Abstract
CRISPR-Cas9 has rapidly emerged as a transformative genome editing platform. To analyze and map scientific research on CRISPR-Cas9 in the context of human disease pathology. A bibliometric study was conducted on original research articles indexed in Scopus between 2010 and 2024, focused on CRISPR-Cas9 applications in human disease pathology. A total of 4,313 publications were identified with an average annual growth rate of 54.5% and an H-index of 146. The top publishing journals were Scientific Reports, International Journal of Molecular Sciences, and Nature Communications. The United States led with 1,727 articles, followed by China (n = 1,355), with 50.9% international collaboration for the United States versus 34.4% for China. The average number of authors per article was 10.8. The top-cited article reported on CRISPR-Cas9 therapy for beta-thalassemia and sickle cell disease, followed by major studies on cystic fibrosis, amyloidosis, muscular dystrophy, and colorectal cancer. The following research themes were identified: (1) cancer pathways and metastasis (2) viral genome targeting; (3) mutation-driven genetic disorders; and (4) hematological disorders. VOSviewer maps indicated that off-target effects, delivery methods, and ethical concerns are major challenges for CRISPR-Cas9 use in human diseases. The analysis reflects exponential scientific growth and translational momentum. Future research will require refined delivery strategies, safety optimization, and ethical integration to fully harness CRISPR-Cas9’s clinical potential.
Plain Language Summary
Scientists have developed a powerful tool called CRISPR-Cas9, which works like molecular scissors to change specific parts of the DNA. This tool will change the way we treat and study human diseases. This study looked at how CRISPR-Cas9 has been used in medical research over the past decade by analyzing more than 4000 research articles published from 2010 to 2024. The study found that CRISPR-Cas9 research is growing extremely fast. Most of the research comes from the United States and China, with top scientists collaborating across many countries and institutions. CRISPR-Cas9 is being used to study and try to treat many illnesses, including cancer, genetic diseases like sickle cell disease and thalassemia, cystic fibrosis, HIV, and Alzheimer’s disease. CRISPR-Cas9 works by targeting faulty genes to achieve a desired effect. For example, scientists have used CRISPR-Cas9 to correct the gene problems in blood disorders and to reduce harmful proteins in the body for diseases like amyloidosis. The study also mapped out research areas: (1) cancers, (2) viruses like HIV, (3) inherited diseases, and (4) blood disorders. These areas are seeing the most progress because CRISPR-Cas9 offers a precise way to fix the underlying genetic diseases. CRISPR-Cas9 is being used now and has the potential to cure the diseases that were previously untreatable. But there are still challenges, like making sure it is safe and works well inside the body. Scientists must also address ethical concerns as the technology gets closer to being used in real patients.
Highlights
CRISPR-Cas9 research in human diseases grew rapidly indicating global interest.
Clinical success in treating sickle cell disease and thalassemia signaling a transition from lab to clinic.
CRISPR-Cas9 is being investigated in different types of cancers, rare diseases, and viral diseases.
Research on CRISPR-Cas9 is team-based requiring geneticists, clinicians, and bioengineers.
The findings help researchers, policymakers, and physicians track progress and set priorities for future breakthroughs.
Research progress must proceed with careful regulation and ethical consideration.
Introduction
Gene therapy represents a paradigm-shifting approach in medicine that aims to correct or modify defective genetic material responsible for disease (High & Roncarolo, 2019; Kaufmann et al., 2013; R. Tang & Xu, 2020; Wirth et al., 2013). Since its inception in 1970s, when recombinant DNA technology enabled gene modification, gene therapy has evolved dramatically, from early viral vector-based systems to contemporary genome editing platforms (Bulcha et al., 2021; Friedmann, 1992; Wirth et al., 2013). The field has witnessed significant breakthroughs, particularly in treating monogenic diseases, by delivering functional genes, or editing disease-causing mutations within target cells (de la Fuente-Núñez & Lu, 2017; Hsu et al., 2014). A landmark in gene editing emerged in 2012 with the application of CRISPR-Cas9 (clustered regularly interspaced short palindromic repeats associated protein 9), a microbial defense system re-engineered to target specific DNA sequences in eukaryotic genomes (de la Fuente-Núñez & Lu, 2017; Hsu et al., 2014). The discovery and development of CRISPR-Cas9 as a genome-editing tool began with the identification of CRISPR in bacterial genomes in the late 1980s, but its immune function was elucidated in 2007 by Philippe Horvath and colleagues (Barrangou & Doudna, 2016; Doudna & Charpentier, 2014; Hsu et al., 2014). The breakthrough came in 2012, when Jennifer Doudna (USA) and Emmanuelle Charpentier (France) demonstrated that the Cas nuclease could be programed with a guide RNA to cut DNA at specific sites, enabling precise gene editing. Their pioneering work laid the foundation for widespread applications in biotechnology and medicine (Doudna & Charpentier, 2014; Hsu et al., 2014; Jinek et al., 2012). In recognition of this transformative advancement, Doudna and Charpentier were awarded the Nobel Prize in Chemistry in 2020, making history as the first all-female team to win a Nobel in the sciences (Charpentier & Doudna, 2020).
CRISPR-Cas9 utilizes a guide RNA to direct the Cas9 nuclease to precise genomic loci enabling targeted DNA double-strand breaks and facilitating gene knockout, repair, or insertion (Doudna & Charpentier, 2014). Compared with previous genomic-editing tools such as zinc finger nucleases (ZFNs) and TALENs, CRISPR-Cas9 is simpler, more cost-effective, highly specific, and amenable to multiplexing (Gupta et al., 2019; Janik et al., 2020; Severi & Akbari, 2024; Tyumentseva et al., 2023). The advantages of CRISPR-Cas9 in human disease applications include its scalability, ease of design, broad applications across cell types and precision in targeting specific genomic sites. These attributes have led to its rapid adoption across biomedical disciplines including oncology, neurology, infectious diseases, and rare genetic diseases (Ma et al., 2023; S. W. Wang et al., 2022). However, challenges persist, including off-target effects, immunogenic response, limited in vivo delivery systems, and ethical concerns regarding germline editing (Gostimskaya, 2022; Guo et al., 2023; X. H. Zhang et al., 2015). Nevertheless, recent clinical successes, such as CRISPR-Cas9 based therapies for sickle cell disease (SCD) and beta-thalassemia, underscore its revolutionary potential to transform medicine (Ansori et al., 2023; Bhowmik & Chaubey, 2022; Hanafy et al., 2020; M. H. Lee et al., 2022; Ma et al., 2023; Yun & Ha, 2020; Zhan et al., 2019; X. H. Zhang et al., 2015; Zhao et al., 2021). CRISPR-Cas9 could transform medicine by letting us diagnose, treat, and prevent many diseases with personalized therapies that aim to cure, not just ease symptoms. As the field rapidly expands, there is an urgent need to understand its trajectory, influential contributors, and emerging thematic areas.
