Abstract
Background
Childhood brain tumours, even though rare, present with significant diagnostic and treatment challenges. Radiomics involves feature extraction that is data-driven from standard imaging modalities, such as magnetic resonance imaging (MRI). In paediatric brain tumour imaging, MRI is often preferred because it is non-invasive and avoids exposure to radiation, making it safer for children. Radiomic features reveal additional information about tumour morphology and heterogeneity. By integrating biological meaning into imaging data, this approach enhances our understanding of tumour biology, thus supporting improved classification, treatment planning and management.
Purpose
This review focuses on MRI-based radiomics for the diagnosis and prognosis of childhood brain tumours. It assesses various approaches used in image pre-processing, tumour segmentation, feature extraction, and predictive model development to understand their accuracy and outcome, while it aims to understand the biological meaning and interpretation of radiomic features.
Methods
A systematic review was conducted, including 559 MRI-based radiomics studies in PubMed and Engineering Village Compendex databases, following preferred reporting items for systematic reviews and meta-analyses guidelines (PROSPERO registration: CRD42024503524). Data extraction focused on age, sample size, tumour type, pre-processing techniques, segmentation methods, feature extraction, and performance metrics.
Results
Nineteen studies were included, primarily addressing ependymoma (EP) and medulloblastoma (MB). Common pre-processing methods included intensity normalisation (
Conclusion
While challenges such as limited datasets and varied imaging protocols exist, we recommend identifying and integrating the most informative radiomic features to enhance diagnostic and prognostic accuracy in childhood brain tumours, ultimately improving patient outcomes.
Keywords
Introduction
Brain tumours are abnormal growth of cells in the human brain, affecting its regular functioning. In children, they are the most prevalent type of solid tumour, often leading to suboptimal health outcomes. 1 Treatment and management of such tumours depend on their size, location, type, growth status, and behaviour. 2 In children specifically, diagnosis becomes more difficult due to the rarity of the disease, the complexity of the symptoms, ongoing brain development, the need for specialised imaging, and the precision of accurately locating the tumour. 3 Radiomics is an emerging field which enhances conventional image analysis by extracting detailed quantitative data from various radiological images. It provides insights into tumour heterogeneity and its microenvironment. It can be applied to diverse imaging modalities such as computed tomography (CT) and magnetic resonance imaging (MRI), among others. 4 MRI is often preferred for paediatric brain tumour imaging because it is non-invasive, provides highly detailed images of soft tissues, allows multi-planar imaging capabilities, and requires no exposure to radiation, while it also prioritises patient safety.5,6
Integrating radiomic analysis into clinical routine practice has the potential to improve tumour subtyping, predicting treatment responses, and monitoring disease progression. 7 However, as this area of research has shifted towards more machine learning (ML) driven feature sets, the biological meaning and interpretation of radiomic features have often been lost. This could be one of the factors limiting clinical translation. Recent efforts are being made to reconnect radiomics with biological insights into tumour characteristics to enhance its clinical relevance. 8 Thus, the complexities of childhood brain tumours, combined with radiomics’ advanced imaging approach, present an opportunity to improve diagnosis, refine prognosis, and optimise treatment options. 9
While recent reviews in this area of research provide valuable overviews of radiomics and artificial intelligence in paediatric neuro-oncology, our systematic review distinguishes itself in many ways. First, it offers an in-depth analysis of the radiomics pipeline, including image pre-processing, tumour segmentation, feature extraction and selection, and predictive model development. Second, it emphasises the importance of understanding the biological meaning of the extracted radiomic features and their clinical relevance. We also aim to identify any gaps in the existing literature to propose recommendations for future research and clinical implementation. Finally, our review also considers emerging challenges such as hardware implementation offering a more comprehensive and clinically relevant perspective.
Methods
This study has been carried out according to the following stages: (i) search strategy; (ii) study protocol and selection criteria; (iii) data extraction; (iv) study quality assessment; and (v) data synthesis.
