Abstract
Objective
The practice of evidence-based medicine can be challenging when relevant data are lacking or difficult to contextualize for a specific patient. Large language models (LLMs) could potentially address both challenges by summarizing published literature or generating new studies using real-world data.
Materials and Methods
We submitted 50 clinical questions to five LLM-based systems: OpenEvidence, which uses an LLM for retrieval-augmented generation (RAG); ChatRWD, which uses an LLM as an interface to a data extraction and analysis pipeline; and three general-purpose LLMs (ChatGPT-4, Claude 3 Opus, Gemini 1.5 Pro). Nine independent physicians evaluated the answers for relevance, quality of supporting evidence, and actionability (i.e., sufficient to justify or change clinical practice).
Results
General-purpose LLMs rarely produced relevant, evidence-based answers (2–10% of questions). In contrast, RAG-based and agentic LLM systems, respectively, produced relevant, evidence-based answers for 24% (OpenEvidence) to 58% (ChatRWD) of questions. OpenEvidence produced actionable results for 48% of questions with existing evidence, compared to 37% for ChatRWD and <5% for the general-purpose LLMs. ChatRWD provided actionable results for 52% of questions that lacked existing literature compared to <10% for other LLMs.
Discussion
Special-purpose LLM systems greatly outperformed general-purpose LLMs in producing answers to clinical questions. Retrieval-augmented generation-based LLM (OpenEvidence) performed well when existing data were available, while only the agentic ChatRWD was able to provide actionable answers when preexisting studies were lacking.
Conclusion
Synergistic systems combining RAG-based evidence summarization and agentic generation of novel evidence could improve the availability of pertinent evidence for patient care.
Keywords
Background and significance
Evidence-based medicine, in which decisions about patient care are purposefully made using the best available evidence, has been the standard for the last three decades. 1 However, in some specialties less than 20% of daily medical decisions are supported by quality evidence.2,3 The gap between the needed and available evidence in care decisions is driven by two issues. First, clinical trials often lack generalizability 4 to complex patients who often fail to qualify for trials.5,6 This creates a need for timely and relevant real-world evidence (RWE) to guide care and treatment decisions.7,8 Second, even when studies exist, they can have conflicting findings arising from heterogeneous patient populations, study designs of variable quality, or nonstandard endpoints. These factors make it difficult to compile research findings into specific recommendations for a given patient.9,10 As a result, physicians often require either summarized evidence from reliable sources or custom evidence generated specifically for the patient in front of them.8,11
Large language models (LLMs) are increasingly studied for their ability to answer questions in various medical domains.12,13 Large language models have displayed impressive performance summarizing relevant literature14–16 and responding to natural language queries. However, they are prone to hallucinating reference materials or treatment guidelines17,18 and may produce “recommendations” that are satirical, inappropriate, 19 or misaligned with evidence-based guidelines. 20
One approach to adapt LLMs for summarizing evidence is the use of retrieval-augmented generation (RAG), where an LLM is used to compile information retrieved from curated knowledge sources. 21 A clinician could submit a clinical question to an LLM and receive summaries of relevant research articles and practice guidelines retrieved from the knowledge base. This approach is used by OpenEvidence (https://www.openevidence.com) to answer clinical queries via RAG using an LLM. In such a RAG system, the answers are limited to preexisting evidence sources.
Alternatively, an agentic system for on-demand evidence could use an LLM as a natural language interface to an evidence generation platform with access to medical record data. 22 In such an agentic system, the LLM would serve as a copilot to pass clinical intent to an underlying purpose-built engine for generating evidence.8,11 ChatRWD™ (https://www.atroposhealth.com/chatrwd) is one such system. Its LLM-driven user interface translates clinical queries into a structured population–intervention–control–outcome (PICO) 23 study design for processing through a causal inference engine that can generate on-demand RWE 8 in response to the question.
In this study, we assessed the ability of OpenEvidence and ChatRWD to provide treatment recommendations in response to clinical questions that might arise in the context of care delivery. As a baseline, and to simulate how healthcare professionals may conveniently turn to the more widely available LLMs, we also assessed the ability of three off-the-shelf LLMs (ChatGPT-4, Claude 3 Opus, and Gemini Pro version 1.5) to answer the same questions. We submitted 50 clinical questions to each of the five systems. The generated answers were evaluated by a panel of nine clinicians. We hypothesized that OpenEvidence and ChatRWD would outperform the LLMs. We further hypothesized that OpenEvidence would perform well on questions where existing evidence was likely present, while ChatRWD would be the only system capable of providing RWE when relevant published literature was lacking.
