Abstract
Background
Skin lesion segmentation plays a critical role in computer-aided diagnosis systems, serving as a foundation for the early detection and treatment of skin cancer. Nonetheless, obtaining accurate segmentation remains difficult because of inconsistencies in lesion visual features, texture, image sharpness, and the presence of indistinct edges.
Objective
To develop and evaluate a novel deep neural network (DNN)-based approach for robust and accurate segmentation of skin lesions from dermoscopic images using advanced pre-processing and post-processing techniques.
Methods
The proposed method integrates a DNN architecture with specialized pre-processing and post-processing modules. The pre-processing step enhances image quality by denoising and normalizing the lesion intensities. The DNN framework extracts hierarchical features, while the post-processing module refines segmentation masks by correcting boundary irregularities and removing artifacts. The model was tested using three widely recognized dermoscopic International Skin Imaging Collaboration (ISIC) image databases from the years 2016, 2017, and 2018 without extensive data augmentation. Statistical analysis, including the Wilcoxon signed-rank test, was conducted to compare performance with existing methods.
Results
The proposed method achieved Jaccard index scores of
Conclusions
This study presents a high-performing, scalable solution for automated skin lesion segmentation. The proposed method effectively addresses critical challenges by integrating robust feature extraction and boundary refinement, making it well-suited for real-world clinical applications in skin cancer diagnosis and management.
Keywords
Introduction
Skin diseases are a significant global health challenge, with melanoma, a type of skin cancer, recognized as among the most severe and dangerous variants. Based on data from the American Cancer Society 1 and the National Cancer Institute, 2 it was projected that approximately 1.9 million cancer cases would be identified in the United States, equating to roughly 5,250 diagnoses per day. Of these, skin cancer was estimated to account for around 108,000 new cases and nearly 12,000 deaths. 3 On a global scale, each year sees an estimated 2 to 3 million occurrences of non-melanoma skin cancer, along with about 132,000 new diagnoses of melanoma. 4
Skin cancers are broadly classified into two categories: melanoma and non-melanoma. Non-melanoma skin cancers, which include squamous cell carcinoma and basal cell carcinoma, are generally less aggressive and carry a lower mortality risk. Despite this, they represent a substantial healthcare burden due to their high prevalence. Melanoma, on the other hand, is notorious for its ability to metastasize rapidly, making early detection and treatment critical for patient survival. 5
The rising incidence of skin cancers can be attributed to factors such as increased ultraviolet radiation exposure, aging populations, and heightened public awareness, leading to better detection rates. 6 However, this trend also highlights the urgent need for innovative diagnostic tools, preventive strategies, and targeted therapies to combat the growing burden of skin cancer. 7 Non-melanoma skin cancers, although less fatal compared to melanoma, often necessitate invasive treatment procedures that can be both painful and distressing 8 for patients. By comparison, cutaneous melanoma represents an exceptionally aggressive as well as a malignant type of skin cancer, associated with elevated mortality rates due to its potential for rapid progression and metastasis. The early detection and timely intervention of skin cancer are critical for improving patient outcomes and survival rates. 9 However, diagnostic practices that rely solely on visual inspection by dermatologists are prone to subjectivity, leading to inconsistencies in evaluations even among seasoned experts. This subjectivity underscores the pressing need for advanced diagnostic methods that can provide accurate and reproducible results. 8
Automated segmentation techniques have emerged as a pivotal solution in addressing these diagnostic challenges. These methods enable precise and objective analysis of skin lesions, which is essential for enhancing the consistency and reliability of diagnostic processes. In computer-aided diagnosis (CAD) systems, precise segmentation of cutaneous abnormalities serves as an essential foundational step to enable efficient disease diagnosis and therapeutic planning. However, the segmentation of dermoscopic images presents unique challenges due to variations in lesion appearance, including diverse chromatic characteristics and interfering elements such as body hair, ruler marks, and ink stains. These complexities demand sophisticated algorithms capable of navigating the intricate nature of skin lesions and delivering accurate segmentation outcomes. 10
Dermoscopy has become a cornerstone in dermatological diagnostics, offering a non-invasive method for examining skin lesions with enhanced clarity. By employing optical magnification and specialized lighting, dermoscopy allows for detailed visualization of subsurface structures, facilitating the identification and analysis of pigmented lesions. Despite its advantages, dermoscopic images often exhibit irregular and poorly defined lesion boundaries, making accurate delineation difficult. 11
Moreover, the subtle contrasts between lesions and surrounding healthy skin, coupled with the irregular shapes and color patterns of lesions, further complicate the segmentation process. Artifacts such as hair strands, blood vessels, and measurement markings introduce additional challenges, necessitating the development of robust segmentation methods. To overcome these obstacles, recent advancements in artificial intelligence (AI) and deep learning have been leveraged to create innovative segmentation algorithms. These techniques offer promising capabilities for addressing the diverse challenges associated with dermoscopic image segmentation. By integrating these advanced methods into CAD systems, healthcare practitioners can achieve more accurate and reliable diagnoses, ultimately improving the efficacy of skin cancer detection and treatment strategies. Conventional image segmentation techniques often depend on manually designed features, which can struggle to perform effectively when applied to complex dermoscopic images. These features require specialized domain knowledge and often fail to adapt to the diverse variations in lesion appearances. Our proposed methodology addresses these limitations by combining traditional enhancement methods with advanced automated approaches. The pre-processing module improves image quality by reducing noise and managing contrast variability through morphological operations and Wiener filtering, ensuring that essential features are emphasized for segmentation. The post-processing module uses a deep neural network (DNN)-based approach supported by Otsu’s thresholding for accurate binarization. This integrated approach combines the strengths of manual enhancements and automated learning, providing robust segmentation results suitable for clinical application, where reliability and precision are paramount.
U-shaped Convolutional Neural Network (U-Net) is a widely adopted architecture for medical image segmentation, particularly recognized for its encoder–decoder design with skip connections that capture intricate details effectively. Over time, several variants such as Attention U-Net, Recurrent Residual U-Net, and others have been introduced to address specific segmentation tasks. While these models have shown promising results, their reliance on large numbers of trainable parameters often leads to redundancy and inefficiency. 12
Advanced techniques, including DAGAN with dual adversarial discriminators, attention-scale aggregation network (AS-Net) with spatial and channel attention, and feature adaptive transformer network (FAT-Net) utilizing transformer-based encoders, have enhanced segmentation accuracy (Acc) and contextual understanding. However, these approaches face challenges with computational demands and limited generalization, especially when applied to complex datasets such as International Skin Imaging Collaboration (ISIC) 2017. To address these limitations, we propose a hybrid methodology that integrates traditional pre-processing techniques with modern neural network advancements. The pre-processing module improves image quality by reducing noise and enhancing contrast through morphological operations and Wiener filtering, making the images more suitable for segmentation. The post-processing module employs a DNN-based Otsu’s thresholding, which automates the binarization process for lesion extraction.
Our methodology effectively balances parameter complexity and segmentation performance, avoiding issues such as overfitting. This balance enables the model to achieve robust and accurate results. The comparative analysis of the Jaccard metric alongside precision against the total count of adjustable model weights underscores the advantages of our approach.
