Abstract
Objectives
This study aimed to develop a disease-free survival (DFS) prediction model incorporating radiomics and intratumor heterogeneity (ITH) scores for locally advanced nasopharyngeal carcinoma (LANPC), and to establish an anti-epidermal growth factor receptor (EGFR) therapy risk model.
Materials and Methods
A retrospective analysis was conducted on 950 pathologically confirmed LANPC patients (training cohort: n = 632, including 81 receiving anti-EGFR therapy; test cohort: n = 318, including 47 receiving anti-EGFR therapy). All patients underwent 1.5 T MRI (T2-Weighted Imaging and contrast-enhanced T1-Weighted Imaging). Feature selection was performed using the Least Absolute Shrinkage and Selection Operator (LASSO) and support vector machine (SVM) survival algorithms. Subsequently, five predictive models were developed and compared: a Comprehensive Risk Model (CRM) integrating clinical features, ITH score, and radiomics score; an ITH-radiomics model (IRM); a standalone ITH model (ITHM); a standalone radiomics model (RM); and a clinical model (CM). Model performance was evaluated using the area under the curve (AUC) and the concordance index (C-index), and clinical utility was assessed with decision curve analysis. Finally, Kaplan-Meier analysis with the log-rank test compared survival between the model-defined risk groups. Additionally, the DeepSurv deep neural network was employed to simulate personalized treatment recommendations based on the patient's risk profile.
Results
With median follow-ups of 73 months (training) and 68.1 months (test), disease progression occurred in 34.2% (216/632) and 36.5% (116/318) of cases, respectively. The CRM achieved the highest C-index value for assessing DFS in patients with LANPC, with values of 0.829 and 0.760 in the training and test cohorts, respectively. Patients who met the DeepSurv treatment recommendations had better DFS.
Conclusion
The superior performance of the CRM supports its potential to enhance DFS prediction in LANPC and to inform anti-EGFR therapy selection.
Keywords
Introduction
Nasopharyngeal carcinoma (NPC) is an epithelial malignancy originating in the mucosa of the nasopharynx. NPC exhibits considerable geographical differences, with more than 70% of new cases occurring in Southeast and East Asia. 1 Owing to its onset location, approximately 70% of patients are already at a locally advanced stage of NPC at the time of diagnosis. 2 Although current mainstream treatments can increase the 5-year overall survival rate of patients with locally advanced NPC (LANPC) to more than 80%, approximately 20% experience recurrence. 3 Recurrence and metastasis are the main causes of NPC-specific mortality. 4 The current Tumor, Node, Metastasis (TNM) staging system for NPC is based primarily on anatomical features.5,6 In the 8th edition of the American Joint Committee on Cancer (AJCC) staging system, the 5-year survival rates of patients with clinical stages III and IVa were 83% and 71%, respectively. Thus, some patients, despite being at the same clinical stage, have different clinical outcomes. 5 The plasma level of Epstein-Barr virus (EBV) DNA is a well-established prognostic biomarker for NPC and reflects tumor burden. 6 However, it cannot fully capture the intricate spatial heterogeneity within the tumor. Most cancers are genetically heterogeneous. The selective pressure exerted by targeted therapies on tumors leads to the proliferation of resistant clones, ultimately resulting in cancer drug resistance. 7 Therefore, anatomical staging alone does not reflect tumor heterogeneity. An updated tool is required to stratify the risk of NPC. Additionally, based on the genetic heterogeneity of cancer, patients may have varying sensitivities to targeted therapies. An assessment tool is also required to evaluate whether patients can benefit from targeted treatments.7,8
In recent years, radiomics has significantly progressed in oncology research. Radiomics analyzes images and clinical information differently from traditional “semantic” features. It can decode images into quantitative features and capture information from image textures that is undetectable by the eye. 9 Image features can provide physiological information about cancer and can be transformed into actionable data through machine learning or deep learning methods, providing a basis for clinical diagnosis and assessment of survival status.10,11 Radiomics has demonstrated good stability and accuracy across various cancer types and imaging modalities.10,11 Intratumor heterogeneity (ITH) is a common feature of solid tumors 9 and is often caused by a combination of factors, such as vascular abundance, local metabolism, proliferative activity, and the uneven distribution of different cell populations and tumor tissues within different subregions. 12 While the TNM staging system provides a fundamental anatomical framework, it may not fully capture the complex biological heterogeneity inherent to tumors. Similarly, conventional radiomics approaches, which typically analyze the tumor as a whole, could overlook critical prognostic information encoded in tumor heterogeneity. To explore whether a more granular analysis could address these limitations, our study implemented a subregional assessment strategy. This involved partitioning the tumor region of interest (ROI) into multiple subregions, extracting features from each, and applying clustering techniques to identify recurrent phenotypic patterns. The resulting quantitative ITH descriptor was then integrated into our prognostic model to assess its potential to enhance predictive performance beyond conventional staging and radiomic methods.13–17
Therefore, the aim of this study was to construct a Comprehensive Risk Model (CRM) based on clinical features, ITH score, and radiomics score and to determine whether this model improved the assessment of DFS in patients with LANPC. A secondary aim was to leverage this model to identify which patients were most likely to benefit from anti-epidermal growth factor receptor (EGFR) therapy.