Materials and methods
Figure 1 outlines the evaluation process. First, we selected 50 questions as the basis for evaluation (Question Selection) then submitted them to ChatGPT-4, Claude 3 Opus, Gemini 1.5 Pro, OpenEvidence, and ChatRWD. All of the LLM-based systems except ChatRWD provided supporting citations that were checked for hallucinations (Citation Review). Because ChatRWD performs a new study on demand, the intermediate code generated by ChatRWD for patient cohorts was reviewed by trained clinical informaticians (Study Integrity Review). The output of each system was graded according to a standard medical rubric by a panel of clinicians (Clinical Review).

Evaluation process.
Question selection
We selected 50 questions (Supplemental Table 1) that were either submitted to Atropos Health by physicians requesting evidence for clinical decisions or inspired by such questions. All questions met the following criteria:
Answerable using a cohort study design; All four PICO components can be fully defined; Control group is an active comparator rather than absence of treatment; Question is not evaluating treatment route, dosage, duration, or line of therapy; Intervention does not require washout from prior intervention.
Because we hypothesized that performance of the models may differ based on whether the questions had existing answers in the literature, we oversampled novel questions in selecting our question set. Novelty was determined by a consensus vote among three independent reviewers who searched the medical literature for keywords from each question (Supplemental Table 1).
Response generation
Large language models
We used three general-purpose LLMs (ChatGPT-4 [2 May 2024], Claude 3 Opus [20240229], and Gemini 1.5 Pro [001]) to produce answers to the selected clinical questions. OpenAI's ChatGPT-4 24 is an instruction-tuned pretrained transformer model designed to produce human-like text in response to natural language instructions. Anthropic's Claude 25 is a suite of AI language and image models trained within a governance framework of prespecified rules and principles for generative output. Gemini 26 from Google is a suite of natively multimodal AI models designed to interact with and generate multiple data types including text, audio, and images.
We submitted questions to ChatGPT, Claude, and Gemini via their REST APIs. We provided a standardized prompt followed by the clinical question (Supplemental Table 2). Inspired by previous research, 27 the prompt directed the model to serve as a helpful assistant with medical expertise. Additionally, to facilitate evaluation and understand LLM reasoning, we specifically asked the models to cite any referenced studies and respond with “I do not know the answer” when that were the case. Large language model responses are found in Supplemental Table 3.
OpenEvidence
OpenEvidence uses an LLM for RAG with different types of medical literature, including PubMed articles and FDA drug labels to answer clinical questions submitted via the web or its API. 28 Using RAG on existing medical literature in this manner reduces the likelihood of hallucinating information because OpenEvidence can summarize the relevant literature retrieved and present the conclusions to the requester. The final output includes references to the papers identified in the literature search. To evaluate OpenEvidence, we submitted each of the 50 plain English clinical questions through their API and checked the citations provided (Citation Review).
ChatRWD
The core of ChatRWD is an advanced cohorting engine for selecting patients and data 29 coupled with a causal inference engine for automated statistical analysis. 8 ChatRWD adds four steps to this system: (1) Chain-of-Thought prompting to convert plain English questions into PICO format, classify the study design, and perform named entity recognition, (2) semantic search of a curated phenotype library, (3) generation of Temporal Query Language (TQL) 29 code to do the cohort selection and invoke an underlying purpose-built platform for statistical analyses, and (4) summarization of findings from statistical analyses (https://www.atroposhealth.com/chatrwd). Users can confirm and modify the inferred PICO as well as the retrieved phenotypes via a web interface (Supplemental Figure 1) before the study is executed.
