Calibration curve. (C) Decision curve analysis.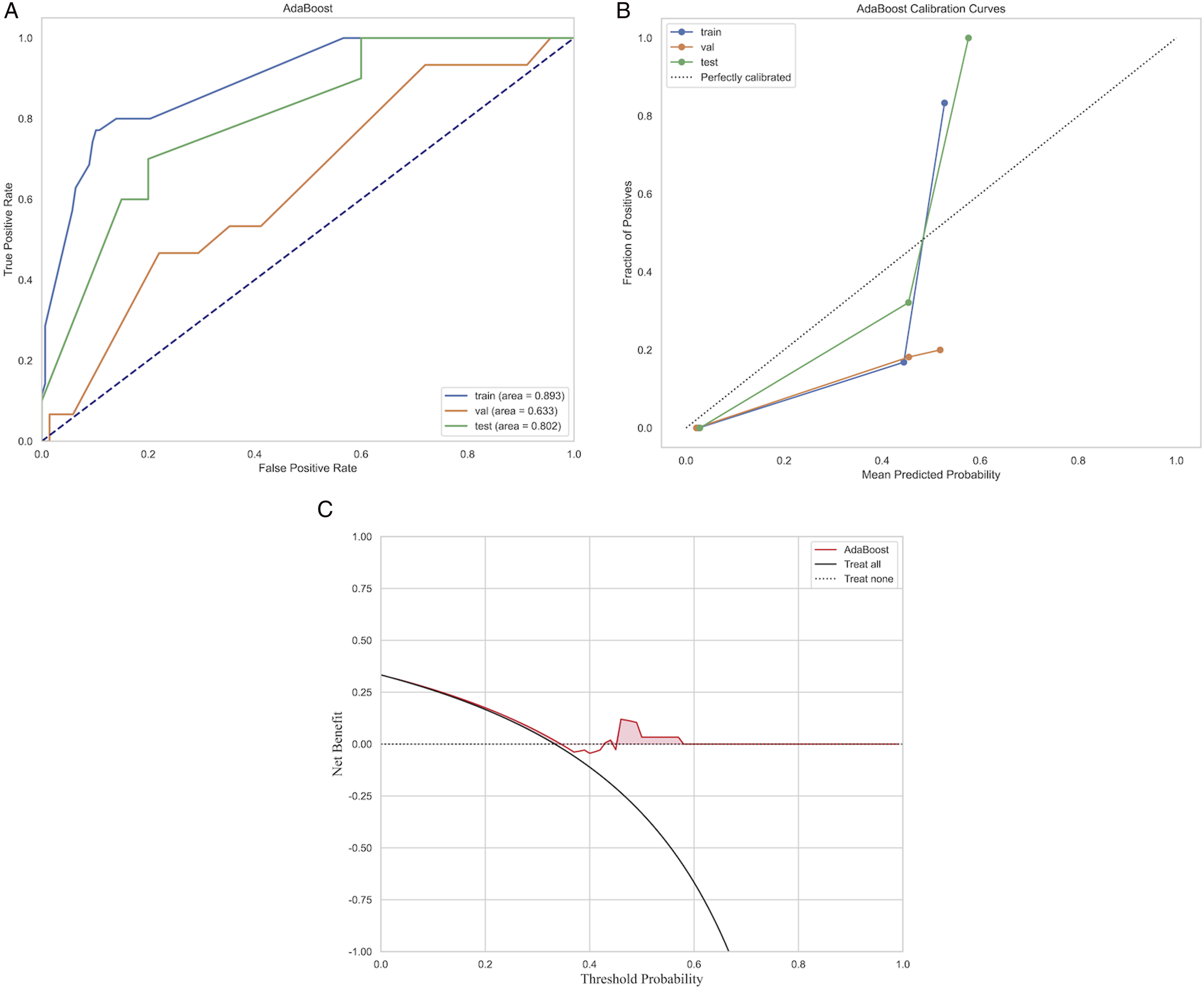
Combined Model Based on Baseline Clinical Data, Rad-Sign, and DL-Sign
We established a combined model using baseline clinical data of patients alongside the previously extracted Rad-sign and DL-sign. Among all models, the most effective machine learning algorithm was the MLP, achieving AUCs of 0.993, 0.936, and 0.835 for the internal training, internal validation, and external test sets, respectively (Figure 10A). Performance of the machine learning model based on the MLP algorithm. (A) ROC curve. (B) Feature importance ranking based on the LASSO-selected features using the GBM. (C) Calibration curve. (D) Decision curve analysis.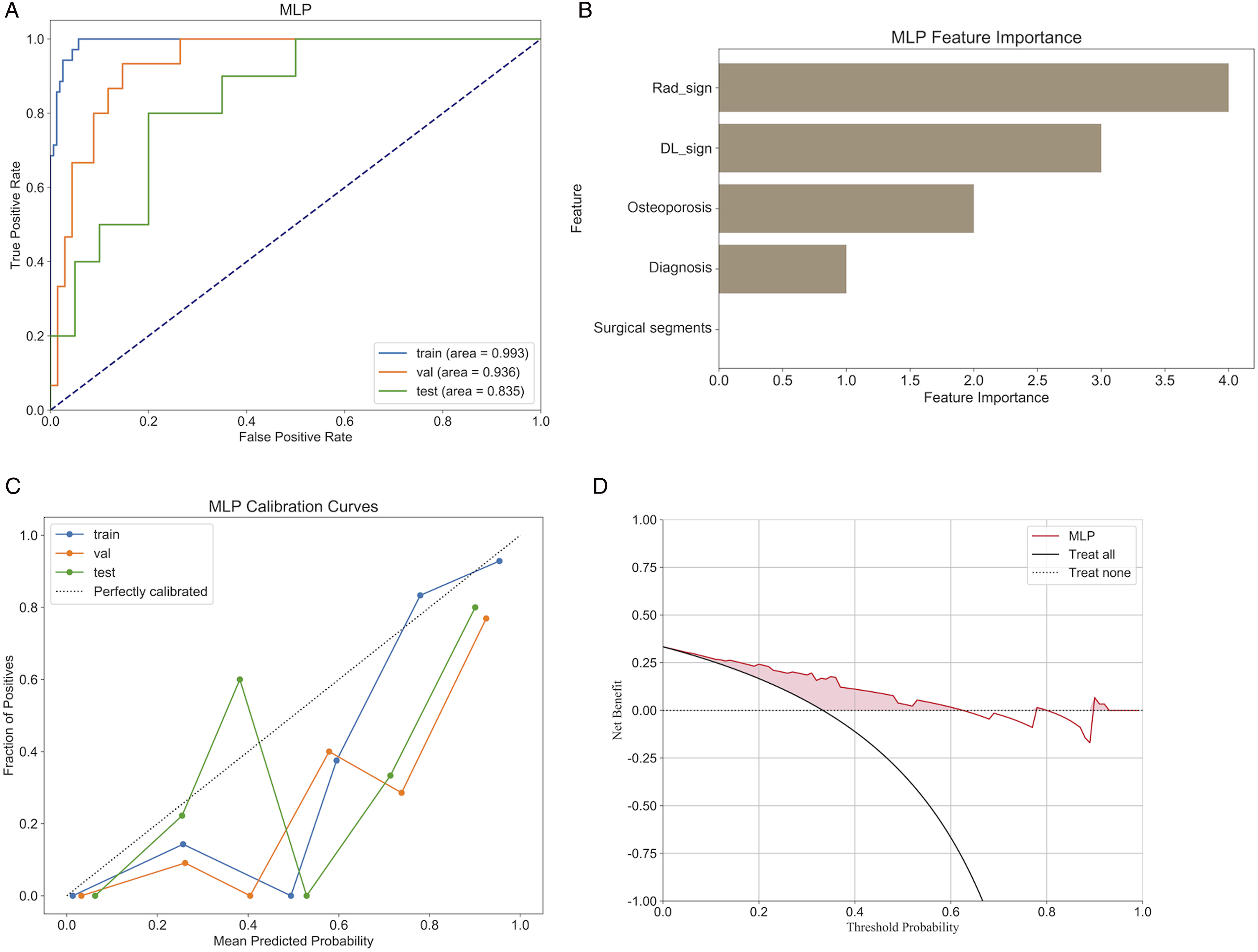
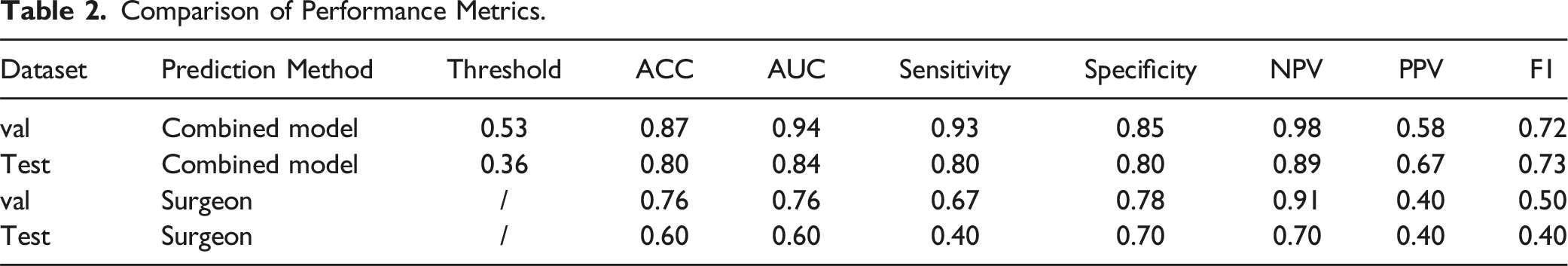