Materials and methods
Study cohort
This retrospective study was approved by the review boards of Centers A and B, and the requirement for written informed consent was waived. This study utilized datasets from both centers, with data collected from January 2014 to June 2019. Inclusion criteria for patients were as follows: (1) Patients with pathologically confirmed NPC. (2) Patients classified as locally advanced according to the 9th edition of the AJCC, 6 including clinical stages II and III. (3) Patients with available MRI scans, including axial T2-weighted imaging (T2WI) and contrast-enhanced T1-weighted imaging (T1C). (4) Availability of complete clinical, treatment, and follow-up data for DFS analysis. Exclusion criteria were as follows: (1) Poor image quality precluding accurate segmentation (e.g. significant artifacts, incomplete tumor coverage). (2) History of recurrent NPC or any prior malignancy. (3) Presence of distant metastases at initial diagnosis. (4) Incomplete treatment course or loss to follow-up immediately after treatment.
After screening 1105 patients, we excluded 155 cases that failed to meet the inclusion criteria. The remaining 904 eligible Center A patients were randomly allocated to training (n = 632, 70%) and testing (n = 272, 30%) cohorts. To enhance validation, we supplemented the testing cohort with 46 additional eligible patients from Center B, resulting in a final composite testing cohort of 318 cases (Center A: 272 + Center B: 46). This achieved a final training-to-test ratio of approximately 2:1 (632:318) (Figure 1).

Study workflow comprising (a) patient screening and selection process, and (b) analytical pipeline integrating clinical data with radiomic features (including intratumoral heterogeneity metrics) for risk prediction modeling and treatment recommendation system development.
Hematological and clinical features
The following patient data were collected from clinical medical records: marital status, age, sex, height, weight, history of hypertension, smoking history, drinking history, and pathological examination results. Blood test indicators were collected within 1 week before treatment, including white blood cell count, hemoglobin, platelet count, neutrophil count, monocyte count, lymphocyte count, total protein, albumin, alkaline phosphatase, lactate dehydrogenase, and EBV DNA. Additionally, the study included several inflammatory indicators: platelet-to-lymphocyte ratio (PLR) = platelets/lymphocytes; systemic immune-inflammation index (SII) = (neutrophils × platelets)/lymphocytes; systemic inflammation response index (SIRI) = (neutrophils × monocytes)/lymphocytes. 18
Clinical outcome and follow-up
Each patient underwent follow-up examinations every 3 months during the first 2 years after radiotherapy. Subsequently, follow-up was conducted every 6 months for years 2–5, and annually thereafter. Each examination included an assessment of clinical symptoms, NPC-related blood tests, MRI scans of the nasopharynx, fiberoptic nasopharyngoscopy, and chest and abdominal imaging. If patients experienced recurrence or metastasis, disease progression was confirmed and evaluated using nasopharyngoscopy, CT, MRI, ECT, positron emission tomography/computed tomography, or biopsy.
Image processing
Patients from both centers underwent imaging examinations and evaluations using a 1.5 T MRI (Magnetom Avanto, Siemens Healthcare, Germany) with a combined head and neck coil. For details of the MRI scanning sequences, please refer to Additional File 1. All images underwent N4 bias field correction and were resampled to a voxel size of 1 × 1 × 1 mm to reduce MR intensity inhomogeneity and errors caused by different voxel sizes. T2WI and T1C images were aligned using affine transformation. The 3D tumor regions were annotated by two radiologists (LL and CC, with 5 and 8 years of experience in head and neck tumor diagnosis, respectively). For disputed tumor regions, LL or CC consulted HH (with 17 years of experience in head and neck tumor diagnosis) for a decision. A randomly selected group of 100 patients was used to evaluate interobserver consistency. ITK-SNAP software (version 3.8.0; http://www.itksnap.org/pm.s/pm.s.php) was used for the analysis.
