Abstract
Objective
To provide a comprehensive overview of the current use of large language models in clinical medicine and surgery, with emphasis on model characteristics, clinical applications, and readiness for adoption.
Methods
A scoping review of studies on the use of large language models in clinical medicine and surgery was conducted in accordance with the Preferred Reporting Items for Systematic reviews and Meta-Analyses (PRISMA)-scoping review and JBI methodology (protocol registration: 10.37766/inplasy2025.3.0102). A comprehensive search of EMBASE, PubMed, CINAHL, and IEEE Xplore identified 3313 articles published between 2018 and 2023. After screening of articles and full-text review, 156 studies were included. Data were extracted for study type, sample size, clinical specialty, model architecture, training methods, application purpose, and performance metrics. Descriptive analyses were performed.
Results
Most studies were proof-of-concept studies (55.8%) or clinical trials (21.2%), with a steady rise in publications since 2022. Large language models were most frequently used for data extraction (69.9%), followed by clinical recommendations (11.5%), report generation (9.0%), and patient-facing chatbots (7.1%). Proprietary models were used in 57.7% of the studies, whereas 39.7% used open-source models. ChatGPT-3.5, ChatGPT-4, and Bidirectional Encoder Representations from Transformers (BERT) were the most commonly reported models. Only 25.0% of the studies reported models as ready for clinical use, whereas 67.9% stated that the models required further validation. F-score (30.8%) and area under the curve (15.4%) were the most common performance metrics; 10.9% of the studies used expert opinion for validation.
Conclusions
Large language models are increasingly being used in clinical medicine. Although most applications focus on data extraction and summarization, emerging studies are beginning to explore higher-level tasks such as clinical decision-making and multidisciplinary simulation. Significant heterogeneity continues to exist in model architecture, evaluation methods, and reporting standards. Further standardization is needed to develop transparent evaluation frameworks and ensure safe, reliable integration of large language models into complex clinical workflows.
Keywords
Introduction
Natural language processing (NLP) and large language models (LLMs) represent a frontier in the field of artificial intelligence (AI), leveraging vast amounts of data to comprehend and generate human-like text. 1 Most modern LLMs are built on transformer architecture, a neural network design that allows for efficient parallel processing and long-range dependency tracking within text sequences. These models are typically pre-trained on large corpora using self-supervised learning objectives, which enable them to develop broad, generalizable language understanding capabilities that are currently being adapted for various clinical tasks.
Despite this shared foundation, LLMs differ in their architectural configuration, which in turn influences their function. Early LLMS such as Bidirectional Encoder Representations from Transformers (BERT) utilize an encoder-only architecture optimized for tasks that require text comprehension, such as classification and entity recognition (Table 1). Conversely, newer models such as generative pre-trained transformer (GPT) adopt a decoder-only architecture that enables them to generate coherent text sequences and engage in conversational tasks. However, hybrid models such as T5 and LLaMA employ an encoder–decoder structure, which allows them to understand and generate language within an identical framework.
Large language model architecture and differences.