Search strategy
The literature search was conducted in two academic databases, PubMed and Engineering Village Compendex, known for their comprehensive coverage in this field of research. Search strings used were: for PubMed, (“radiomics” OR “radiomic”) AND (“brain tumour” OR “brain tumor” OR “brain neoplasm” OR “brain neoplasms”) AND (“MRI” OR “magnetic resonance imaging”), with a filter to remove systematic reviews; and for Engineering Village Compendex ((“radiomics” OR “radiomic”) AND (“brain tumour” OR “brain tumor” OR “brain neoplasm” OR “brain neoplasms”) AND (“MRI” OR “magnetic resonance imaging”)) WN ALL. Results from Inspec and GEOBASE were excluded. Keywords such as ‘childhood’ or ‘paediatric’ were not used; instead, studies were manually reviewed at a later stage to remove those that did not primarily focus on children, while retaining information about studies that included >50% paediatric data. Only studies involving human subjects and published in English language were considered. The search process was conducted according to the guidelines mentioned in the 2020 preferred reporting items for systematic reviews and meta-analyses (PRISMA) protocol. 10
Study protocol and selection
This study was registered in the Prospective Register of Systematic Reviews (PROSPERO) with protocol number CRD42024503524. The PRISMA flowchart (Figure 1) illustrates the selection process, from initial study identification to final inclusion.
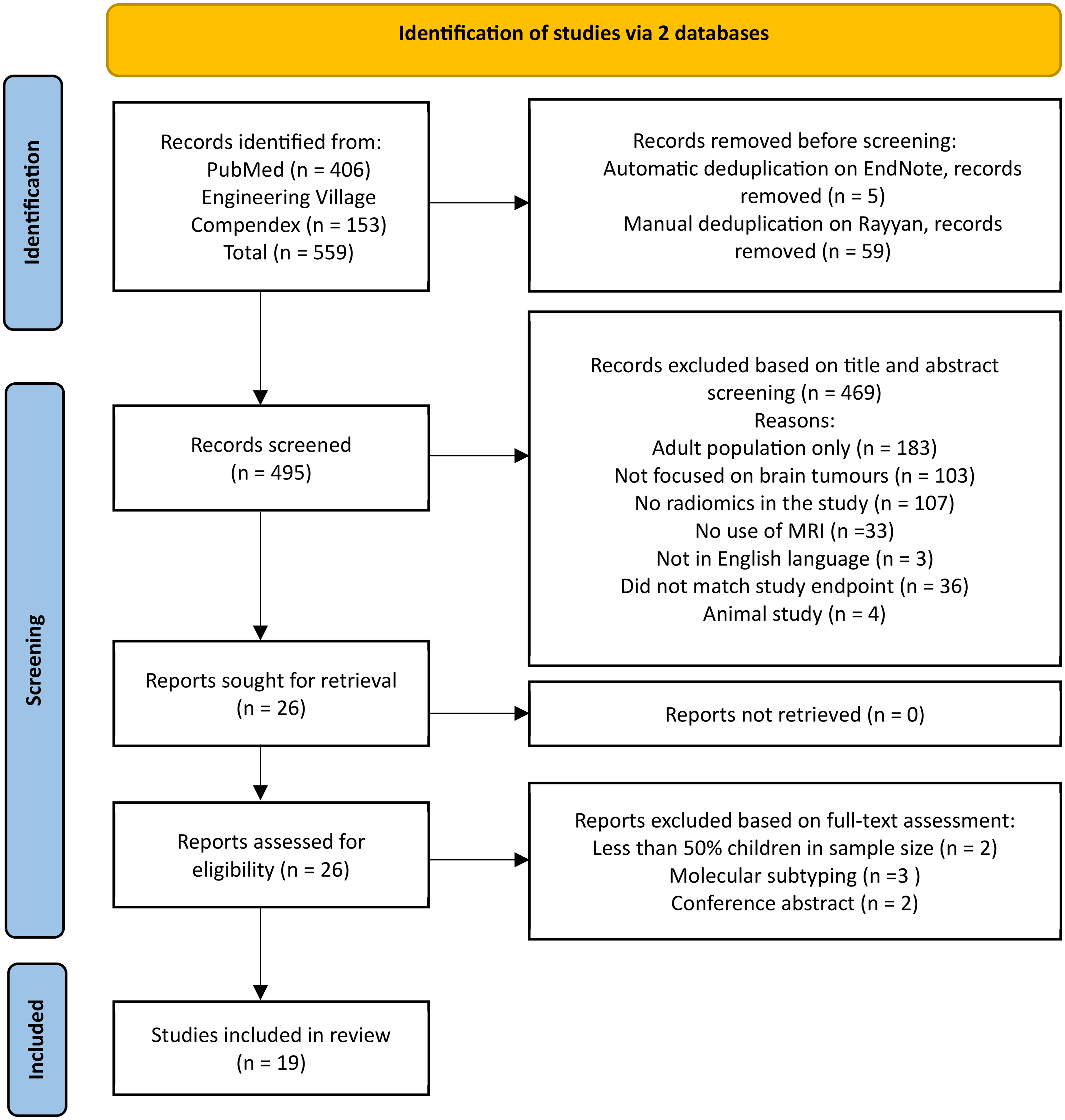
Preferred reporting items for systematic reviews and meta-analyses (PRISMA) flow diagram for study selection.