Extraction of radiomic score and ITH score
PyRadiomics (version 3.1; https://pyradiomics.readthedocs.io) was used to extract textural features from the tumor regions. For the ITH score, simple linear iterative clustering supervoxel segmentation was conducted by dividing the tumor ROI into roughly 100 uniform pixel regions, followed by feature extraction. Finally, regions with similar radiomic characteristics were clustered using a Gaussian mixture model. The number of clusters for intratumoral subregions was determined in an unbiased manner using the Bayesian Information Criterion, with the clustering range set from one to ten. 15
Feature dimensionality reduction and feature selection
Data were centralized and standardized using the “preProcess” function from the “caret” package. The “findCorrelation” function was then used for preliminary screening of texture and clustering features, removing highly correlated features. Finally, the features were incorporated into the Least Absolute Shrinkage and Selection Operator (LASSO) model to fit four types of features: T2WI radiomics score, T1C radiomics score, T2WI ITH score, and T1C ITH score. This study included 32 features, including clinical features, treatment regimens, anti-EGFR therapy (Additional File 2), pathological characteristics, blood biomarkers, MRI radiomics scores, and the ITH score.
Constructing machine learning models
Two representative machine learning algorithms—LASSO and support vector machine (SVM) for survival analysis—were used for the initial screening of all included features. Hyperparameter tuning was performed using a combination of grid search and manual adjustments. The top 15 features from each machine-learning model were intersected to form a set of common features. Five predictive models were constructed using multivariable Cox regression: a CRM incorporating clinical features, ITH score, and radiomics score; a combined ITH-Radiomics Model (IRM) based on both the ITH score and radiomics score; a standalone ITH Model (ITHM) using only the ITH score; a standalone Radiomics Model (RM) using only the radiomics score; and a Clinical Model (CM) based on T classification, clinical stage, treatment modality, targeted therapy, and EBV DNA status. The models were validated using a test cohort.
Constructing an evaluation model for anti-EGFR therapy
To balance baseline differences between patients undergoing and not undergoing anti-EGFR therapy, propensity score matching (nearest-neighbor matching) was performed within the training cohort. Specifically, the covariates for matching were those previously identified by the CRM, namely clinical features, the ITH score, and the radiomics score. The propensity score model derived from this matched training cohort was then applied to the test cohort, resulting in final matched cohorts of 243 patients for training and 141 for testing. The DeepSurv model was used to calculate the medication risk for each patient and to evaluate the anti-EGFR treatment plan. Hyperparameter tuning was performed using a combination of grid search and manual adjustments, focusing on whether newly diagnosed patients should choose an anti-EGFR regimen.
Statistical analysis
Interobserver consistency was evaluated using the Dice Similarity Coefficient. Continuous variables were summarized as mean and standard deviation (normal distributions) or median and interquartile range (non-normal distributions). Categorical variables were presented as frequencies and percentages. Baseline tables for the training and test cohorts were generated using the “CreateTableOne” function. A Cox model was used to identify the features of patients with LANPC and develop a risk model by calculating hazard ratios, 95% confidence intervals, and p-values. The “surv_cutpoint” function was used to divide patients into high-risk and low-risk groups. Kaplan-Meier survival analysis and log-rank tests were used to compare the differences in survival between high- and low-risk patients. The discriminative ability of each model was evaluated using the area under the curve (AUC) and the concordance index (C-index). The calibration was assessed visually with calibration curves. The bootstrap method was used to compare the C-index differences between the models (p < 0.05). The clinical net benefit of each risk model was assessed using decision curve analysis. Statistical analyses were performed using R version 4.3.3. Python version 3.7.1 was also used.
Results
Patient characteristics and data quality control
The median age of the 950 patients was 45.5 years [IQR: 37–53 years], and 73.9% were men. The median follow-up time for the training cohort was 73 months [IQR: 38.3–95 months], and for the test cohort, it was 68.1 months [IQR: 29.4–92 months]. There were 216 (34.2%) and 116 (36.5%) cases of disease progression in the training and test cohorts, respectively. As detailed in Table 1, the baseline characteristics showed generally balanced distributions. In a random sample of 100 cases, the Dice Similarity Coefficients for the ROI drawn by the two observers were all greater than 0.8 (Figure S1
Comparative analysis of clinical-radiological features between the training and test cohort.
