Abstract
Objectives
In the world, skin cancer is a significant health concern, and early diagnosis of this cancer plays a key role in improving patient outcomes. The early detection of this cancer reduces the death rate, but due to the complexity of the diagnosis, incorrect detection and prediction are provided by the experts. Therefore, it is essential to propose a computer-aided diagnostic system based on deep learning and explainable Artificial Intelligence (XAI) techniques that can be used as a second opinion in clinics and help physicians more accurately detect and predict this type of cancer.
Methods
This work presents the proposed deep learning architecture consisting of two modules—skin lesion segmentation and lesion type classification. The proposed architecture is interpreted using XAI techniques to better evaluate the black-box model. In the skin lesion segmentation phase, we implemented DeepLab V3 architecture for semantic segmentation. The ResNet-18 model was used as the backbone, and later hyperparameters were optimized using Bayesian Optimization (BO). In the classification phase, we design a FusedNet architecture called Inverted self-attention with Vision Transformer (ISAwViT). The proposed fused network combines an inverted self-attention residual architecture with a vision transformer. The proposed fused network extracted feature information more deeply than performing an accurate prediction in a later stage. The design model is trained, and later in the testing phase, extracted features are classified using Softmax and several other classifiers.
Results
The lesion segmentation and classification experiment was conducted on the HAM10000 dataset. The accuracy achieved by the HAM10000 dataset was 95.16% for lesion segmentation and 97.5% for lesion classification.
Conclusion
Compared with recent techniques, the proposed model is more effective and efficient. In addition, the interpretation of the proposed model was performed using LIME and Grad-CAM, which show how the fused model makes correct classifications.
Keywords
Introduction
In the United States, skin cancer is the most common type of cancer to be diagnosed. 1 The most serious form of skin cancer, melanoma, has emerged as a major public health concern in recent years. 2 A skin cancer is an abnormal growth of skin cells that often develops on the skin as a result of exposure to UV light or sunshine. 3 Skin cancer is a deadly condition that falls into two categories: benign (basal cell and squamous cell carcinoma) and malignant (melanoma). Benign tumors rarely spread to other skin tissues and are always resistant to therapy. 4 A deadly kind of skin cancer that begins in the pigment cells is called melanoma. Malignant lesions give rise to skin cancer, which is responsible for around 75% of all deaths. 5
According to reports, there were 207,390 cases reported in the United States in 2021 that were diagnosed with skin cancer. 6 From these, 106,110 are noninvasive, and 101,280 are invasive, with 62,260 men and 43,850 women. 7 In the United States (US), 7180 deaths are anticipated in 2021, comprising 4600 males and 2580 females. In 2020, there were 100,350 instances recorded in the US, comprising 60,190 men and 40,160 women. 8 Over 16,221 unique cancer cases, comprising 9480 men and 6741 women, are expected to have been studied in Australia in 2020. Of these, 1375 deaths were recorded, with 891 men and 484 women dying. 9 According to dermatologists, melanoma can spread to neighboring tissues or the entire body if not detected in its early stages. However, there is a good possibility of survival if it is found early. The high death rate of melanoma has drawn significant attention from the scientific community. 10
In the past, dermatologists have employed several techniques, such as laser technology, a seven-point checklist, the ABCDE rule, and a few others, for the diagnosis of skin cancer. 11 However, an experienced dermatologist is needed for these procedures. Furthermore, it is costly, time-consuming, and challenging to manually evaluate and diagnose skin cancer using these techniques. 12 Thus, computerized techniques are widely required for the segmentation and classification of skin cancer, providing a quick and second opinion for dermatologists. 13 A recent technological advancement in the diagnosis of skin cancer is dermoscopy. 14 Dermoscopy uses RGB imaging to capture skin images, which dermatologists then examine. By employing dermoscopic images, several computerized models have been introduced in the literature. 15
A computerized model consists of a few important steps, including preprocessing dermoscopic images, segmenting skin lesions, extracting features, and classification. 16 Preprocessing is the stage in which noise and hair artifacts can be eliminated using various image processing techniques. 17 Further, the low-contrast images are improved by employing multiple filtering techniques. After that, segmentation is performed where lesion and healthy regions from the dermoscopic images are extracted into two parts and a boundary is drawn that is later compared with the ground truth value. 18 Lesion classification is another important and challenging part that relies on the extraction of useful features. The extracted features are classified using machine learning classifiers. 19
Convolutional neural networks (CNNs) are a well-known deep learning (DL) method used in several medical imaging tasks, especially for skin cancer. 20 There are large collections of labelled dermoscopic images used to train CNNs. Several hidden layers extract image features, and even minor changes are easily predicted. The most commonly used layers in a CNN architecture are convolutional, pooling, ReLU, fully connected, and Softmax. 13 They can now identify and interpret subtle visual cues that may indicate the presence of carcinoma, thanks to the CNN model's training. The computer-aided diagnostic system is a powerful tool for the early and more accurate detection of skin cancers, but it cannot replace the expertise of dermatologists. The performance of the CNN model or the diagnosis and classification of skin cancer depends on the accurate training; however, sometimes, the training is not performed well due to the complex structure of the dermoscopy image, similarity in colors of different types of lesions, and high intra- and inter-class similarity (can be seen in Figure 1).