The literature search was conducted up until the end of January 2025, without any initial date restrictions, to identify all relevant papers in this area of research. The inclusion criteria comprised: (i) studies focusing on radiomics, (ii) studies assessing primary brain tumours only, (iii) studies involving paediatric populations (age <18 years) or studies with >50% children cases, and (iv) studies primarily using MRI.
Exclusion criteria encompassed studies on adults, conference abstracts, and studies utilising any other imaging modality. Additionally, radiogenomics, molecular subtyping, animal-based investigations, standardisation of radiomic processes, and non-primary brain tumour studies were also excluded.
The selection process aimed at identifying studies focusing on diagnosis and prognosis in paediatric primary brain tumours. To identify studies relevant to this research question, two screening phases were undertaken: title/abstract screening followed by full-text screening. Any inconsistencies encountered during the selection process were resolved by discussion among all reviewers. When consensus could not be reached, the most senior reviewer T.N. Arvanitis (TNA) was consulted to make the final decision on whether the article met the eligibility criteria for inclusion or not.
Data extraction
Two reviewers, T. Mukherjee (TM) and O. Pournik (OP) with expertise in radiomics and systematic review methodologies, independently conducted data extraction using a pre-determined proforma. Supervision of the study selection was provided by the senior author (TNA). Variables used for extraction of data comprised the following information: age, sample size, type of brain tumour, data collection methods, MRI sequence, image processing techniques, tumour segmentation approaches, feature extraction process, feature reduction and selection methods, predictive model development procedure, performance of the best model, and any limitations/challenges identified during the study.
Quality assessment
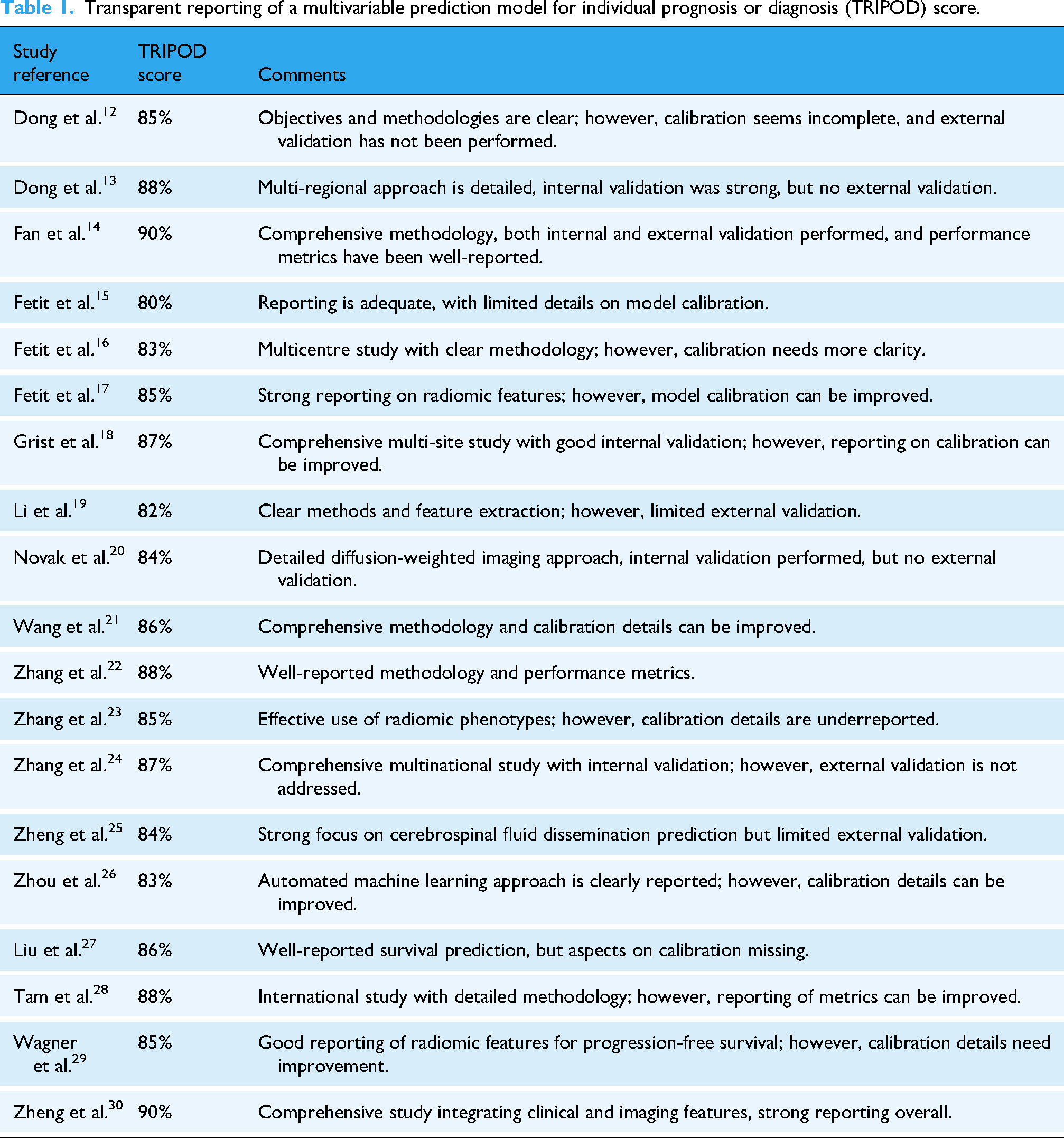
Transparent reporting of a multivariable prediction model for individual prognosis or diagnosis (TRIPOD) 11 assessment was conducted to evaluate the quality and completeness of reporting in the included studies and have been listed in Table 1. It summarises each study's TRIPOD adherence score, and comments on study objectives, internal validation, and methodologies. It also highlights areas for improvement, including insufficient calibration details and the absence of external validation in some cases. This assessment provides additional value and insights into the methodological rigour of the studies included in this systematic review.