The phenotype library defines nearly 2000 diseases and conditions based on procedure codes, diagnosis codes, medication prescriptions, and/or laboratory tests and results. The definitions are curated by physicians and subject matter experts; where possible, definitions are sourced from published frameworks such as the Observational Health Data Sciences and Informatics initiative. For example, the phenotype “metastatic solid tumor” is defined based on ICD9 codes 196-199 or ICD10 codes C77-C80. The phenotype “amputation lower extremity leg” is defined based on CPT codes 27290, 27295, 27590, 27592, 27594, 27596, 28598, 27880-27882, 27884, 27886, 27888, 27889, 28800, 28805, 28810, 28820, 28825, 28111-28113, and 27591. The phenotype “stage 3a chronic kidney disease” is defined based on values between 45 and 59 mL/min/172 m2 for LOINC codes 33914-3, 48642-3, 48643-, 50210-4, 62238-1, 69405-9, 98979-8, 76633-7, 77147-7, 98980-6, 88293-6, 88294-4.
The data source used with ChatRWD (Eversana's Electronic Health Record Integrated Database) consisted of electronic health records of 159 million patients from outpatient and inpatient providers in the United States, including structured medication, laboratory, procedure, and diagnosis data.
Evaluation of AI-generated responses
Citation review
Four of the LLM systems tested (ChatGPT, Claude, Gemini, and OpenEvidence) can cite from the medical literature. We checked the validity of these citations before passing the responses on to our clinical reviewers (Clinical Review). Since OpenEvidence provides links for each citation, we were able to verify the citations by confirming that each link directed to the appropriate study. For the general LLMs, we verified citations by determining if the relevant article could be located on PubMed. For any citations that were still unmatched, we checked the provided URL from the LLM, if any. Citations still unmatched at that point were considered to be hallucinations.
Study integrity review
ChatRWD does not return citations because the study is run on-demand using RWD. Therefore, we conducted a Study Integrity Review instead. ChatRWD involves rule-based generation of TQL code 29 defining a study cohort. For each of the 50 questions, two medical informaticists reviewed the phenotypes and the TQL code for each PICO element to ensure their appropriateness, selecting from one of three grades: incorrect, not ideal but acceptable, good. The phenotypes were scrutinized to ensure there were no inappropriate inclusions or omissions in order to accurately answer the research question. The study's overall appropriateness and the primary cause of failure to answer the research question if any was noted.
Clinical review
Nine physicians across several specialties graded the responses from all five LLM systems using a standardized rubric (Supplemental Table 4). None of the reviewers were employees of either Atropos Health or OpenEvidence. Before grading, all reviewers underwent training on the use of the rubric using several case studies. Reviewers graded answers along with three primary dimensions (response generation, relevance, and evidence quality) to which reviewers had to rate: “yes,” “no,” or optionally “mixed” for the relevance and evidence quality dimensions. Because we specifically prompted “If you do not know the answer, respond with ‘I do not know the answer’,” such a response was deemed as “no response.” To judge relevance, reviewers had to determine if the response explicitly answered the question at hand. For evidence quality, the reviewers had to determine if the cited studies exist (see Citation Review) and if so, were they appropriate and of high quality (e.g., sufficient cohort size, appropriate PICO, and analysis).
We also asked reviewers to evaluate the actionability of each response. As described in the rubric, a response was considered actionable if it was of sufficient relevance and quality to justify or change clinical practice. Additionally, after reviewing all five responses to each question, reviewers were asked which was the best response.
Because the various LLM systems all have easily identifiable response structures, the physicians were not blinded to the system that generated the answer. The physicians reviewed the content independently and were not able to see the responses of their peers.
Data analysis
We first aggregated the ratings along with the three primary dimensions from the nine clinical reviewers based on a majority vote. From the combination of these aggregated ratings, we then binned each response into one of the five exhaustive and mutually exclusive response categories according to the logic shown in Table 1.
Clinical review results: response category.

*Yes: when 5+ Reviewers responded with “Yes.”
Mixed or Yes: when 5+ Reviewers responded with “Yes” or “Mixed.”
No: when <5 Reviewers responded with “Yes” or “Mixed.”
N.A.: not applicable.
We considered a given response to be actionable if at least five reviewers classified it as high enough quality to justify or change practice. The response receiving the highest number of votes from the nine reviewers was deemed the ‘best’, with ties being allocated to both equivalent LLM systems.
To test the hypothesis that ChatRWD and OpenEvidence would outperform the other models, we defined the highest response category (“Relevant & Evidence-Based,” Table 1) as a success and compared success proportions between the five models using a chi-square test. If this test indicated a significant difference between models, we performed Fisher's exact test between each pair of models, using the Holm–Bonferroni correction for multiple comparisons. We performed the same process for actionable results.