Figure 1 illustrates the tradeoff between segmentation performance and model complexity by comparing the Jaccard index (JI) against the number of trainable parameters for various state-of-the-art (SOTA) models. The proposed method achieves the highest Jaccard score while maintaining a significantly lower parameter count. This highlights the efficiency of the hybrid framework in achieving precise segmentation without the overhead of over-parameterized architectures.

Jaccard index versus number of parameters: This plot compares the segmentation accuracy (Jaccard index, %) of various models with respect to the number of trainable parameters (in millions). The proposed model achieves top performance with significantly fewer parameters, demonstrating superior efficiency.
Unlike models such as DAGAN and FAT-Net, which require tens of millions of parameters to achieve moderate performance, the proposed design leverages classical image enhancement steps alongside a compact DNN to deliver competitive results. The lower parameter burden not only improves inference speed but also reduces memory consumption, making the framework more adaptable for real-time or resource-limited clinical applications.
These observations underscore the method’s balance between performance and computational efficiency, reinforcing its value for practical deployment.
The contributions of the proposed method are as follows:
The hybrid approach integrates traditional pre-processing with DNN-based post-processing. In the pre-processing stage, noise is minimized, and contrast is enhanced through morphological operations and Wiener filtering. The post-processing stage uses DNN-based coherence filtering to refine lesion boundaries and applies Otsu’s thresholding for accurate binary segmentation. The method achieves a balance between parameter complexity and segmentation performance, avoiding redundancy and overfitting. Comparative evaluations using the JI show that the proposed method outperforms SOTA models, such as DAGAN and FAT-Net, achieving better Acc with fewer trainable parameters. Precision evaluations across various models demonstrate the method’s consistent ability to accurately segment lesions, even with a reduced parameter count. This efficiency ensures its suitability for real-world scenarios with limited computational resources. The proposed methodology exhibits strong generalization across multiple datasets, including ISIC 2016, ISIC 2017, ISIC 2018, and Pedro Hispano Hospital dataset (PH2). Notably, the model achieves high performance without relying on data augmentation, highlighting its robustness and adaptability for diverse clinical applications.
The structure of the paper is as follows: Section “Related work” provides a detailed review of related work, highlighting significant advancements and existing approaches in dermoscopic image segmentation. Section “Proposed method” describes the proposed methodology, presenting the hybrid framework, including the pre-processing module and DNN-based coherence filtering. Section “Skin lesion segmentation algorithm” elaborates on the experimental setup, detailing the datasets, evaluation metrics, and implementation specifics. Section “Dataset description and performance evaluation” presents the experimental results, showcasing the performance of the proposed method on multiple publicly available datasets. Section “Results and analysis” discusses the findings, including an analysis of segmentation performance, processing speed, along with limitations associated with the proposed methodology. Finally, section “Ablation study” concludes the study, summarizing the key contributions and outcomes while offering insights for future research directions.
Related work
In the past, traditional methods for segmenting skin lesions relied on designing handcrafted features to identify distinct patterns in images. These features were developed to separate skin lesions from surrounding tissues, often using histogram thresholding algorithms to establish intensity-based thresholds. Although such approaches provided foundational capabilities by recognizing intensity variations, they were heavily reliant on domain expertise and struggled to generalize across datasets with varying lesion appearances.13,14
Recent advancements in neural networks have significantly enhanced the efficiency and robustness of segmentation tasks. Convolutional neural networks (CNNs), for instance, have demonstrated the ability to autonomously extract meaningful features from datasets, thus improving segmentation performance for skin lesions. 15 Unlike traditional approaches, CNN-based methods eliminate dependency on manual feature crafting, making them more adaptable and precise.16,17
Maji et al. 18 proposed a generator architecture tailored to improve feature learning in decoding layers through leveraging multiple loss functions. This approach produces feature maps with higher semantic value and precision. Furthermore, the integration of attention gates allows selective processing of critical lesion regions, further improving segmentation Acc.
To improve skip connections, researchers have introduced various enhancements to refine feature representation. Spatial enhancement modules within skip connections enable networks to capture and utilize spatial details more effectively, improving segmentation Acc. 19 Attention gates have been applied to address semantic ambiguities between encoder and decoder layers. For example, the attention U-Net selectively highlights important encoder features, providing refined guidance during decoding. 20
BCDU-Net enhances segmentation by combining U-Net with BConvLSTM and dense convolutions in skip connections, enabling nonlinear fusion of feature maps and capturing temporal dependencies. However, these additions lead to increased computational complexity, higher memory demands, and a greater risk of overfitting. Cross-scale parallel fusion network (CPFNet) addresses integration challenges by introducing a GPG module embedded within skip pathways to incorporate high-level contextual information along with a hierarchical-aware fusion component for multi-scale feature fusion. Nevertheless, CPFNet is limited by its heavy parameter requirements, which increase computational and storage demands. 21
In Hafhouf et al., 22 the authors presented modifications to the U-Net architecture, applying enhancements to both encoding and decoding pathways. The encoding process integrated 10 convolutional layers from VGG16, a dilated convolutional block, and pyramid pooling, preserving spatial resolution and improving feature reliability. The decoding pathway was enhanced with dilated residual blocks, which improved the extraction of intricate features and generated more precise segmentation maps.
Recent studies have introduced advanced attention mechanisms and architectural innovations to improve skin lesion segmentation. For example, a self-attention mechanism was employed within the encoder–decoder framework to enhance contextual understanding and segmentation Acc. 23 Similarly, RA-Net applied region-aware attention to better focus on lesion-relevant areas during segmentation. 16 The attention-based dilated residual network (AD-Net) model, also proposed by Naveed et al., 24 integrates dilated convolutional residual blocks with an attention-based spatial feature enhancement block and a guided decoder strategy. This architecture enables robust feature extraction and improves segmentation performance across multiple skin lesion datasets, even without relying on data augmentation. In another approach, contextual feature fusion network (CFF-Net) combined global and local feature representations through a dual-branch encoder that integrates CNN and MLP modules. 25 SUNetDCP focused on efficient feature fusion and model compression to reduce parameter load while maintaining performance. 15 Rolling matrix multi-scale local pattern (RMMLP) utilized adaptive matrix decomposition and rolling tensors to effectively integrate multi-scale contextual features for improved segmentation outcomes. 26
To address generalization limitations, recent studies have turned to transformer-based architectures, such as TransUNet V2 and MedNeXt,27,28 which integrate convolutional backbones with attention mechanisms to capture long-range dependencies. An enhanced multi-scale attention network for skin lesion segmentation was proposed in Wang et al., 17 while a multi-domain transformer segmentation model was introduced by Selvakumar and Sundararaj. 29 These models improve contextual understanding and adaptability across datasets. Vignesh and Rajalakshmi 30 designed a deep attention transformer network optimized for lesion complexity, and Fatima et al. 31 proposed a hybrid classification framework combining self-attention with CNNs.
Additionally, Alhudhaif et al. 14 developed a multipath fusion network with a specialized fusion loss that showed promising results on high-resolution lesion data, demonstrating strong boundary preservation and clarity.