Transparent reporting of a multivariable prediction model for individual prognosis or diagnosis (TRIPOD) score.
Data synthesis
The selected studies showed diversity in their sample sizes, radiomic methodologies, and techniques used for segmentation and feature extraction, which did not allow for conducting a meta-analysis. Instead, this review offers a comprehensive narrative alongside a semi-quantitative summary, highlighting the application of radiomic analysis in the diagnosis and prognosis of children's brain tumours.
Results
Our assessment of the included studies focused on various aspects related to MRI-based radiomics in the diagnosis and prognosis of paediatric brain tumours. We identified techniques used in image pre-processing, tumour segmentation, feature extraction, feature reduction and selection, and predictive model development. Accuracies achieved by the best-performing model and the biological significance of radiomic features were also evaluated. Additionally, we identified the limitations highlighted in the existing literature. These aspects have been discussed in the following sections.
Search results
The literature search identified 559 studies from two databases: 406 from PubMed and 153 from Engineering Village. Following automatic de-duplication in EndNote, 31 five studies were removed, and an additional 59 duplicates were removed manually. In total, 495 articles remained for title and abstract screening. Rayyan, 32 an online tool for systematic reviews, was used to manage and screen these studies. Following this screening, 469 articles were excluded based on the pre-defined inclusion and exclusion criteria. Of the remaining 26 studies assessed in full-text, seven studies were excluded for not meeting the inclusion criteria. Thus, 19 articles were included in this systematic review.
Study characteristics
All studies included in this review were full-text articles (100%) and were retrospective (100%) in their design. The review was conducted without any time restrictions, to include all relevant studies in this field until January 2025. The distribution of publications over time and their geographical origins are shown in Figure 2(a) and (b), respectively.

Demographics of included studies: (a) distribution of published studies over the years; (b) distribution of study geographies.