Sample images of the dataset (a) melanocytic nevus, (b) actinic keratosis, (c) dermatofibroma, (d) basal cell carcinoma, (e) vascular lesions, (f) benign keratosis, (g) melanoma.
Recently, DL has made impressive progress in medical imaging. 21 In medical imaging, skin cancer is among the leading cancer types, and numerous computerized techniques have been introduced in the literature.13,22 The techniques presented in the literature are based on traditional and DL methods; however, due to the large number of datasets, DL techniques have achieved significant success. 23 The CNN architectures presented in the literature not only performed well on lesion-type classification but also showed improved precision in lesion segmentation. 24 In literature,25–29 several DL techniques such as CNNs, vision transformers (ViTs), hybrid CNN–transformer architectures, advanced training optimization strategies, ensemble learning, and U-Net-based segmentation networks are being explored to enhance model performance. These studies investigate architectural improvements, feature extraction mechanisms, learning rate scheduling, and model interpretability to achieve better accuracy, efficiency, and generalization. Collectively, these advancements indicate that diverse DL methodologies are continually being developed and evaluated to advance medical imaging and computer-aided analysis. Masni et al. 30 presented an integrated DL technique for the segmentation and classification of skin lesions. For lesion boundary extraction, a DL model with full resolution was presented. Later, for the classification of skin lesions in the appropriate category, four pretrained DL models were trained and performed experiments on three datasets: International Symposium on Biomedical Imaging 2016 (ISBI2016), which contains two classes; International Skin Imaging Collaboration 2017 (ISIC2017), which includes three classes; and ISBI2018 (which consists of seven classes), respectively. Sikkandar et al. 31 presented an automated system that uses the GrabCut algorithm in conjunction with an adaptive Neuro Fuzzy classifier (NeFy) to classify skin lesions. In this method, the GrabCut algorithm was used to perform segmentation after applying the top-hat filter to enhance the contrast of the original images. The NeFy classifier was then used to extract and classify the DL features. They conducted experiments on the ISIC dataset and achieved a sensitivity of 93.47%. Philipp et al. 32 presented a method for classifying and segmenting skin lesions using pretrained fully convolutional network encoders. They primarily used pre-trained CNN models for segmentation and classification. Hemsi et al. 33 presented a group of DL models for classifying skin cancer into multiple classes. They used InceptionV3, DenseNet201, Inception-ResNetV2, GoogLeNet, and MobileNetV2, and trained them using transfer learning. They used the HAM10000 dataset in their experiments and achieved an accuracy of 87.7%. Based on the results, the Densenet model performs better on the balanced dataset. Nawaz et al. 34 presented a DL-based faster region convolution neural network. Fuzzy k-means clustering is also used in this method to diagnose benign and malignant melanoma regions. They conclude that the preprocessing techniques for image enhancement and noise reduction have improved the segmentation accuracy. Singh et al. 35 presented a transfer learning (TL)-based system called Transfer Constituent Support Vector Machine (TrCSVM) for melanoma classification. Two key components of the framework are embedded, including Transfer AdaBoost (TrAdaBoost) and Support Vector Machine. Farhat et al. 36 presented a DL algorithm for classifying skin cancer. The HAM10000 and ISIC 2018 databases are used in this work. They extracted DL features, which were then selected by a metaheuristic algorithm. Srinivasa et al. 37 presented a hybrid approach combining an LSTM and MobileNetV2 for lesion classification. They show that the fusion process improved the overall system accuracy. Chandrahaas et al. 38 explored DL-based computer vision techniques for the automated analysis of skin lesions, which often exhibit similar visual symptoms. They employed TL with CNNs to enable rapid preliminary screening and lesion classification. Their findings demonstrated that such models could achieve promising accuracy and assist in early detection and clinical decision-making.
In summary, these studies focused on contrast enhancement, pre-trained CNN architecture, feature fusion, and feature selection. Overall, they attempted to improve accuracy, but they continue to encounter errors in lesion segmentation and class prediction. Therefore, this challenge can be resolved through a fusion mechanism in which multiple CNN architectures are designed, and their outputs are fused in a single layer (network-level fusion) or in a matrix (feature-level fusion). However, another challenge noted in recent studies is that the fusion process increases computational time and complexity and introduces redundant features. These challenges reduced the model's predictive performance and increased overall computational time during testing. The novelty of this study lies in four key innovations that address existing limitations in skin cancer detection systems. First, we introduce an inverted self-attention residual (ISAR) architecture in which attention mechanisms are embedded within inverted residual blocks rather than applied as post-processing layers, enabling simultaneous local-global feature learning with fewer parameters (5.3 M vs 25 M + in comparable models). Second, our network-level fusion strategy using depth concatenation preserves independent gradient paths for both ISAR and ViT components, preventing gradient interference that commonly occurs in early feature fusion approaches a critical advantage given the different operational principles of CNNs (local relationships) and Transformers (global dependencies). Third, we propose a Bayesian-optimized DeepLabV3 + framework with ResNet18-SelfAttention backbone for semantic segmentation, representing the first application of hyperparameter optimization for lesion boundary detection in dermoscopy. Fourth, we provide comprehensive interpretability through dual explainable Artificial Intelligence (XAI) techniques (LIME and GradCAM) with quantitative validation metrics (IoU scores, localization accuracy), going beyond typical qualitative visualization to ensure clinical trustworthiness. These innovations collectively address gaps in computational efficiency, gradient optimization, segmentation accuracy, and interpretability in existing literature. In this work, we proposed a computerized framework based on DL to segment and classify skin cancer from dermoscopy images. Our significant contributions are listed as follows:
Implemented a semantic segmentation-based skin lesion detection framework that is based on DeepLab V3 + with ResNet18-SelfAttention backbone architecture. The hyperparameters are initialized using Bayesian Optimization (BO), which improves the lesion segmentation workflow and accuracy. Design a new CNN architecture named Inverted self-attention Residual that includes the residual blocks in an inverted mechanism. Also, a ViT architecture is implemented to reduce the number of learnable parameters. Fused information of ISAR and ViT, called ISAwViT, and trained on the selected skin cancer dataset HAM10000 for lesion type classification. The trained model is interpreted in the testing phase using explainable AI techniques such as GradCAM and LIME (Local Interpretable Model-agnostic Explanations).
Proposed methodology
In this work, we propose a fully automated DL framework for skin lesion segmentation and classification from dermoscopy images. The HAM10000 skin cancer dataset is used in this work for experiments. The dataset consists of two parts: the segmentation part, which includes ground truths, and the classification part, which consists of labeled images of seven classes. The proposed segmentation task is based on semantic segmentation. The semantic segmentation task is based on the DeeplabV3 and ResNet-18 pre-trained models. We designed two CNN architectures in the classification phase and later fused them for the final classification. The fused model is tested on labeled images and then interpreted using XAI techniques. Figure 2 illustrates the complete framework for skin lesion segmentation and classification. The description of each step is given in the subsections below.

Proposed fusion-based deep learning architecture for skin cancer segmentation and classification using dermoscopic images.
Dataset collection
International Skin Imaging Collaboration (ISIC) HAM10000 39 “Human Against Machine with 10,000 training images” is among the largest datasets available to the general public through the ISIC repository. The dataset consists of 10,015 dermoscopy images to identify skin lesions with pigmentation. 40 The dataset consists of seven distinct classes: melanocytic nevus (6705 images), actinic keratosis (327 images), dermatofibroma (115 images), basal cell carcinoma (514 images), vascular lesions (115 images), benign keratosis (1099 images), and melanoma (1113 images). A certain number of images are assigned to each class. Male skin lesions are shown in 54% of the collection's images, while female skin lesions are in 45%. This challenging-to-classify dataset comprises many skin lesion images with substantial intra-class and low inter-class variation, which increases the risk of misclassification. A few sample images for the classification task are shown in Figure 1, and ground truth images for the segmentation task are illustrated in Figure 3. For the segmentation task, a total of 10,015 images are available from experts.

Sample original and ground truth images of dermoscopic images.
Dataset augmentation
To improve the robustness of the CNN design and reduce bias in the dataset, the best approach for handling significant class imbalances is to use minority oversampling. The DL models performed better using large-scale available datasets. The selected dataset is highly imbalanced in this work, as presented in Table 1. To improve the robustness of the design CNN model and reduce bias in the dataset, we first performed a 50–50 train-test split on the original 10,015 images, then applied augmentation exclusively to the training set (5008 images), expanding it to 49,881 images, while the test set (5007 images) remained completely unseen and unaugment to ensure valid performance evaluation without data leakage. The images in each class are presented in Table 1, which includes the original images, training images (50%), testing images (50%), and training images after augmentation. A 50-50 train-test split was chosen to maintain a substantial test set (5007 images) for robust evaluation, given the severe class imbalance in HAM10000. This approach ensures adequate representation of minority classes (e.g. dermatofibroma, with only 115 total images) in both training and testing. Additionally, we employed 5-fold cross-validation to validate model robustness and ensure reliable performance assessment comparable to literature standards.
Summary of the selected HAM10000 dataset for the classification process.