While AD-Net 24 introduces an attention-guided architecture based on dilated residual blocks and an attention-based spatial feature enhancement module, our approach takes a fundamentally different route. Instead of incorporating deep attention mechanisms or guided decoding, we present a lightweight framework that combines classical image enhancement techniques—specifically morphological operations and Wiener filtering—with a streamlined DNN for segmentation. Furthermore, our post-processing stage employs Otsu thresholding and morphological reconstruction to refine lesion boundaries. This hybrid methodology reduces the need for large parameter sets while maintaining competitive segmentation Acc. Unlike AD-Net, which emphasizes architectural complexity to enhance learning, our method prioritizes computational efficiency, making it well-suited for real-world scenarios where resources may be constrained.
Despite these improvements, challenges remain, including high computational costs, memory-intensive architectures, and reliance on large annotated datasets. Over-parameterized models often face redundancy and generalization issues, especially in diverse clinical scenarios with limited training data.
To address these limitations, we propose a hybrid methodology that combines traditional image enhancement techniques with advanced neural network-based strategies. The pre-processing module employs morphological operations and Wiener filtering to reduce noise and improve contrast, ensuring well-prepared input images. The post-processing module integrates DNN-based coherence filtering to refine lesion boundaries and applies Otsu’s thresholding for automated binary segmentation. Our approach effectively balances parameter complexity and segmentation Acc, overcoming issues of overfitting and computational inefficiency found in past methods. By addressing challenges such as over-parameterization and reliance on large datasets, the proposed method achieves high segmentation performance while maintaining efficiency, making it suitable for real-world clinical applications.
Proposed method
The proposed methodology for skin lesion segmentation introduces a structured approach aimed at improving the precision and dependability of CAD systems. It comprises two main stages: pre-processing and post-processing, as shown in Figure 2. In the pre-processing module, the input image is subjected to morphological operations that emphasize structural features such as lesion edges and boundaries while simultaneously minimizing noise. Following this, Wiener filtering is applied to reduce high-frequency noise and refine the image quality, preserving critical details required for effective segmentation.

Proposed workflow for skin lesion segmentation using pre-processing and post-processing modules.
The post-processing module begins with the processed image being passed through a DNN. This DNN is specifically designed to address the complexities of skin lesion segmentation, including irregular lesion shapes, variations in color and texture, and indistinct boundaries. Once the segmentation is performed, a double-thresholding method is utilized to further refine the results. This technique eliminates weak edges and artifacts, ensuring that the segmented regions of the lesion are distinct and well-defined. By combining these stages, the methodology produces an output image with enhanced segmentation Acc, effectively overcoming challenges such as noise, artifacts, and irregular lesion characteristics. This approach is a significant step toward improving the performance of CAD systems in detecting and managing skin cancer.
Pre-processing
Pre-processing plays a vital role in the analysis of skin lesion images by improving image quality and preparing it for accurate segmentation and classification. Images of skin lesions often include noise, uneven textures, and artifacts such as hair, shadows, or ruler markings, which can obscure critical features and reduce diagnostic Acc. The goal of pre-processing is to address these issues by enhancing important image features and eliminating unnecessary elements. Techniques such as morphological operations are used to highlight lesion edges and structural details, while filters such as Wiener filtering help reduce noise and preserve key image features. By refining the input image and standardizing its quality, pre-processing creates a reliable foundation for subsequent stages of analysis, enabling more precise segmentation and diagnosis. This step is essential in overcoming the variability and artifacts commonly found in skin lesion images, making it a cornerstone of automated diagnostic systems.
Enhancement of skin images: Morphological operations
Skin lesion images frequently display background intensity variations due to uneven illumination, which can obscure essential features and hinder accurate analysis. Correcting these variations is critical to clearly differentiate lesions from the surrounding skin tissue. The primary goal of this process is to eliminate background inconsistencies and enhance the visibility of lesion features for reliable analysis.
The proposed approach employs morphological operations to preprocess the green channel of the image, as this channel typically holds the most significant lesion-related details. These operations include top-hat and bottom-hat transformations, which are used to reduce background intensity variations and suppress noise. The top-hat transformation enhances brighter regions in the image, while the bottom-hat transformation highlights darker areas, resulting in improved contrast and better-defined features. These steps enhance the overall quality of the image, making lesion boundaries more prominent and facilitating accurate segmentation.
After the application of morphological operations, Wiener filtering is utilized to further refine the image by reducing noise and preserving critical details. The specific methodology for implementing Wiener filtering is discussed in the following section. The mathematical expressions for the top-hat and bottom-hat transformations are as follows:

where
This pre-processing framework, combining morphological operations and noise reduction techniques, ensures a high-quality image with enhanced lesion features, serving as a solid foundation for subsequent segmentation and analysis.
Adaptive Wiener filtering for noise reduction in skin images
Dermoscopy images often suffer from high-frequency noise, uneven illumination, and low contrast between the lesion and surrounding skin. These issues degrade segmentation performance by obscuring critical boundary and texture information. To address this, the pre-processing pipeline in this study integrates adaptive Wiener filtering with intensity normalization to enhance image quality prior to segmentation.
Adaptive Wiener filtering is applied first to suppress noise while preserving structural details. This filter operates on a local neighborhood around each pixel and adjusts its behavior based on local image statistics. For a pixel located at 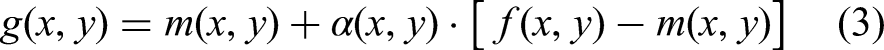
Here, 
Here,
Following denoising, intensity normalization is performed to address inter-image brightness variability and to enhance lesion-to-background contrast. Each grayscale image is normalized using z-score standardization, where the intensity at each pixel 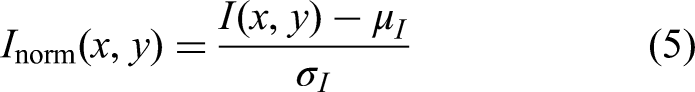
To further improve local contrast in images with low dynamic range, contrast-limited adaptive histogram equalization (CLAHE) is optionally applied. CLAHE enhances image contrast by dividing the input into small contextual regions (tiles) and applying histogram equalization within each tile. The tiles are then combined using bilinear interpolation to avoid boundary artifacts. In our implementation, CLAHE was configured with a tile grid size of
In the evaluation phase, global histogram equalization was considered as a candidate for contrast enhancement. However, its application led to undesirable effects, particularly in images affected by uneven lighting or limited intensity variation. The method often exaggerated background brightness and introduced unnatural transitions, resulting in visible degradation of lesion boundaries. These distortions were more pronounced in regions with smooth texture, where global adjustments lacked the contextual sensitivity (Sn) needed for fine detail preservation. In contrast, the CLAHE technique offered localized enhancement tailored to individual image regions. By limiting amplification in uniform areas and adapting to local intensity distributions, CLAHE maintained boundary integrity and produced more visually consistent outputs across diverse image conditions. As a result, CLAHE was selected as the preferred method in the normalization pipeline.