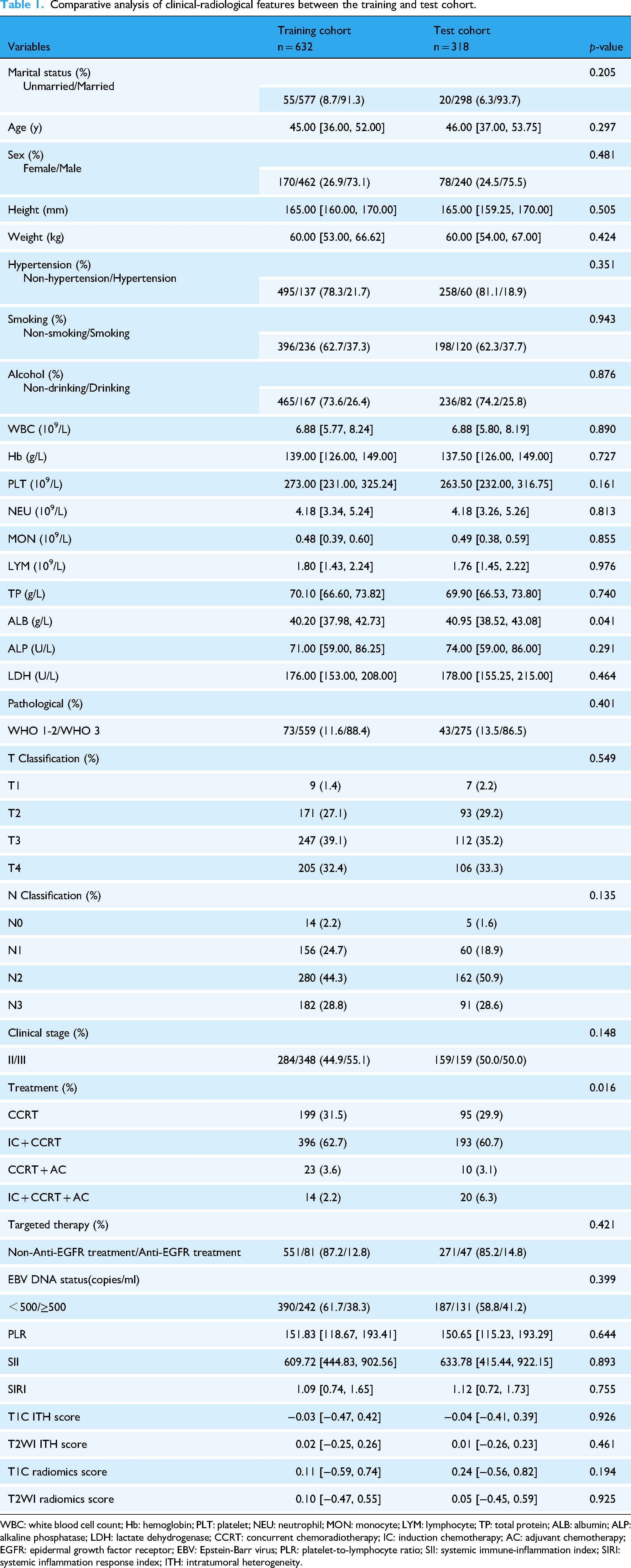
WBC: white blood cell count; Hb: hemoglobin; PLT: platelet; NEU: neutrophil; MON: monocyte; LYM: lymphocyte; TP: total protein; ALB: albumin; ALP: alkaline phosphatase; LDH: lactate dehydrogenase; CCRT: concurrent chemoradiotherapy; IC: induction chemotherapy; AC: adjuvant chemotherapy; EGFR: epidermal growth factor receptor; EBV: Epstein-Barr virus; PLR: platelet-to-lymphocyte ratio; SII: systemic immune-inflammation index; SIRI: systemic inflammation response index; ITH: intratumoral heterogeneity.
Feature selection and model development
Following data preprocessing, the correlation matrix indicated a strong correlation (≥0.8) between neutrophil count and both white blood cell count and SIRI, leading to the removal of the neutrophil feature and retention of 31 features for subsequent analysis (Figure S2). From this set, the top 15 most important features were identified from both the LASSO and SVM-survival models (Figure 2). The intersection of these two lists yielded a consensus set of 10 features. These 10 features were subsequently entered into a multivariable Cox regression analysis to identify independent prognostic factors for DFS. The final model retained the following significant variables: T Classification, Clinical Stage, Treatment, Targeted Therapy, EBV DNA Status, T1C ITH Score, T1C Radiomics Score, and T2WI Radiomics Score. Variables with p-values <0.1 in screening were retained for subsequent predictive modeling (Table 2, Figure 3, Supplemental Table 1).