A total of 10 studies (52.6%) were single centre12–14,17,19,21,24,25,29,30 and nine studies (47.4%) were conducted across multiple centres.15,16,18,20,22,23,26–28 Of the total studies analysed, 15 studies (78.9%) were diagnosis-focused,12–26 while four studies (21.1%) were prognosis-focused.27–30
The studies in this review have been categorised as either diagnosis-focused or prognosis-focused. Table 2(a) and (b) covers diagnosis: Table 2(a) includes age, sample size, tumour type, MR imaging sequence, and study limitations; while Table 2(b) contains information about image pre-processing, segmentation, feature extraction, feature selection/reduction, and model performance. Table 3(a) and (b) provides the same for prognosis.
Radiomics approach for diagnosis of paediatric brain tumours.

ADC: apparent diffusion coefficient; ATRT: atypical teratoid/rhabdoid tumour; BT: brain tumour; CE: contrast-enhanced; CSF: cerebrospinal fluid; DSC: dynamic susceptibility contrast; EP: ependymoma; ET: embryonal tumour; FLAIR: fluid-attenuated inversion recovery; HGG: high-grade glioma; MB: medulloblastoma; ML: machine learning; PA: pilocytic astrocytoma; PFT: posterior fossa tumour; T1C (or T1CE): T1-weighted contrast-enhanced; T1W: T1-weighted; T2W: T2-weighted; AdaBoost: adaptive boosting; ANN: artificial neural network; AUC (95% CI): area under the curve (95% confidence interval); C-SVM: C-support vector machine; Entropy-MDL Discretisation: entropy minimum description length discretisation; FCBF: fast correlation-based filter; GLCM: gray level co-occurrence matrix; GLDM: gray level dependence matrix; GGCM: gray level gradient co-occurrence matrix; GLH: gray level histogram; GLRLM: gray level run-length matrix; GLSZM: gray level size zone matrix; IBSI: image biomarker standardisation initiative; IM: image metrics; k-NN: k-nearest neighbour; LASSO: least absolute shrinkage and selection operator; LR: logistic regression; mRMR: minimum redundancy maximum relevance; MLR: multiple logistic regression; NB: Naïve Bayes; NN: neural network; NA: not available; PCA: principal component analysis; RFE: recursive feature elimination; RF: random forest; SVM: support vector machine; TPOT: tree-based pipeline optimisation tool; UA: univariate analysis; UAS: univariate analysis screening; XGB: xtreme gradient boosting.
Radiomics approach for prognosis of paediatric brain tumours.

CE-T1W: contrast-enhanced T1-weighted; DIPG: diffuse intrinsic pontine glioma; FLAIR: fluid-attenuated inversion recovery; MB: medulloblastoma; PFS: progression-free survival; T1C (or T1CE): T1-weighted contrast-enhanced; T1W: T1-weighted; T2W: T2-weighted; C-index: concordance index; GLCM: gray level co-occurrence matrix; GLDM: gray level dependence matrix; GLRM: gray level run-length matrix; GLRLM: grey-level run-length matrix; GLSZM: gray level size zone matrix; LASSO: least absolute shrinkage and selection operator; NA: not available.
Table 4 provides additional information on the extracted radiomic feature classes, their biological meaning, and clinical relevance.
Feature classes, biological meaning, and their clinical relevance.

GLCM: grey-level co-occurrence matrix; GLRLM: grey-level run-length matrix; GLSZM: grey-level size zone matrix.
Evaluation metrics
In this systematic review, the best model performance from each selected study was recorded, expressed as the area under the receiver operating characteristic curve (AUC) (unless otherwise specified) which evaluates a model's diagnostic performance integrating both sensitivity and specificity across varying classification thresholds. A higher AUC represents superior discriminative ability and reporting a 95% confidence interval (CI) around the AUC also helps with comparisons between different radiomics models or study findings. Other metrics were also reported in certain studies such as accuracy (proportion of all correctly classified cases), sensitivity/recall (ability to correctly identify positive instances), specificity (capacity to accurately rule out negative cases), precision (proportion of predicted positives that are true positives), and F1-score (harmonic mean of precision and recall).
Radiomics in paediatric neuro-oncology
The distribution of studies focused on various types of paediatric brain tumours such as medulloblastoma (MB) (11), ependymoma (EP) (5), diffuse intrinsic pontine glioma (DIPG) (2), pineal region germinoma (1), pineoblastoma (1), along with six studies covering other brain tumour types, including astrocytomas, gliomas, and unspecified tumours. Sample sizes varied across studies and ages ranged from infants (<1 month old) to 18 years old.