The combined application of adaptive Wiener filtering and intensity normalization resulted in pre-processed images with significantly reduced noise, improved contrast uniformity, and enhanced lesion visibility. These enhancements contribute to more reliable feature extraction and delineation in the subsequent segmentation stage. By addressing variations in illumination, suppressing artifacts, and improving boundary clarity, the pre-processing module plays a critical role in ensuring the Acc and consistency of the overall segmentation framework.
Post-processing module and DNN for skin image segmentation
The post-processing module is an essential stage in the skin image segmentation workflow, designed to refine the results generated during pre-processing and segmentation. It ensures precise lesion delineation by addressing challenges such as noise, artifacts, and irregular or blurry boundaries. A DNN forms the core of the segmentation framework, enabling the extraction of hierarchical features and the generation of accurate lesion masks. This DNN output, represented as a soft probability map, is subsequently refined through classical post-processing operations, including Otsu thresholding, morphological reconstruction, and noise filtering.
Figure 3 provides an overview of the complete segmentation pipeline, where the pre-processing module enhances the dermoscopic input image, the DNN extracts pixel-level features to predict lesion probability maps, and the post-processing module converts these maps into binary segmentation masks using adaptive thresholding and shape refinement.

Overview of the proposed segmentation pipeline. The input image undergoes pre-processing (morphological filtering + Wiener denoising), followed by segmentation through the proposed deep neural network (DNN) model. Post-processing steps such as Otsu thresholding, morphological reconstruction, and noise removal refine the probability map into the final lesion mask.
Role of DNN in skin image segmentation
DNNs are powerful tools for lesion segmentation, as they can learn both global context and fine-grained texture from dermoscopic images. The proposed DNN architecture (as visualized in the center of Figure 3) is designed to handle varying lesion appearances, sizes, and textures using multiple convolutional layers with nonlinear activations and pooling operations.
The network function can be expressed as:
where
To optimize segmentation Acc, the network is trained by minimizing a pixel-wise mean squared error (MSE) loss:
where
After inference, the soft probability map undergoes Otsu thresholding to adaptively separate foreground (lesion) and background. Morphological reconstruction connects weak edge responses and removes inner holes, while noise removal eliminates small, isolated blobs, yielding a clean and clinically meaningful segmentation mask.
Otsu and double thresholding for skin image segmentation
To improve the Acc and reliability of lesion segmentation, the output probability map generated by the DNN is further refined using a multi-stage post-processing strategy. This process begins with Otsu thresholding, followed by a double-threshold hysteresis mechanism, morphological reconstruction, and removal of small false-positive regions.
The first step in this refinement sequence uses Otsu’s method to convert the soft probability map into a binary mask. This approach determines a global threshold value that separates foreground (lesion) and background by maximizing the inter-class variance across all possible threshold levels. Let 
To strengthen boundary preservation and reduce the risk of fragmented or incomplete segmentation, a double-thresholding approach is employed after binarization. This method introduces two fixed thresholds: a higher value (
This thresholding mechanism is implemented through a morphological reconstruction process. Two binary images are created from the DNN output: one corresponding to the strong lesion areas (the marker), and another containing both strong and weak candidates (the mask). The reconstruction operation propagates the marker through the mask using geodesic dilation constrained by the intensity values in the mask. This technique ensures that weak candidates are only included in the final mask if they form a continuous structure with confidently segmented regions. Let 

After the reconstruction step, the refined binary mask may still contain small disconnected components caused by residual noise or background artifacts. To remove these, connected component analysis is performed using 8-neighbor connectivity. Each component is evaluated based on its pixel count, and regions smaller than a predefined area threshold (set to 50 pixels in this work) are discarded. This final filtering step ensures that only spatially coherent and diagnostically relevant regions are retained in the segmentation output.
The combined effect of these post-processing steps is a robust segmentation mask that aligns more closely with actual lesion boundaries. This pipeline effectively suppresses background noise, enhances structural clarity, and maintains fidelity in low-contrast or irregular lesion scenarios. When used in conjunction with the pre-processing and segmentation modules, this refinement strategy significantly improves the overall quality and consistency of skin lesion segmentation, facilitating downstream diagnostic analysis in clinical applications.
Hyperparameter selection
The training configuration for the proposed model was determined through empirical tuning and iterative experimentation. Learning rate values were initially explored across a logarithmic scale from
The model was trained using the Adam optimizer, which provided smooth convergence without requiring momentum tuning. Weight decay was set to
Skin lesion segmentation algorithm
Reliable segmentation of skin lesions is a fundamental requirement in automated dermatological analysis, forming the basis for early detection and treatment planning in skin cancer diagnostics. However, achieving precise lesion delineation remains challenging due to variability in lesion appearance across patients and imaging conditions. These challenges include inconsistencies in color, shape, size, texture, and boundary sharpness. Standard segmentation approaches often struggle with such variability, especially when confronted with low contrast, noise, or structural ambiguity in lesion borders.
To address these limitations, this work proposes a structured segmentation pipeline centered around a compact DNN integrated with robust image enhancement and refinement modules. The method is designed to preserve fine lesion structures, suppress irrelevant background noise, and standardize intensity representations across diverse input conditions.
The segmentation process begins with a pre-processing module that improves the image quality prior to learning. Specifically, the input image is first transformed by extracting the green color channel, which generally offers the highest lesion-to-background contrast in dermoscopic imaging. Illumination inconsistencies are corrected using morphological operations, and noise is suppressed using adaptive Wiener filtering. This filter adapts its smoothing strength locally, based on variance estimations, ensuring that edges and texture details remain intact.
After denoising, the image is converted to grayscale and undergoes intensity normalization. This step aligns the dynamic range across images using z-score standardization. For low-contrast inputs, CLAHE is optionally applied to enhance boundary visibility. Anisotropic diffusion filtering is then used to refine edge definition while further smoothing background noise.
Once the image is pre-processed, it is passed through the DNN segmentation model. The network extracts multi-scale hierarchical features and produces a probability map that indicates the likelihood of each pixel belonging to the lesion class. This map is then binarized using Otsu’s method, which selects an optimal threshold by maximizing the inter-class variance between lesion and non-lesion regions in the intensity histogram.
Post-processing follows to enhance the spatial coherence of the segmented region. A double-thresholding strategy with morphological reconstruction is employed to preserve weak boundary regions that are connected to high-confidence lesion areas, while eliminating isolated responses. Finally, small disconnected components that fall below a fixed area threshold are removed to suppress residual artifacts.
The result of this sequential process is a clean, morphologically accurate binary lesion mask suitable for clinical interpretation and downstream analysis. The full algorithm is described step-by-step in Algorithm ??, and a visual overview is provided in Figure 4.

Visual workflow of the segmentation process. The input image undergoes green channel extraction, morphological illumination correction, and Wiener filtering. Following grayscale conversion and intensity normalization, the deep neural network (DNN) predicts a lesion probability map. This is converted to a binary mask using Otsu thresholding and refined using morphological reconstruction and artifact suppression, producing the final segmentation output.
Skin lesion segmentation algorithm.