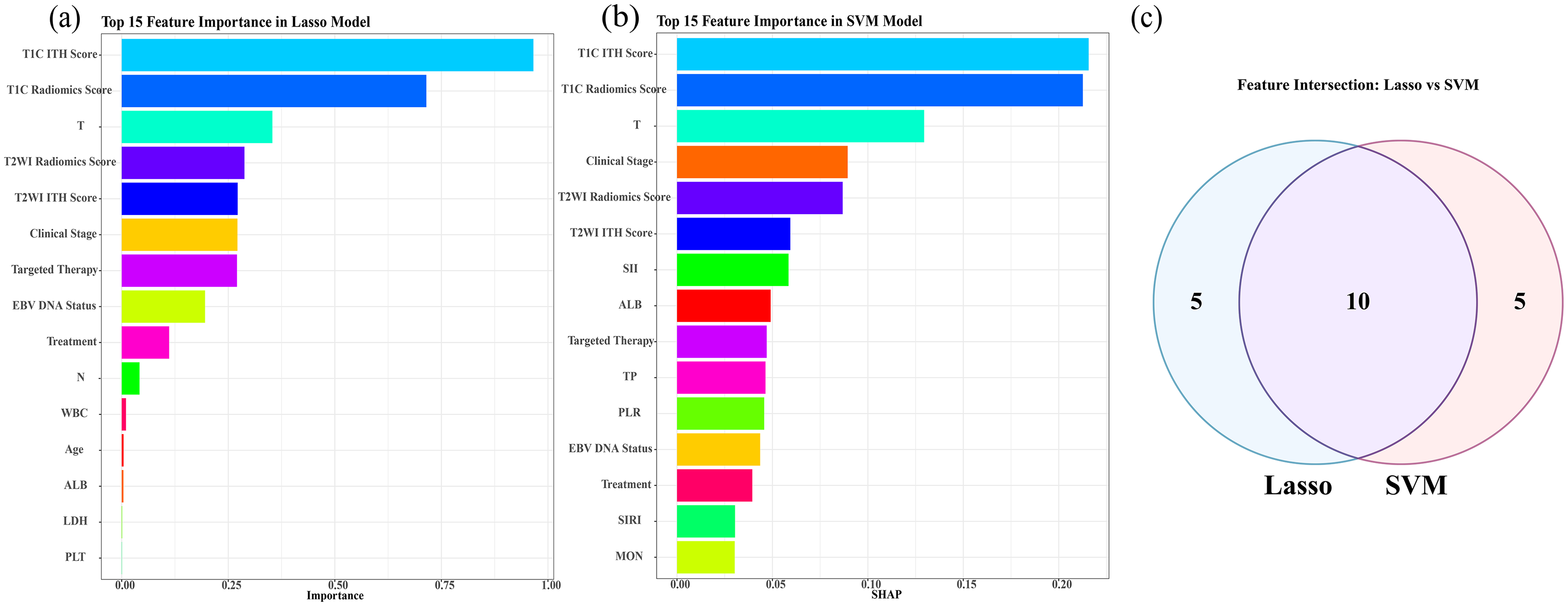
Feature selection analysis. (a) Top 15 most important features identified by LASSO regression. (b) Top 15 most important features selected by SVM. (c) Venn diagram illustrating the ten consensus features shared between the LASSO and SVM survival analysis methods. Abbreviations: LASSO: Least Absolute Shrinkage and Selection Operator; SVM, support vector machine for survival analysis; ITH, intratumoral heterogeneity; WBC, white blood cell count; LDH, lactate dehydrogenase; PLT, platelet count; SII, systemic immune-inflammation index; TP, total protein; PLR, platelet-to-lymphocyte ratio; SIRI, systemic inflammation response index; ALB, albumin; EBV DNA, Epstein-Barr virus deoxyribonucleic acid.

Multivariable Cox regression analysis of DFS in LANPC. (a) Forest plot of prognostic factors. (b) Nomogram for 1-/3-/5-year DFS prediction. *In both panels, asterisks indicate statistical significance (*< 0.05, ** < 0.01, *** < 0.001). Abbreviations: DFS, disease-free survival; LANPC, locoregional nasopharyngeal carcinoma; ITH, intratumoral heterogeneity; ALB, albumin; ITH, intratumor heterogeneity; EBV DNA, Epstein-Barr virus deoxyribonucleic acid.
