Abstract
Study Design:
Retrospective study.
Objectives:
Huge amounts of images and medical reports are being generated in radiology departments. While these datasets can potentially be employed to train artificial intelligence tools to detect findings on radiological images, the unstructured nature of the reports limits the accessibility of information. In this study, we tested if natural language processing (NLP) can be useful to generate training data for deep learning models analyzing planar radiographs of the lumbar spine.
Methods:
NLP classifiers based on the Bidirectional Encoder Representations from Transformers (BERT) model able to extract structured information from radiological reports were developed and used to generate annotations for a large set of radiographic images of the lumbar spine (N = 10 287). Deep learning (ResNet-18) models aimed at detecting radiological findings directly from the images were then trained and tested on a set of 204 human-annotated images.
Results:
The NLP models had accuracies between 0.88 and 0.98 and specificities between 0.84 and 0.99; 7 out of 12 radiological findings had sensitivity >0.90. The ResNet-18 models showed performances dependent on the specific radiological findings with sensitivities and specificities between 0.53 and 0.93.
Conclusions:
NLP generates valuable data to train deep learning models able to detect radiological findings in spine images. Despite the noisy nature of reports and NLP predictions, this approach effectively mitigates the difficulties associated with the manual annotation of large quantities of data and opens the way to the era of big data for artificial intelligence in musculoskeletal radiology.
Introduction
The years between 2015 and 2020 have seen a sharp increase in the use of artificial intelligence (AI) and machine learning in medicine, which is expected to continue in the next future. 1 Radiology is the medical discipline which has been most radically impacted by the AI revolution 2 ; a recent study showed that, among the 222 AI-based medical devices approved in the United States of America or the European Union between 2015 and 2020, 129 relate to this medical specialty. 3 This trend may be explained, at least to some extent, by the constantly increasing productivity of radiologists reflected in the availability of huge databases which can potentially be employed to train AI models. 3
However, in most cases the textual data generated in radiological departments cannot be directly exploited for AI applications. Medical reports are typically unstructured texts in natural languages, e.g. English, potentially rich in valuable information which is however difficult to access, and do not lend well to be used as annotations to train and test AI models. 4 While structured reports, which may overcome these limitations, are gaining popularity, they are not free from disadvantages 5 and the vast majority of hospitals worldwide still employ unstructured reporting. 6 Even in the case of a future wider adoption of standardized reports, all the unstructured data collected in the past would still pose the same problems of information accessibility.
A possible solution lies in the use of natural language processing (NLP) tools, a class of algorithms designed to extract semantic information, i.e. understanding the meaning of natural text in various languages. NLP techniques were first presented in the 1950s and, in the first decades of their history, exploited a predefined set of words and rules to extract and encode information from texts. 7 In the last 2 decades, NLP widely benefited from the recent AI advances, which allowed for a vast performance improvement. 8
The successful use of NLP to process medical reports has been widely reported.9-12 NLP is indeed now considered a cornerstone of the radiological workflow, and several applications have been documented 4 : improving the quality of speech recognition in dictation systems, 7 raising alerts in case of symptoms or pathologies which have been reported but not acted on, building cohorts for epidemiological studies, retrieving cases from databases based on specific symptoms or findings, and assessing the quality of radiological reporting.
The vast majority of AI studies with a radiological focus still rely on relatively small sets of manually annotated data, and the potential impact of NLP-based training data from medical reports is enormous. While this possibility has already been reported,13,14 the topic appears to be under-investigated as there are no studies quantifying its advantages and limitations. As a matter of fact, training deep learning models with NLP-generated data, which may contain errors, should be considered as a weakly-supervised approach 15 and can potentially result in poor performances depending on the quality and quantity of the data.
The aim of this study is therefore to test the potential of training deep learning models to detect radiological findings based exclusively on data extracted by NLP tools from the medical reports associated with the images. As a benchmark case, we used coronal and sagittal radiographs of the lumbar spine and some of the most commonly reported findings such as loss of lordosis, osteophytosis, scoliosis, etc. Furthermore, we compared the performance of the NLP-based deep learning models with that of neural networks trained on a smaller set of human-annotated data.
Materials and Methods
Overview
The hypothesis of this work is that existing large databases of radiological reports can successfully replace or integrate manually annotated images to train deep learning models aimed at the automatic diagnosis of spinal disorders from radiological images. To test this hypothesis, we performed the following tasks: (1) NLP models able to extract structured information from radiological reports written in the Italian language were developed and tested; (2) the NLP models were used to assemble a large set of radiographic images of the lumbar spine with the corresponding annotations automatically extracted from the medical reports; (3) exploiting this dataset, deep learning models aimed at detecting radiological findings directly from the radiological images were trained and (4) tested against a set of manually annotated set of images.
Data, Images and Outputs
The study was approved by the ethical committee of IRCCS Ospedale San Raffaele (protocol “RETRORAD”). All patients provided written informed consent for the use of images and anonymized data for scientific and educational purposes.
14777 couples of planar radiographic images of the lumbar spine, i.e. one coronal and one sagittal view of the same patient in the standing posture acquired during the same session, were downloaded from the Picture Archiving and Communication System (PACS) of IRCCS Istituto Ortopedico Galeazzi with the associated medical report written by the radiologist at the time of the exam (Figure 1); all data and images were anonymized before any processing. The radiological reports were written inthe Italian language and, although based on a rather homogeneous vocabulary, were unstructured.