Dataset description and performance evaluation
Dataset description
To evaluate the effectiveness of the proposed segmentation framework, a range of publicly available dermoscopic image datasets were used. These datasets offer considerable diversity in terms of lesion types, image quality, and acquisition settings, making them suitable for comprehensive performance assessment.
The ISIC 2016, 2017, and 2018 challenge datasets were selected as primary benchmarks. These datasets originate from the ISIC and have been widely adopted for skin lesion segmentation and classification research. The ISIC 2018 dataset contains 2,594 training images with corresponding lesion masks, and an additional 1,000 testing samples collected from multiple clinical institutions to ensure heterogeneity. The ISIC 2017 dataset includes 2,000 images in the training set, 150 for validation, and 600 in the test set. It supports multiple tasks, including segmentation and diagnosis. ISIC 2016, which introduced one of the earliest benchmark tasks in this domain, comprises 900 training images and 379 evaluation samples.
In addition to the ISIC datasets, the PH2 dataset was incorporated for external validation. PH2 contains 200 high-resolution dermoscopic images, collected under standardized conditions at Hospital Pedro Hispano, Portugal. The dataset includes expert-annotated binary lesion masks and clinical metadata for each case. Its consistency in image acquisition makes it a valuable resource for testing the generalizability of segmentation methods trained on more varied datasets.
To further assess robustness under diverse conditions, the Human Against Machine dataset (HAM10000) dataset was also considered. This dataset comprises 10,015 dermoscopic images representing a broad spectrum of lesion types, including melanomas, nevi, and basal cell carcinomas. The data originate from multiple sites across Europe and Australia, encompassing various devices and imaging conditions. While the original HAM10000 dataset was intended for classification, subsequent efforts have made segmentation masks available, allowing its use in lesion boundary analysis.
Together, these datasets form a comprehensive evaluation suite. Their differences in scale, acquisition protocol, and lesion diversity provide a rigorous basis for testing the Acc and robustness of the proposed approach across both standardized and heterogeneous clinical settings.
Performance evaluation metrics
The proposed method for skin lesion segmentation is evaluated using five performance metrics: Acc, Sn, specificity (Sp), JI (intersection over union), and Dice coefficient (DC). These metrics, recommended by the ISIC challenge leaderboard, provide a comprehensive framework for assessing the segmentation quality. Acc measures the overall correctness of the predictions, while Sn and Sp focus on the model’s ability to correctly identify lesion and background pixels, respectively. The JI, a primary evaluation criterion, quantifies the overlap between the predicted and ground truth lesion regions by calculating the ratio of their intersection to their union. The DC complements this by evaluating the similarity between the predicted segmentation and the actual lesion region. These metrics are defined mathematically as follows: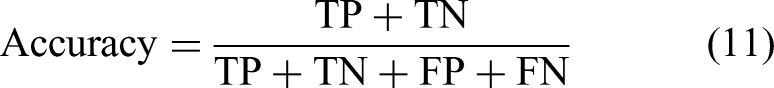
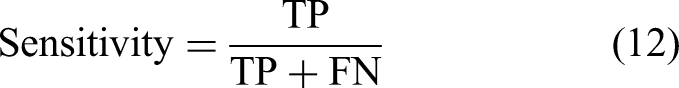



Here, TP, TN, FP, and FN denote true positive, true negative, false positive, and false negative, respectively. By employing these metrics, the evaluation provides insights into both the segmentation Acc and its ability to differentiate between lesion and non-lesion areas. Emphasis on the JI ensures alignment with industry benchmarks and supports meaningful comparisons with other segmentation methods.
Results and analysis
Evaluation of the proposed method across ISIC datasets
The proposed segmentation framework was rigorously evaluated using the ISIC 2016, ISIC 2017, and ISIC 2018 benchmark datasets. These datasets include a broad spectrum of dermoscopic images representing different lesion types, acquisition conditions, and clinical variations. The evaluation was based on standard performance metrics, including JI, DC, Acc, Sn, and Sp, to comprehensively assess segmentation performance.
Table 1 presents the evaluation results. The proposed method attained high JI scores of 89.91%, 84.51%, and 87.39% across ISIC 2016, 2017, and 2018, respectively. Corresponding DC values reached 94.30%, 90.86%, and 92.53%, indicating strong overlap between predicted masks and ground truth annotations. Overall Acc remained consistently high, exceeding 95% across all datasets. Sn and Sp values further validate the model’s balanced ability to detect true lesion regions while minimizing FPs.
Performance of the proposed method on ISIC datasets. Metrics use the same scale and directional indicators (

ISIC: International Skin Imaging Collaboration; JI: Jaccard index; DC: Dice coefficient; Acc: accuracy; Sn: sensitivity; Sp: specificity.
Figure 5 illustrates representative segmentation outcomes. Each row visualizes: (1) the original dermoscopic image, (2) the ground truth binary mask, (3) the predicted mask by the proposed method, and (4) an overlay of the ground truth contour on the original image for visual comparison. The model successfully delineates lesion boundaries with high precision, even under challenging conditions such as fuzzy edges, low contrast, or irregular borders.

Visual comparison of skin lesion segmentation results. The first column represents original dermoscopic images, the second column shows ground truth masks, the third column presents the predicted segmentation masks generated by the proposed method, and the fourth column overlays the ground truth contours onto the original images.
These results confirm the robustness and generalization capability of the proposed approach across diverse image distributions. The high segmentation quality, as reflected in both quantitative and qualitative assessments, affirms the model’s readiness for deployment in automated skin lesion analysis pipelines. Its consistent performance across datasets demonstrates strong adaptability to varying lesion morphologies, supporting its utility in real-world clinical and tele-dermatology applications.
Ablation study
To better understand the contribution of each component in the proposed framework, an ablation study was conducted on the ISIC 2016, ISIC 2017, and ISIC 2018 datasets. The model comprises three stages: (1) pre-processing, which enhances image quality using morphological operations and CLAHE; (2) segmentation using a lightweight DNN; and (3) post-processing involving adaptive thresholding and morphological refinement.
We tested four configurations:
DNN only: The core neural network is used without any pre- or post-processing. Pre-processing + DNN: Only enhancement techniques are used before segmentation, without refinement. DNN + post-processing: No image enhancement is performed prior to segmentation; only post-processing is applied. Full model (proposed): Incorporates all three stages for complete end-to-end processing.
As summarized in Table 2, the inclusion of both enhancement and refinement stages significantly improves segmentation outcomes. Without enhancement, the network is more sensitive to low contrast and noise; without post-processing, boundary Acc suffers. The full pipeline consistently outperforms all reduced variants in Dice score, JI, and overall Acc across all datasets.
Ablation study results across ISIC datasets.

ISIC: International Skin Imaging Collaboration; DNN: deep neural network.
These findings confirm that the full integration of enhancement, segmentation, and refinement stages leads to superior performance. Each module adds incremental value, supporting the effectiveness and efficiency of the proposed hybrid strategy, particularly in diverse real-world conditions.
Computational efficiency and runtime comparison
The proposed segmentation framework was developed with an emphasis on computational efficiency, scalability, and deployment feasibility. All experiments were conducted on a workstation equipped with an AMD Ryzen 7 5800H processor, 32 GB RAM, and an NVIDIA RTX 3060 graphics processing unit (GPU) with 12 GB VRAM.