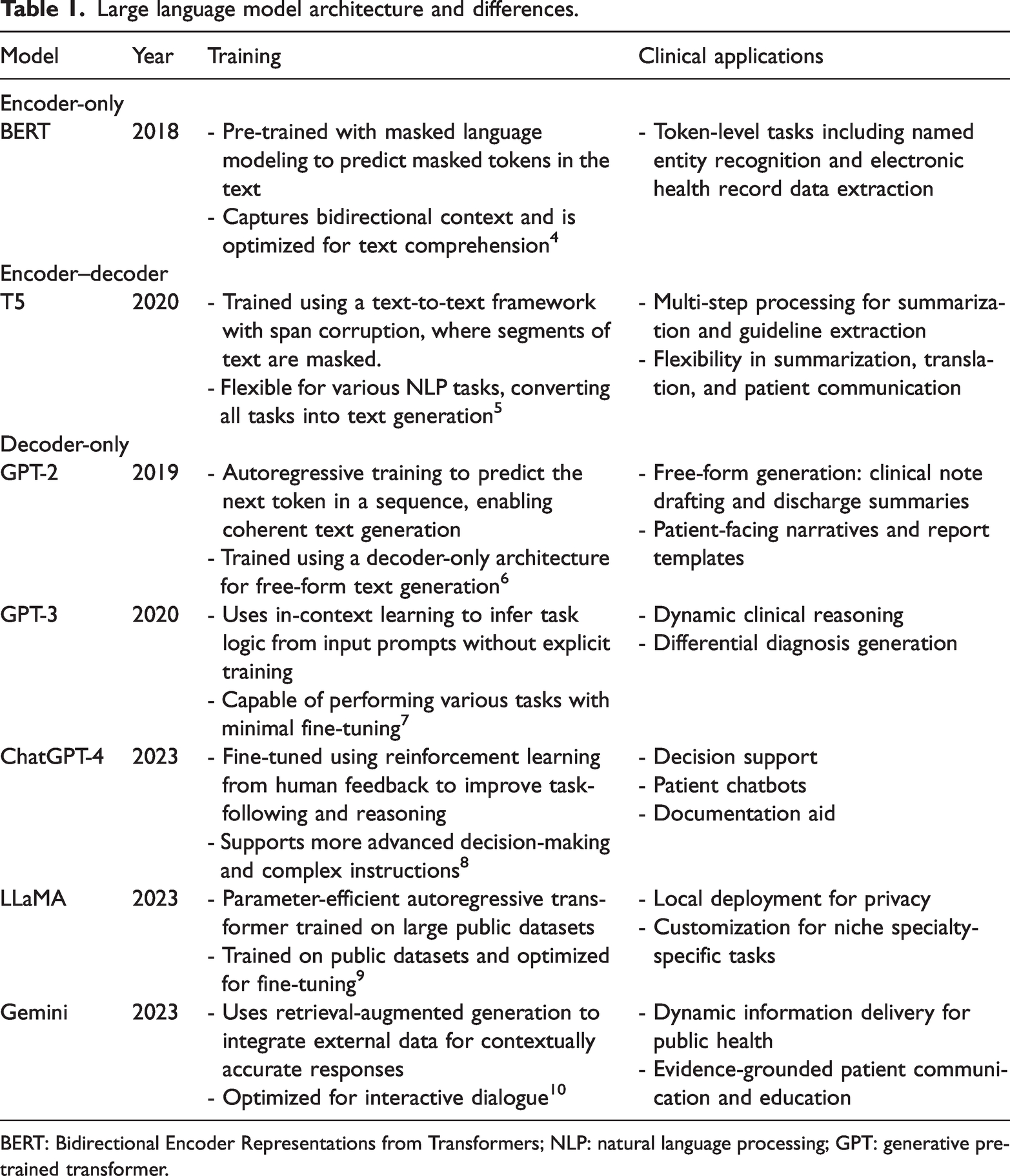
BERT: Bidirectional Encoder Representations from Transformers; NLP: natural language processing; GPT: generative pre-trained transformer.
Model architecture in LLMs has a direct impact on their real-world applications and potential for improving clinical medicine and surgery. Encoder-only models such as BERT are well-suited for structured tasks such as extracting electronic health records. Decoder-only models such as GPT are better suited for generative applications such as medical note transcription. Encoder–decoder models can combine these computational properties and assist with complicated tasks such as clinical decision support. Additionally, retrieval-augmented generation has further extended model capabilities by integrating external databases, enabling the generation of contextually grounded, evidence-based responses in real time.2,3
Validated models such as ClinicalBERT and BioBERT are available for medical applications, but they are typically used for medical chart extraction and medical exam administration.11–13 These tasks do not take advantage of the computational potential of modern LLMs, and preliminary studies have shown that these models can be applied to complicated tasks such as diagnostics and clinical decision support.14–16 At present, however, LLM use in medical diagnostics and clinical practice is challenged by the lack of standardized protocols and reporting guidelines for LLM development, training, and performance evaluation. To address this knowledge gap, we conducted a scoping review to (1) describe the adoption of LLMs in clinical medicine and surgery, (2) evaluate the role of LLMs in advanced medical application, and (3) detail modern LLM training and reporting practices in medicine.