We calculated the inter-rater agreement on response category, best model, and actionable metrics using Fleiss’ Kappa. 30 For each of the five LLM systems, we assessed LLM-specific inter-rater reliability of the nine clinical reviewers across 50 questions. We also computed overall inter-rater agreement across all 250 combinations of the 50 clinical questions and five LLM systems.
As a
Results
Relevance, evidence, and actionability
The RAG and agentic systems produced answers for 86% (OpenEvidence) and 94% (ChatRWD) of the questions (Table 1, Supplemental Figure 2), compared to 58–78% of questions in the three general-purpose LLMs (Supplemental Table 3) used for benchmarking. The physician reviewers found that OpenEvidence and ChatRWD produced relevant, evidence-based answers for 24% and 58% of the questions, respectively. For comparison, the physician reviewers rarely judged answers from the general-purpose LLMs to be relevant and evidence-based, meeting this standard for 2–10% of the responses. This pattern of differences was statistically significant (
For the stricter criterion of actionability, where the reviewers judged the answers to be of sufficient quality to justify or change clinical practice, the reviewers rarely found the LLM answers (2–4%) to be actionable, while answers from OpenEvidence (30%) and ChatRWD (44%) were more often judged to be actionable (Table 2). This pattern of differences was also statistically significant (
Clinical review results: actionable and best answers.

Failure analysis
We tallied the reasons the clinical reviewers rated answers as lacking either relevance or evidence (Table 3). For ChatRWD, the most common problem was misspecification of the study design (44.7%, Table 3). Misspecification by ChatRWD usually stemmed from ill-defined phenotypes and sometimes from logical errors (e.g., misinterpreting “and” for “or” logic in drug combinations). Examples of ChatRWD phenotyping errors were including antibiotics in a question about migraine medications or including upper extremities in questions about surgery for lower extremities.
Clinical review results: failure analysis by issue.

Issue details were optional, allowed for multiple selection, and tallied only if 5+ reviewers noted that issue.
Relevant issues were classified minor if reviewers rated relevance as “mixed—still useful, but not exact.”
The general-purpose LLMs’ most common failure was the inclusion of hallucinated or irrelevant citations. Systematic searching of PubMed and LLM-generated URLs identified citation errors in 40–80% of relevant answers (Table 3). Over 40% of all citations from Claude and Gemini could not be located on PubMed, as well as 25.5% of ChatGPT's citations (Supplemental Table 6). As an example, in response to a question on body mass index after hormonal contraceptive use, Gemini responded with a plausible combination of article author, title, and journal (Bonny A.E.; American Journal of Obstetrics and Gynecology; Weight gain in adolescents receiving depot medroxyprogesterone acetate) which does not exist.
When the answers by LLMs were deemed less relevant, it was often due to minor study misspecification such as the study population (e.g., ulcerative colitis) not being fully relevant to the question (e.g., Crohn's disease). Of note, OpenEvidence rarely misinterpreted the questions and did not hallucinate citations.
Qualitatively, the reviewers found little value in the responses from the general-purpose LLMs (Supplemental Table 7). As one reviewer noted, “Claude, Gemini, and ChatGPT often produce hallucinations and factually incorrect information, necessitating independent verification of all data provided.” Two reviewers noted that the advice from these LLMs could be potentially fatal. The reviewers were more favorable toward ChatRWD and OpenEvidence: “ChatRWD and OpenEvidence generated the most consistently relevant/robust responses.”
Inter-rater reliability
Inter-rater reliability across the key metrics was found to be fair to moderate (Supplemental Table 8): Fleiss’ Kappa statistics of 0.38 and 0.31 (fair) for actionability and best answer, respectively, and 0.56 (moderate) for response categories. Within each LLM system, the inter-rater reliability for response category ranged from fair (ChatRWD: 0.30; OpenEvidence: 0.38) to good (Gemini: 0.63).
The role of question novelty
We hypothesized that ChatRWD would outperform OpenEvidence on novel questions for which ChatRWD can perform on-demand studies. We stratified questions by their novelty and compared the relative performance of ChatRWD and OpenEvidence (Figure 2, Table 4). Among the novel questions, ChatRWD could produce answers that were actionable (52.2%) as well as answers that were relevant and evidence-based (65.2%). When faced with novel questions, OpenEvidence was rarely able to produce actionable answers (8.7%) or answers that were relevant and evidence-based (8.7%).