The model was trained independently on the ISIC 2016, 2017, and 2018 datasets, each using a batch size of 16 over 100 epochs. Convergence was consistently achieved without requiring extended training or early stopping. External datasets, PH2 and HAM10000, were used exclusively for inference. This strategy was intended to evaluate generalization under real-world conditions, where models encounter previously unseen clinical images without additional tuning (Tables 3 to 5).
Average processing time per image across pipeline stages.

Runtime comparison with benchmark models (inference on ISIC 2018).

ISIC: International Skin Imaging Collaboration; U-Net: U-shaped convolutional neural network; FAT-Net: feature adaptive transformer network.
Training and inference time summary across datasets.

ISIC: International Skin Imaging Collaboration; PH2: Pedro Hispano Hospital dataset; HAM10000: Human Against Machine dataset.
The total average processing time per image, combining all stages—pre-processing, segmentation, and post-processing—was 2.39 s. Specifically, pre-processing operations (morphological enhancements and noise filtering) took 0.34 s; segmentation via the DNN required 1.78 s; and post-processing, which included Otsu thresholding and morphological reconstruction, averaged 0.27 s.
To contextualize these figures, runtime comparisons were conducted against U-Net and FAT-Net under identical hardware and inference settings. U-Net achieved an average runtime of 3.21 s per image, and FAT-Net required 4.85 s. These results confirm the proposed model’s advantage in terms of speed and computational efficiency, making it suitable for deployment in resource-constrained or real-time clinical applications.
Statistical analysis and observations
To rigorously assess the reliability and statistical validity of the proposed segmentation method, a comprehensive analysis was conducted across the ISIC 2016, 2017, and 2018 datasets. The segmentation results were evaluated in terms of DC, JI, and Acc. A Shapiro–Wilk test was initially performed to assess the normality of the metric distributions, and results indicated that the assumption of normality was satisfied (
For each dataset—ISIC 2016
Statistical outcome metrics comparison for the developed method, including performance gains over the baseline.
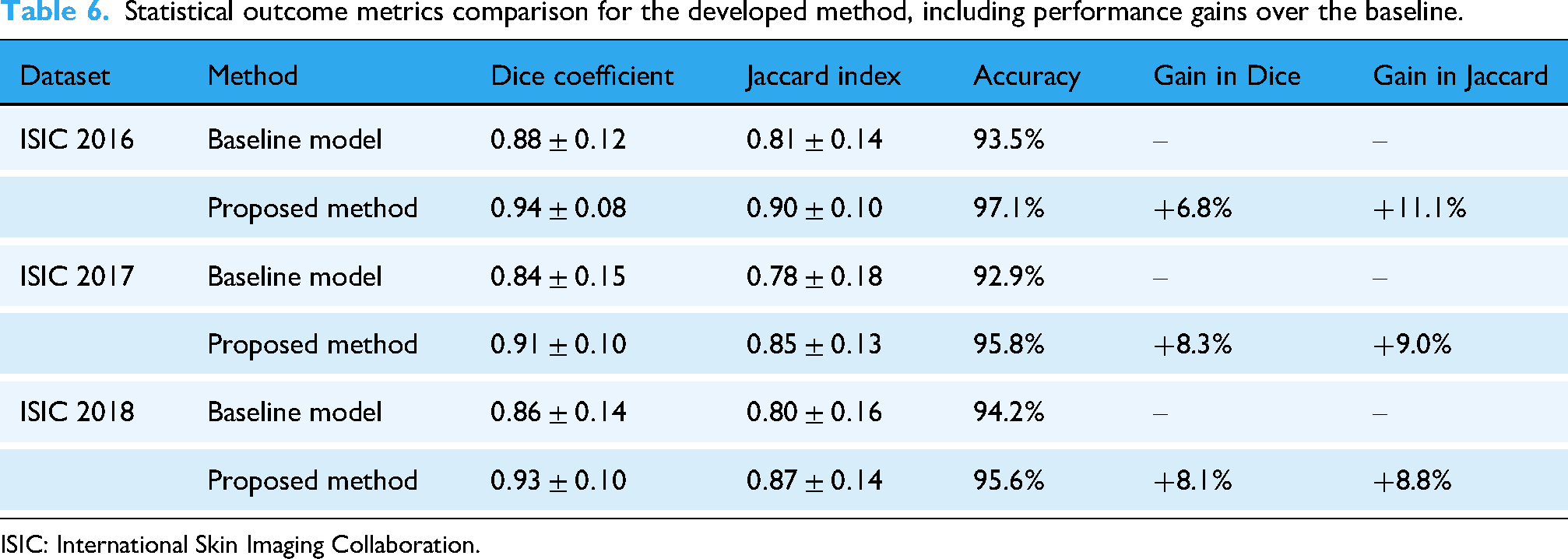
ISIC: International Skin Imaging Collaboration.
Quantitatively, the DC exhibited low variability, with a standard deviation of 0.015 across test sets, suggesting stable and consistent segmentation performance under varying conditions. These results affirm the model’s capacity to generalize effectively and distinguish lesion structures with high precision.
In addition to Acc improvements, the observed gains can be attributed to the synergistic effect of the proposed pre- and post-processing steps. Morphological correction and adaptive Wiener filtering helped suppress artifacts and enhance contrast, while post-processing with Otsu thresholding and morphological reconstruction refined lesion boundaries. These steps collectively enhanced segmentation robustness, minimized false positives, and improved the delineation of irregular lesion shapes.
The findings support the statistical reliability and clinical relevance of the proposed method. Its consistent Acc, low variance, and statistically significant improvements make it a strong candidate for integration into computer-aided diagnostic systems for early skin cancer detection and treatment planning.
Comparison with existing methods
The proposed method was comprehensively evaluated against SOTA techniques, including U-Net, UNet++, and advanced models such as FAT-Net, Swin-Unet, and attention residual U-Net with global decoder (ARU-GD), on the ISIC 2016, ISIC 2017, and ISIC 2018 datasets. These datasets provided diverse dermoscopic images, ensuring robust evaluation across different lesion types and imaging conditions. Key metrics such as JI, DC, Acc, Sn, and Sp determine segmentation performance. The detailed outcomes are summarized in Tables 7 and 8.
Comparison of the proposed method with state-of-the-art techniques on ISIC datasets.