Final multivariable cox regression model for DFS in LANPC after backward stepwise selection.

Note: The final model was derived from an initial multivariable model containing all candidate predictors, by applying a backward stepwise selection procedure with a retention criterion of p < 0.1.
DFS: disease-free survival; HR: hazard ratio; CI: confidence interval; EBV DNA: Epstein-Barr virus deoxyribonucleic acid; ITH: intratumoral heterogeneity; LANPC: locally advanced nasopharyngeal carcinoma.
Model performance and clinical validation
The tROC curves (Figure 4

Time-dependent ROC curves of the CRM for predicting 1, 3, and 5-year DFS in patients with LANPC. (a) Training cohort. (b) Test cohort. Abbreviations: ROC, receiver operating characteristic; CRM, Comprehensive Risk Model; DFS, disease-free survival; LANPC, locoregional nasopharyngeal carcinoma.

Decision curve analysis comparing the clinical utility of different models for predicting DFS in LANPC. (a) Training cohort. (b) Test cohort. Abbreviations: CRM, Comprehensive Risk Model (integrating clinical features, radiomics score, and ITH score); IRM, Integrated Radiomics-Intratumoral Heterogeneity Model; ITHM, Intratumoral Heterogeneity Model; RM, Radiomics Model; CM, Clinical Model; DFS, disease-free survival; LANPC, locoregional nasopharyngeal carcinoma.
Bootstrap comparison of C-index differences among predictive models in training and test cohorts.

C-index: concordance index; CRM: comprehensive risk model (integrating clinical features, radiomics score, and ITH score); IRM: integrated radiomics-ITH model; ITHM: intratumoral heterogeneity model; RM: radiomics model; CM: clinical model; CI: confidence interval.
Time-dependent AUC and C-index performance of predictive models in training and test cohorts.

AUC: area under the receiver operating characteristic curve; C-index: concordance index; CI: confidence interval; CRM: comprehensive risk model (integrating clinical features, radiomics score, and ITH score); IRM: integrated radiomics-intratumoral heterogeneity model; ITHM: intratumoral heterogeneity model; RM: radiomics model; CM: clinical model.
The CRM risk score stratified patients into high- and low-risk groups. Survival analysis demonstrated that in both the training and test cohorts, patients in the low-risk group had a significantly longer DFS than those in the high-risk group (Training cohort: p < 0.001; Test cohort: p < 0.001) (Figure S4).
Treatment recommendation efficacy
After propensity score matching, the standardized mean differences of the covariates were all less than 0.2 (Supplemental Table 2), indicating that the balance quality of the covariates in the matched data was acceptable. To evaluate the association between model-derived recommendations and patient outcomes, we employed the DeepSurv algorithm to simulate treatment assignments for anti-EGFR therapy. Retrospective analysis compared patients whose treatment was consistent with the model's recommendation (i.e. receiving anti-EGFR therapy when recommended, or not receiving it when not recommended) to those with inconsistent treatment. The results showed that the consistent group had significantly superior survival DFS (p < 0.001). This survival advantage remained consistent across subgroups of patients who either received (p = 0.01) or did not receive (p = 0.006) anti-EGFR therapy (Figure 6).

(a) KM curves of patients who adhered to or deviated from Deepsurv model treatment recommendations; (b) KM analysis of DFS in anti-EGFR treated LANPC patients, stratified by consistency with the Deepsurv model recommendation, showed a significant association between model-adherent recommendations and longer DFS; (c) KM curves comparing standard treatment patients who followed recommendations (and were anti-EGFR insensitive) versus those who were potentially anti-EGFR sensitive but did not receive treatment. Abbreviations: EGFR, epidermal growth factor receptor; KM, Kaplan-Meier.
Discussion
This study developed and validated a CRM that integrates clinical features, radiomics, and ITH scores for predicting DFS in LANPC. The CRM demonstrated superior prognostic performance compared to models relying on clinical features or radiomics score alone, underscoring the incremental value of capturing tumor heterogeneity. Our findings demonstrate that multimodal data integration provides a more nuanced approach for personalized prognosis.