In the augmentation process, we first performed contrast stretching through Bi-Histogram Equalization (BiHE)
41
and then applied three mathematical operations: left flip, right flip, and a 90-degree rotation. Mathematically, these mathematical operations are defined as follows: Consider the notation 



Visual illustration of the data augmentation process.
Proposed lesion segmentation task
In this work, we presented a hybrid skin lesion segmentation architecture based on semantic segmentation. The DeepLab V3 + architecture is implemented using the ResNet18-SelfAttention CNN, trained on 70% of the training images, which consist of the original and corresponding ground-truth images. After training, testing was performed, and the resulting images were converted to binary for comparison with the ground-truth images.
Improved DeepLab V3 architecture
An enhanced version of the Deeplabv3 architecture is called DeepLab V3+. 42 It includes a more effective and efficient decoding module, leading to better performance in semantic segmentation and feature fine-tuning. Generally, a pre-trained CNN serves as the basis for DeepLab V3. Well-liked choices include ResNet, MobileNet, and Xception. This backbone network is responsible for extracting high-level information from an input image. ASPP (Atrous Spatial Pyramid Pooling) is a key component of DeepLab V3, allowing the model to combine local and global context information effectively. Next, this module uses global average pooling in conjunction with three simultaneous 3 × 3 convolutions with dilation rates of 6, 12, and 18 to capture high-level semantic information. The low-level features from the backbone network's input layer are downsampled via 1 × 1 convolutions by the decoder and mixed with the encoder's high-level data. After performing multiple 3 × 3 convolutions to extract spatial information from the feature maps, accurate modification of the target objects’ limits is achieved through bilinear up-sampling.
Optimized backbone ResNet18-SelfAttention model
The optimized ResNet18-SelfAttention CNN model is used in this work as the backbone for DeepLab V3 + . ResNet-18 is selected as the main network for its strong feature-extraction capabilities. The network has been updated to include a self-attention mechanism for deeper information extraction. A multi-scale feature fusion-enabling dual-path, dual-feature pyramid structure is provided as an expansion of the DeepLab V3 + architecture. This technique effectively uses feature maps produced by the backbone network at different scales. ResNet's residual structure with identity mapping, in contrast to Xception's architecture, enables smooth gradient propagation from shallow to deep layers. Using this residual architecture to train very deep neural networks enhances their ability to extract features and capture the nuances of the input data. The ResNet model, which integrates residual connections, helps better preserve attributes across different scales in the later layers. The Residual Network (ResNet) architecture, which includes residual connections to address vanishing gradients in very deep neural networks, is widely used in the ResNet-18 variant. Initially, we downsampled the images in the HAM10000 dataset from 650 × 450 to 240 × 240 pixels to prepare them for segmentation. The 240 × 240 resolution was chosen to balance computational efficiency with preservation of diagnostically critical dermoscopic features (pigment networks, border irregularities, color variations), which remain distinguishable at this resolution due to the 10×–20× magnification of dermoscopic imaging. The model depth ranges from 64 to 512, yielding a total of 512 features for the segmentation task.
Bayesian optimization
In the training of the Resnet18 architecture as a backbone on the selected dataset, we opted for BO
43
for the hyperparameter tuning. Typically, hyperparameters are selected manually or through literature review; however, this approach is not effective for complex datasets in segmentation tasks. BO allows us to optimize specific learning parameters, such as the initial learning rate, momentum, batch size, and dropout rate. Mathematically, the BO is defined as follows: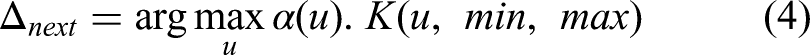
Where, 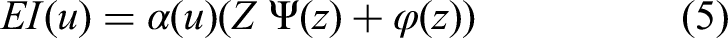

Where
Fine-tuning and training
As mentioned above, we fine-tune the learning rate, momentum, batch size, and dropout value of the ResNet architecture during the training for the lesion segmentation task. In the training phase, the model relied on BO; hence, during this validation process, it was evaluated using mean absolute error (MAE).
Where
The visual process of lesion segmentation training and testing is shown in Figure 5. This figure shows that the proposed ResNet18-SelfAttention CNN architecture is trained on the training images and their corresponding ground-truth images. In the training phase, the hyperparameters are tuned using BO. Following this, the testing process is conducted, where the test image is fed into the network, yielding a semantic segmentation image. This image is converted to binary and compared with the ground-truth image, which in turn displays the output DeeplabV3 image. Furthermore, the resulting image is applied to the original image, and the proposed segmented and ground-truth images are mapped to the final Dice score and Jaccard index.

Proposed lesion segmentation task using ResNet-SelfAttention architecture.
Skin lesion classification task
This section explains the proposed lesion segmentation task, which uses information fusion from DL architectures. The proposed fused architecture classifies skin lesions into appropriate classes, such as melanoma, benign keratosis (bkl), benign, and a few others. The fused CNN architecture combines a CNN with a Self-Attention module and a ViT. The detailed description of this architecture is given as follows.
Modified vision transformer
A neural network model called ViT uses the transformer architecture to turn image inputs into feature vectors. 44 The head and the backbone are the two essential components of the ViT. The network's encoding process places restrictions on the backbone. After receiving the input images, the backbone creates an output feature vector. The prediction scores are produced by the head using the encoded feature vectors.
ViT introduced transformers into vision tasks by splitting input samples into several patches that were tokenized into u tokens. Learnable positional embeddings are applied to each token to capture 2D relations among image patches while preserving positional information. Mathematically, the tokens and positional embeddings are defined as follows:

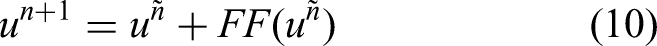

Visual architecture of vision transformer for skin lesion classification.
Proposed inverted self-attention architecture
An inverted residual block is one of the most important components for building effective CNNs. Inverted residual blocks begin with fewer channels than regular residual blocks, which increases the number of channels at the beginning of the block. A depth-wise convolutional layer is added to efficiently collect and process data. Because it reduces processing costs while preserving and improving the network's representational capacity, inversion is a common option in contemporary DL architectures for object identification and image classification.
We presented an innovative, inverted self-attention-based architecture in this study. The term “Inverted Self-Attention” in our architecture refers to the integration of multi-head self-attention mechanisms within inverted residual blocks, where attention is computed at the expanded feature dimension rather than at the input or output stages. Unlike standard approaches that apply attention after residual connections (e.g. MobileNetV2 with post-block SE attention) or before expansion (e.g. Squeeze-and-Excitation networks), our method embeds self-attention at the peak of channel expansion within each inverted block, allowing the attention mechanism to operate on richer feature representations (e.g. 512 channels) rather than compressed inputs (e.g. 64 channels). This differs from MobileViT and EfficientNet variants, which append attention modules as separate layers; our inverted self-attention fuses spatial attention computation directly within the depthwise convolution stage of inverted residuals, enabling simultaneous local feature extraction and global context modeling in a single, unified block with fewer parameters. With an input size of 224 × 224 × 3, the suggested network consists of five parallel and one series inverted block. Batch normalization is the starting point of the first block, which consists of two parallel blocks of convolutional layers, RELU activation, and a grouped convolution layer with a depth size of 16, a kernel size of 3 × 3, a stride of 1 × 1, and the same padding. The second parallel block begins with batch normalization, followed by two convolution layers and RELU activation. It is then attached to a grouped convolution layer with a depth size of 32, a stride of 1, and a kernel size of 3 × 3. The third block starts with batch normalization and has ten layers: two convolutional layers, two RELU activations, two batch normalizations, and a grouped convolutional layer. The 3 × 3 convolutional layer kernel has a depth of 64 and a stride of 1. This architecture's fourth block comprises 3 × 3 convolution layers with batch normalization and RELU activation. It is also connected to the grouped convolution layer, which has a depth size of 128 and a kernel size of 3 × 3 with the same padding. The fifth block design starts with RELU activation and includes a 3 × 3 convolution layer with batch normalization, followed by 256-depth convolution layers with a 3 × 3 kernel size and a 1 × 1 stride. The final and fifth series of block architecture begins with RELU activation. It is connected to one convolution layer, one batch normalization layer, and a grouped convolution with a stride of 1 × 1, a kernel size of 3 × 3, and a depth of 512. The inverted self-attention mechanism operates through a three-stage computational process within each residual block. First, the inverted expansion phase uses 1 × 1 convolutions to expand the number of channels from the input (e.g. 16→64), in contrast to standard residual blocks, which compress features. Second, depthwise separable convolutions efficiently extract spatial features along the expanded dimension. Third, before the residual addition, the self-attention module computes query (Q), key (K), and value (V) matrices through 1 × 1 convolutions on the expanded features, generating attention weights via softmax(QK^T/√d_k) that are then applied to V, allowing the network to selectively emphasize discriminative lesion patterns. The attention-weighted features are then projected back via a 1 × 1 convolution and added to the input via a residual connection, preserving gradient flow while incorporating global context. This differs fundamentally from post-processing attention by enabling simultaneous local feature extraction (via depthwise convolution) and global context modeling (via self-attention) within the same residual unit. In the fused architecture, feature flow operates as follows: dermoscopic images (224 × 224 × 3) are processed independently through ISAR and ViT pathways until their penultimate layers. ISAR produces spatial feature maps (7 × 7 × 512) that retain local texture and boundary information through its inverted residual hierarchy, while ViT generates patch-based token embeddings (196 tokens of dimension 768) that capture global lesion morphology via multi-head self-attention across all image patches. At the fusion point, ISAR features are flattened to 1D vectors (25,088 dimensions) and ViT class tokens (768 dimensions) are extracted, then concatenated via depth-wise concatenation, producing a combined feature vector (25,856 dimensions) that maintains independent gradient paths—gradients backpropagate separately through ISAR and ViT branches without interference. This architectural design ensures ISAR's local discriminative patterns (edges, texture, color variations) and ViT's global semantic understanding (overall lesion shape, spatial relationships) are jointly leveraged without the gradient dilution typical of early feature fusion strategies.
To transform the 2D data into 1D, a self-attention layer and a flattening layer are added. The output of the self-attention layer is routed to a fully connected layer, which is then connected to a classification layer and a softmax activation. Mathematically, the attention map is generated as:
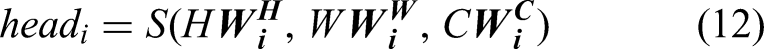