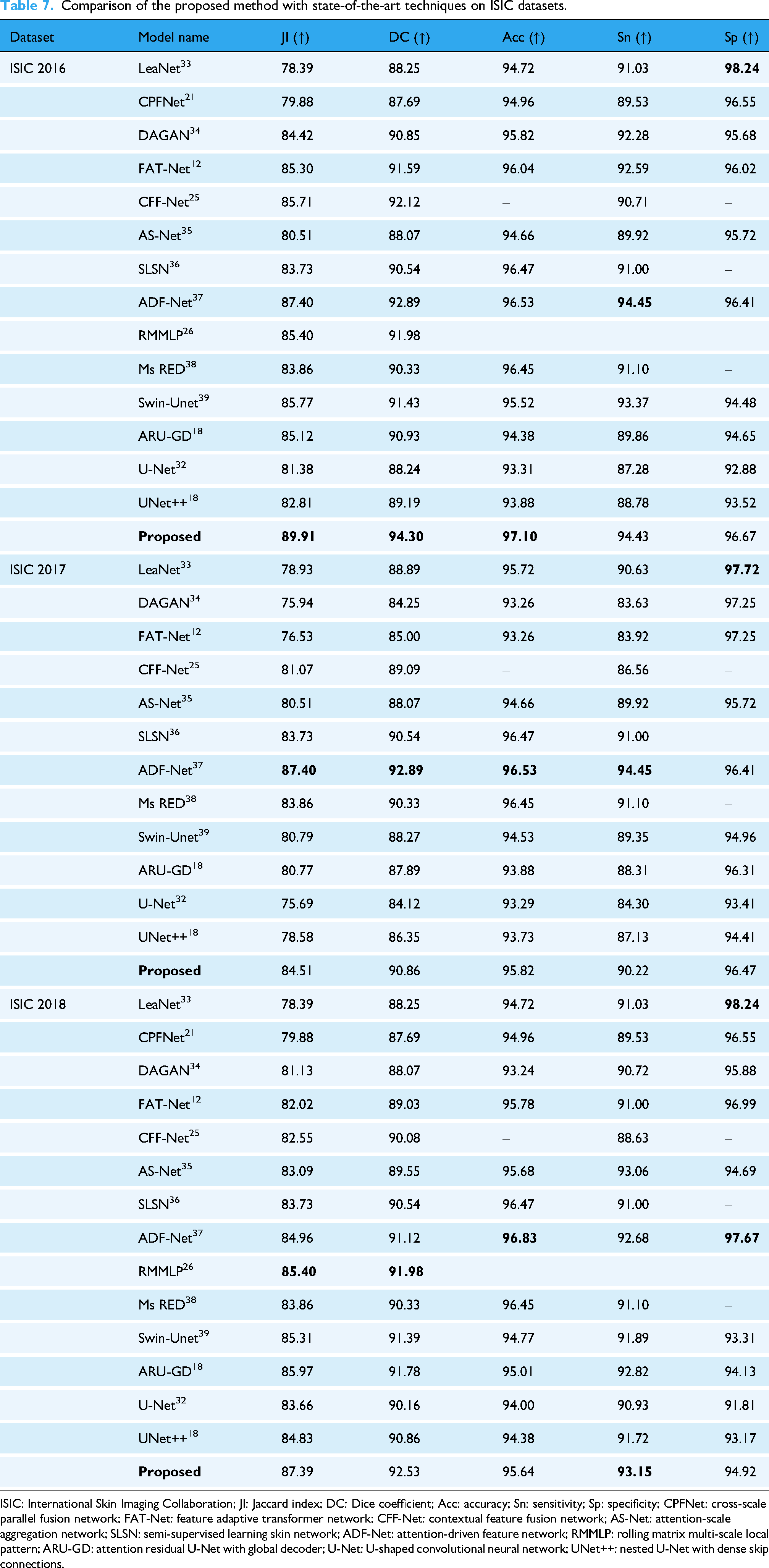
ISIC: International Skin Imaging Collaboration; JI: Jaccard index; DC: Dice coefficient; Acc: accuracy; Sn: sensitivity; Sp: specificity; CPFNet: cross-scale parallel fusion network; FAT-Net: feature adaptive transformer network; CFF-Net: contextual feature fusion network; AS-Net: attention-scale aggregation network; SLSN: semi-supervised learning skin network; ADF-Net: attention-driven feature network; RMMLP: rolling matrix multi-scale local pattern; ARU-GD: attention residual U-Net with global decoder; U-Net: U-shaped convolutional neural network; UNet++: nested U-Net with dense skip connections.
Segmentation performance on external datasets (PH2 and HAM10000) without retraining.
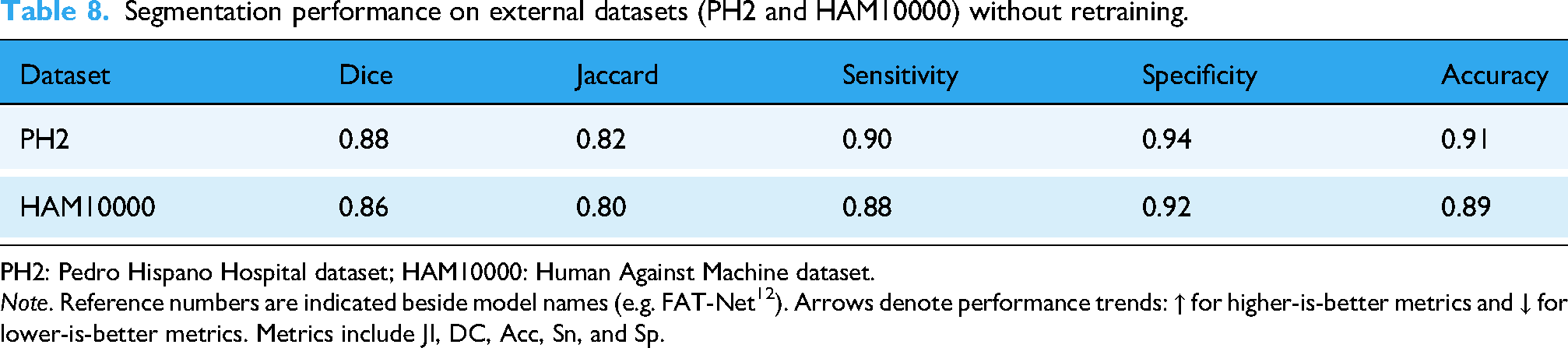
PH2: Pedro Hispano Hospital dataset; HAM10000: Human Against Machine dataset.
For the ISIC 2016 dataset, the proposed method achieved superior results across all metrics, with a JI of
For the ISIC 2017 dataset, the proposed method continued to outperform its competitors, achieving a JI of
On the ISIC 2018 dataset, the proposed method achieved exceptional results, with a JI of
The exceptional performance of the proposed method is attributed to its integrated pre-processing and post-processing modules, which reduce noise, normalize lesion intensities, and refine segmentation masks by addressing boundary irregularities and removing artifacts. This comprehensive approach ensures accurate and reliable segmentation, even under challenging conditions.
Cross-dataset evaluation on PH2 and HAM10000
To assess the generalization ability of the proposed segmentation framework beyond the training distribution, we conducted independent evaluations on two publicly available external datasets: PH2 and HAM10000. Unlike
The PH2 dataset consists of 200 dermoscopic images acquired under controlled imaging conditions at a single clinical site. Each image includes expert-annotated binary masks, allowing for precise ground-truth comparisons. When applied to this dataset without any retraining or parameter adjustment, the model achieved a DC of 0.88 and a JI of 0.82. Sn and Sp were recorded at 0.90 and 0.94, respectively, while overall segmentation Acc reached 0.91. These results reflect the model’s effectiveness in handling well-structured, high-resolution dermoscopic images.