Performance of large language model (LLM) systems stratified by question novelty.
OpenEvidence and ChatRWD performance overall and stratified by question novelty, separately, and combined.
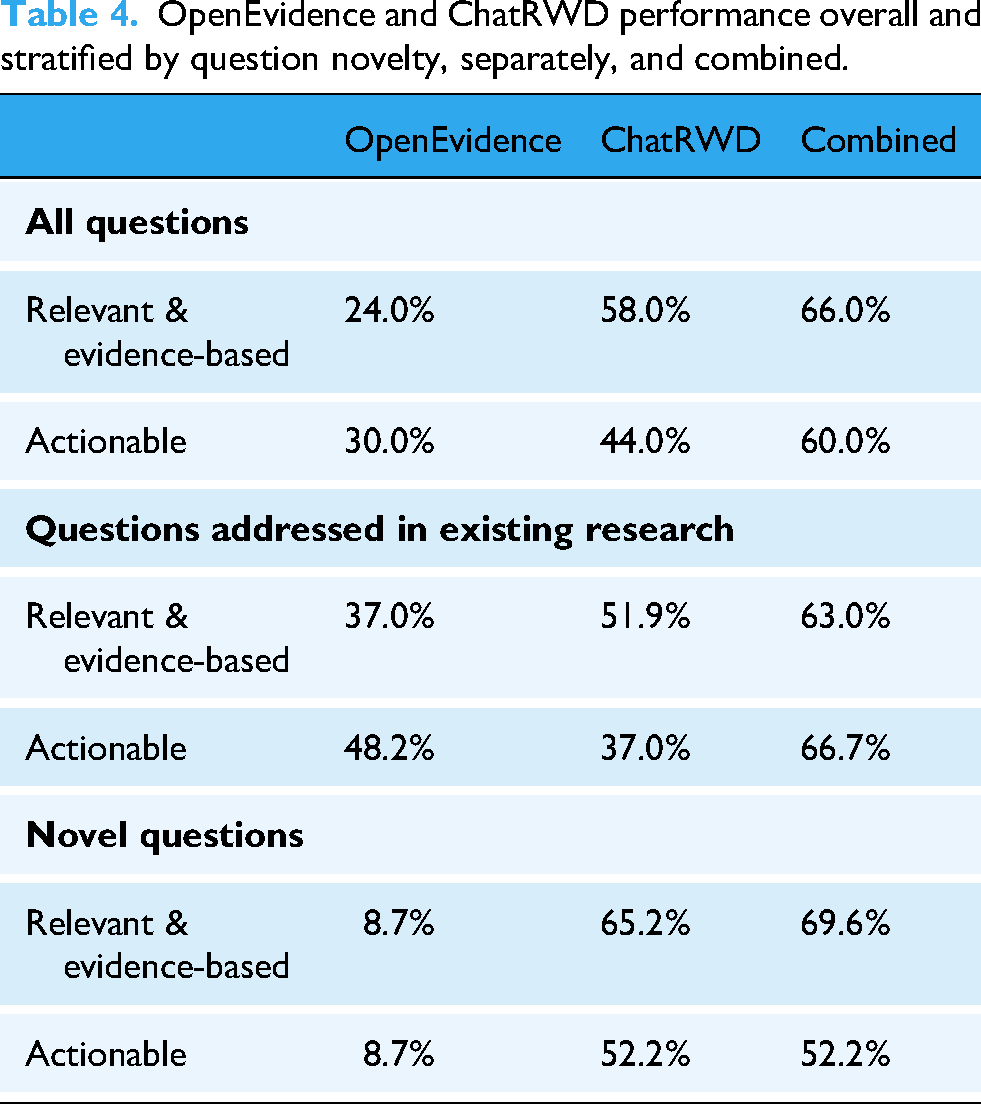
Conversely, on questions that have existing literature, the comparative gap narrowed (37% relevant and evidence-based, 48.2% actionable by OpenEvidence vs. 51.9% relevant and evidence-based, 37.0% actionable by ChatRWD). ChatRWD was more likely to generate answers of varying quality while OpenEvidence would, at worst, provide partially relevant and partially evidence-based answers (Figure 2, Table 4, Supplemental Figure 3). ChatRWD and OpenEvidence were complementary: the frequency of actionable answers increased from 30% for OpenEvidence alone and 44% for ChatRWD alone to 60% for the two combined (Table 4).