Knowledge mapping and bibliometric analysis are two complementary methodologies used to systematically evaluate the landscape of scientific research on a specific topic. Knowledge mapping refers to the process of visualizing structure, development, key thematic areas within a body of literature, helping researchers identify how knowledge is generated, interconnected, and evolving over time. Meanwhile, bibliometric analysis quantitatively assesses publications using metrics such as average annual growth rate (AAGR), citation counts, and key contributors (Donthu et al., 2021). Bibliometric analysis is different from traditional or systematic review. Traditional narrative reviews can be subjective and selective. Systematic reviews use strict criteria and narrow focus on interventions or clinical outcomes. Bibliometric analysis takes a wider view: it offers an objective, data-driven map of a field’s structure and how it evolves (Grant & Booth, 2009; Sutton et al., 2019). Bibliometric analysis is particularly useful in fast evolving fields such as the use of CRISPR-Cas9 technology in human disease pathology, where the research volume and diversity are rapidly growing.
Over the past decade, numerous bibliometric studies have explored different topics in the rapidly expanding landscape of CRISPR-Cas9 research areas. One study focused on agricultural biotechnology where CRISPR-Cas9 is applied to enhance crop yield, nutritional benefit, and resistance to stress. This area also addresses regulatory and ethical concerns surrounding gene-edited organisms (Kolkur et al., 2024). A second study focused on environmental biotechnology, where CRISPR-Cas9 is employed for ecological monitoring, enhancing plant tolerance to environmental stress, developing biofuels, and controlling invasive species, all contributing to climate change mitigation and sustainability goals (Verdezoto-Prado et al., 2025). A third study focused on disease detection application focusing on CRISPR-Cas9-based diagnostics for infectious diseases and point-of-care detection for pathogens such as SARS-CoV-2 and HIV (Kumar Sachan et al., 2024; Leta et al., 2024). A fourth study focused on stem cell research in which CRISPR-Cas9 is utilized to model human diseases, investigate gene regulation, and explore therapeutic gene correction without inducing double-strand DNA breaks (Yoon et al., 2024). Closely related to the previous topic is the growing body of research on the induced pluripotent stem cells (iPSCs), where CRISPR-Cas9 enables disease modeling, drug testing, and chromosomal therapy for hereditary disorders (J. Xu et al., 2025). A fifth study focused on extracellular vesicle research, where CRISPR-Cas9 is studied for gene and protein delivery systems in gene therapy applications (J. Lin et al., 2023). Finally, disease-specific studies addressed CRISPR-Cas9-based therapies in neurodegenerative diseases, oncology, and hereditary diseases (He et al., 2025; Wan et al., 2024).
Unlike previously published bibliometric studies, the current study offers a focused and integrated mapping of CRISPR-Cas9 research strictly within the domain of human disease pathology by focusing specifically on how CRISPR-Cas9 is used to model, study, and treat human disease mechanism at the molecular and cellular levels. The current study includes only publications addressing molecular, genetic, or therapeutic mechanisms in human diseases.
Based on above-mentioned explanation, the objective of the current study is to provide a comprehensive bibliometric and knowledge mapping analysis of global research activities focused on application of CRISPR-Cas9 technology in human disease pathology. The current study is significant in that it not only quantifies the research trajectory and global collaboration pattern, but also highlights how gene editing is shifting from experimental models to clinical application, which is strategically informative, horizon-scanning, and translationally valuable.
By synthesizing data-driven indicators with thematic analysis, the study provides a foundational resource for researchers, clinicians, and policymakers to understand emerging hotspots, assess knowledge gaps, and prioritize research directions in precision medicine and therapeutic genome editing. The study holds significant scholarly, strategic, and translational value. By examining the scientific literature in the field, the study aims to uncover publication trends, identify influential contributors, explore thematic clusters, and evaluate the scholarly impact of CRISPR-Cas9 within the biomedical domain. Given the explosive growth and translational breakthroughs of CRISPR-Cas9, the current study offers a critical insight into how the field is evolving and diversifying.
Methods
Study Design
This bibliometric study was designed to systematically map and evaluate the global research landscape surrounding the application of CRISPR-Cas9 technology in human disease pathology. The analysis followed the Guideline for Reporting Bibliometric Reviews (BIBLIO) ensuring transparency, replicability, and methodological rigor throughout the research process (Montazeri et al., 2023). The study aimed to provide insight into publication trends, key contributors, international collaboration, thematic developments, and citation dynamics within this rapidly evolving field.
Data Source
The Scopus database, maintained by Elsevier, was selected as the sole source for data retrieval due to its comprehensive coverage of peer-reviewed biomedical literature, its advanced search capabilities, and its compatibility with bibliometric tools such as VOSviewer (van Eck & Waltman, 2010). Scopus is particularly well-suited for studies of this nature because of its detailed metadata (e.g., keywords, author affiliation, and citations) necessary for constructing accurate bibliometric and visualization maps. Its strong indexing in the field of genetics, molecular biology, medicine, and biotechnology made it ideal for capturing the breadth of CRISPR-Cas9 applications in the context of human disease.
Search Query
A tailored search query was executed on May 01, 2025 to identify relevant publications between 2010 and December 31, 2024. In the current study, human disease pathology refers to the molecular, cellular, and genetic mechanisms underlying the onset, progression, and therapeutic targeting of diseases in humans. This encompasses monogenic genetic diseases (e.g., SCD), oncogenic pathways in cancer, neurological conditions with known genetic etiologies, infectious diseases where CRISPR-Cas9 is applied to disrupt viral genome, and immune related diseases involving gene regulation or editing. The search query was designed to ensure that each retrieved article met three essential criteria: (1) inclusion of CRISPR-Cas9 technology, (2) relevance to human disease, (3) emphasis on molecular, genetic, or therapeutic mechanisms. To achieve this, the search strategy included CRISPR-Cas9 terms in the title/abstract field while an extensive list of terms related to human diseases was required to appear in the title field to restrict the focus to human disease pathology. The search query included more than 50 different terms directly related to specific diseases or terms related to human diseases such as “cardiovascular diseases” or “inherited diseases.” The terms used in the search query included the following:
Inclusion and Exclusion Criteria
We included (1) peer-reviewed original research articles in English that apply CRISPR-Cas9 to study, model, or therapeutically modify molecular/cellular mechanisms of human disease; (2) involve gene correction/knockout/regulation/screening in human disease contexts; and/or (3) clinical trials, human cell lines, animal models, or pre-clinical validation targeting disease-related genes/pathways. The following studies were excluded using the Boolean “AND NOT”:
(1) plant or animal CRISPR-Cas9 studies; (2) basic CRISPR-Cas9 technology development not linked to disease application; and (3) studies using CRISPR-Cas9 solely as a gene expression tool without disease focus.