As illustrated in the figure, combining radiomics and deep learning prediction models with clinical baseline features to create a combined model revealed a feature importance bar chart, which indicates that Rad-sign, DL-sign, osteoporosis, diagnosis, and surgical segments are the most significant contributors to the prediction results (Figure 10B). The combined model demonstrated a significant improvement in predictive performance compared to the previous 2 models based solely on radiomics features or DLR features, both in the internal validation and external test sets. DCA indicated that the combined model provided greater net benefits to patients than single-feature models (Figures 11H and 11I). The DeLong test results indicated a statistically significant advantage of the combined model over the single-feature model (P < .05). Additionally, the combined model outperformed the predictions of the 2 spinal surgeons, with significant differences between the machine and clinician comparisons (Table 2). Therefore, we believe that this combined model can more intuitively and effectively identify high-risk patients for ASDeg. Performance of 3 models—the radiomics model, the deep learning radiomics model, and the combined model. Comparison of Performance Metrics.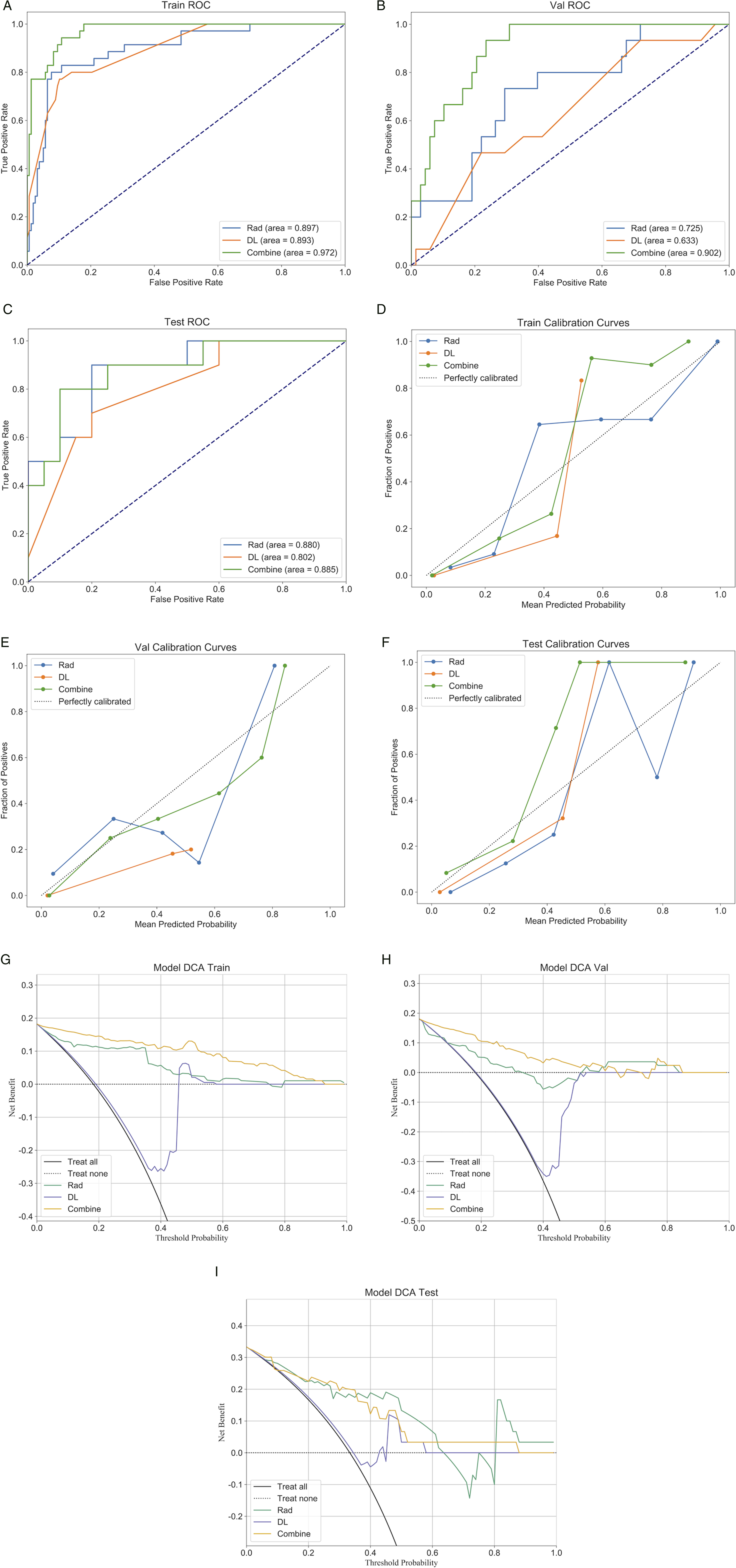
Discussion
In clinical practice, imaging examinations during postoperative follow-up are essential for diagnosing ASDeg. 8 However, since the occurrence of adjacent segment disease (ASDis) can lead to increased patient returns for additional treatment, identifying high-risk patients during preoperative evaluation is crucial for improving patient prognosis. Recent studies have identified various risk factors for predicting the possibility of ASDeg after spinal fusion surgery, including age, higher BMI, and the degree of preoperative ASDeg.9,10 Additionally, paraspinal muscle degeneration has been linked to ASDeg. 9 However, there is a scarcity of studies employing algorithms derived from preoperative imaging to make pertinent predictions. In this study, we used datasets from 3 hospitals to construct a combined predictive model that integrates deep learning radiomics (DLR), traditional radiomics from CT and multi-sequence MRI, and baseline clinical data, to preoperatively identify patients at high risk of developing ASDeg, thereby assisting surgeons in optimizing surgical strategies and clinical decision-making.