Tumor heterogeneity is one of the reasons for treatment failure and disease recurrence. 7 Tumor heterogeneity is prevalent in solid tumors, and the assessment of heterogeneity includes genetic heterogeneity, histological heterogeneity, and imaging heterogeneity. The complexity and high cost of whole-genome sequencing of tumor tissues, the difficulty in obtaining tumor tissue samples, and the cumbersome process of assessing heterogeneity all limit the implementation of these evaluation methods. 19 The assessment of tumor heterogeneity through imaging is a new measure in tumor imaging evaluation in recent years, adding a new dimension to radiomics for evaluating solid tumors. For example, Xu et al. divided the primary tumor lesions of patients with NPC into three subregions using clustering methods named S1, S2, and S3. The correlation of the S3 region with survival status was stronger than that of the entire tumor. 20 Yuan et al. found that the ITH score combined with a clinical model demonstrated excellent performance in assessing the early response of patients with LANPC to concurrent chemoradiotherapy. 14 In contrast, our study implemented a patient-specific heterogeneity assessment. Instead of aggregating tumor subregions from all patients into a common set of clusters, we quantified the heterogeneity of each texture feature individually for every tumor. This finer-scale approach is designed to capture the distinct, multi-dimensional heterogeneity pattern unique to each case, thereby moving beyond a population-averaged representation. Notably, our patient-specific ITH score metric complements conventional radiomics. Where conventional radiomics typically yields an averaged, global signature of the tumor, our method captures the heterogeneity in the spatial distribution of internal texture features. Integrating these two perspectives—the global profile and the localized heterogeneity—enables a more comprehensive representation of the tumor phenotype. This integration supports the observed performance improvement of our composite risk model compared to models using clinical features or conventional radiomics alone, contributing to a potentially more individualized prognostic assessment.
In the current study, the CRM based on multivariable Cox regression incorporated two radiomic score indicators: T1C and T2WI. Using radiomics scores alone also achieved good assessment performance, which is consistent with previous studies.21,22 For example, Du et al. demonstrated that a machine learning model trained with pretreatment MRI radiomic features effectively assessed the 3-year risk of disease progression after intensity-modulated radiotherapy. 21 In addition, Ming et al. found that the textural features of primary NPC lesions and metastatic lymph nodes improved the accuracy of clinical TNM staging strategies for assessing the survival status of NPC. 22
Increasing evidence suggests that EGFR is highly expressed in nearly 85% of patients with NPC and is associated with poor outcomes.23,24 Several studies have evaluated whether anti-EGFR treatment should be added to chemoradiotherapy. A meta-analysis using aggregated data concluded that cetuximab combined treatment can improve efficacy compared with conventional treatment. 25 Nimotuzumab combined with concurrent chemoradiotherapy or intensity-modulated radiation therapy has promising efficacy in patients with LANPC.26,27 Moreover, nimotuzumab has been found to have lower toxicity than cetuximab. 28 However, anti-EGFR treatments remain controversial, and there is a lack of largescale multicenter studies to support the reliability of these conclusions. 28 The identification of patients who are most likely to benefit from anti-EGFR therapy remains a significant challenge in the management of LANPC. Current clinical indicators lack the precision to reliably stratify patients based on their potential treatment response, highlighting an urgent need for more sophisticated predictive tools. Adding targeted drugs to traditional treatment regimens, such as nimotuzumab, requires patients to bear an additional cost of approximately $6000 for a complete treatment process. This poses a heavy burden on people in developing countries. 29 Anti-EGFR targeted therapy fails to improve DFS in all patients with LANPC. Some individuals incur substantial financial burdens from these treatments with limited clinical benefit. However, some patients who could benefit significantly from targeted therapy miss treatment opportunities. This study preliminarily explores this issue. The integrated model incorporating clinical features, the ITH score, and the radiomics score was used to evaluate the potential therapeutic benefit for patients. The results suggested that patients whose treatment aligned with the model-derived recommendations experienced a significantly better DFS. This analytical framework holds the potential to inform future clinical decision-making regarding medication selection.
Limitations
This study has several limitations that should be acknowledged. First, its retrospective nature inherently introduces the potential for selection bias and unmeasured confounding factors. Although we implemented stringent inclusion and exclusion criteria, the findings must be interpreted with caution and require validation in prospective, multi-center cohorts to confirm their generalizability. Second, manual tumor delineation was employed, which can introduce inter-observer variability. While this was mitigated by having two experienced observers and a senior physician adjudicating disputed regions, this process remains subjective. Future research would benefit from automated or semi-automated segmentation algorithms to enhance reproducibility and minimize manual bias. Third, an imbalance in case numbers between the two participating centers exists, which may impact the external validity of our model. The primary cohort (Center A) contributed the majority of cases, whereas the external validation cohort (Center B) was relatively small. To strengthen the validation, we combined the internal test cohort from Center A with the entire cohort from Center B to form a larger, composite test cohort. Nonetheless, future studies should aim for larger and more balanced external cohorts to more robustly verify the model's performance across diverse populations and clinical settings. Finally, other potentially significant biological variables, such as long non-coding RNA, 30 degree of lymphocyte infiltration, and specific immune targets, 31 were not incorporated. Including these factors in future investigations could further improve prognostic accuracy and provide a more comprehensive biological understanding of the ITH and radiomics scores.