Validation of the Search Query
To validate the robustness and accuracy of the search strategy, several steps were undertaken. A manual screening of the most cited articles (n = 30) confirmed their relevance to CRISPR-Cas9 technology in human diseases. The most prolific authors, journals, and institutions were checked to ensure alignment with the biomedical CRISPR-Cas9 gene editing. These measures confirmed that the search query successfully captured literature targeting human disease pathology and excluded unrelated content, ensuring the credibility of the dataset used in the bibliometric and knowledge mapping analysis. The inclusion of disease specific terms in the “title” field restricted the dataset to studies directly addressing human disease pathology. This ensured relevance to therapeutic or mechanistic research in human diseases. This filtering strategy ensured that the final dataset reflected CRISPR-Cas9 research with direct relevance to understanding, modeling, or treating human disease pathology at the molecular or cellular level.
Data Export and Analysis
All bibliographic metadata were exported in CSV format. The exported fields include research title, abstract, keywords, author names, affiliations, citation count, journal name, publication year, and funding acknowledgments. These data were used to compute standard bibliometric indicators such as average annual growth rate, leading authors, prolific countries, institutions, and journals. In addition, the most highly cited articles were also presented. Citation metrics such as citation per article and the H-index of the retrieved articles were also presented. The value of H-index is an indication of the impact of publications in the field. The H-index is defined as the “highest number of publications (e.g., H) that have received at least H number of citations” (Hirsch, 2005).
VOSviewer Mapping
For visual and network analysis, the dataset was processed using VOSviewer (version 1.6.20), a specialized software for creating bibliometric maps (Arruda et al., 2022; Bukar et al., 2023). Several types of network visualization maps were created. Keyword co-occurrence maps were created to highlight dominant research topics in the retrieved articles. Additionally, a co-occurrence map of terms in the titles/abstracts was developed to visualize the conceptual and intellectual structure of the field. In VOSviewer, node size reflects frequency (occurrence/weight), edge thickness indicates link strength, and shorter distances denote stronger relatedness; clusters (colors) represent densely connected themes (Wong, 2018). Overlay visualizations color nodes by average publication year, while density maps emphasize local concentration of weighted items.
Glossary
A brief glossary of key bibliometric and clinical terms is provided in Supplemental Appendix 1.
Results
Flow Diagram
The flow diagram (Figure 1) showed that the net total number of original research articles retrieved through the search strategy was 4,313. Of the 24,700 articles on CRISPR-Cas9, approximately 17.5% (n = 4,313 articles) were dedicated for human disease pathology.

Flow diagram of the number of articles retrieved through search strategy.
Growth of Publications
The annual growth of publications in the field (Figure 2) showed a rapid rise between 2014 and 2022 followed by a slight decline (2023–2024). The peak number of publications was achieved in 2022 (n = 654 articles). The AAGR during the study period was 54.5%. While the growth of publications showed an S-shaped pattern, the growth of citations showed a steep upward linear pattern with an AAGR of 79.0%, outpacing the growth of publications over time.

Growth of publications and citations for CRISPR-Cas9 in human disease pathology (2010–2024).
After 2019 to 2022’s rapid expansion, the modest plateau in 2023 to 2024 likely reflects maturation into regulated translation rather than fatigue in basic research (N. K. Lee & Chang, 2024).
Most Prolific Journals
The top 5 most prolific journals pointed that Scientific Reports leads with 155 publications (3.6%), reflecting its multidisciplinary nature, open access, and rapid publications. The International Journal of Molecular Sciences (n = 128 publications; 3.0%) ranked second by offering a venue for studies emphasizing molecular mechanisms and experimental models for disease pathology. Nature Communications (n = 97 publications; 2.2%) ranked third and represents a high impact venue. The presence of this journal indicates that a significant portion of the retrieved articles meets the standards of novelty and showed adequate mechanistic depth to be published in this high impact journal. Table 1 shows the top 5 prolific journals with their SJR value.
Top 5 Prolific Journals in Publishing Research Articles on CRISPR-Cas9 in Human Disease Pathology.

Scimago Journal Ranking; obtained from scimagojr.com.
IF = impact factor, obtained from JCR published by Clarivate.
Most Prolific Countries and International Research Collaboration
The analysis of authorship and collaboration patterns among the top 5 most prolific countries in CRISPR-Cas9 research related to human disease pathology reveals notable differences in both research output and the structure of scientific collaboration. Among the 4,313 articles examined, the United States (US) emerged as the leading contributor, accounting for 1,727 publications (40.0% of total output). Strikingly, US-based research was almost evenly split between intra-national (n = 847 articles, 49.1%) and international collaborations (880 articles, 50.9%), reflecting a balanced model of domestic capacity and global scientific engagement. China, with 1,355 publications (31.4%), ranks second in overall output. However, it demonstrates a contrasting collaboration profile, with the majority of its work (889 articles, 65.6%) conducted within national institutions and a relatively smaller share (466 articles, 34.4%) involving international partners. Germany, by comparison, produced 403 publications (9.3%) but showed a clear preference for international engagement. Table 2 shows the top 5 prolific countries and the extent of international research collaboration for each country. At the institutional level, most prolific institution is Ministry of Education of the People’s Republic of China (n = 174; 4.0%), followed by Harvard Medical School with 164 publications, representing 3.8% of the research output. This is followed by, the Chinese Academy of Sciences, the Dana-Farber Cancer Institute (United States), and Inserm (France).
Top 5 Prolific Countries and Their Research Collaboration In Publishing Research Articles on CRISPR-Cas9 in Human Disease Pathology.

Most Prolific Institutions
Of the 4,313 retrieved articles, the Ministry of Education of the People’s Republic of China emerged the most prolific institutions, contributing 174 publications (4.0%). This dominance can be attributed to China’s centralized and vertically integrated research ecosystem, where several top-tier universities and national research laboratories are administratively linked to the Ministry of Education. The aggregation of affiliated output under the Ministry of Education umbrella in Scopus indexing database further amplifies its apparent productivity. This also reflects China’s national strategic investment in biotechnology and high-impact technologies in gene therapy. Harvard Medical School ranked second with 164 publications (3.8%). The presence of Harvard Medical School exemplifies the clinical translational model in the US. Table 3 represents the top prolific institutions in the field. The institutions represent geographically diverse centers of excellence. In the prolific list, the Dana-Farber Cancer Institute represents the only specialized institute in which CRISPR-Cas9 research is being carried out in the context of oncology.