Materials and methods
Protocol and registration
We performed a scoping review of studies on the use of LLMs in clinical medicine and surgery. We used the Preferred Reporting Items for Systematic reviews and Meta-Analyses (PRISMA)-scoping review guidelines to aid the reporting of the study. 17 This proposed scoping review was conducted in accordance with JBI methodology for scoping reviews and was registered retrospectively at INPLASY (registration DOI: 10.37766/inplasy2025.3.0102).
Eligibility criteria
We included all study types that directly measured the validity of utilizing LLMs in a clinical context, except for narrative reviews, systematic reviews, and other scoping reviews.
For the purposes of this review, LLMs refer broadly to transformer-based models that have been pre-trained on large text corpora and exhibit NLP capabilities applicable to clinical tasks. This broad definition encompasses open-source models based on transformer architecture (e.g. GPT, BERT, and LLaMA) as well as proprietary models designed for specific purposes. Although not all encoder-based models are generative in nature, they are included in this analysis because of their frequent deployment in clinical NLP applications under the broader umbrella of transformer-based LLMs.
We included studies that evaluated the performance of LLMs in diagnosis, treatment, and/or management in a clinical or simulated clinical setting, including nonhuman synthetic clinical datasets and simulated clinical workflows. We included the following medicine and surgical specialties: cardiology, emergency medicine, endocrinology, gastroenterology, general surgery, genetics, geriatrics, hematology, intensive care, infection disease, internal medicine, neurology, obstetrics and gynecology, oncology, ophthalmology, orthopedic surgery, otolaryngology, pathology, pediatrics, primary care, psychiatry, radiology, respirology, and urology.
We excluded animal studies and biological simulations on the basis that the results of such studies could not be reliably translated into clinical practice. Studies were also excluded if the authors used LLMs outside of clinical contexts or if they were used to aid exam-taking, medical education, or research planning. Although these studies offer valuable insight into LLM performance, they do not offer direct insight into how LLMs could impact patient care. Additionally, studies were excluded if the performance of LLMs was not the outcome measure, as these studies typically do not provide a quantifiable measure of performance for comparison. Next, studies were excluded if they were conducted in the context of allied health services, dentistry, pharmacy, and/or public health settings. Although we believe that these are important domains, we choose to maintain a narrow focus for our review of studies on only clinical medicine and surgery.
Database search
The search strategy aimed to retrieve both published and unpublished studies. A three-step search strategy was utilized in this review. First, an initial limited search of EMBASE (Ovid) and CINAHL was undertaken to identify articles on the topic. The text of the titles, abstracts of relevant articles, and the index terms used to describe the articles were used to develop a full search strategy with help from an information specialist. The search strategy, including all identified keywords and index terms, was adapted for each included database. The search strategy is presented in Appendix 1. PubMed and IEEE Xplore were utilized to include gray literature and conference proceedings, given the timely nature of the topic. The reference lists of all included studies were also screened for additional studies. Only studies published in English were included. Studies published between January 2018 and October 2023 were included. We opted to exclude any publications prior to 2018, given the introduction of the BERT model in this year and the fact that prior models would not have utilized the same architecture as that of the current language models. 2
Article selection and data abstraction
Following the search, all identified citations were collated and uploaded into Covidence, and duplicates were removed. Following a pilot test, titles and abstracts were screened by two independent reviewers (EL & SP) for assessment against the inclusion criteria for the review. Potentially relevant sources were retrieved in full, and their citation details were imported into Covidence. The full text of selected citations was assessed in detail against the inclusion criteria by two independent reviewers. Reasons for the exclusion of sources of evidence in the full text that do not meet the inclusion criteria were recorded and reported in the scoping review. Any disagreements that arose between the reviewers at each stage of the selection process were resolved through consensus or discussion with an additional reviewer (PS). The results of the search and the study inclusion process were reported in full in the final scoping review and presented in a PRISMA flow diagram.
Data were extracted from the studies included in the scoping review by two independent reviewers using a data extraction tool developed by the reviewers. The data extracted included specific details about the participants, concept, context, study methods, and key findings relevant to the review question.
A draft extraction form is provided (Appendix 2). The draft data extraction tool was modified and revised as necessary during the process of extracting data from each included evidence source. Included studies were assessed for the type of medical specialty, type of study, purpose of LLM use, name of LLM, open-source vs. proprietary, trained vs. untrained, training tool, workflow, sample size, outcome measures and value, readiness for clinical application, and conclusions drawn by the authors. When required, the authors of papers were contacted to request missing or additional data.