Conclusion
This study developed a CRM that integrated clinical features, ITH score, and radiomics score for assessing DFS in LANPC. The CRM demonstrated robust prognostic performance, with both the ITH and radiomics scores individually providing superior value to the clinical model alone. Importantly, their incorporation offered significant incremental value, substantially enhancing the predictive capability of the traditional clinical model. Furthermore, our retrospective analysis suggested that the CRM-based risk stratification holds promise for informing anti-EGFR treatment strategies, a potential that warrants further investigation in prospective studies.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076251397749 - Supplemental material for MRI-based intratumoral heterogeneity analysis for prognostic evaluation of disease-free survival in locally advanced nasopharyngeal carcinoma
Supplemental material, sj-docx-1-dhj-10.1177_20552076251397749 for MRI-based intratumoral heterogeneity analysis for prognostic evaluation of disease-free survival in locally advanced nasopharyngeal carcinoma by Yixin Lu, Yang Liu, Zhichao Zuo, Binling Liang, Caiyun Huang, Lijuan Liu, Zhenyu Chen, Danke Su and Hai Liao in DIGITAL HEALTH
Supplemental Material
sj-docx-2-dhj-10.1177_20552076251397749 - Supplemental material for MRI-based intratumoral heterogeneity analysis for prognostic evaluation of disease-free survival in locally advanced nasopharyngeal carcinoma
Supplemental material, sj-docx-2-dhj-10.1177_20552076251397749 for MRI-based intratumoral heterogeneity analysis for prognostic evaluation of disease-free survival in locally advanced nasopharyngeal carcinoma by Yixin Lu, Yang Liu, Zhichao Zuo, Binling Liang, Caiyun Huang, Lijuan Liu, Zhenyu Chen, Danke Su and Hai Liao in DIGITAL HEALTH
Supplemental Material
sj-docx-3-dhj-10.1177_20552076251397749 - Supplemental material for MRI-based intratumoral heterogeneity analysis for prognostic evaluation of disease-free survival in locally advanced nasopharyngeal carcinoma
Supplemental material, sj-docx-3-dhj-10.1177_20552076251397749 for MRI-based intratumoral heterogeneity analysis for prognostic evaluation of disease-free survival in locally advanced nasopharyngeal carcinoma by Yixin Lu, Yang Liu, Zhichao Zuo, Binling Liang, Caiyun Huang, Lijuan Liu, Zhenyu Chen, Danke Su and Hai Liao in DIGITAL HEALTH
Footnotes
ORCID iDs
Ethical approval
This retrospective study was approved by the Institutional Review Board of Guangxi Cancer Prevention and Treatment Research Institute (Approval No. KY-2022-303), with a waiver of informed consent. The study represents one of the research directions within this ethically approved project.
Author contributions
Conceptualization: D.S., H.L., Y.L. Software: Y.L., Y.L., Z.Z. Data curation: Y.L., Y.L., Z.Z., C.H., Z.C. Formal analysis: Y.L., Y.L., L.L. Supervision: D.S., H.L. Investigation: Y.L., Y.L., Z.Z., B.L. Validation: Y.L., Y.L., B.L., C.H. Methodology: Y.L., Y.L., B.L. Writing—original draft: Y.L. Writing—review & editing: D.S., H.L. All authors read and approved the final manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by the Guangxi Natural Science Foundation (Grant No.2023GXNSFAA026249), Guangxi Science and Technology Program (Grant No. AB23026018), National Natural Science Foundation of China (Grant No. 82460338). The funders had no role in study design, data collection, analysis, interpretation, manuscript writing, or the decision to submit the article for publication.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Guarantor
The scientific guarantor of this publication is Guangxi Medical University Cancer Hospital.
Statistics and biometry
Python 3.7.1 and R 4.3.3 kindly provided statistical advice for this manuscript. One of the authors has significant statistical expertise. No complex statistical methods were necessary for this paper.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