Top 5 Prolific Institutions in Publishing Research Articles on CRISPR-Cas9 in Human Disease Pathology.

Authorship Analysis
Authorship analysis indicated that a total of 46,629 author names appeared across the 4,313 retrieved articles, yielding an average of approximately 10.8 authors per article, a median of 9 authors. Approximately 88% (n = 3,795 articles) involved five or more authors underscoring the multidisciplinary and inter-institutional collaboration in the field. Eric N. Olson, Rhonda Bassel-Duby, and Kamel Khalili are among the leading scientists in the field. Olson focuses on genetic editing for muscular and cardiac diseases, particularly Duchenne muscular dystrophy. Bassel-Duby, in collaboration with Olson, investigates the molecular mechanism of muscle development and the correction of mutations in skeletal and cardiac muscles using CRISPR-Cas9. Khalili has pioneered the use of CRISPR-Cas9 technology to eliminate cells infected with HIV.
Citation Metrics
A total of 135,970 citations were accrued by the 4,313 articles included in this bibliometric of CRISPR-Cas9 research applied to human disease pathology. The mean number of citations per article was 31.5, indicating a moderate to high average impact across the dataset. However, the median citation count was notably lower (median = 10), with an interquartile range (IQR) of 27, suggesting a positively skewed distribution. This pattern is characteristic of emerging and high-impact scientific fields where a small subset of publications garners disproportionately high citations, while many others contribute to the field with modest recognition. The citation range spanned from 0 to 1,185 citations, underscoring significant variability in influence across studies. The H-index of the retrieved articles on CRISPR-Cas9 in human disease pathology was 146. The average annual growth rate (AAGR) in citation frequency was estimated at 79% over the 2013 to 2024 period, reflecting rapid and accelerating scholarly engagement with CRISPR-Cas9 technologies. This steep growth trajectory parallels the increasing clinical relevance and global research investment in gene editing, particularly as preclinical models transition toward therapeutic application.
Top Five Cited Articles
Among the most highly cited studies in CRISPR-Cas9 research applied to human disease, several landmark publications stand out for their translational significance and impact (Table 4). The most cited paper, published by Frangoul et al. in The New England Journal of Medicine (2021), with 1,184 citations, represents the first clinical application of CRISPR-Cas9 in treating beta-thalassemia and SCD by ex vivo editing of the erythroid-specific enhancer of BCL11A (B-Cell Lymphoma/Leukemia) in autologous hematopoietic stem cells (Frangoul et al., 2021). The edited cells restored fetal hemoglobin expression, resulting in transfusion independence and clinical remission. Similarly, Gillmore et al. (2021), also in The New England Journal of Medicine, reported on NTLA-2001, the first systemic in vivo CRISPR-Cas9 therapy for transthyretin amyloidosis (Gillmore et al., 2021). Delivered via lipid nanoparticles, this therapy achieved up to 87% reduction in serum TTR protein levels with minimal adverse effects, underscoring the feasibility of non-viral gene editing in humans. Schwank et al. (2013) provided an early proof-of-concept study in stem cell, demonstrating correction of the CFTR gene in intestinal stem cell-derived organoids from cystic fibrosis patients using CRISPR-Cas9-mediated homologous recombination-paving the way for personalized regenerative medicine approaches (Schwank et al., 2013). Nelson et al. (2016), publishing in Science, advanced the field of neuromuscular gene therapy by demonstrating systemic AAV-mediated delivery of CRISPR-Cas9 to correct the dystrophin gene in a mouse model of Duchenne muscular dystrophy, restoring partial muscle function and biochemical integrity (Nelson et al., 2016). Lastly, Matano et al. (2015) in Nature Medicine showcased the use of CRISPR-Cas9 to introduce and functionally model multiple driver mutations in colorectal cancer using human intestinal organoids, providing mechanistic insight into tumor initiation and progression (Matano et al., 2015). Collectively, these studies exemplify the breadth of CRISPR-Cas9 applications-from monogenic blood and metabolic disorders to complex diseases such as cancer—and mark key milestones in translating genome editing from experimental models to human therapy.
The Top 5 Cited Articles on CRISPR-Cas9 in Human Disease Pathology.

Keyword Co-occurrence Map
The keyword co-occurrence map (Figure 3) of the 100 most frequent author keywords included several dominant central terms such as CRISPR-Cas9, gene editing, and cancer. Keywords such as TP53, apoptosis, proliferation, migration and invasion cluster alongside specific malignancies like colorectal cancer, hepatocellular carcinoma, melanoma, pancreatic cancer, and breast cancer, reflecting a strong focus on the dissection of cancer pathways and metastasis using CRISPR-Cas9-based approaches. For example, a study demonstrated the use of CRISPR-Cas9-mediated VHL gene knockout to model renal cell carcinoma, highlighting the technology’s utility in functional cancer genomics (Schokrpur et al., 2016). A study employed genome-wide CRISPR-Cas9 screens to identify MEST as a metastasis driver in esophageal carcinoma emphasizing its role in identifying novel therapeutic targets (W. W. Xu et al., 2023). Another study reviewed the role of CRISPR-Cas9 in colorectal cancer shedding light on the mechanism of tumorigenesis and chemoresistance (Hu et al., 2023).

Network visualization of keywords co-occurrence map. The top 100 frequent author keywords were visualized.