CT and MRI are crucial diagnostic tools for lumbar degenerative diseases, capable of revealing microenvironmental changes in the spine and surrounding tissues. Based on previous studies, 11 we believe that lumbar CT images of the vertebrae, sagittal MR images of the vertebrae and intervertebral discs, and axial MR images of muscle tissue collectively provide comprehensive information related to the spine and its microenvironment. Therefore, we segmented and analyzed the vertebral bodies on CT images and the vertebral bodies and intervertebral discs on multi-sequence sagittal MR images (T1 and T2 sequences). Besides, we examined the quadratus lumborum, psoas major, and paraspinal muscles on axial images. Recently, radiomics based on machine learning has been widely applied in the medical field. Researchers have used machine learning algorithms, including image segmentation, feature extraction and selection, and predictive modeling, enabling automated medical imaging analysis and providing intelligent diagnostic support to clinicians, thereby improving diagnostic accuracy and efficiency. 12 Some studies have used deep learning to process medical images, developing effective screening methods for ASDis based on deep learning and cervical MRI. 13 However, the poor interpretability of deep learning models poses limitations for clinical applications. DLR has rapidly developed with the emergence of network architectures like ResNet.14,15 ImageNet is a dataset consisting of millions of natural images, and successful transfer learning relies on the feature similarity between the source and target tasks. 16 After pre-training on ImageNet, these models have been widely applied in medical imaging analyses, including pneumonia detection (X-ray) 17 and COVID-19 pneumonia assessment (CT). 18 RadImageNet, an open-source medical imaging dataset specifically designed for medical applications, theoretically outperforms ImageNet in medical tasks. 19 In this work, we used lumbar CT and MR images from local medical centers to construct DLR predictive models based on RadImageNet and compared their performance in predicting ASDeg after lumbar fusion surgery. By integrating deep learning methods, this approach not only enhances the interpretability of medical imaging that utilizes deep learning but also innovatively extends traditional radiomics, establishing a solid foundation for future research endeavors.
In this study, the optimal model with feature fusion retained 21 radiomics features, 12 DLR features, and 3 clinical features. The Dependence Variance feature from the gray-level dependence matrix, following wavelet transformation (wavelet_HHH_gldm_Dependence Variance), exhibited the highest correlation coefficient with the occurrence of ASDeg, with higher values indicating increased heterogeneity in tissue textures. 20 Clinical imaging manifestations of lumbar degenerative diseases typically include degenerative changes in the vertebral bodies and surrounding soft tissues, which increase the complexity of texture features. This feature may reflect microstructural changes in the tissues surrounding the vertebral bodies and their risk for future degeneration. This finding supports the strong correlation of wavelet_HHH_gldm_DependenceVariance and aligns with previous research results.21,22 Overall, the feature importance analysis indicates that the texture, morphology, and 3D structural features of the fused regions and adjacent segments in preoperative MRI significantly contribute to the model. Collectively, these features suggest that the complex mechanical and biological states of adjacent segments before fusion have multidimensional impacts. The DLR features with the highest correlation coefficients originated from the intervertebral discs. This, combined with previous research findings,2,23-25 suggests that pathological changes in the intervertebral discs significantly impact postoperative ASDeg. While the interpretability of current DLR features requires further investigation, integrating them with traditional radiomics features provides preliminary insights into lesion-specific characteristics. Additionally, it should be noted that the majority of radiomics features selected for the model and all DLR features were derived from MR images. This highlights the greater importance of MR imaging over CT imaging in the preoperative imaging-based method proposed in this study for predicting high-risk patients of ASDeg. Finally, our findings indicate that osteoporosis significantly increases the risk of ASDeg, consistent with previous studies.26-28 The inclusion of patients undergoing three-segment fusion was limited due to a small sample size, resulting in discrepancies in the odds ratio (OR) for surgical segments compared to previous studies. 29 Significantly, in the combined model constructed using the MLP algorithm, the surgical segment factor received the lowest importance weight, indicating its minimal contribution to the predictive outcome of the model and reducing potential bias in the study. Future studies should include more patients in this category to further reduce the bias of this study.