Diagnosis
Fifteen studies were diagnosis focused. Table 2(a) and (b) lists key characteristics of each study, including age, sample size, tumour type, imaging modality, image pre-processing techniques, segmentation approach, feature extraction and selection methods, and best model performance.
A thorough analysis of the selected studies revealed the use of a diverse range of radiomic approaches to extract quantitative features from images, and ML techniques that use algorithms to analyse these features and develop predictive models for the diagnosis of paediatric brain tumours. Dong et al. 12 differentiated EP and MB, with an AUC of 0.910 using identified imaging biomarkers, while Dong et al. 13 classified Paediatric Posterior Fossa Tumours (PPFTs), with an AUC of 0.977 by extracting features from multiparametric MRI. Replacing contrast-enhanced T1W images with apparent diffusion coefficient (ADC) maps improved performance, highlighting the importance of using additional MRI sequences. Similarly, ML's efficacy in PPFT differentiation was also studied by 19 targeting pre-operative differentiation between EP and PA. Multicentre use of ADC for classifying PFTs achieved high accuracy across diverse hospital settings and scanners. 20 Wang et al. 21 also classified PPFTs, achieving 93.8% accuracy using a random forest (RF) classifier. It is important to note that, the best features for classification came from the ADC sequence. This highlights the value of ADC sequences in extracting quantitative information in differentiating between different types of childhood brain tumours. Zhang et al. 22 also used ML to differentiate embryonal tumours (ETs), High-Grade Gliomas (HGGs), and EPs in paediatric supratentorial tumours. Zhang et al. 23 used radiomic phenotypes derived from T2 W and gadolinium-enhanced T1W MRI images to distinguish atypical teratoid/rhabdoid tumours from MBs. Zhang et al. 24 proposed a sequential ML classifier to improve the diagnosis of PPFTs across multinational cohorts. Zhou et al. 26 also focused on differentiated different types of PPFTs using ML approaches applied to routine MR imaging. They compared manual expert optimisation with automatic ML using the Tree-Based Pipeline Optimisation Tool. The automatic ML model achieved significantly higher accuracy in classification.
To differentiate between pineal region germinoma and pinealoblastoma, a clinic–radiomic model was used. 14 Using both radiomic and clinical features, the model achieved high discrimination with an AUC of 0.950 in the training set and 0.940 in the validation set. This non-invasive approach demonstrated high sensitivity and performed well in differentiating pineal region tumours.
Studies15–17 focused on improving the diagnostic classification of childhood brain tumours using texture analysis (TA). While Fetit et al. 15 conducted a multicentre study and achieved good model performance across different centres and scanners, Fetit et al. 16 evaluated the efficacy of three-dimensional (3D) TA compared to 2D TA, showing improved classification performance with 3D textural features. Fetit et al. 17 assessed the efficacy of both 3D TA and 2D TA, re-confirming classifiers trained with 3D textural features show improved classification performance compared to those trained with conventional 2D features alone.
The integration of multicentre diffusion and perfusion imaging with ML resulted in a classifier with >80% predictive precision. 18 Finally, in Zheng et al., 25 multivariable logistic regression was used to combine clinical and radiomic features for prediction of pre-operative cerebrospinal fluid (CSF) dissemination in children with MB. Combining one clinical and nine radiomic features demonstrated higher predictive performance with an AUC of 0.89.
Prognosis
Four studies were prognosis-focused. Table 3(a) and (b) summarises key characteristics and results for each study.
In Liu et al., 27 a radiomics signature and nomogram were developed to predict progression-free survival (PFS) in MB patients. It can be used to stratify patients for postoperative radiotherapy. Tam et al. 28 focused on DIPGs, using MRI-based radiomics to create a prognostic model for overall survival (OS), which outperformed other models that were based only on clinical variables. In contrast, Wagner et al. 29 evaluated the prognostic value of radiomics in DIPG subsets, demonstrating the potential of MRI-based radiomics in progression-free survival. Finally, in predicting overall survival in MB patients, Zheng et al. 30 highlighted that an integrative model combining radiomics, clinical, and conventional MRI features, outperformed other models. All these studies showed emphasis on the prognostic value of radiomics in paediatric brain tumours.