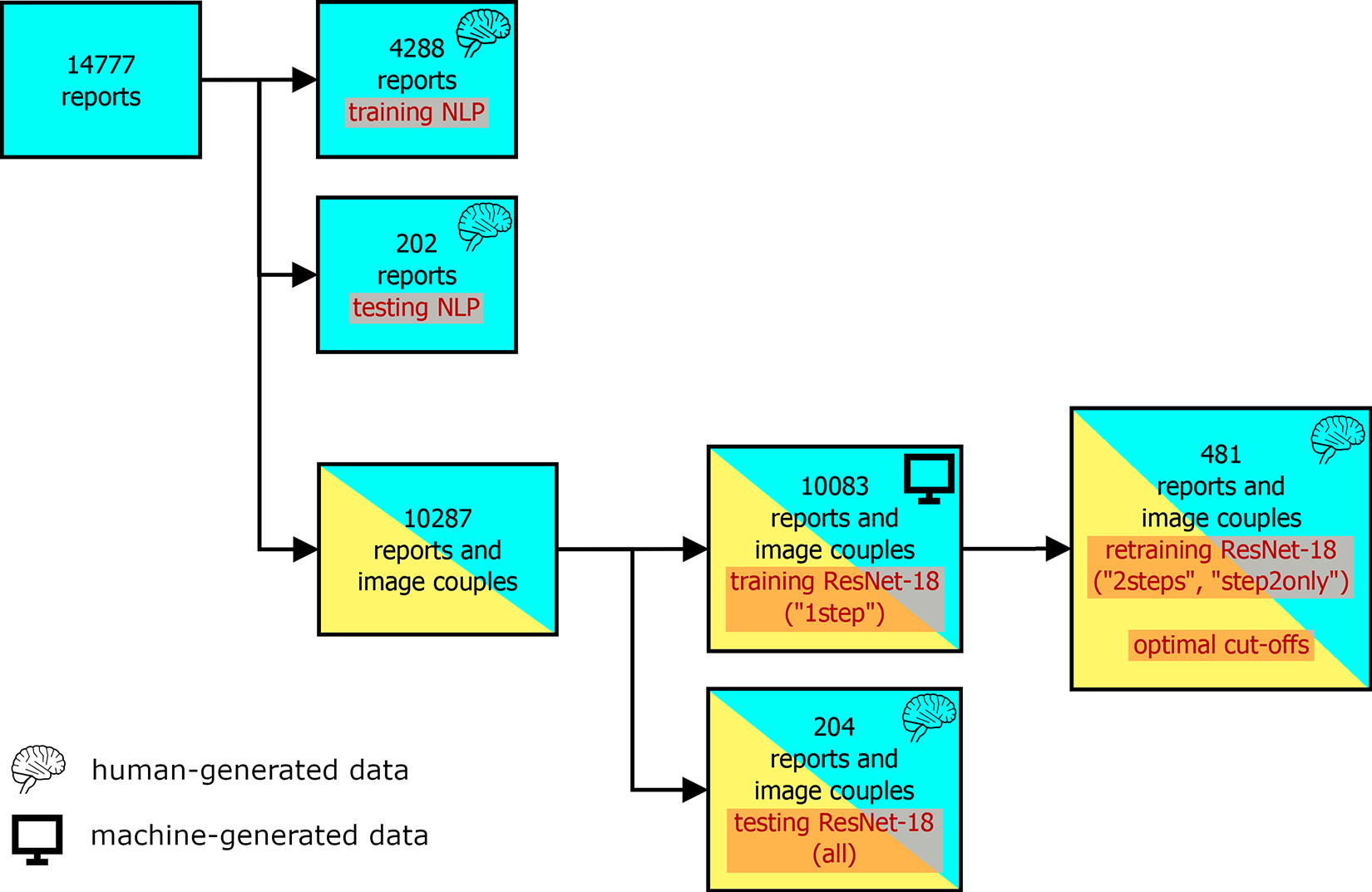
Schematic diagram showing the use of the available data (medical reports, couples of coronal and sagittal radiographs of the lumbar spine) for training and testing the NLP and ResNet-18 models.
The images referred to consecutive patients who underwent radiological investigation in 2016 and 2017, with no exclusion criteria based on the diagnostic query. Therefore, the images covered subjects in a wide age range and with various types of disorders such as low back pain, instability, traumatic and osteoporotic fractures, spinal deformities, tumors, as well as patients undergoing post-operative radiological examination and follow-up. The scope of the present automated diagnosis was limited to the following radiological findings: (1) presence of spinal implants such as pedicle screws, posterior rods, cages, artificial discs (“instrumentation”); (2) reduced, absent or inverted lumbar lordosis (“loss of lordosis”); (3) sclerosis, hypertrophy or degeneration of the facet joints (“facet sclerosis”); (4) sclerosis or degeneration of the sacroiliac joints (“SIJ degeneration”); (5) presence of osteophytes on the thoracolumbar vertebral bodies (“osteophytosis”); (6) osteopenia or osteoporosis (“osteoporosis”); (7) diffuse spondyloarthritis and degeneration (“diffuse degeneration”); (8) scoliosis or scoliotic attitude (“scoliosis”); (9) presence of osteoporotic or traumatic vertebral fractures (“fractures”); (10) narrowing of the intervertebral space in at least one lumbar level (“disc narrowing”); (11) anterolisthesis and (12) retrolisthesis of at least one vertebra (“anterolisthesis” and “retrolisthesis” respectively). All these findings are routinely listed in the radiological reports and, therefore, could in principle be extracted by means of NLP techniques.
Extracting Structured Information From Radiological Reports
Among the 14777 available reports, 4490 were allocated to the development and testing of the NLP models; specifically, 4288 reports were employed in the training phase whereas the remaining 202 reports were used for model validation (Figure 1).
A simple application aimed at facilitating the annotation of the medical reports was developed in the Python language using the graphical user interface toolkit Qt (https://www.qt.io/) and the relative bindings PyQt (https://riverbankcomputing.com/software/pyqt/intro). The application was used by a native Italian speaker to classify the 4490 medical reports in the training set about the 12 radiological findings; for example, if a report mentioned the presence of any vertebral fracture, the variable “fracture” for that report was set to 1, and 0 if not. The output of the manual annotation phase was a CSV file listing the presence or absence of all 12 radiological findings in the 4490 reports.
Text classifiers based on the Simple Transformers library (https://github.com/ThilinaRajapakse/simpletransformers) were then trained, one for each radiological finding, starting from the pre-trained model “bert-base-italian-uncased” (https://huggingface.co/dbmdz/bert-base-italian-uncased). This model is based on the Bidirectional Encoder Representations from Transformers (BERT) NLP approach, 16 originally developed for the English language, and utilizes a corpus with a size of 13 GB. After pre-processing the reports by converting upper-case characters into the lower case and replacing line terminators with spaces, the NLP models were trained for a maximum of 20 epochs; automatic weight balancing was enabled to take into account the unbalanced nature of the training set. The NLP models were then used to automatically classify the 202 reports in the test set.