Reviewers also noticed a difference between the ChatRWD and OpenEvidence based on the novelty of the questions. One reviewer remarked, “I tended to favor OpenEvidence as its cited studies were both peer-reviewed and generally had more robust study designs (e.g., systematic reviews, RCTs). ChatRWD was most helpful when no studies existed in the published literature for the exact query” (Supplemental Table 7).
Discussion
To practice evidence-based medicine, physicians are best supported with rapid access to reliable summaries of trusted literature and tools for generating on-demand evidence to support the decision at hand. We demonstrated that special-purpose LLM systems, when augmented with specialized knowledge (24%, OpenEvidence) or a causal inference engine (58%, ChatRWD), far outperformed off-the-shelf LLMs (2–10%) in producing relevant and evidence-based answers for the clinical questions examined.
Several factors contributed to the poor performance of general-purpose LLMs in this setting of generating RWE. First, LLMs hallucinate 31 and use noncredible sources. Large language models struggle with appraising sources for relevance, quality, and trustworthiness, a critical task for RWE. Particularly harmful is when LLMs are so adept at mimicking the corpora they have been trained on that it becomes difficult to distinguish fact from fiction. We observed that probable authors, journals, and article titles were often composed together into nonexistent citations, making up nearly 40% of citations reported. Further, general-purpose LLMs are not designed for the complex tasks required for RWE generation: study design classification, PICO extraction, and clinical named-entity recognition. Even LLMs that perform well on medical benchmarks such as the United States Medical Licensing Examination 32 may not generalize across all medical tasks. 33 Finally, an LLM cannot provide responses to novel medical questions whose answer is in content created after the LLM's most recent training completion date. One solution is using RAG to augment an LLM with external data sources like the PubMed knowledge base to provide recent evidence sources to draw from, as OpenEvidence has demonstrated.
However, because there is a considerable time lag between asking a clinical question and publishing a comparative study to answer the question, relying only on past studies is inadequate. Given that almost half of our selected questions were novel at the time of this study, there is a need to rapidly conduct new studies on demand. These new studies are motivated by patients with specific preexisting conditions and medications who do not qualify for clinical trials. It is precisely because clinicians struggle to find relevant and high-quality evidence for such real-world patients that studies have to be performed on demand. 34
Thus, it is unsurprising that ChatRWD, specifically designed to conduct comparative studies on demand, outperformed the general LLMs and the RAG-based OpenEvidence for novel questions. This makes OpenEvidence and ChatRWD complementary. OpenEvidence can provide relevant, evidence-based responses to questions where literature already exists, often making use of high-quality sources such as randomized controlled trials and meta-analyses. On the other hand, ChatRWD can generate new evidence for questions that have not previously been studied in the published literature. In combination, these two tools provided relevant, evidence-based answers to 66% of the questions and were deemed actionable 60% of the time.
Several limitations of this study are worth noting. Foremost is that we restricted our clinical questions to those potentially answerable using RWD: specifically, treatment comparisons that could be assessed with a cohort study design and that used phenotypes already in our phenotype library. As a result, this study focused mainly on questions that lend themselves to comparative effectiveness studies. An analysis focused on different types of questions, such as listing drug–drug interactions, could give different results. Second, it was not possible to blind the reviewers to the models because each model has a distinctive output format. This means we could not control for possible reviewer preference for a given model. Additionally, the novelty rate of the questions may differ in other contexts. We also note that we observed only fair inter-rater reliability across the core measures; this indicates that a different group of reviewers might change the specific percentages. However, the overall trends will likely hold. Also, we did not consider domain-specific LLMs finetuned on medical knowledge which may fare better than the general LLMs we used. 35 ChatRWD was allowed access to a single data source from the United States due to data rights considerations, and results may be different if evaluated using other RWD sources, including data from another country.