Feature classes, biological meaning, and clinical relevance
Based on the studies, texture-based features12–17,19,20,22–24,28 were seen as a preferred choice for brain tumour classification and characterisation, because of their higher discriminative power and ability to capture fine-grained spatial patterns, and variations within the tumour region. These patterns are believed to be biologically linked to cellular variability, angiogenesis, and necrosis, which are all important for tumour progression and understanding response to treatment. 33 Thus, they offer a more comprehensive and detailed characterisation of tumour heterogeneity and microstructure, compared to histogram-based features.
In contrast, histogram-based features12,15–21,25,26 primarily describe the overall distribution of voxel intensities without considering spatial relationships or textural patterns. Features, such as skewness and variance,12,15,21,25,26 describe the variability of intensities within a tumour, which may reflect the areas of necrosis or differing tissue densities, both of which have biological and prognostic relevance. 34
Shape-based features12,14,19,22–25,28–30 provide information about tumour morphology, invasion, and geometry. These correlate with the aggressiveness of a tumour and thus have an impact on surgical planning and treatment decisions. However, they may not always capture the subtle differences in tissue texture and composition captured by texture features. 34
Challenges
Various challenges associated with MRI-based radiomics in the diagnosis and prognosis of children's brain tumours were identified across studies, as listed in Tables 2(a) and 3(a). The most frequently reported limitation was a small sample size, highlighted in 14 studies. Additionally, the use of limited MR sequences, heterogeneity in image acquisition, and retrospective study design were prevalent, mentioned in 10 studies. Imbalanced datasets, lack of data from multiple centres, and the need for prospective validation were also commonly cited limitations, identified in seven studies each.
Discussion
It has been seen in this review that in the context of diagnosis, radiomics and ML techniques have been widely used to improve the classification of various types of paediatric brain tumours. Studies12,13,21 showed how effective ML algorithms are in accurately differentiating EP, MB, and other PPFTs. The integration of advanced MRI sequences, particularly ADC maps, was seen as one of the most important factors in enhancing diagnostic accuracy.13,20 The use of ML models across multiple centres highlights their robustness and potential for real-world clinical application.19,26 Also, the integration of radiomic and clinical features outperformed conventional models in brain tumour characterisation. 14
In the prognosis-focused studies, the prognostic value of radiomics in predicting PFS and OS, in paediatric brain tumour patients, was seen. The development of radiomics signatures, which quantify imaging features to predict tumour behaviour, and nomograms, which visually represent statistical models for estimating patient outcomes, provides valuable tools for stratifying patients and guiding treatment decisions.27,28,30 Integrating radiomic (e.g. texture and shape), clinical (e.g. patient demographics and laboratory results), and conventional MRI features (e.g. tumour morphology and anatomical information) in prognostic models, showed improved performance compared to models based only on clinical variables, 30 emphasising the importance of a multimodal approach to prognostic assessment.
Radiomic features quantitatively describe tumour characteristics such as shape, texture, and intensity.8,33 In this review, the biological meaning of these features is also mentioned to understand the underlying biological mechanisms that these features reflect, which is essential for easier clinical translation of radiomics (Table 4). These image characteristics are also used to see if they can be connected to a disease status (helpful in the prediction of an outcome).
However, the data-driven nature of radiomics and the heavy use of ML techniques do not provide insights into the biological mechanisms of these observed relations. This often leads to slower translation into clinical practice as the biological meaning and interpretation of these features are often missing.8,33 This is an attempt to reconnect radiomic findings to biological meaning.
Features such as texture, shape, histogram, and wavelet reveal additional information about tumour biology and can be used to accurately classify tumour types, subtypes, and grades. Texture features such as GLCM, GLRLM, and GLSZM reveal patterns and spatial relationships within the image, showing consistency or heterogeneity of the tumour.
However, despite these advancements, challenges still exist. To mitigate the identified challenges effectively, innovative solutions are essential. For example, collaborative efforts among multiple centres can facilitate data pooling, overcoming limitations such as small cohorts and enabling validation across diverse patient populations.15,20,22,26 Integrating advanced imaging techniques and molecular data into radiomic analysis can improve the accuracy and robustness of models13,20,23,27–30 and make them more generalisable. Additionally, standardising imaging protocols and using fully automatic segmentation techniques can reduce image acquisition variability and enhance the reproducibility of the radiomics approach used.