Automated Diagnosis From Radiographic Images
The aforementioned 4490 manually-annotated reports and their corresponding images were excluded from the training and testing of the deep learning models performing the diagnosis directly on the radiographic images; in this way, we ensured that only machine-generated data was employed in this stage. Among the remaining 10287 image couples, 10083 were used for training the deep learning models, whereas 204 were used exclusively for testing the model performance (Figure 1).
The NLP models were used to automatically generate annotations for the 10083 medical reports associated with the image couples in the training set. The 204 image couples in the test set were processed by an expert human operator, who manually detected the radiological findings by examining both the images and the original medical report in natural language.
Image classifiers were trained based on the ResNet-18 convolutional neural network architecture 17 pre-trained on the ImageNet database (http://www.image-net.org/), using the 10083 image couples and the relative machine-generated annotations as the training set (models “1step”). In order to process the coronal and sagittal projections together, the 2 views were combined into a single 3-channel image in which the former constitutes the red channel while the latter was the blue channel (Figure 2). Prior to merging, the 2 projections were resized to 512x512 without preserving the original aspect ratio.
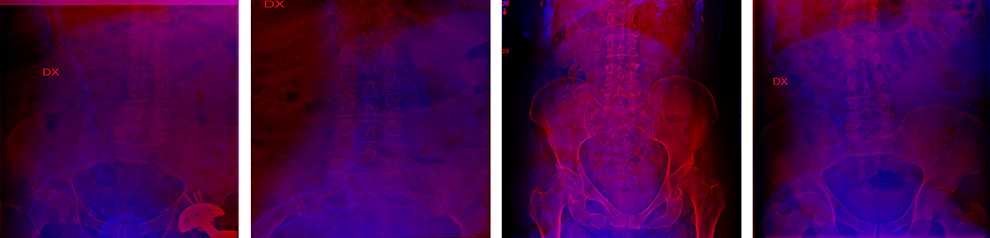
Four examples of the images combining the coronal projection as the red channel and the sagittal projection as the blue channel used to train and test the ResNet-18 models.
The neural networks were trained for a maximum of 200 epochs within the PyTorch framework (https://pytorch.org/), using the Stochastic Gradient Descent optimizer. The values of the hyperparameters (batch size: 32; learning rate: 0.001; momentum: 0.9) were selected after a preliminary investigation exploiting the grid search function of scikit-learn (https://scikit-learn.org/). Random image augmentation was performed with the imgaug library (https://imgaug.readthedocs.io/en/latest/); random left-right flipping, gaussian noise, gaussian blur and contrast adjustment were implemented.
Additionally, 481 image couples were randomly extracted from the 10083 items in the training set and manually reevaluated by a human observer as done for the test set (Figure 1). This set was used both to (1) refine the training obtained with the original test set (models “2steps”) and (2) retrain the models starting from the ImageNet weights (models “step2only”). This approach allowed us to assess the added value of a large set of machine-generated training data with respect to a smaller set of annotations made by a human observer.
Data Analysis
The performance of the NLP models was assessed by comparing the model predictions with the manual annotations on the test set of 202 reports; the number of true positives, true negatives, false positives and false negatives was calculated for each radiological finding, along with confusion matrices, sensitivity and specificity, F1 score.
The ResNet-18 models processing the images were evaluated in an analogous manner on the test set of 204 image couples. In addition to confusion matrices and the other metrics, receiver operating characteristic (ROC) curves and the relative areas under the curve (AUC) were calculated for each radiological finding based on the outputs of the top layers of the models, exploiting the “roc_curve” and “roc_auc_score” functions provided by scikit-learn. Furthermore, we attempted to optimize the predictions of the neural networks by calculating the optimal values of the cut-offs used to classify the output of the top layers. The calculation was performed on the outputs obtained on the 481 image couples used as the second training set (Figure 1) and was based on Youden’s J statistics. 18
Occlusion sensitivity maps 19 were used to visualize which areas of the images had the highest importance in determining the predictions. A square patch with a size of 80 pixels moving across the image was used to occlude a portion of the image; the resulting outputs of the top layer of the network were then plotted in a heatmap in which the highest values represented the locations more important for a positive outcome (i.e. the presence of the radiological finding). By occluding the red and blue channels separately, the importance of the regions of the coronal and sagittal projections could be independently determined.
Results
The NLP models had generally high accuracies, ranging from 0.88 to 0.98 (Table 1, Figure 3). Sensitivity and specificity, in particular the former, tended to show lower values; this finding can be attributed to the unbalanced nature of the dataset, in which the radiological findings are more frequently absent than present. Whereas the specificity ranged between 0.84 to 0.99, lower sensitivities such as 0.5 (osteoporosis) and 0.63 (fractures) were calculated. However, 7 out of 12 radiological findings had sensitivity greater than 0.90. A qualitative analysis of the false negative cases demonstrated that the errors were mostly associated with complex wording, long sentences or typographical/speech recognition errors.
Performance of the NLP Models for the 12 Radiological Findings.
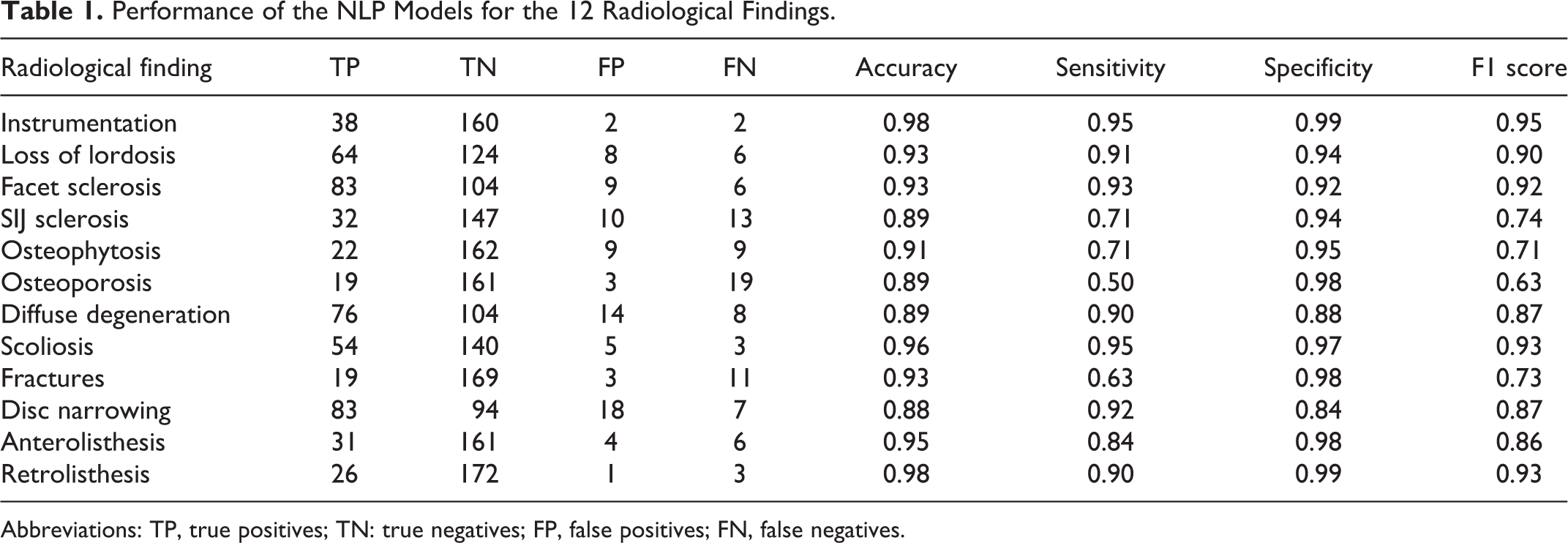
Abbreviations: TP, true positives; TN: true negatives; FP, false positives; FN, false negatives.