The designed architecture comprises 81 layers, including 43 convolutional layers. The facts about this architecture that make it significantly different from prior methods are as follows:
Unlike traditional ResNet expansion used in previous studies, this architecture utilizes inverted residual blocks that start with fewer channels and expand, thereby improving computational efficiency. Attention mechanisms are embedded within residual blocks rather than applied as post-processing. Strategic use of grouped convolutions reduces parameters while maintaining representational capacity.
Detailed architecture is shown in Figure 7. In addition, the model has 5.3 million trainable parameters.

Proposed inverted self-attention architecture for skin lesion classification.
Unlike existing hybrid architectures, ISAwViT introduces several distinctive design choices. TransUNet employs skip connections between the encoder and decoder in the standard U-Net architecture, whereas our approach uses depth concatenation at the network level to preserve independent gradient paths. Swin-Transformer-based models use shifted-window attention with hierarchical feature maps but require significantly more parameters (88M + for medical imaging variants). Standard CNN-ViT hybrids typically apply attention as a post-processing layer after convolutional feature extraction, whereas ISAwViT embeds self-attention directly within inverted residual blocks, enabling simultaneous local and global feature learning. Furthermore, our inverted residual mechanism (expanding from fewer channels) contrasts with TransUNet's standard residual expansion, reducing computational overhead while maintaining representational capacity.
Proposed fused architecture
We fused the deep models in this subsection using a depth concatenation layer. Network fusion aims to obtain richer information about skin lesion types, enabling accurate classification into relevant classes. The implemented ViT architecture has 140 layers, whereas the proposed inverted self-attention architecture has 81 layers. Also, this architecture is lighter than ViT. However, the ViT often outperforms the other due to its deeper architecture, though at the expense of greater computational complexity. To address this drawback, we combined it with a lightweight inverted self-attention architecture, yielding a total of 10.4 M trainable parameters.
To remove the impact of erroneous results from individual models, the final three layers are removed from both networks, and the architectures are merged using a depth-wise concatenation layer, thereby strengthening the network. To transform the 2D data into 1D, a flatten layer is introduced to the network after fusion. Finally, fully connected softmax and classification layers are added to complete this network. Figure 8 depicts the complete fused architecture of skin lesion classification.

Proposed FusedNet CNN architecture for skin lesion classification.
Results and discussion
Experimental setup
The experimental procedure for the proposed skin lesion segmentation and classification is presented in this section. The HAM10000 dataset was used for segmentation and classification experiments. The dataset details are given in the section “Dataset Augmentation.” In the first phase, we trained the proposed models for both segmentation and classification. In the segmentation phase, we passed 70% of the images, along with their ground-truth labels, to the model, and the remaining 30% were used for testing. In the classification training, 50% of the images were used for training and 50% for testing. The recall rate (TPR), accuracy, time, precision rate (PPV), and sensitivity rate were calculated for each classifier during the evaluation procedure. All experiments were conducted using MATLAB 2024b on a system equipped with a 20 GB graphics card (A4500), 128 GB of RAM, and an Intel(R) Core(TM) i5-7200U CPU operating at 2.50 and 2.7 GHz.
Proposed segmentation results
The proposed segmentation results for the HAM10000 dataset are presented in Table 2 and Figure 9. In Table 2, the HAM10000 training results are reported for different epochs. Each time, training accuracy and error are noted. As mentioned in the section “Experimental Setup”, 70% of images are used for training the segmentation task, and 30% are used for testing. Hence, based on the results in this table, it is observed that increasing the epoch value gradually improves training accuracy. For example, 10 epochs yielded an accuracy of 87.56%, which improved to 88.24% after 15 epochs. The best accuracy is achieved after 50 epochs, at 93.97%, with an error rate of 6.03%. After that, we tried more epochs, such as 55 and 60, but accuracy did not improve; the results were 93.84% and 93.90%, respectively. Hence, 93.97% accuracy is the best in the training phase for the proposed segmentation model.
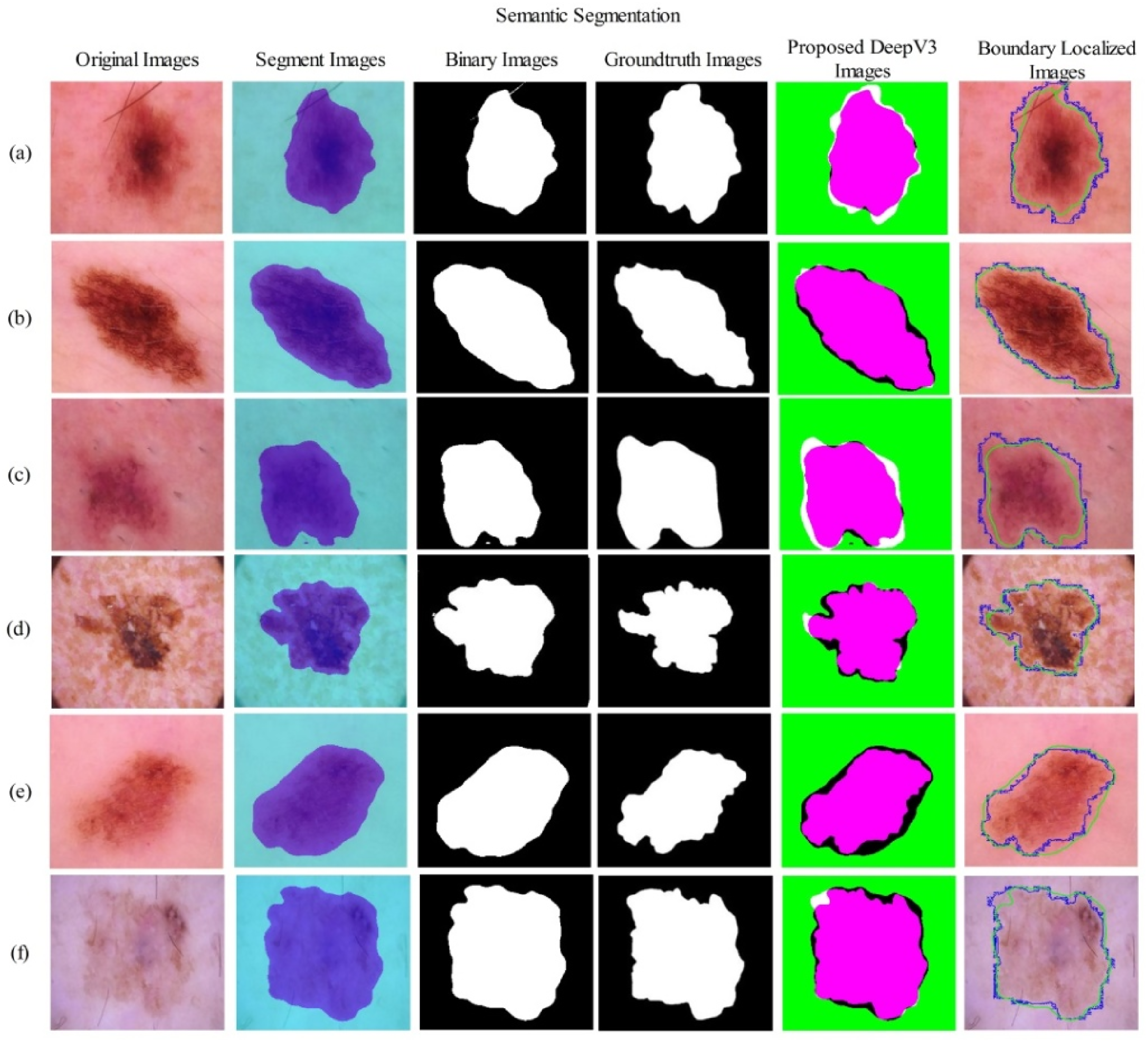
Proposed lesion segmentation results using testing images of the HAM10000 dataset.
Training accuracy of the proposed segmentation model using the HAM10000 dataset.