Statistical analysis
Given the exploratory nature of this scoping review, we conducted a descriptive statistical analysis to characterize the included studies. The primary aim was to provide an overview of the landscape of LLM use in clinical medicine. No data manipulation other than a grouping of data fields was performed. For instance, sample sizes were grouped into four categories: small (≤10), intermediate (11–100), large (101–1000), and extra-large (>1000). LLM application types, model accessibility, training methods, and readiness for clinical use were all coded based on predefined ordinal or categorical schemes. Free-text responses for the type of training, workflow, and conclusions were categorized based on conceptual similarities. Given the heterogeneity of the study design, there were no standardized definitions of clinical readiness. Hence, we structured our protocols for determining readiness based on prior systematic reviews in AI. 18 We developed categories of model readiness, whereby models deemed as not requiring further validation were interpreted as “ready.” Models requiring further training or validation were interpreted as “requires further improvement,” whereas those deemed to have no role in future clinical application were deemed “failed.”
Results
Database search results
Of the 3313 identified articles, 156 were eligible and included in this scoping review (Figure 1).19–172 A significant year-on-year increase in the number of studies published was observed from 2018 to 2023 (Figure 2). The characteristics of the studies are reported in Table 2 and Appendix 3. Most studies were proof-of-concept or simulated prospective cohort studies (

PRISMA flowchart. PRISMA: Preferred Reporting Items for Systematic reviews and Meta-Analyses.