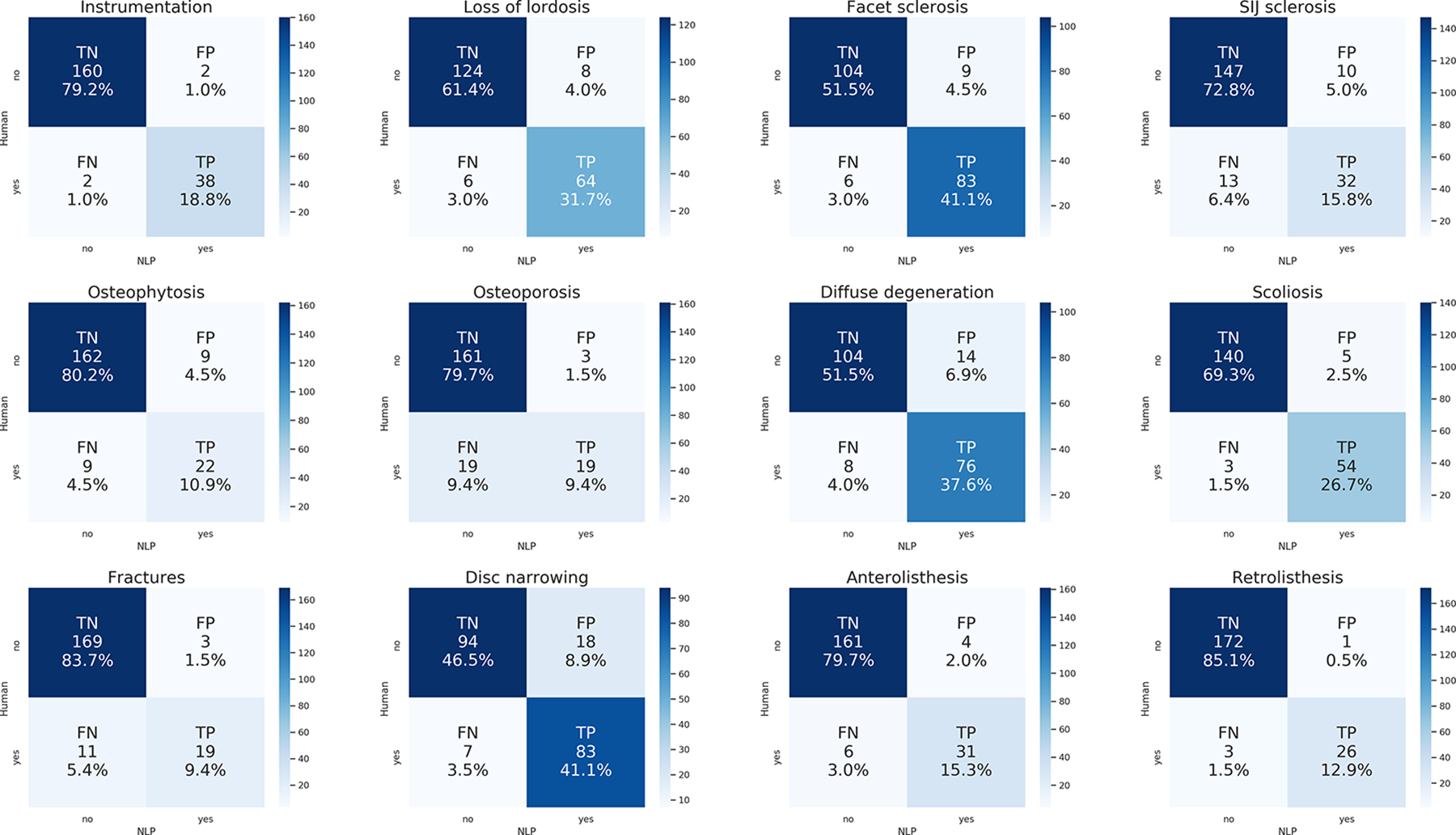
Confusion matrices showing the performance of the NLP models for each radiological finding. The numbers and the color scale indicate the occurrences of each class (TP: true positives; TN: true negatives; FP: false positives; FN: false negatives), both in absolute and relative terms.
The ResNet-18 models trained on the machine-generated data (“1step”) showed a lower performance with respect to the NLP models, with generally higher numbers of false negatives, i.e. missed detections (Table 2, Figure 4). The accuracies ranged between 0.61 (SIJ sclerosis) to 0.99 (instrumentation), the specificity was between 0.41 (disc narrowing) and 1.00 (instrumentation), whereas the sensitivity ranged between 0 (retrolisthesis, for which no true positives were detected) to 0.91 (disc narrowing). In general, the radiological findings for which the models performed better based on the F1 score were: instrumentation (0.94), facet sclerosis (0.86), diffuse degeneration (0.82), osteophytosis (0.78), disc narrowing (0.77), and loss of lordosis (0.76). The models which showed the poorest performance were characterized by high rates of false negatives and few true positives, and were: retrolisthesis (F1 score 0.00), anterolisthesis (0.12), and fractures (0.30).