Another prominent cluster captures research into non-oncological genetic disorders, including Duchenne muscular dystrophy, cystic fibrosis, HIV-1 and Alzheimer’s disease, indicating CRISPR-Cas9’s growing role in therapeutic development for monogenic and neurodegenerative conditions. A study reviewed the early progress and challenges in applying CRISPR-Cas9 to eliminate latent HIV proviruses, emphasizing its curative potential (Xiao et al., 2019). Another study validated this approach by targeting HIV-1 tat/rev genes, which resulted in a significant suppression of viral replication in infected T-cells (Ophinni et al., 2018). Most recently, a study demonstrated that guide RNAs targeting the TAR region of HIV achieved >95% inhibition in dual reporter systems, reinforcing CRISPR-Cas9’s therapeutic efficacy (Allen et al., 2023). Parallel breakthroughs have occurred in the correction of mutation driven genetic disorders, most notably cystic fibrosis (CF). A recent review of 40 studies on the use of CRISPR-Cas9 to correct CFTR mutations in CF was published (Harris & Kittur, 2025). A study expanded on this by demonstrating functional restoration of CFTR activity in patient derived intestinal stem cell organoids following precise gene correction (Li et al., 2023). Earlier work by Schwank et al. provided proof-of-concept for CRISPR-Cas9-mediated homologous recombination in CFTR laying the foundation for patient-specific regenerative therapies (G. Wang, 2023). Hematological disorders represent another fertile for CRISPR-Cas9 research translation. Frangoul et al. reported the first clinical success using CRISPR-Cas9 to disrupt BCL11A enhancer in CD34+ hematopoietic stem cells, restoring fetal hemoglobin and eliminating transfusion dependence in patients with SCD and beta-thalassemia (Ahmed et al., 2025; Frangoul et al., 2021).
Finally, a portion of the literature addresses advancements in gene editing technologies and delivery systems, with terms like Cas9, base editing, mRNA editing, and adeno-associated virus reflecting continued innovation in precision genome engineering.
Co-Occurrence Map of Terms in Titles and Abstracts
Figure 4 visualizes the frequency and co-occurrence relationships between terms that appear in the retrieved articles. The total number of terms in the map was 1,498 terms. The map included four clusters of related terms that represent four research themes.
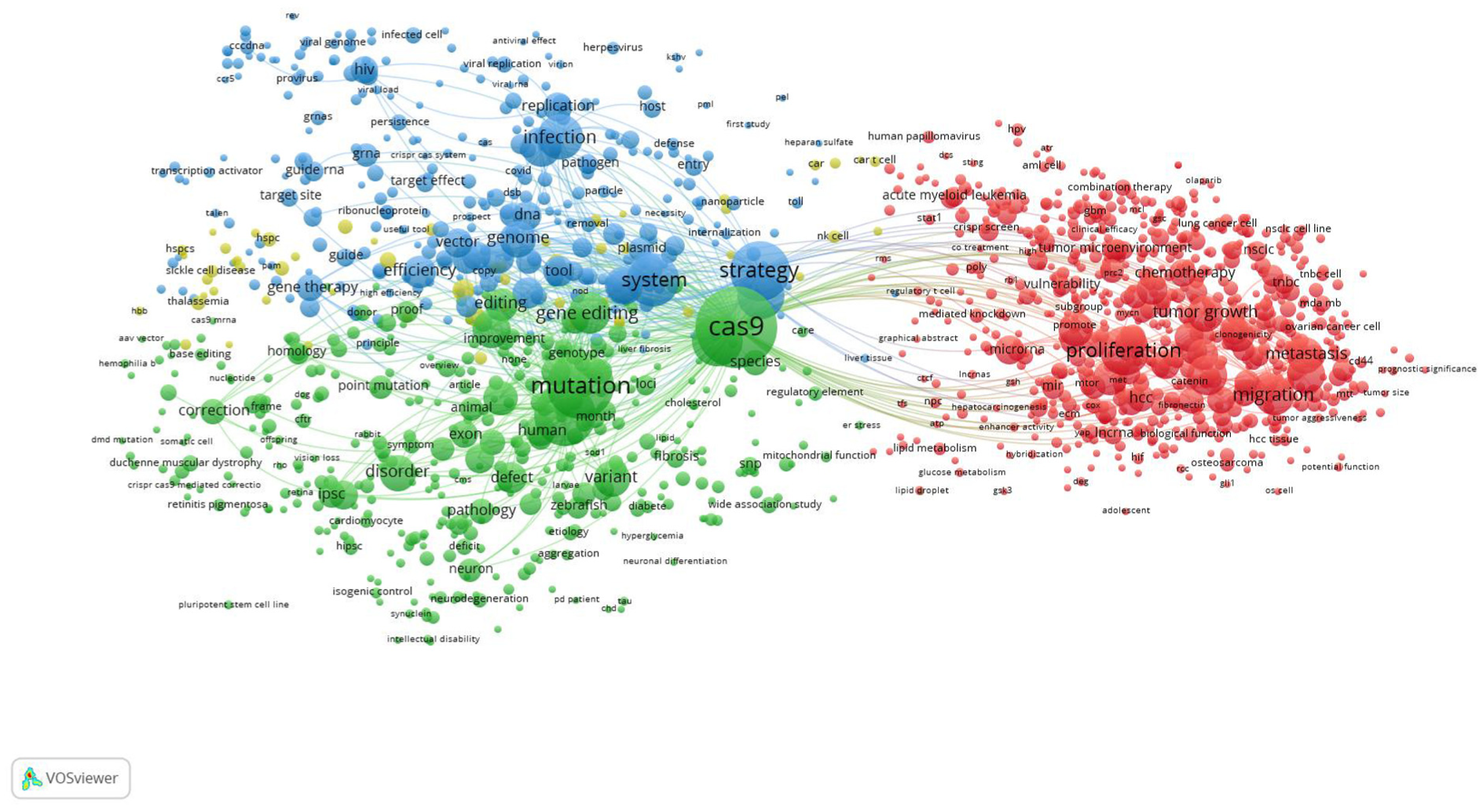
Network visualization map of title/abstract term co-occurrence map. Terms with a minimum occurrence of 10 times were included in the map (n = 1,498).
The Red Cluster
The red cluster centers on the application of CRISPR-Cas9 technology to unravel the molecular mechanisms underlying cancer development, progression, and therapeutic response within the broader context of human disease pathology. Dominated by high-frequency terms such as proliferation, cell proliferation, tumor growth, migration, invasion, metastasis, therapeutic target, tumorigenesis, and drug resistance. This cluster clearly reflects a strong focus on oncological research. The presence of keywords like overexpression, poor prognosis, and chemotherapy suggests that researchers are not only using gene editing tools to dissect cancer biology but also to model and potentially reverse mechanisms of chemoresistance and poor treatment outcomes. Furthermore, disease-specific terms such as colorectal cancer, hepatocellular carcinoma, glioblastoma, and triple-negative breast cancer indicate that this body of research is highly translational, targeting high-burden or therapeutically challenging cancer types.