Bold values show the most significant results.
After that, we tested the proposed trained model on testing images (30% images) and computed Dice Score, Jaccard Index, and Jaccard Distance.
Table 3 presents the test results, showing that the dice score is 95.16%, the Jaccard distance is 9.24, and the Jaccard index is 90.76%. Furthermore, Figure 9 shows the segmentation region and labeling produced by the proposed architecture.
Proposed testing lesion segmentation results for the HAM10000 dataset.

Proposed FusedNet testing classification results
This section presents the classification results of the proposed FusedNet model on test data. The trained model presented in the section “Ablation study 4: evaluate pre-trained models for classification” is utilized to extract deep features. In the feature extraction phase, we selected a depth-concatenation layer. The extracted features are passed to the neural network classifiers, and the accuracy, precision, sensitivity, F1-Score, and test time are obtained. 5-fold cross-validation is performed to assess the model's robustness, and the results are presented as standard deviations across all metrics.
Five different neural network classifiers, including narrow neural network (NNN), medium neural network (MNN), wide neural network (WNN), bi-layered neural network (BNN), and tri-layered neural network (TNN), are used to evaluate the model's performance. Each classifier has its distinct properties that affect the feature processing and the model's complexity level. The NNN classifier contains one hidden layer with 10 neurons and provides baseline performance at minimal computational cost. However, it is better suited to linearly separable features due to its limited ability to transform features. On the other hand, the MNN classifier consists of two hidden layers with 25 and 10 neurons, respectively. It offers balanced complexity with moderate feature transformation and provides enhanced non-linear feature mapping as compared to NNN. WNN also has a single hidden layer with 100 neurons, which supports parallel feature processing. Because it has the maximum feature representation capacity within a single layer, it can capture diverse feature combinations, ultimately resulting in high classification accuracy. BNN and TNN contain two and three hidden layers, respectively, each with 10 neurons. Their sophisticated architecture allows for deep hierarchical feature extraction for complex pattern recognition. Note that BNN is better suited to sequential feature refinement and abstraction because of its balanced depth, whereas TNN performs better for multi-level feature abstraction due to its greater depth. However, this increased depth often comes with the risk of vanishing gradient.
Results on all these selected NN classifiers are presented in Table 4 in the form of numerical values. In this table, the WNN classifier obtained the maximum accuracy of 97.5%, with a standard deviation (std) of 0.6, while the computation time is 274.6 (sec). The precision rate of this classifier is 97.7%±0.6, the sensitivity rate is 97.8%±0.7, and the F1-score value is 97.7%±0.5, respectively. The narrow standard deviation on all metrics confirms the model's consistent performance across different data partitions. A confusion matrix illustrated in Figure 10 can further confirm these computed values. This figure shows that the nevi (nv) class has the highest error rate, whereas the Actinic keratosis (AKIEC) class has the lowest error rate for correct predictions. The rest of the classifiers, such as NNN, obtained an accuracy of 96.5 ± 0.8, the MNN classifier achieved 97.0 ± 0.9, BNN achieved 96.4 ± 0.5, and TNN obtained 96.4%±0.7, respectively. A detailed analysis of minority-class performance reveals critical clinical implications for rare lesion types. For melanoma (MEL), the most clinically dangerous class with 1113 images, our model achieved 96.2% sensitivity with 21 false negatives (3.8% miss rate). These misclassifications were predominantly melanocytic nevi (14 cases) and benign keratoses (7 cases), representing a moderate clinical risk, as both require follow-up monitoring. More concerning is dermatofibroma (DF) with only 115 images, achieving 88% precision and exhibiting seven false negatives (12.3% miss rate), the highest among all classes. These DF misclassifications were distributed as: four cases misidentified as basal cell carcinoma (BCC), two as melanocytic nevus (NV), and one as melanoma, the latter representing a critical diagnostic error. The severe class imbalance (DF represents only 1.1% of the dataset vs NV at 67%) directly contributes to this performance gap, as the model has insufficient exposure to DF's characteristic morphological patterns (central scar-like area, peripheral pigment network). Conversely, Melanoma's relatively better performance (96.2% sensitivity) despite being a minority class (11.1%) suggests that its distinct dermoscopic features (asymmetry, irregular borders, color variegation) are more discriminative. From a clinical risk perspective, the 3.8% melanoma false-negative rate could delay critical cancer diagnosis for approximately 4 patients per 100 cases, while the 12.3% DF miss rate, though less immediately life-threatening, may lead to unnecessary biopsies when misclassified as BCC or melanoma.

Confusion matrix of the proposed FusedNet architecture for HAM10000 dataset.
Proposed FusedNet architecture classification accuracy on the selected dataset HAM10000.