Performance of the ResNet-18 Models (“1step”) for the 12 Radiological Findings.
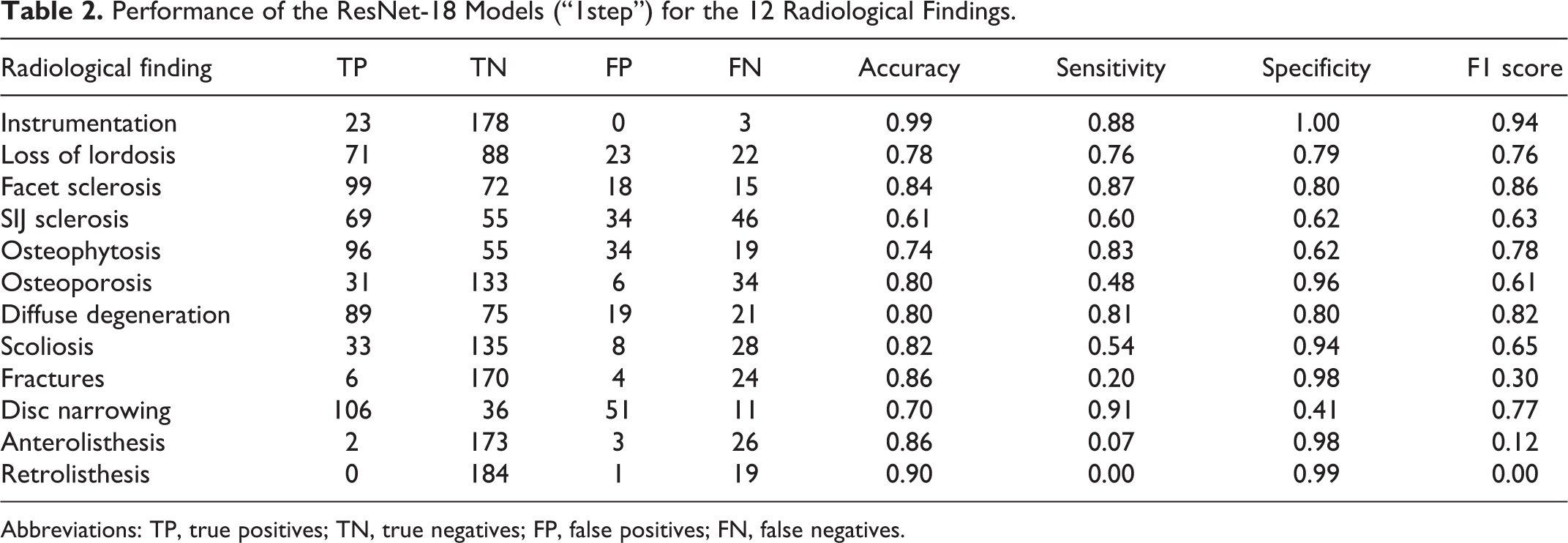
Abbreviations: TP, true positives; TN, true negatives; FP, false positives; FN, false negatives.
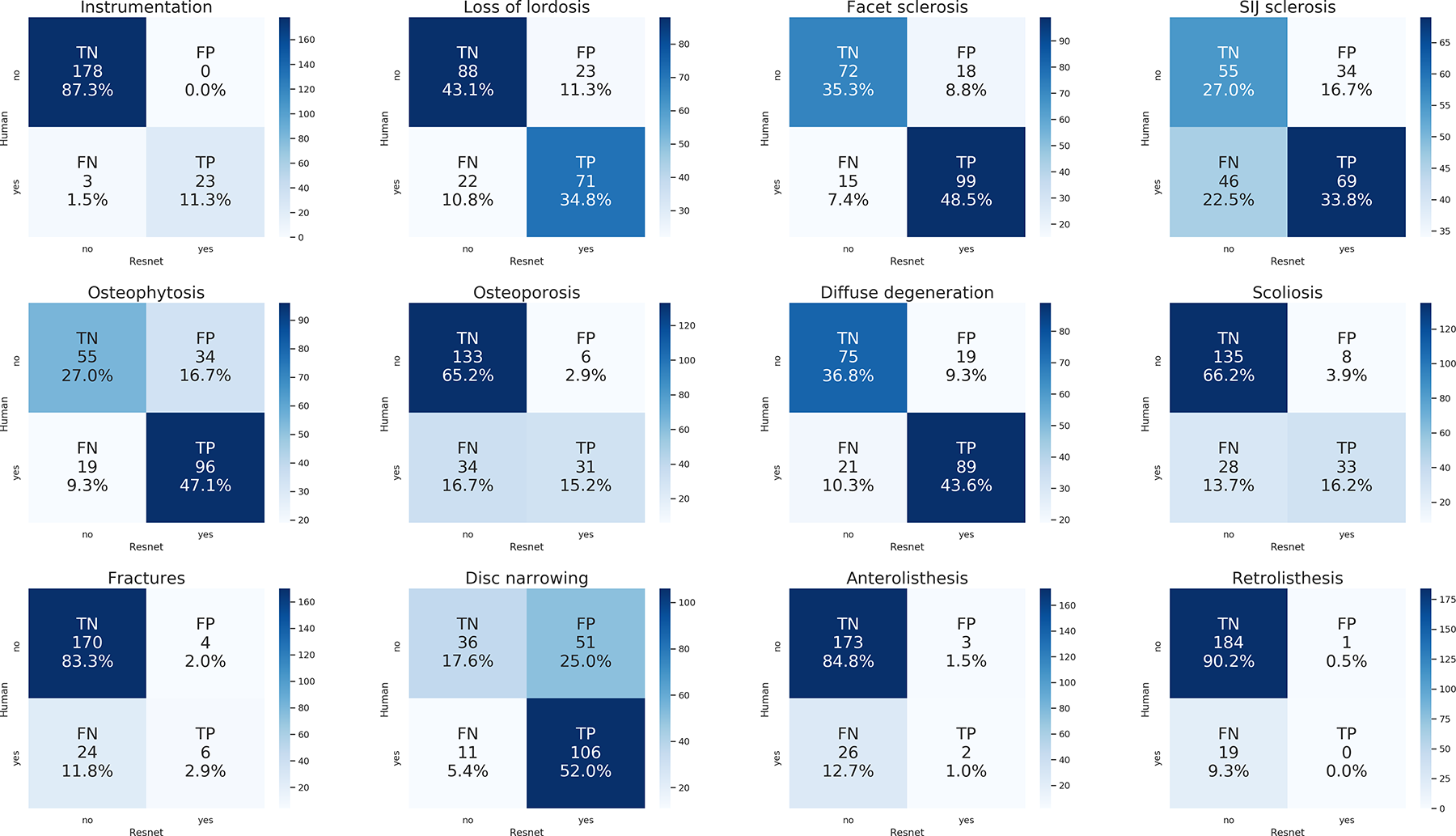
Confusion matrices showing the performance of the ResNet-18 models (“1step”) for each radiological finding. The numbers and the color scale indicate the occurrences of each class (TP: true positives; TN: true negatives; FP: false positives; FN: false negatives), both in absolute and relative terms.