The Blue Cluster
The blue cluster represents a distinct research theme at the intersection of CRISPR-Cas9 technology and viral infections, with a particular emphasis on HIV, viral replication, and gene therapy for infectious diseases. The most prominent terms in this cluster include infection, virus, HIV, HIV infection, replication, viral vector, cleavage, mutagenesis, immune response, and restriction factor, indicating a thematic focus on the application of CRISPR-Cas9 systems for studying, inhibiting, or editing viral genomes within host cells. The core theme of this cluster revolves around using CRISPR-Cas9 as functional tool to dissect host-virus interactions, disrupt viral life cycles, and develop gene therapy-based antiviral strategies. In particular, the dense connection of terms such HIV, viral replication, integration, guide RNA, cleavage, and target site suggest intensive efforts to utilize genome editing to eradicate latent viral reservoirs, especially in the context of HIV/AIDS cure research. Moreover, the cluster includes references to HBV, herpes virus, retrovirus, and viral latency indicating that the scope of this research is not limited to HIV but extends to other chronic and latent viral infections.
The Green Cluster
The green cluster focused on the application of CRISPR-Cas9 technology to investigate and therapeutically address genetic disorders and mutation-driven human diseases. Central keywords such as mutation, disease, disorder, variant, gene editing, genome editing, Cas9, and point mutation highlight the cluster’s core emphasis on identifying, characterizing, and correcting pathogenic genetic variations. This thematic area is deeply rooted in understanding the molecular basis of hereditary conditions, with frequent references to terms like pathology, defect, dysfunction, and polymorphism that signal a strong interest in linking specific mutations to disease phenotypes. The presence of terms such as animal model, pluripotent stem cell, and human disease indicates the use of a variety of in vivo and in vitro models, including patient-derived induced pluripotent stem cells (iPSCs) and transgenic organisms to simulate human disease mechanisms and test gene-editing strategies. Additionally, disease-specific terms such as amyotrophic lateral sclerosis, cardiomyopathy, obesity, Alzheimer’s disease, and Fabry disease underscore the broad application CRISPR-Cas9-based tools across diverse pathological contexts.
The Yellow Cluster
The yellow cluster focused on hematological genetic disorders and CRISPR-Cas9-based gene therapy. These nodes cluster around key terms including SCD, thalassemia, hematopoietic stem cell (HSPC), fetal hemoglobin, erythroid cell, BCL11A, and gene therapy, highlighting a well-established research trajectory that aims to develop durable curative treatments for monogenic blood disorders. This thematic area reflects the use of CRISPR-Cas9 technology to edit hematopoietic stem and progenitor cells (HSPCs), particularly through the disruption of the BCL11A enhancer, a known transcriptional repressor of fetal hemoglobin (HbF). The presence of terms such as insertion, target site, and nuclease further reinforce the precision-editing strategies employed in this context, emphasizing both ex vivo and in vivo approaches to gene correction.
Discussion
Interpretation of Growth and Key Contributors
The current study aimed to map and analyze research on CRISPR-Cas9 in the context of human disease pathology. The study identified a striking AAGR and an exceptional growth expansion due to multiple converging factors. The growth expansion was initiated by the demonstration made by Jennifer Doudna and Emmanuelle Charpentier in 2012 that CRISPR-Cas9 system could be harnessed for precise genome editing (Doudna & Charpentier, 2014). The simplicity, affordability, and programmability allowed the adoption of CRISPR-Cas9 system in various disciplines including oncology, neurology, molecular genetics, and virology (Gupta et al., 2019). Soon after this, research on CRISPR-Cas9 moved from basic science to clinical application in various disciplines especially genetic disorders (Frangoul et al., 2021; Germino-Watnick et al., 2022; Gillmore et al., 2021). This explains the sharp rise in research output seen between 2015 and 2021. Thematic diversification and global investment further fueled this growth.
After 2019 to 2022’s rapid expansion, the modest plateau in 2023 to 2024 likely reflects maturation into regulated translation, which typically yields fewer, slower publications due to longer study timelines, manufacturing scale-up, and regulatory requirements (N. K. Lee & Chang, 2024). First-in-class clinical and marketing approvals (e.g., exagamglogene autotemcel) required extensive chemistry, manufacturing and controls and long-term safety planning; such activities often displace rapid paper output while expanding clinical evidence bases (Hoy, 2024). Regulators now expect risk-based long-term follow-up (often up to 15 years) for gene therapy/editing, which lengthens programs and can compress near-term publication counts. Additional bottlenecks include delivery limits (AAV/LNP trade-offs), immunogenicity of nucleases, conditioning toxicity for ex vivo HSC editing, data-governance constraints that complicate cross-border studies (e.g., China’s human genetic resources rules; GDPR rules on genetic data), and geopolitics that dampen some US–China collaborations (Kabadi et al., 2024; Kazemian et al., 2022). Together, these factors plausibly explain a short-term plateau without implying reduced field momentum (M. Chen et al., 2019; X. Cheng et al., 2020; Lino et al., 2018; Pineda et al., 2019; F. Zhang et al., 2014).
The current study showed that the Ministry of Education of the People’s Republic of China was the most prolific contributor. The appearance of the MOE as an active institution is due to several administrative and structural factors related to the centralized research model in China, where several top tier universities are affiliated or administered by the MOE (C. Cao et al., 2006). In addition, several national research labs, including ones in molecular biology, are funded and recognized by the MOE. Research output from these research laboratories is aggregated under the MOE in indexing databases.
Interpretation of Citation and Authorship Metrics
The citation metrics reflect not only the immense scientific interest in the field but also its profound impact across disciplines. An H-index of 146 demonstrates that a substantial body of literature has achieved a widespread recognition and scientific utilization. The relatively high H-index is significant because of the short timeframe since the first application of CRISPR-Cas9 to human diseases. Since its repurposing as a genome editing tool in 2012, CRISPR-Cas9 has transitioned from a basic molecular tool to a central component of translational medicine with applications in genetic correction, disease modeling, and clinical therapy. The fact that so many studies crossed the 100-citation threshold is indicative of the rapid maturation and wide adoption of the CRISPR-Cas9 technology across biomedical research sectors. Several key reasons help explain the high H-index of the retrieved literature. First, the CRISPR-Cas9 represents a paradigm shift in molecular medicine compared to earlier gene editing methods such as ZFNs and TALENs. Second, the pressing need of new methodological treatment for certain diseases such as SCD, thalassemia, HIV, and hepatitis. Results of CRISPR-Cas9 clinical trials in the field of monogenic disorders have been published in highly prestigious academic scientific journal. Third, the cross-disciplinary nature of the field broadened the citation base by attracting citations by researchers in the field of genetics and researchers from adjacent areas related to various diseases. In comparison to other fields in biomedicine, such as infectious diseases (Sweileh et al., 2017), the H-index of the CRISPR-Cas9 in human disease pathology is striking, making the field among the most influential in the 21st century.