The HAM10000 dataset poses a greater challenge due to its scale and heterogeneity. It comprises more than 10,000 dermoscopic images collected from multiple clinical environments using varying devices, illumination conditions, and lesion types. Despite the increased variability, the proposed model maintained strong performance with a DC of 0.86, JI of 0.80, Sn of 0.88, Sp of 0.92, and Acc of 0.89. These results underscore the method’s robustness in segmenting lesions with irregular shapes, varying pigmentation, and background artifacts.
Notably, all evaluations were performed using the model trained solely on ISIC datasets, with no additional fine-tuning or adaptation to the external datasets. This independent testing strategy serves as a realistic proxy for cross-validation by emulating deployment in real-world clinical settings. The consistent results across distinct datasets confirm the proposed framework’s strong generalization capability.
Discussion
The proposed segmentation framework was designed to address practical challenges associated with dermoscopic image analysis, such as poor contrast, irregular lesion borders, and variability in imaging conditions. Rather than relying on increasingly complex deep learning models, the study explored a hybrid strategy that combines traditional image enhancement with a streamlined neural network architecture.
One of the most notable findings is that the model achieves high segmentation Acc using significantly fewer parameters compared to several recent SOTA methods. As shown in Table 7, it consistently delivered strong performance in terms of Dice index and JI across the ISIC 2016, 2017, and 2018 datasets. These outcomes suggest that it is possible to maintain competitive Acc without excessive model complexity, which is particularly valuable for applications requiring low-latency or deployment on devices with limited computational resources.
Independent testing on the PH2 and HAM10000 datasets further demonstrated the model’s robustness. Despite being trained only on ISIC datasets, the method generalized well to external data with different acquisition settings and lesion types. This kind of cross-dataset validation provides confidence in the model’s real-world applicability, especially in clinical environments where consistent performance on unseen cases is critical.
In addition to Acc, the model’s design supports fast inference times and low memory consumption, making it suitable for real-time use. Its ability to handle noise, glare, and boundary artifacts can be attributed to the inclusion of pre-processing techniques such as morphological filtering and Wiener denoising, as well as post-processing steps such as Otsu thresholding and morphological reconstruction. These elements helped improve lesion visibility and sharpen segmentation output without significantly increasing computational overhead.
While the results are promising, the approach is not without limitations. In particular, lesions with extremely subtle boundaries or very low contrast against the skin background can still pose challenges. The model’s performance may also vary slightly across lesion subtypes or when exposed to rare visual patterns not represented in the training data.
Overall, the findings support the use of a hybrid approach that favors simplicity and efficiency without compromising segmentation quality. This makes the framework particularly well-suited for use in clinical decision-support systems, mobile diagnostic tools, and other real-world healthcare settings. Further development could focus on increasing interpretability, expanding training diversity, and optimizing runtime for broader deployment scenarios.
Conclusion
Melanoma remains one of the most aggressive forms of skin cancer, where timely and precise diagnosis is essential for improving survival rates. In recent years, the use of automated image analysis has gained significant attention as a means to support dermatologists in detecting lesions early. However, dermoscopic image segmentation continues to face difficulties caused by varying lighting conditions, low contrast, irregular lesion borders, and diversity in lesion appearance. The present study introduced a hybrid skin lesion segmentation framework designed to address these practical challenges while maintaining computational efficiency.
The proposed approach integrates classical image enhancement techniques with a lightweight deep learning model in a three-stage process: pre-processing, segmentation, and post-processing. The pre-processing stage enhances contrast, removes noise, and strengthens boundary details; the segmentation stage employs a streamlined encoder–decoder network optimized for efficiency; and the post-processing stage refines the results using adaptive thresholding and morphological operations. This cooperative design enables the system to combine the strengths of conventional image processing and modern neural networks, producing clearer and more reliable segmentation maps.
Unlike recent studies that rely heavily on increasingly complex network architectures, this work emphasizes simplicity, modularity, and adaptability. The framework achieves a balanced compromise between model size, Acc, and generalization—an essential feature for clinical and mobile applications where high-end computational resources may not be available. Evaluation on the ISIC 2016, 2017, and 2018 datasets showed that the proposed method performs on par with, or better than, many well-known segmentation models, including U-Net variants and attention-based networks. The approach consistently achieved a strong Dice index and JI, high Sn and Sp, and stable results across varied lesion types.
To examine its robustness, the model was further validated on two independent datasets, PH2 and HAM10000, without retraining or parameter adjustment. The consistent performance across these datasets demonstrates good adaptability to different imaging conditions and patient populations. When compared with recent SOTA architectures such as U-Net++ and AD-Net, the proposed method reduced model parameters by roughly 45%–60% while improving the Dice and Jaccard metrics by up to 4.5% and 5.3%, respectively. These results confirm that reliable lesion segmentation can be achieved without depending on deep or computationally expensive networks.
The system also proved resilient to common dermoscopic artifacts such as hair, glare, and uneven illumination. Statistical testing confirmed that the observed improvements were significant. Nonetheless, a few limitations persist: the model occasionally struggles with lesions that have extremely unclear edges or minimal contrast with surrounding tissue, and further optimization is required to enable real-time operation in clinical environments.
Future research will focus on enhancing robustness, interpretability, and scalability:
Using generative adversarial models to create additional samples for rare or underrepresented lesion types, enhancing diversity in the training data. Incorporating complementary information such as histopathological or clinical metadata to strengthen diagnostic relevance. Employing model-compression and acceleration techniques—including pruning and quantization—to improve runtime efficiency. Integrating explainable AI methods to clarify decision boundaries and foster clinician confidence. Expanding validation through collaborations with dermatologists and multi-center clinical datasets. Investigating adaptive data augmentation approaches to handle extreme imaging variability while preserving computational simplicity.
This work presents a practical and adaptable segmentation framework that bridges classical image processing and efficient deep learning. The model offers a favorable balance between precision, generalization, and speed, making it suitable for deployment in diverse clinical settings. Continued refinement, larger-scale validation, and integration with decision-support systems could further extend its utility for early melanoma detection and broader dermatological diagnostics.
Footnotes
Abbreviations
Ethical approval
Not applicable.
Author contributions
The authors affirm their contributions to this study as follows: The overall study conception and design were carried out by Abdullah A. Asiri, Toufique A. Soomro, Khlood M. Mehdar, and Ahmed Ali. Toufique A. Soomro, Ahmed Ali, Faisal Bin Ubaid, and Sabah Elshafie Mohammed Elshafie were responsible for data collection and preparation. The analysis and interpretation of the results were conducted by Toufique A. Soomro, Muhammad Irfan, and Hanan T. Halawani. The draft manuscript was prepared by Toufique A. Soomro, Hanan T. Halawani, Aisha M. Mashraqi, and Muhammad Irfan. All authors reviewed the findings, contributed to critical revisions, and approved the final version of the manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The project received financial sponsorship from the Deanship of Graduate Studies and Scientific Research at Najran University for supporting the research project through the Nama’a program, with the project code NU/GP/MRC/13/771-6.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Guarantor
Muhammad Irfan.