The recalculation of the optimal cut-offs based on Youden’s J statistics allowed for equilibrating the values of sensitivity and specificity for each radiological finding (Table 3), permitting a straightforward interpretation and comparison of the predictive power of the neural networks. While the F1 scores did not show major changes with respect to those shown in Table 2, the values of sensitivity and specificity were largely affected. The best performing models were those assessing instrumentation (sensitivity 0.96, specificity 0.96), facet sclerosis (0.84, 0.84), diffuse degeneration (0.81, 0.85), and osteoporosis (0.82, 0.81); the worst performances were again found for retrolisthesis (0.53, 0.53). Fractures (sensitivity 0.77, specificity 0.76) and anterolisthesis (0.78, 0.72) largely benefited from the recalculation of the cut-offs which allowed reducing the rate of false negatives.
Performance of the ResNet-18 Models (“1step”) for the 12 Radiological Findings, After Recalculating the Optimal Cut-Offs.
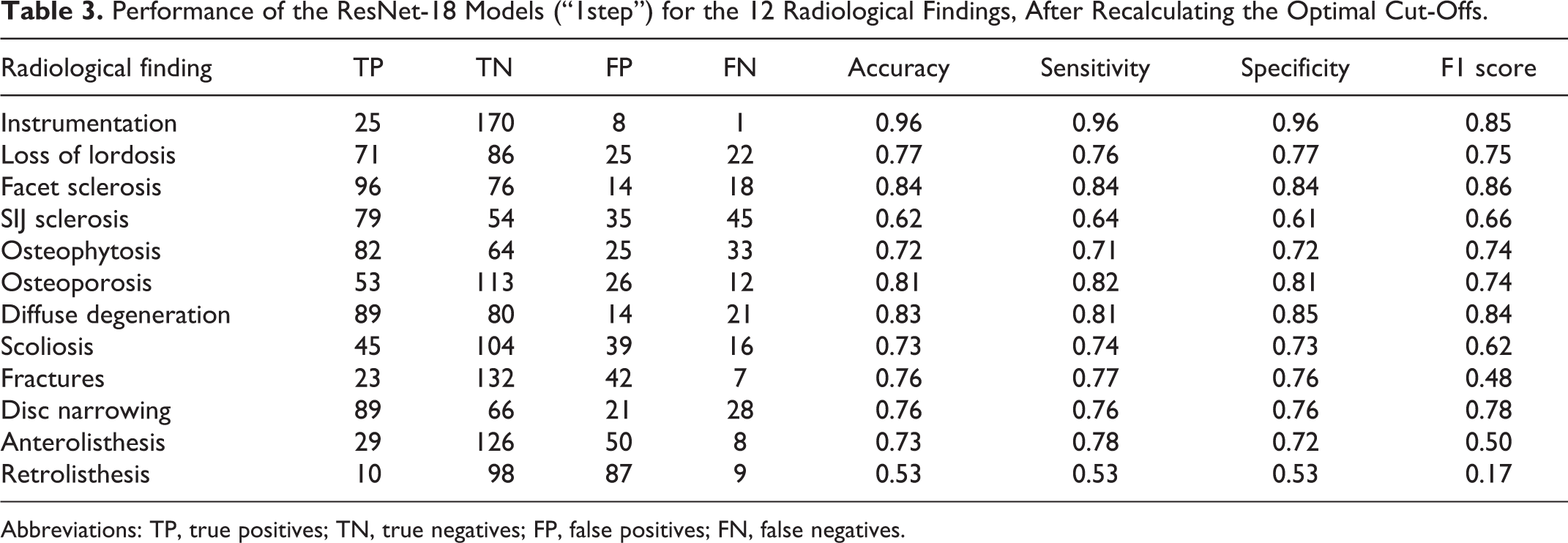
Abbreviations: TP, true positives; TN, true negatives; FP, false positives; FN, false negatives.
The models for which the training was refined exploiting the set of 481 human-annotated image couples (“2steps”) showed marginally improved performance with respect to the “1step” models (Figure 5). The improvement was reflected by the AUC, which increased in all cases with the exception of some cases in which it did not change: instrumentation (1.00), facet sclerosis (0.91), osteoporosis (0.89), and diffuse degeneration (0.91). The largest improvement was observed for osteophytosis, which had AUC 0.71 with the “1step” model and 0.78 with the “2steps” model. The models trained with the 481 image couples only (“step2only”) showed a consistently and widely lower performance with respect to the other models, with the exception of instrumentation (“1step”: 1.00; “step2only”: 0.96), SIJ sclerosis (“1step”: 0.66; “step2only”: 0.66) and osteophytosis (“1step”: 0.71; “step2only”: 0.66).