Bold values show the most significant results.
To validate the reliability of our accuracy measurements, we computed 95% confidence intervals using bootstrap resampling with 1000 iterations. The highest accuracy of 97.5% (95% CI: 96.9–98.1%) was achieved by the WNN classifier. The narrow confidence interval indicates its consistent performance across different data partitions. Some classifiers (e.g. BNN and TNN, both at 96.4%) show overlapping confidence intervals, indicating statistically similar performance. In contrast, WNN's non-overlapping interval with lower-performing classifiers confirms its statistically significant superiority. Hence, based on these results, it is observed that the fused features from ISAwViT are sufficiently rich and benefit more from parallel processing offered by WNN than deep hierarchical transformation. All reported computation times represent total processing time for the complete test set (5007 images). WNN classifier achieves 54.8 ms/image (18.2 FPS), NNN: 76.0 ms/image (13.2 FPS), MNN: 52.7 ms/image (19.0 FPS), BNN: 88.7 ms/image (11.3 FPS), TNN: 82.6 ms/image (12.1 FPS). These metrics demonstrate real-time capability suitable for clinical deployment (>10 FPS threshold).
Receiver operating characteristic curve analysis
The receiver operating characteristic curve analysis shown in Figure 11 demonstrates the discriminative performance of the neural network classifiers (WNN, NNN, MNN, BNN, and TNN), which were applied to features from our proposed ISAwViT. The WNN classifier achieved the highest area under the curve (AUC) of 0.985, indicating it better discriminates between skin cancer classes than the other classifier architectures. The NNN and MNN classifiers had AUC values of 0.965 and 0.970. In contrast, the BNN and TNN had nearly identical AUC values of approximately 0.963. All classifiers outperformed a random baseline (AUC = 0.500), indicating that ISAwViT feature extraction effectively captures the discriminative patterns required for skin cancer classification. WNN achieved the best overall performance, likely due to its wider architecture, including a hidden layer with 100 neurons that provides greater capacity to process the rich feature representation from the fused ISAwViT.

Receiver operating characteristic (ROC) curves comparison: neural network classifiers ISAwViT feature extraction on HAM10000 dataset.
Statistical significance analysis
To confirm statistical significance, we used paired t-tests to compare WNN (the best-performing classifier) with each other classifier, based on the 5-fold cross-validation results. Table 5 shows the mean difference, t-statistic, p-value, and significance (£) of each pair. Here, the mean difference represents the average performance difference between WNN and the compared classifier. At the same time, the t-statistic indicates the significance of this difference, with higher values indicating greater confidence that the difference is real and not random. On the other hand, the p-value indicates the probability that this difference occurs by chance. A p-value less than 0.05 means there is less than a 5% chance that the difference occurred by chance. This p-value defines the significance, where p < 0.05 means significant (*), p > 0.05 means not significant (-), and p < 0.01 means highly significant (**). The statistical results in Table 5 indicate that WNN outperforms NNN, with a 1.0% improvement and a p-value of 0.008. Additionally, it likely surpasses MNN, given the small but non-negligible difference. A 1.1% improvement and a p-value of 0.002 indicate that WNN is significantly superior to both TNN and BNN. These results demonstrate differences in performance among the classifiers and validate our conclusion that WNN performs much better and is statistically superior to the other classifiers.
Statistical significance analysis of classifier performance.

Ablation study 1: proposed inverted self-attention architecture results
We performed several ablation studies to validate the proposed classification architecture, FusedNet, including evaluating separate models, comparing them with pre-trained models, and cross-dataset testing. In the first ablation study, we selected the first inverted self-attention CNN architecture from FusedNet. The inverted self-attention CNN architecture is trained on the training set used in this work, and the extracted features are obtained from the self-attention layer. The extracted features are passed to the neural network classifiers, and evaluation protocols are noted in Table 6.
Proposed inverted self-attention architecture classification results.
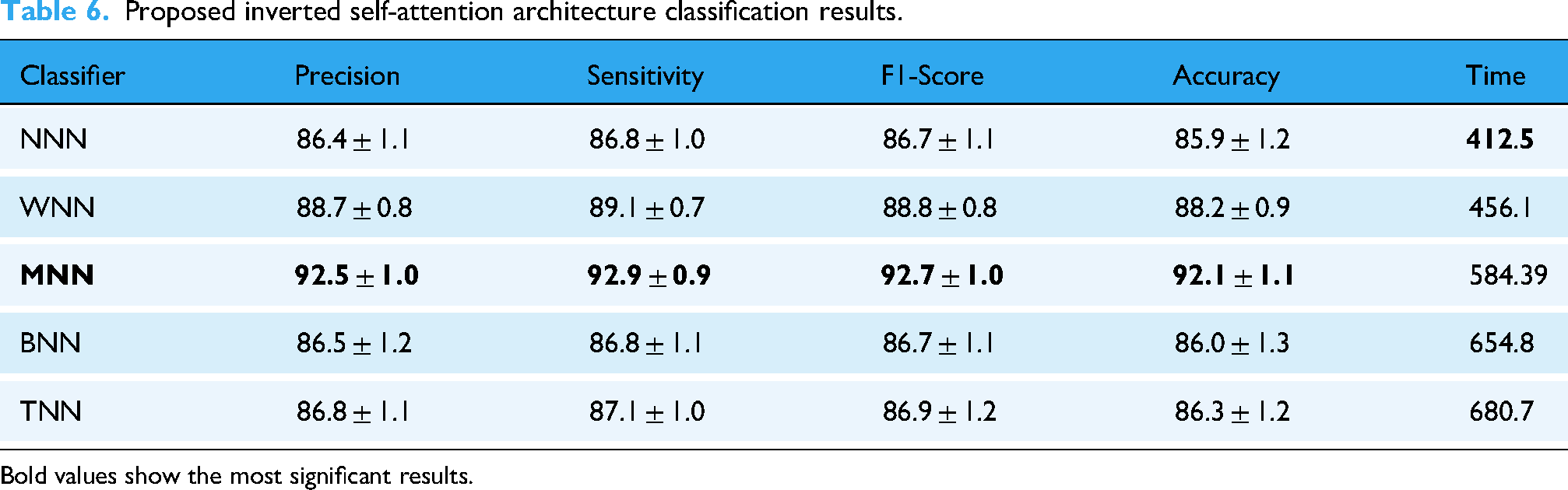
Bold values show the most significant results.
This table shows that the MNN performed better, with an accuracy of 92.1%±1.1, whereas the other classifiers achieved 85.9%±1.2, 88.2%±0.9, 86.0%±1.3, and 86.3%±1.2, respectively. Computational time is also noted for all classifiers, and the minimum noted time is 412.5 s for the NNN classifier.
Ablation study 2: proposed vision transformer architecture results
In the second ablation study, we selected a ViT model from FusedNet architecture and added a global average pool layer for the feature extraction. The extracted features are passed to the Neural Network classifiers, and the results obtained are presented in Table 7. This table shows that the WNN classifier achieves a maximum accuracy of 95.6% ± 0.7. There are other calculated measures, including a precision of 95.7%±1.0, a sensitivity of 95.9%±0.9, and an F1-score of 95.8%±1.2. The NNN classifier consumed a minimum time of 278.21 s for the execution, whereas the WNN classifier required 355.6 s. When comparing results such as accuracy, precision, and sensitivity, it is observed that the ViT architecture contributed more to the FusedNet's final accuracy.
Proposed vision transformer-based architecture classification results.

Bold values show the most significant results.
Ablation study 3: features-level fusion results
In this ablation study, we extracted features from the individual models discussed in ablation studies 1 and 2 and fused them in a serial manner. The fused features are passed to Neural Network classifiers, and the highest accuracy of 94.1%±1.0 is achieved with the BNN classifier, as listed in Table 8. The precision rate of this classifier is 94.5 ± 1.2, the sensitivity rate is 94.6 ± 0.6, and the F1-Score value is 94.5 ± 0.9%, respectively. Also, the remaining classifiers achieved 92.1 ± 0.9, 93.4 ± 1.0, 92.1 ± 1.1, and 91.9 ± 0.7% accuracy, respectively.
Features-level fusion results of self-attention and vision transformer architecture using the HAM10000 dataset.