The authorship metrics indicated a high mean number of authors per article reflecting a striking high level of team-based science. The high level of co-authorship can be rationalized by the nature of CRISPR-Cas9 research, which is complex and methodologically diverse. Research in this area often involve multiple stages that require expertise in specific fields that necessitate research collaboration and team planning. The high level of collaboration in the field was also noticed in similar translational fields such as CAR-T cell therapy (Huang et al., 2024; Miao et al., 2022).
Interpretation of Research Collaboration
The varying levels of international research collaboration between the US, China, Europe, and Japan reflect deeper structural, cultural, and geopolitical dynamics (Arakawa & Anderson, 2020; Liu et al., 2023; Pinho & Reeves, 2021). Observed US–China–Europe hubs reflect funding scale and institutional density, but collaboration frictions persist. Data-sovereignty and security rules (e.g., China’s Human Genetic Resources measures; EU GDPR Article 9 on genetic data) complicate cross-border data sharing and ethics approvals (J. Chen & Li, 2025; Clunie et al., 2025). Language and editorial norms also matter: English dominance in high-impact journals can dampen participation from non-English teams and may under-index regional outputs (B. Yao, 2021). Periodic policy shifts (export controls, visa regimes, security reviews) further modulate co-authorship networks. These structural factors help explain cluster boundaries and asymmetries in the co-authorship maps
The United States exhibits a high level of international research collaboration due to its longstanding leadership in biomedical research. The research environment in the U.S is highly diverse and inclusive, with significant portion of its scientific workforce comprising foreign-born researchers. This diversity naturally fosters cross-border collaboration. The U.S. funding agencies also support international partnership, especially in translational projects. In contrast, China demonstrated relatively low levels of international collaboration. This is highly due to its inward-focused search strategy that emphasizes self-reliance and national innovation (Zhou & Leydesdorff, 2006). Furthermore, language barriers, differing research practices, concerns about data sharing, intellectual property and geopolitical tension with western countries contribute to the gap in the extent of international research collaboration (Matthews et al., 2020; J. L. Tang, 2010). European countries, on the other hand, show remarkably high level of international research collaboration. This is driven by the structural integration of research across the continent, facilitated by EU-funded initiatives, which require international consortia.
Interpretation of Highly Cited Articles
The top 5 cited articles reveal certain common elements that help their exceptional scholarly impact. First and foremost, the top 5 cited articles were published in high impact and reputable journals known for their rigorous peer-review, increasing the likelihood of dissemination, visibility and citations. Second, the top cited articles introduced first-in-field advancements that fundamentally shifted the direction of CRISPR-Cas9 research. Whether it was demonstrating the feasibility of systemic in vivo gene editing in humans, correcting disease-causing mutations in patient-derived organoids, or modeling complex cancer mutations using genome editing tools, each study of the top cited ones broke new ground and set a new precedent for future work in its respective domain. A third key factor for contributing to their citation success lies in their direct clinical relevance. These studies addressed high priority, burdensome diseases, including SCD, CF, thalassemia, transthyretin amyloidosis, Duchenne muscular dystrophy, colorectal cancer, for which effective treatments have long been lacking. By offering not just mechanistic insight, but tangible therapeutic strategies, the top cited studies resonated deeply with both basic researchers and translational scientists seeking to bridge the gap between bench and bedside. Finally, the top cited articles reflect broad, cross-disciplinary appeal of CRISPR-Cas9 technology. By intersecting with fields such as hematology, molecular genetics, oncology, regenerative medicine, and virology, these studies attracted citations from a wide spectrum of disciplines. This multidimensional relevance amplified their visibility and positioned them as cornerstone publications in the rapidly evolving landscape of genome editing.
Interpretation of Research Themes
The prominence of the specific research themes identified in the current study, namely cancer biology/pathology, viral infections, genetic disorders, and hematological diseases, can be attributed to both scientific opportunity and clinical urgency, shaping the trajectory of CRISPR-Cas9 research in human disease pathology. Firstly, oncology emerged as a dominant theme due to the complexity, prevalence, and genetic underpinnings of various cancers. CRISPR-Cas9 offers a powerful platform for dissecting oncogenic pathways (e.g., TP53, KRAS), modeling tumor progression, and overcoming therapy resistance. The inclusion of terms such as proliferation, migration, metastasis, and chemoresistance reflects this focus. Cancer’s high global burden, combined with unmet therapeutic needs in aggressive types such as glioblastoma and triple negative breast cancer makes it a natural area for CRISPR-Cas9 application (Al-Sammarraie & Ray, 2021; Begagić et al., 2024; Deepak Singh et al., 2021; Tiwari et al., 2023). Secondly, viral infections, specifically HIV, constituted the second largest thematic cluster because of the longstanding quest for an effective cure for HIV infection. CRISPR-Cas9’s ability to disrupt latent viral genomes and modulate host-pathogen interactions fuels intense research interest, especially in chronic and incurable infections like HIV and hepatitis B (De Silva Feelixge et al., 2018; H. Lin et al., 2021; Salimi-Jeda et al., 2022; White et al., 2015; Z. Q. Yao et al., 2024). The high density of terms such as replication, cleavage, and latency reflect this translational intent with potential to revolutionize antiviral therapy. The third cluster, the genetic disorder theme, encompassing cystic fibrosis and Duchenne muscular dystrophy, reflects the core promise of CRISPR-Cas9: precision correction of disease-causing mutations. These diseases often have well-characterized genetic causes and lack curative options, positioning CRISPR-Cas9 as a beacon of hope for durable gene correction therapies. The fourth and final research cluster represents hematological disorders like SCD and thalassemia. These diseases were presented in a separate cluster, research theme, due to recent clinical successes and the accessibility of hematopoietic stem cells for ex vivo editing (Y. Chen et al., 2022; Dever et al., 2016; Frangoul et al., 2021; Ma et al., 2023; Park & Bao, 2021; Ugalde et al., 2023). The disruption of the BCL11A enhancer to reactivate fetal hemoglobin, for instance, showcases a clear therapeutic mechanism with measurable outcomes, making this an attractive and clinically validated application for CRISPR-Cas9.