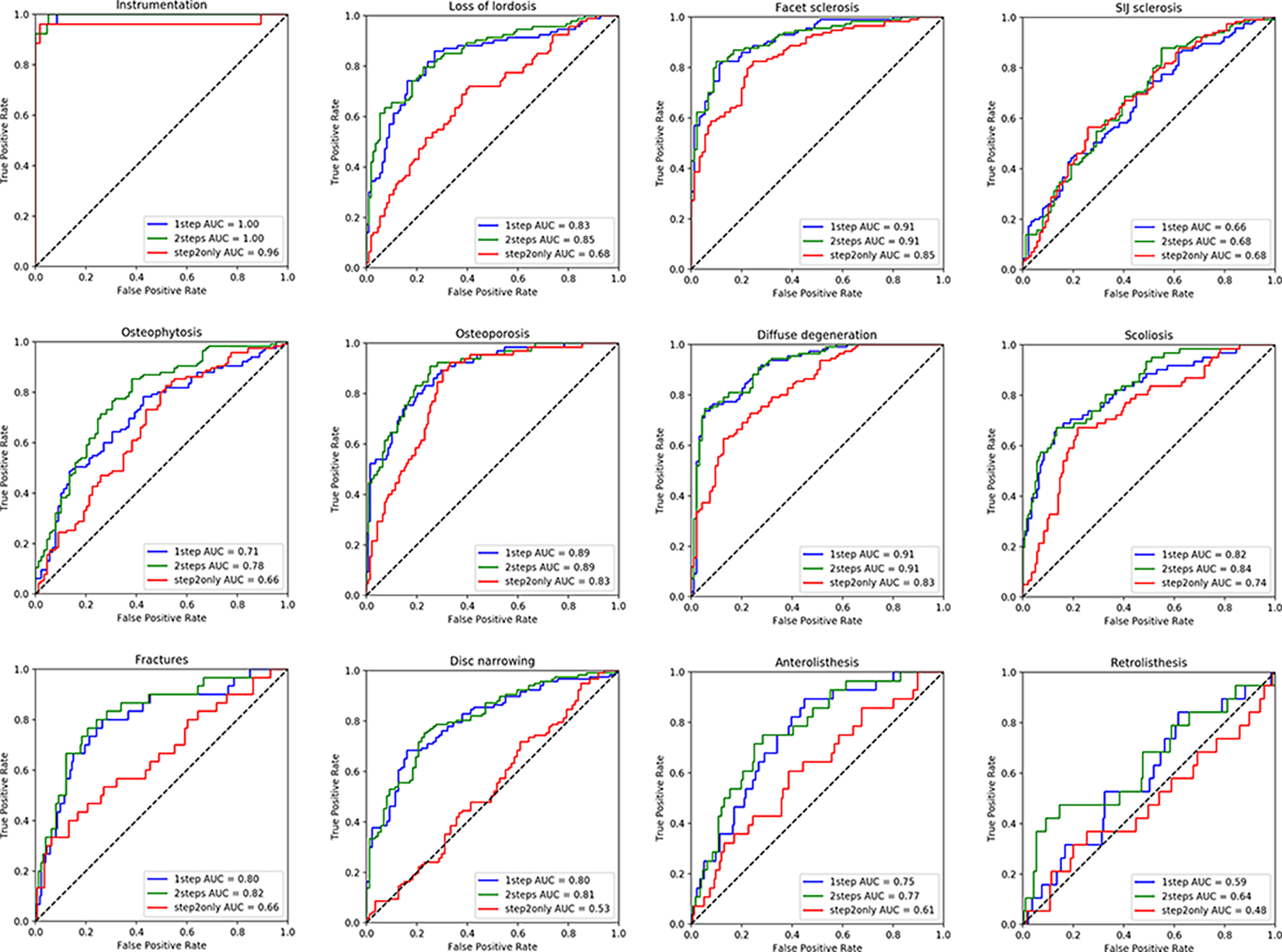
ROC curves showing the performance of the 3 types of ResNet-18 models (“1step,” “2steps,” “step2only”) and the relative AUC for each radiological finding.
Occlusion sensitivity maps did not always offer a human-readable interpretation of the behavior of the neural networks, possibly due to associations between different radiological findings and their frequent simultaneous presence (Figure 6). Regarding true positive detections, in some cases the interpretation of the sensitivity map was straightforward, i.e. the colors indicated regions such as fractures, osteophytes etc. clearly responsible for the positive prediction of the model. In several other cases the maps highlighted areas apparently not involved with the radiological finding of interest.
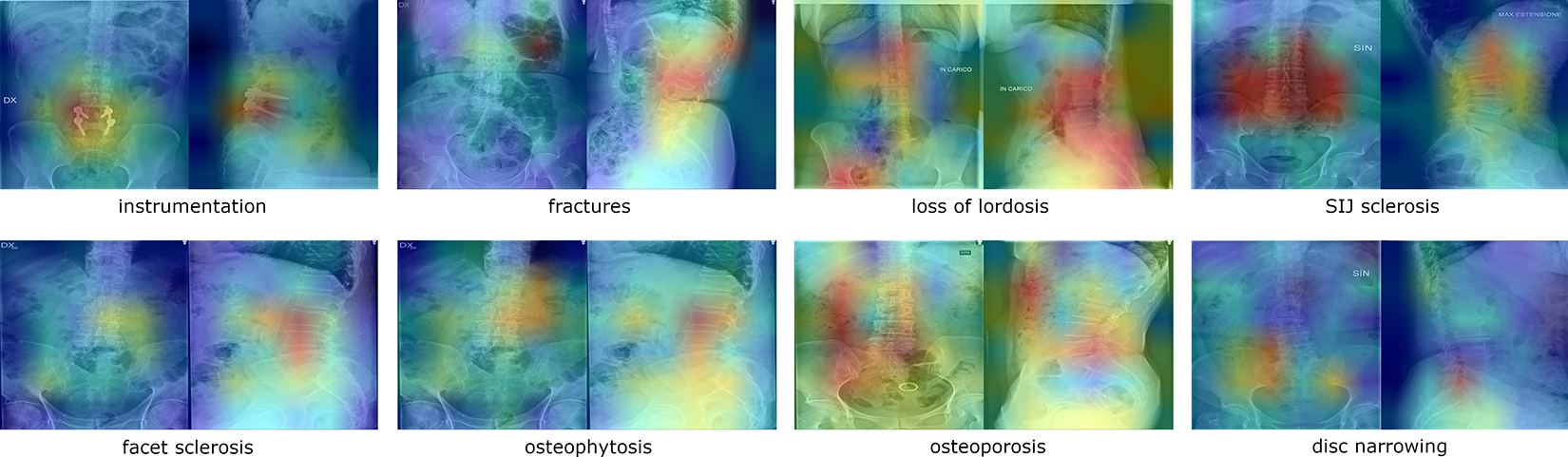
Exemplary occlusion sensitivity maps for true positive cases of various radiological findings, either relatively easy to interpret (first row) or rather obscure (second row). The color scale indicates the areas with low (blue) and high (red) importance in determining the prediction of the ResNet-18 model.