With the rise in the integration of artificial intelligence, multi-omics data, and novel imaging protocols, new challenges have emerged. These include interpretability of increasingly complex models, data governance and regularity oversight, data storage issues, and computational infrastructure for real-time analyses, among others. 35 Addressing these issues will require cross-disciplinary collaborations, the development of secure data-sharing and storage platforms, and refining ethical frameworks.
The majority of radiomics-related studies in paediatric neuro-oncology including the papers included in this review primarily discuss algorithmic approaches (such as segmentation, feature extraction, and ML) and workflow optimisation (such as standardising pipelines). None explicitly discuss hardware implementation strategies. Yet, hardware considerations are equally important when improving computational workflows, especially given the large volume of high-resolution MRI data and the complexity of radiomic pipelines. In general, using graphics processing units (GPUs) and high-performance computing (HPC) environments, including clusters and cloud-based solutions, when performing radiomics computations can accelerate data processing, and significantly reduce processing times for feature extraction and model training. 36 However, specialised hardware comes at a cost, especially for smaller research groups and healthcare institutions with limited resources. Also, different hardware setups could lead to variations in performance and reproducibility. It is therefore necessary to report hardware specifications and computational runtimes to facilitate replication and comparison across studies.
While previous reviews37,38 have significantly advanced our understanding of radiomics and artificial intelligence in paediatric neuro-oncology, our review extends this foundation as it includes not only a detailed analysis of the entire radiomics pipeline but also considers the biological meaning of radiomic features and discusses emerging challenges such as hardware implementation. This offers a comprehensive and clinically relevant perspective to guide future research in the field.
Conclusion
This systematic review highlighted key techniques used in all the phases of a radiomics pipeline, including image pre-processing, segmentation, feature extraction, feature selection and reduction, and model building. It also highlighted the biological meaning and clinical relevance of the extracted radiomic features. The identification and integration of the most informative radiomic features are important, as they hold the potential to significantly improve diagnostic and prognostic accuracy. Research should focus on validating and standardising these features to facilitate easier translation into clinical practice.
Despite advancements in radiomics, challenges such as limited datasets and inconsistent image acquisition protocols still exist. Addressing these challenges will further enhance radiomic analysis. Data-sharing among institutions and the use of transfer learning to adapt models trained on larger datasets to smaller, more specialised datasets, such as paediatric brain tumours, are recommended. Ultimately, refining radiomic analysis, using more generalisable models, and integrating them with clinical data will be essential in advancing the field of paediatric neuro-oncology.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076251336285 - Supplemental material for Magnetic resonance imaging (MRI) radiomics in paediatric neuro-oncology: A systematic review of clinical applications, feature interpretation, and biological insights in the characterisation and management of childhood brain tumours
Supplemental material, sj-docx-1-dhj-10.1177_20552076251336285 for Magnetic resonance imaging (MRI) radiomics in paediatric neuro-oncology: A systematic review of clinical applications, feature interpretation, and biological insights in the characterisation and management of childhood brain tumours by Teesta Mukherjee, Omid Pournik and Theodoros N Arvanitis in DIGITAL HEALTH
Footnotes
Abbreviations
Guarantor
Theodoros N Arvanitis is the guarantor for this systematic review.
Ethical considerations and informed consent
This study is a systematic review of previously published literature and does not involve human participants, identifiable personal data, or direct patient interaction. As such, ethical approval and informed consent were not required.
Author contributions/CRediT
All authors have reviewed this manuscript. TM and OP simultaneously performed the inclusion/exclusion process and agreed on the final selection of the studies in this review. TM prepared the data extraction pro forma, analysed data, and wrote the manuscript. OP validated the data extraction results and edited the manuscript. TNA verified the data extraction results, reviewed the manuscript, and secured TM's funding for this work. All authors have complete access to all the data in this review.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: TM received an Engineering and Physical Sciences Research Council (EPSRC) Studentship Scholarship for her PhD study at the Department of Electronic, Electrical and Systems Engineering, School of Engineering, University of Birmingham.
Conflicting interests
The author(s) have declared the following potential conflict of interest regarding the authorship, and/or publication of this review: Theodoros N. Arvanitis serves as the Editor-in-Chief of
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