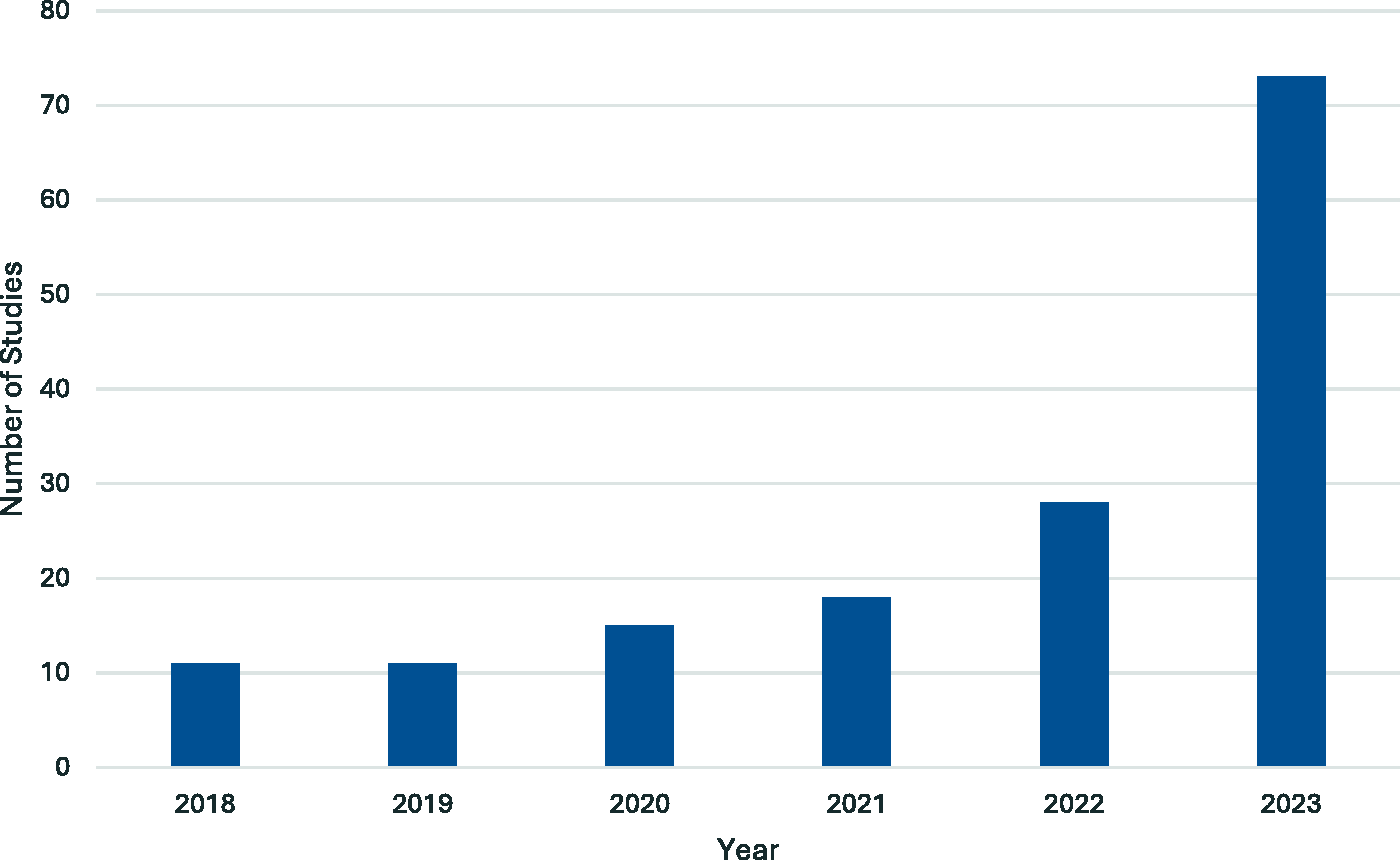
Number of studies published by year.
Baseline characteristics of the included studies.

Characteristics of LLM studies and tools
All studies reported on the function and performance of LLMs in various clinical scenarios. The most common use for LLMs was data extraction (
Summary of large language models used in clinical settings.
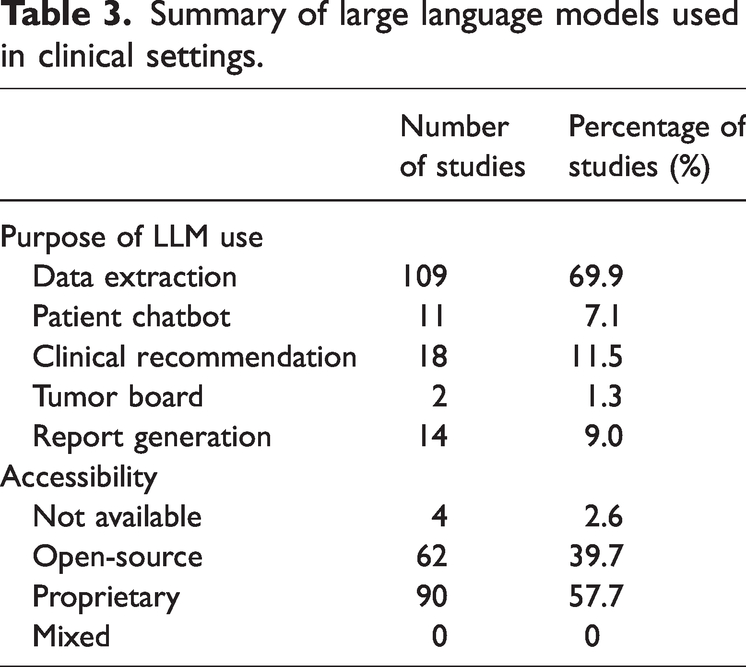
Frequency of large language model training methods and readiness for clinical application.

Description of clinical workflow
Despite the varying specific parameters and instructions, the studies examined can be largely grouped into two distinct workflow regimes. Most studies (
A smaller group of studies (
Measures of accuracy
A wide range of outcome measures were used to evaluate the performance and accuracy of LLMs across the included studies. The most frequently reported metric was the F-score, which was cited in 48 studies (30.8%), followed by the area under the curve (AUC), which was reported in 24 studies (15.4%). Diagnostic accuracy was reported in 25 studies (16.0%), whereas agreement with expert opinion was reported in 17 studies (10.9%). Measures such as sensitivity and specificity were reported together in 11 studies (7.1%) and positive predictive value in 3 studies (1.9%). A smaller number of studies utilized alternative or task-specific measures, including accuracy of data extraction (
Analysis of LLM use by medical specialties
The type of LLM utilized varied across medical and surgical specialties, reflecting differing clinical needs, access to data, and stages of adoption. A detailed report of LLMs used by different medical specialties is provided in Appendix 4.