Discussion
In this study, we attempted to leverage the availability of large databases of images and medical reports to train deep learning models exploiting only the information contained in the reports, without any manual annotation of the images. The models trained for detecting radiological findings on the images showed a variable performance depending on the specific findings; whereas 4 in 12 models showed sensitivities and specificities above 0.8 and several others above 0.7 (Table 3), the detection of some specific findings such as retrolisthesis and, to a lower degree, anterolisthesis, fractures and SIJ sclerosis was more problematic. On the contrary, the NLP models which extracted information from the medical reports performed consistently well for all radiological findings, although with some variability. The errors of the NLP models were associated with long sentences containing several pieces of information, clause chaining, typographical mistakes as well as errors of speech recognition software used for dictating the reports.
The performance of the NLP models was in line with those reported for models used for similar purposes. Zech et al reported accuracy, sensitivity and specificity of 0.92, 0.90 and 0.92 respectively for a binary classification task similar to those conducted in the present study, i.e. detecting whether a report contains a critical finding 9 ; such results are in good agreement with several ones of the current models (Table 1). Wang et al developed a NLP model to detect osteoporotic fractures, and the relative fracture sites, from radiological reports 20 ; the authors reported an excellent performance with sensitivity ranging from 0.675 (for vertebral fractures) to 1.00, and specificity of 1.00 for all fractures sites. Ong and coworkers used NLP to extract information about ischemic stroke, its acuity and location from radiological reports, 21 and obtained sensitivities of 0.92, 0.90 and 0.92 for the 3 outcomes respectively, and specificities of 0.75, 0.70 and 0.69 respectively. It should be noted that the 3 literature studies described NLP models purposely developed for the specific application, whereas we decided to use an off-the-shelf model, although at the state-of-the-art, such as BERT; the comparison therefore confirms the value of the recent major advances of general-purpose NLP.
Despite the good performance, the predictions of the NLP models may not correspond exactly with the human interpretation of the text; as mentioned above, the annotations should indeed be considered noisy, resulting in a weak supervision of the deep learning models processing the images. Besides, the original radiological reports are not free from errors and misdiagnoses, which have been quantified as affecting 4% of the reports, 22 further weakening the quality of the annotations. Although it has been shown that deep learning models trained on large datasets are robust with respect to noisy labels, 23 a degradation of their performance should anyway be expected.
Besides, other sources of label noise exist. One of them, possibly even more important than the limited performance of NLP, comes from the fact that radiologists may report only findings which are clinically relevant with respect to the clinical question, discarding observations with negligible, or believed so, clinical value. It is indeed well known that providing clinical information to the radiologist has an effect, reportedly beneficial, on the quality of the report.24,25 However, the availability of clinical information may also negatively affect the report completeness; a clear example is given by the post-operative reports, which tended to be short in the present study as they focused on describing the success of the intervention and the positioning of the instrumentation, disregarding other findings which have been noted in previous examinations. Another example of a commonly unreported finding is given by osteophytes, which are visible in the vast majority of the images of degenerative spines but are not reported unless believed to be clinically relevant, i.e. potentially associated with symptoms and disability. This issue is clearly highlighted by the performance improvement of the “2steps” model (AUC 0.78) with respect to “1step” (AUC 0.71); when providing complete information about osteophytes, i.e. after providing complete annotations based on the images and not only on the reports, the quality of the predictions significantly increased.
It should also be noted that some of the variables are qualitative in nature and therefore subjective. Loss of lordosis is a typical example; most elderly subjects tend to show a reduction of the physiological lordosis with a relatively strong interindividual variability, 26 a direct consequence of the large differences even in the young adult population. 27 The assessment of the loss of lordosis, especially when conducted with no reference to previous examinations and without considering the spinopelvic parameters, 28 assumes therefore a rather subjective nature. Other examples are diffuse degeneration, which has no precise definition, and osteoporosis, which has quantitative measures but they cannot be assessed on planar radiographic projections. 29
The detection of anterolisthesis and retrolisthesis on the radiographs, especially the latter, showed an evidently poorer performance with respect to the other radiological findings. Although being correctly identified by the NLP models in most cases (F1 scores of 0.86 and 0.93 respectively), the ResNet-18 models were generally not able to detect them in the images. The inspection of exemplary reports by a human observer highlighted a possible reason; while only the coronal and sagittal projections obtained in standing were processed by the neural networks, the radiological reports often covered also flexion-extension radiographs acquired in the same session, in which the olistheses were more evident. In the majority of the false negative findings there are indeed no vertebral displacements visible on the images.
Some limitations of the present study should be highlighted. Although the hyperparameters of the ResNet-18 were tuned, we did not perform an extensive optimization of the model architectures, neither for the NLP nor for the models processing the images. We also did not attempt using solutions specifically designed for noisy labels, which may improve the robustness of the predictions.30,31 It should be expected that a higher degree of optimization may determine better performances; however, the use of off-the-shelf models was evidently sufficient to demonstrate the validity of the study hypothesis, i.e. NLP-generated annotations are a valuable resource for training models able to detect radiological findings on images. Another limitation is the relatively small size of the datasets used, 14777 reports and 10083 image couples; although these numbers are larger than those in most AI studies in the field of musculoskeletal radiology,32,33 the use of NLP instead of manual annotations would in principle allow for much larger sizes, even millions of images, which are indeed available in the PACS of our institution. In the present study, the choice of limiting the dataset size was due to practical considerations about the availability of hardware resources for training the models, storage space, as well as the lack of convenient software tools for retrieving images and reports, which are currently being developed. Again, this limitation did not prevent proving the validity of the study hypothesis, which is expected to become even more evident if larger sets of images and reports were available.
In conclusion, this study demonstrated that NLP can generate valuable training data for deep learning models able to detect radiological findings in spine images. Although with limitations associated with the noisy nature of the NLP predictions and the reports themselves, this approach is effective in mitigating the difficulties associated with the manual annotation of large quantities of data, and opens the way to the era of big data for AI tools in musculoskeletal radiology.
Footnotes
Authors’ Note
The study was approved by the ethical committee of IRCCS Ospedale San Raffaele (protocol “RETRORAD”). All patients provided written informed consent for the use of images and anonymized data for scientific and educational purposes.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