Bold values show the most significant results.
Compared with ablation studies 1 and 2, this ablation study shows that the time increases after the fusion process; however, accuracy is slightly improved for NNN, MNN, BNN, and TNN. However, the highest accuracy is individually achieved by the ViT architecture. In addition, the proposed FusedNet achieves higher accuracy, precision, and sensitivity than the three experiments (see Table 4). Hence, ViT architecture contributes first to inverted self-attention. The fusion of these architectures at the network level increased the precision rate and reduced the model's computational time. The accuracy of the best classifiers and other performance measures can be further confirmed by the confusion matrix shown in Figure 12. From these confusion matrices, it is clear which key classes can be improved to enhance the classifiers’ accuracy and precision. This ablation study validates our choice of Network-level fusion over feature-level fusion. The reason lies in the fact that network-level fusion maintains independent gradient paths for both the ISAR and ViT components, preventing the gradient interference that can occur in early feature fusion. Moreover, ISAR and VIT operated on different principles. ISAR captures local relationships via inverted residuals, while ViT captures global dependencies via self-attention. Hence, late fusion allows each architecture to complete its specialized processing before integration.
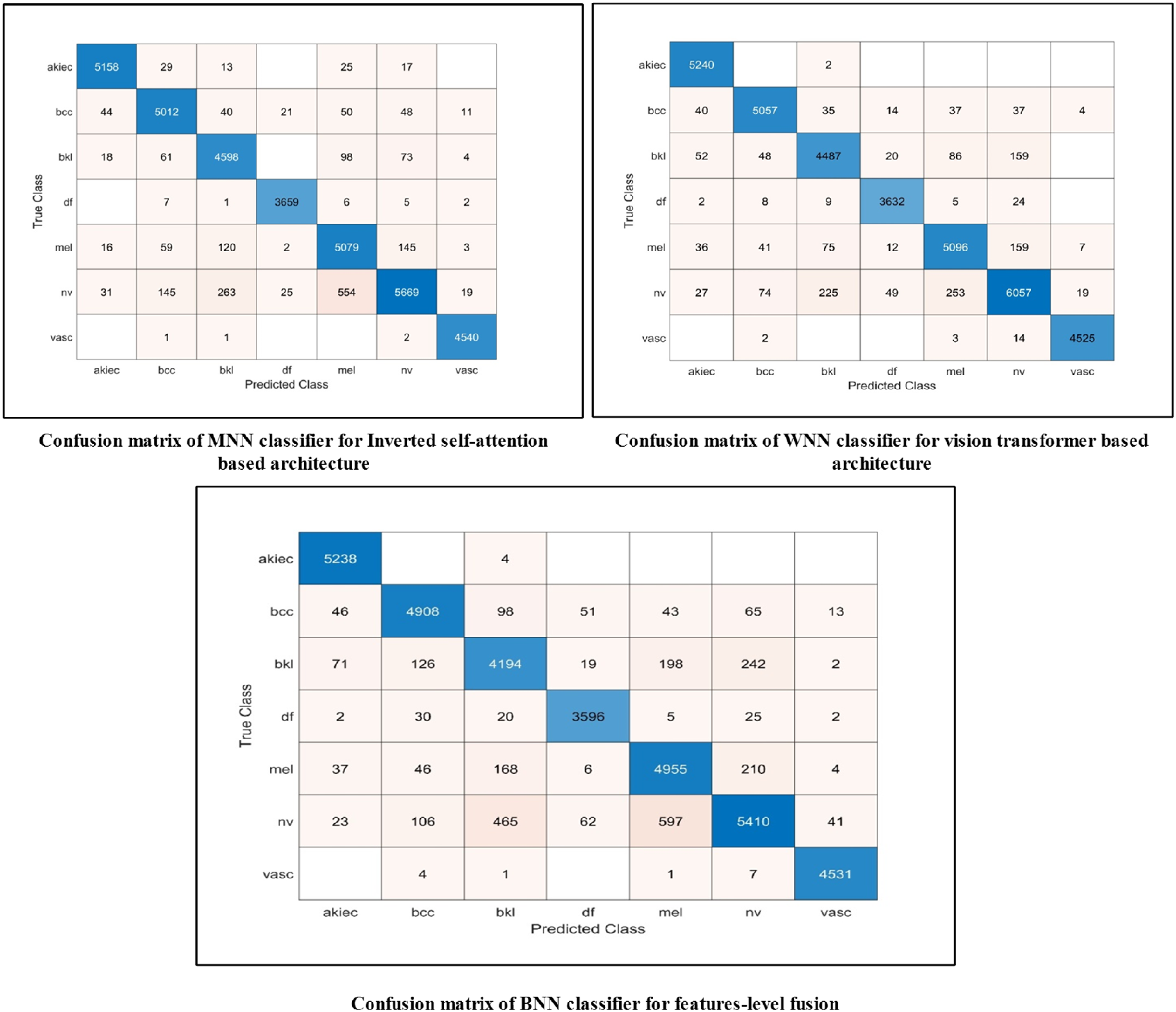
Confusion matrices of MNN, WNN, and BNN classifiers for ablation study 1, 2, and 3.
Ablation study 4: evaluate pre-trained models for classification
This ablation study compared the proposed FusedNet and individual CNN architectures with several pre-trained models. Figure 13 illustrates a visual illustration of this ablation study. In the first half of this image, we trained several pre-trained models, extracted features, and then passed the features to the WNN classifier to obtain accuracy. The proposed inverted self-attention architecture achieved 92.1% accuracy (as noted in Table 5), while the modified ViT obtained 95.6% (as indicated in Table 6), and the proposed FusedNet obtained 97.5% (as reported in Table 4). The pre-trained models, such as AlexNet, achieved an accuracy of 86.4%; VGG19 attained 85.9%, whereas InceptionV3 achieved the highest value of 92.5%. Overall, the proposed FuseedNet obtained improved accuracy for skin lesion classification.

Ablation study to evaluate pre-trained and proposed models based on accuracy and trainable parameters.
In the second half of this image, we compare the proposed models with pre-trained architectures based on the number of learnable parameters (in millions). The VGG19 model has the highest number of learnable parameters, 144 M, whereas the proposed Inverted Self-attention architecture contains only 5.3 M. Also, the proposed FusedNet model contains 10.4 M parameters, fewer than the other listed pre-trained models. Hence, the proposed FusedNet architecture achieves improved accuracy and a larger number of learnable parameters. To ensure a fair and rigorous comparison, all pre-trained baseline models (AlexNet, VGG19, InceptionV3, ResNet, etc.) underwent the same BO hyperparameter tuning process as our proposed architectures. Each baseline model was optimized using identical BO configurations (learning rate range [0.0001–0.01], optimizer selection {SGD, Adam, SGDM}, momentum [0.5–0.95], batch size {8, 16, 32, 64}, dropout [0.1–0.5]) with Expected Improvement acquisition function over 100 iterations, ensuring that performance differences reflect genuine architectural advantages rather than unfair hyperparameter configurations.
Dataset bias analysis and clinical deployment considerations
While HAM10000 is one of the largest publicly available dermoscopy datasets, several inherent biases must be acknowledged when assessing the model's real-world clinical applicability. The dataset exhibits notable demographic bias, with 54% male and 45% female representation, while lacking diversity in skin phototypes, predominantly Fitzpatrick types I–III (lighter skin tones), with limited representation of darker skin types (IV–VI). This imbalance may reduce diagnostic accuracy for underrepresented populations, as melanin-rich skin exhibits dermoscopic patterns our model has insufficient exposure to during training. Acquisition protocol bias is another critical concern. HAM10000 images were collected primarily using standardized dermoscopy equipment under controlled clinical conditions, creating a domain gap when applied to images captured with varying devices, lighting conditions, or imaging angles common in real-world practice. Our aggressive data augmentation (geometric transformations, contrast enhancement) partially addresses appearance variation but cannot fully replicate the diversity of acquisition protocols across different clinical settings, potentially leading to a 5–12% performance degradation due to domain shift. Class imbalance significantly impacts clinical utility, particularly for rare but critical conditions. Dermatofibroma (115 images, 1.1% of the dataset) achieves only 88% precision, compared with 97%+ for abundant classes like melanocytic nevus (6705 images, 67%). This disparity is clinically problematic as minority classes often represent diagnostically challenging cases requiring accurate detection. The 9% precision gap translates to higher false-negative rates for rare lesions, potentially delaying critical diagnoses. Despite augmentation increasing the training samples to 7359 for DF, the synthetic nature of the augmented images may not capture true morphological variability, leading to overfitting to augmentation artifacts rather than genuine clinical patterns. Annotation bias poses additional challenges. HAM10000 annotations were performed by dermatology experts, but inter-rater variability in lesion boundary delineation and classification labels (reported at 10–15% disagreement in dermoscopy literature) may propagate to our model's learned decision boundaries. Our XAI analysis revealed that in 33 of 127 misclassified cases, the model focused on image artifacts (hair, dermoscopic gel, ruler markings) rather than pathological features, suggesting the model may have learned dataset-specific spurious correlations. For clinical deployment, we recommend several mitigation strategies: (1) prospective validation on multi-institutional datasets with diverse demographics and acquisition protocols before clinical use, (2) ensemble approaches combining multiple models trained on different data distributions to improve robustness, (3) continuous learning frameworks that allow model adaptation to local clinical populations, and (4) mandatory dermatologist oversight with clear guidelines on model limitations for minority classes and underrepresented demographics. The estimated performance degradation of 3.3–5.7% on external datasets (Table 9) underscores the necessity of these precautions. Without rigorous external validation to address these biases, deployment could exacerbate healthcare disparities by underperforming for already underserved populations.
Comparison with existing techniques for skin lesion segmentation and classification.