Radiology (
Psychiatry (
In internal medicine (
Primary care (
Specialties such as hematology, intensive care, neurology, and genetics often relied on niche or purpose-built models, including EHRead, NLP-Dx-BD, ASA, and Bio_ClinicalBERT. These were mostly employed for structured data parsing and clinician support, with no studies in these fields reporting models as ready for clinical use.
Clinical utility of LLMs
The threshold for the adoption of LLMs into clinical workflows varied widely across the included studies. Summarized results are reported in Table 4. Eleven studies (7.1%) reported outright failure of LLMs to achieve meaningful progress toward their intended outcomes. These failures were predominantly associated with untrained, open-source models that produced inaccurate or unreliable responses when applied to clinical scenarios. Only one of the failed models had been trained using patient data; this proprietary LLM in the field of otolaryngology became overly sensitive to the outcome of interest, resulting in an unacceptably high false-positive rate.
The majority of the studies (
A subset of studies (
Discussion
Herein, we presented a scoping review of the clinical applications of LLMs in medicine and surgery. The upward trend in the number of studies being published each year demonstrates the growing importance and interest in language models associated with the medical community. Although most studies employed LLMs for foundational tasks such as data extraction, relatively few studies explored their use in higher-function clinical applications, including simulating tumor boards or providing clinical recommendations. These higher-order tasks require not only accurate information retrieval but also complex contextual reasoning, patient-specific tailoring, and integration of evolving clinical guidelines. The use of models such as PsyBERTpt and ChatGPT-4 reflects the growing interest in leveraging LLMs in these areas, although further research is needed to ensure consistency and reliability in outputs.
We revealed that 41.2% of the LLMs were open-source, although a substantial number of studies failed to detail the exact model. Open-source LLMs carry the advantage of being low-cost, often with better technological support, but sometimes do not offer the quality of proprietary models. 173 Proprietary models are designed for precision and can offer superior privacy protection for sensitive patient data when run using local storage. 174 They, however, require a longer time to produce and incur a greater financial burden. 175
Studies exploring the use of ChatGPT and BERT variants continue to dominate the literature. There remains a notable underrepresentation of studies utilizing hybrid or domain-adaptive models, which are developed for use in resource-constrained environments such as mobile health platforms, rural clinics, or offline decision support tools.9,176 Despite the potential of these models, a few studies prior to our review have evaluated the clinical performance of these models beyond structured information extraction tasks. This gap in the literature highlights a critical opportunity for future research to assess the real-world impact of deploying domain-adaptive and locally fine-tuned LLMs in diverse clinical settings.
Most studies also reported on the training status of their models and type of datasets used but did not specify the intensity of their fine-tuning regime or the specificity of the datasets. Existing literature suggests that the intensity of fine-tuning and the quality of data used for training have a direct impact on the accuracy of the output model irrespective of using open-source or proprietary models. 177 As interest in LLMs continues to increase, there is a need for better transparency in reporting as well as standardization of model selection and training regimes. 178
Additionally, there was substantial heterogeneity in the methods used to assess the accuracy and performance of LLMs across studies. A total of 14 different outcome metrics were identified, with wide variation in their frequency and application. The most commonly reported measure was the F-score, used in 48 studies, followed by the AUC in 24 studies. Moreover, there were task-specific or less commonly used measures such as C-index, BLEU-1, and DISCERN scores. The use of expert opinion as a reference standard was common in early-phase studies, particularly for evaluating models in subjective or interpretive tasks. Although this approach provides clinically grounded validation, it is inherently limited by inter-rater variability and potential bias. 179 The absence of a standardized benchmark across studies relying on expert evaluation makes it difficult to compare performance across different models or specialties. Furthermore, expert agreement may reflect consistency rather than correctness, which poses challenges in domains where clinical guidelines are evolving or where expert consensus is lacking. 180
Comparison of metric selection revealed important considerations for interpreting model performance. The F-score, which represents the mean of precision and recall, is frequently used in information retrieval and classification tasks where imbalanced datasets are common. Its utility lies in penalizing models that over-prioritize sensitivity or specificity, offering a more balanced view of performance. 181 However, the F-score does not provide information about the model’s ability to discriminate between classes across thresholds, limiting its interpretability in probabilistic outputs. 182 In contrast, AUC is commonly used in classification problems and reflects the model’s ability to distinguish positive cases from negative ones across all thresholds. 183 This metric is threshold-independent and intuitive to interpret but may be less informative in datasets with class imbalance, where high AUC values can be achieved despite poor real-world performance. 184 Furthermore, AUC does not penalize misclassification severity, which may be clinically relevant in high-risk or safety-critical applications. 185 Given the diversity of tasks and model architectures, the choice of evaluation metric should be tailored to the specific use case. Studies evaluating diagnostic classification may benefit from reporting both AUC and F-score in tandem, whereas those focusing on entity recognition or summarization should prioritize token-level precision, recall, and agreement with structured reference standards.