Existing Challenges
Despite the exponential growth in CRISPR-Cas9 research within human disease pathology, the field continues to face technical and ethical challenges. One of the primary challenges lies in the potential for off-target effects, which pose a threat to genomic editing and therapeutic safety. Several studies have demonstrated unintended genomic alterations due to insufficient guide RNA specificity, which may lead to deleterious consequences in clinical applications (Doench et al., 2016). Beyond technical risks, CRISPR-Cas9 raises material ethical and governance questions. International expert groups and the 3rd International Summit on Human Genome Editing (London, March 2023) concluded that heritable (germline) editing “is not appropriate at this time,” emphasizing transparency, public engagement, and equitable access for somatic applications (National Academies of Sciences, E, 2023). The WHO’s 2021 governance framework urges national oversight systems, a global registry, and proportionate risk governance to reduce inequities between jurisdictions (World Health Organization, 2021). In Europe, the Council of Europe’s Oviedo Convention (Article 13) prohibits interventions aimed at introducing heritable genomic modifications; the EU GDPR also treats genetic data as a “special category,” with added restrictions and heterogeneous Member State implementations (Vidalis, 2022). In contrast, China has tightened rules on cross-border use of human genetic resources, requiring approvals that can slow or re-route international projects (J. Chen & Li, 2025). These asymmetries, while ethically motivated, create uneven pathways for clinical translation and data sharing, reinforcing the need for harmonized standards focused on safety, consent, and justice.
Clinically, delivery remains the gating factor. AAV vectors offer durable in vivo delivery but face payload limits and pre-existing immunity; LNPs enable transient, re-dosable delivery but require tissue-specific formulations (Severi & Akbari, 2024). Recent advances include engineered AAV capsids with improved tropism, next-gen LNP chemistries, and virus-like particle systems that package Cas9 RNPs, achieving AAV-comparable editing in preclinical models while avoiding vector genomes (Kazemian et al., 2022). In parallel, AI-guided guide/pegRNA design (e.g., Azimuth/Elevation; DeepCRISPR; newer prime-editing models reduces off-target risk and improves efficiency, directly informing clinical construct selection (D. Cao et al., 2025; Ling et al., 2025; J. Wang et al., 2020). Together, delivery and design innovations are likely to shift CRISPR-Cas9 therapies toward in vivo applications and multi-tissue indications over the next cycle.
Future Perspectives
Looking ahead, the CRISPR-Cas9 field is expected to evolve toward enhanced precision, safety, and clinical utility. Emerging innovations such as base-editing and prime editing offer promise by enabling single nucleotide alterations without inducing double-stranded breaks, thereby minimizing off-target risk (Anzalone et al., 2019). In parallel, the development of delivery platforms such as lipid nanoparticles and viral vectors specifically tailored for targeted tissues is anticipated to expand the therapeutic window of CRISPR-Cas9 application (Q. Cheng et al., 2020; Saw et al., 2022; Yan et al., 2021). Artificial intelligence and machine learning are also beginning to play a pivotal role in improving guide RNA design and predicting editing outcomes with greater fidelity (Bhat et al., 2022; Khammampalli & Vindal, 2025). Additionally, the integration of CRISPR-Cas9 with single-cell sequencing technologies and organoid models is projected to open new frontiers in disease modeling and personalized medicine (Adamson et al., 2016; Datlinger et al., 2017; Liang et al., 2023). These advancements will likely catalyze the translation of CRISPR-Cas9-based approaches from bench to bedside.
Managerial Implications
From a managerial standpoint, the findings of the current bibliometric analysis provide actionable insights for academic leaders, research institutions, and policy-makers seeking to optimize their strategic investment in CRISPR-Cas9 research. Institutions can leverage the identified collaboration patterns and prolific author networks to establish targeted partnerships with high impact research clusters, thereby enhancing knowledge exchange and research productivity. Furthermore, understanding trends and institutional output can help funding agencies allocate resources more effectively to regions with emerging expertise or infrastructure gaps. For journal editors and scientific publishers, recognizing citation hotspots, and influential topics may inform editorial strategies and special issue planning. Finally, the evident dominance of certain countries in the publication landscape calls for initiatives that promote inclusivity, such as supporting CRISPR-Cas9 capacity building in low- and middle-income countries to bridge the global scientific divide.
Limitations
Despite the manuscript analytical depth, it acknowledges several limitations. Our reliance on Scopus and a time horizon ending in 2024 may bias coverage and omit fast-moving developments (e.g., late-2024/2025 regulatory decisions, late-stage trials, and gray literature). Comparative studies show Scopus and Web of Science differ in journal/language coverage and indexing cadence; PubMed provides rapid biomedical updates but narrower source types. Consequently, counts, collaboration rates, and citation indicators may shift with database selection (Falagas et al., 2008; Mongeon & Paul-Hus, 2016). Future updates should triangulate Scopus with Web of Science, PubMed, and (optionally) Dimensions, using harmonized de-duplication and sensitivity analyses to quantify database effects, and adopt a living-bibliography workflow to incorporate post-2024 clinical/regulatory milestones.
Conclusions
In conclusion, the present bibliometric review elucidates the expansive and rapidly evolving landscape of CRISPR-Cas9 research in human diseases. From its initial demonstration as a programmable nuclease in 2012 to its clinical deployment in diseases like SCD and transthyretin amyloidosis, CRISPR-Cas9 has transitioned from a disruptive molecular tool to a therapeutic reality. The study reveals a steep growth in publications and an even more pronounced citation growth reflecting both innovation velocity and scholarly impact. The identified research thematic clusters include: (1) oncological research, (2) infectious diseases especially HIV, (3) monogenic genetic disorders, and (4) hematological gene therapy. The US leads the field in both output and international collaboration, contrasting with China’s strong domestic research base. Citation metrics, authorship patterns, and journal quality further illustrate the field’s maturity and interdisciplinarity. However, translational challenges including in vivo delivery, immunogenicity, and ethical concerns remain critical. Future research must refine editing precision, develop next-generation delivery systems, and integrate CRISPR-Cas9 with emerging biotechnologies to realize its full therapeutic potential. Ultimately, CRISPR-Cas9 stands not merely as a gene editing tool but as a cornerstone of next-generation medicine.
Supplemental Material
sj-docx-1-sgo-10.1177_21582440261415669 – Supplemental material for Bibliometric Insights into CRISPR-Cas9 Research Targeting Human Disease Pathology
Supplemental material, sj-docx-1-sgo-10.1177_21582440261415669 for Bibliometric Insights into CRISPR-Cas9 Research Targeting Human Disease Pathology by Moutaz W. Sweileh in SAGE Open
Footnotes
Acknowledgements
The authors acknowledge the use of AI in language editing only. No results or data were obtained or generated by any type of AI.
Ethical Considerations
No IRB is required for the current study since all data are obtained from Scopus database and patients or human subjects are involved directly in the study.
Author Contributions
M.S. initiated the idea, developed the search query, and helped in the interpretation of the results. W.S analyzed the data and wrote the manuscript.
Funding
The author received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
All data, including the search query, are available and can be provided upon request from the authors.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