Visualization of lime interpretation
To further evaluate the proposed FusedNet architecture, we utilized a trained model and applied visualization on the testing images using LIME 45 and GradCAM 46 visualization. Figure 14 represents the visualization results of the proposed FusedNet architecture using Explainable AI techniques. In the first part of this figure, we applied LIME interpretation and obtained output images with highlighted regions (shown in different colors). These colors highlight the most important lesion regions, and the extracted features inform the classification decision, such as whether a lesion is melanoma or another class. Similarly, in the second part of this image, GradCAM is applied. The GradCAM generates a heat map of the important region based on the extracted high-priority features, which aids decision-making. Hence, from these visual images, it is observed that the proposed architecture generated correctly highlighted lesion regions for the final classification. For the quantitative evaluation of XAI performance, we computed localization metrics by comparing XAI visualizations with annotated lesion boundaries. LIME achieved an average Union over Intersection (IoU) score of 76.3 ± 12.4% while GradCAM obtained a superior localization accuracy of 82.1 ± 9.7%. The inter-method agreement of 68.4% between LIME and GradCAM suggests moderate consistency between the techniques in identifying relevant image regions. Moreover, we quantify cases where the overlap between the actual lesion region and the highlighted area exceeds 70%, and the results show a success rate of 78.2% for GradCAM and 65.7% for LIME. These results indicate that explainability techniques provide meaningful interpretations, with GradCAM outperforming LIME.

Visualization of the proposed FusedNet architecture on test images.
Although the XAI visualizations indicate the model focuses on lesion areas, our analysis reveals some important limitations. Of the 127 misclassified instances, 78 are predicted incorrectly by the model, despite favorable XAI visualizations that correctly highlight the lesion area. Moreover, in another 33 cases, the model's attention drifts toward image artifacts (hair, dermoscopic masks) rather than the pathological features. This mismatch between model predictions and XAI visualizations indicates the need for careful interpretation of explainability outputs in clinical settings, as favorable visualizations do not always guarantee the right diagnosis.
Comparison with existing techniques
Lastly, we compare the proposed architecture's performance against several pre-trained models using segmentation accuracy (Dice score) and classification accuracy. A comparison is conducted in Table 9. In this table, He et al. 47 used the HAM10000 dataset in the experiments and achieved an accuracy of 87.2%. Srujan et al. 48 and Gururaj et al. 49 achieved an accuracy of 87.0% and 91.2%, respectively. The segmentation accuracy of the proposed architecture is 95.1%, which is higher than that of these techniques. Similarly, the proposed classification accuracy of the HAM10000 dataset is also compared with some existing techniques, as Rajesh et al. 50 and Islam and Panta 51 obtained an accuracy of 85.94% and 92.3%, respectively. The proposed architecture achieved 97.5% accuracy on the HAM10000 dataset, surpassing the recent state-of-the-art techniques.
Conclusion
In this work, we propose DL architectures for skin lesion segmentation and classification from dermoscopic images. The proposed work is based on two fundamental phases—lesion segmentation and lesion classification. In the lesion segmentation phase, we use a ResNet-18-SelfAttention network as the backbone of the DeepLab V3 + model. The new backbone aims to better learn lesion-region pixels and, in turn, produce improved segmentation output. The hyperparameters are initialized using BO, which significantly enhances the proposed model's learning process. In the classification phase, we proposed a FusedNet architecture that fuses two customized models: Inverted Self-Attention and Modified ViT. Fusing models at the network level increased accuracy, precision, and sensitivity, and reduced the number of learnable parameters. The experimental process was conducted on the HAM10000 dataset, and it achieved improved accuracy of 95.1% and 97.5% for segmentation and classification, respectively.
Despite the strong performance, several major limitations in this study need to be acknowledged and addressed in future work. One of these limitations includes aggressive data augmentation, which increases the sample size from 10015 to 49881 and induces the risk of overfitting. Because synthetic patterns do not correspond to actual clinical variations, this excessive augmentation may cause the model to learn synthetic artifacts rather than generalizable features. Moreover, the proposed ISAwViT architecture, with 10.4 million parameters, requires substantial computational resources, which limits its use in resource-constrained settings and in real-time clinical workflows. Furthermore, class imbalance appears as a consistent challenge for minority classes such as dermatofibroma (115 instances), a rare case as compared to other categories. It achieves a precision of 88%, significantly lower than that for abundant classes, which may lead to misdiagnosing rare but clinically important conditions. These limitations emphasize that the proposed ISAwViT has great segmentation and classification abilities. Still, it comes with a clear risk of overfitting, computationally expensive requirements, and underperformance of minority classes, which need to be addressed before clinical deployment. Another limitation is the comparison with outdated baseline architectures (AlexNet, VGG19, ResNet) rather than current state-of-the-art models. Future work will include comprehensive benchmarking against modern architectures such as EfficientNetV2, ConvNeXt, Swin Transformers, TransUNet, and recent hybrid models (CoAtNet, MaxViT) under identical experimental conditions to validate ISAwViT's competitive standing and architectural advantages against contemporary approaches. Additionally, our statistical significance analysis (Table 5) compares only neural network classifier variants rather than established state-of-the-art architectures such as InceptionV3, DenseNet, or EfficientNet. Future work will include rigorous paired statistical tests (t-tests, Wilcoxon signed-rank) comparing ISAwViT with competitive baselines to determine whether the observed performance improvements are statistically significant beyond experimental variation.
Footnotes
Author contributions
Junaid Aftab, Muhammad Attique Khan, Sobia Arshad, Shrooq Alsenan: conceptualization, software, methodology, original draft writeup, funding, supervision. Amir Hussain, Yongwon Cho, Yunyoung Nam: methodology, project administration, supervision, funding. All authors agree to submit this work in this reputed journal.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported through Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2026R506), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia. This work was supported by the National Research Foundation of Korea(NRF) grant funded by the Korea government(MSIT) (No. RS-2023-00218176) and the Soonchunhyang University Research Fund.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