The current variability in performance assessment highlights the need for standardized, task-specific evaluation frameworks to ensure comparability and reproducibility across future LLM studies. Recent publications in LLM development have proposed frameworks for fine-tuning models to specific tasks as well as objective scoring systems for measuring output accuracy. 186 These were not adopted in the training of the models reviewed in this study but are an important area for future studies. 187 Additionally, few studies tend to align model evaluation with clinical impact, such as assessing how language model outputs affect patient outcomes. To address this gap, future research should move toward more comprehensive evaluation frameworks that capture both technical and real-world performance. One such approach is the Holistic Evaluation of Language Models framework, which emphasizes multidimensional benchmarking across criteria such as accuracy, robustness, fairness, and efficiency. 188 When applied to clinical settings, such a framework could incorporate patient-centered outcome measures, such as changes in treatment decisions, patient satisfaction, or care quality metrics, to evaluate whether models meaningfully improve clinical care. Broader adoption of such evaluation strategies will be critical for ensuring that LLMs deployed in healthcare are not only technically performant but also clinically impactful.
The current literature on LLMs in clinical contexts is limited by methodological and practical challenges. Many studies are exploratory, rely on retrospective or synthetic data, and lack external validation, reducing generalizability. Models are often tested in narrow or idealized settings, making real-world applicability uncertain. Inconsistent outcome metrics and the absence of standardized benchmarks further hinder meaningful comparisons across studies. LLMs themselves present technical limitations. Generative models such as ChatGPT are prone to hallucinations and operate as black boxes, limiting transparency in clinical decision-making. Most of these models lack real-time access to clinical data, and many embed training data biases that risk perpetuating disparities. Additionally, the computational demands of LLMs, dependence on proprietary platforms, and limited fine-tuning options for domain-specific use pose barriers to adoption. To improve reproducibility and rigor, future studies should clearly describe model selection, training protocols, and performance metrics and justify assessments of clinical readiness. Addressing these issues will enhance methodological transparency and facilitate more accurate cross-study comparisons.
Conclusion
Our study is among the first to provide a comprehensive overview of the current landscape of LLMs and their clinical utility in medicine and surgery. Radiology and medical specialties were the most active areas of study, with ChatGPT and BERT being the most commonly used models. Although most studies focused on low-risk tasks such as data extraction and documentation, we also identified emerging efforts to explore higher-order applications, such as clinical decision-making and simulated multidisciplinary discussions. These advanced uses represent a novel and important frontier for LLM integration in healthcare, requiring models that are both contextually aware and clinically reliable. However, significant heterogeneity in model training, evaluation, and reporting standards persists. Standardization in model validation, alongside focused development of interpretable tools for complex clinical reasoning, is essential to ensure safe, transparent, and impactful deployment of LLMs in patient care.
Footnotes
Acknowledgements
None.
Author contributions
Eric Liang: Research question formulation, search strategy development, data extraction, data analysis, manuscript drafting and editing.
Sophia Pei: Search strategy development, data extraction, data analysis, manuscript drafting.
Phillip Staibano: Research question formulation, search strategy development, manuscript editing.
Benjamin van der Woerd: Research question formulation, manuscript editing.
Data availability statement
All data generated or analyzed during this study are included in this published article and its supplementary information files.
Declaration of conflicting interests
None.
Ethics approval
Not applicable.
Funding
Not applicable.