We also note two limitations related to the prompting strategy for the general-purpose LLMs. First, these models were prompted using a standard but generic prompt. Optimizing the prompt for each model could potentially improve the performance of these systems. Second, questions were submitted to the general-purpose LLMs programmatically without any iterative revision. This contrasts with ChatRWD, which uses an interactive interface to help correct the inferred PICO and phenotypes (Supplemental Figure 1). We chose this approach to mimic what we expected to be the typical use case for each model. ChatRWD is interactive by nature, so a user would experience the interactive prompting. In contrast, the typical “null” result from the general-purpose LLMs was along with the lines of, “I’m sorry, but I couldn’t find any study comparing [outcome] between patients with [condition] who were treated with [treatment] or [comparator]” (Supplemental Table 3). We expected that many users would not attempt to reprompt the LLM after such an answer, so did not include that in our study. However, it is possible that reprompting the general-purpose LLMs could improve their performance.
Conclusion
Pertinent evidence remains difficult to obtain for many patient care decisions. Challenges in obtaining evidence stem from two sources: (1) nearly 80% of care decisions lack high-quality evidence due to no specific study being available
2
and (2) difficulty in contextualizing available studies for the specific intricacies of the patient at hand. While LLMs excel at summarizing and contextualizing existing literature, either internalized during training or retrieved from external RAG sources, they cannot perform
Supplemental Material
sj-docx-1-dhj-10.1177_20552076251348850 - Supplemental material for Answering real-world clinical questions using large language model, retrieval-augmented generation, and agentic systems
Supplemental material, sj-docx-1-dhj-10.1177_20552076251348850 for Answering real-world clinical questions using large language model, retrieval-augmented generation, and agentic systems by Yen Sia Low, Michael L Jackson, Rebecca J Hyde, Robert E Brown and Neil M Sanghavi, Julian D Baldwin, C William Pike, Jananee Muralidharan, Gavin Hui, Natasha Alexander, Hadeel Hassan, Rahul V Nene, Morgan Pike, Courtney J Pokrzywa, Shivam Vedak, Adam Paul Yan, Dong-han Yao, Amy R Zipursky, Christina Dinh, Philip Ballentine, Dan C Derieg, Vladimir Polony, Rehan N Chawdry, Jordan Davies, Brigham B Hyde, Nigam H Shah, Saurabh Gombar in DIGITAL HEALTH
Footnotes
Acknowledgments
The authors thank OpenEvidence for technical support and permission to use their platform.
ORCID iDs
Ethical considerations
This study uses only de-identified healthcare data, wherein the data provider has no involvement in the research and the research team has no access to linking data that could identify patients. As such, this study is exempt from institutional review board review.
Author contributions
SG, NHS, BH, YL, and NS conceived of the study, defined the main outcomes and measures. YL, MLJ, RH, and RB drafted the manuscript. NS, JB, and SG selected the questions and ran them through ChatRWD. CD ran the questions through OpenEvidence. RH designed the LLM prompts and ran the questions through the general LLMs. RH and SG medical reviewers searched the literature to corroborate the responses from the LLM systems. SG, JM, and NS reviewed the inferred PICO and TQL code. The PICO and TQL evaluation criteria were designed by SG and NS. The medical review rubric was designed by SG, RH, NS, JB, and YL. SG trained the clinical reviewers, NA, HH, RN, MP, CJP, SV, AY, DY, and AZ who performed the clinician review. RH analyzed and summarized the results from the PICO, TQL, clinical reviews with guidance from YL, MLJ, RB, NS, and SG. CWP, MLJ, and YL classified the questions by novelty based on literature searches. The phenotype library was adapted for the study by JB, RB, JM, SG, NS, VP, YL, and PB. ChatRWD was adapted for the study by NS, YL, RB, JB, RC, and JD. A copy of the data for ChatRWD was prepared for this study by DD. All authors reviewed, edited, and approved the final manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by Atropos Health.
Declaration of conflicting interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: ChatRWD, the LLM system evaluated in this study, is developed by Atropos Health where many of the authors are employed. NHS is not an Atropos Health employee but sits on its board. OpenEvidence, another LLM system evaluated here, is provided by OpenEvidence whom we consulted during the writing of this manuscript. Non-Atropos employees NA, HH, RVN, MP, CJP, SV, APY, D-HY, and ARZ, have nothing to disclose.
Data availability statement
The data that support the findings of this study are available from Eversana but restrictions apply to the availability of these data, which were used under license for the current study and so are not publicly available. However, data are available from the authors upon reasonable request and with permission of Eversana.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