With the rise of deep learning, ViT models have significantly advanced computer vision tasks. Unlike traditional CNNs, ViT models utilize self-attention mechanisms, enabling them to capture global features better. 30 This advantage is crucial in complex medical image analysis, as understanding the overall context can lead to more accurate diagnoses and insights. Developing ViT models, alongside their integration into advanced technologies, has brought new opportunities for advancing medical image analysis and improving diagnostic outcomes. In this study, the combined model achieved significantly higher AUC values for identifying ASDeg compared with models based only on conventional radiomics features in both the internal validation and external test sets. This indicates that the ViT-based combined model can significantly enhance predictive performance, providing superior decision consistency, stability, and improved capability to manage complex features and adapt to varying data.
This study has some limitations. Firstly, despite our efforts to enhance the accuracy of manual MRI image annotations, the inherent subjectivity and potential bias of manual labeling remain, which may impact the consistency of results compared to automated segmentation methods. Therefore, future research should prioritize optimizing automatic MRI image segmentation techniques and aim to combine the strengths of both automation and human expertise. This integration would facilitate a more efficient and precise assessment of spinal structures and pathological features. Secondly, this study incorporated multicenter data; however, the external sample size was limited by stringent inclusion criteria. Moreover, the variability introduced by different scanners across the 3 hospitals was not explicitly analyzed, potentially leading to data discrepancies. Although preprocessing techniques such as resampling and intensity normalization were implemented to reduce scanner-related variations, these adjustments might not fully account for differences in acquisition protocols and scanner types. Additionally, similarities in MRI acquisition settings and demographic characteristics between training and validation datasets could have inadvertently enhanced the model’s performance by minimizing variability. Furthermore, the definition of ASDeg was based on radiological follow-up at 6 months post-surgery, capturing primarily early degenerative changes. Thus, the predictive performance of our model for delayed or progressive degeneration beyond this period remains unclear. Future studies should include larger cohorts from a broader range of centers, incorporate longer follow-up intervals (e.g., at least 1 year), systematically address scanner and protocol heterogeneity, and ensure balanced gender and pathology distributions or explicitly control for confounding factors related to gender and pathology to improve model robustness and generalizability. Additionally, integrating advanced imaging techniques such as SPECT-CT could enhance sensitivity in detecting subtle early-stage degenerative changes, although their limited availability across healthcare systems should also be considered. Addressing these factors will facilitate rigorous multicenter validation and improve the generalizability and clinical applicability of our predictive model. Finally, radiomics often faces criticism for limited feature interpretability and the labor-intensive process of data annotation—challenges that our study also encounters. However, similar to our work on automated segmentation of vertebral bodies in CT images, advancements in artificial intelligence and machine learning present opportunities to enhance the utility of radiomics in both scientific research and clinical practice. By improving tools and methodologies, refining automatic image segmentation techniques (including automated measurement of pelvic parameters), and integrating multiple data sources, we can potentially overcome these obstacles and achieve more efficient and accurate assessments.
Conclusion
In conclusion, this study developed a composite model that integrates radiomics, DLR, and clinical features. Compared with single-modality models, this combined model approach enhanced the identification performance of patients at high risk of ASDeg.
Supplemental Material
Supplemental Material - Developing a Deep Learning Radiomics Model Combining Lumbar CT, Multi-Sequence MRI, and Clinical Data to Predict High-Risk Adjacent Segment Degeneration Following Lumbar Fusion: A Retrospective Multicenter Study
Supplemental Material for Developing a Deep Learning Radiomics Model Combining Lumbar CT, Multi-Sequence MRI, and Clinical Data to Predict High-Risk Adjacent Segment Degeneration Following Lumbar Fusion: A Retrospective Multicenter Study by Congying Zoua, Tianyi Wanga, Baodong Wanga, Qi Feib, Hongxing Songc, and Lei Zang in Global Spine Journal
Footnotes
Authors’ Contributions
C.Z., T.W. contributed equally to this work. Conception and design: L.Z.; Acquisition of data: C.Z., T.W., H.S., Q.F.; Analysis and interpretation of data: C.Z. and T.W.; Drafting the article: C.Z. and T.W.; Critically revising the article: L.Z.; All authors have read and agreed to the published version of the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical Statement
Data Availability Statement
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
