Abstract
Background
Automated breast ultrasound analysis is hindered by limited annotated data, institutional heterogeneity, and strict privacy regulations. This study proposes FAME (Federated Attention-guided Multi-task Ensemble Network), a privacy-preserving and data-efficient framework for joint segmentation and classification of breast ultrasound images in decentralized clinical environments.
Methods
Federated Attention-guided Multi-task Ensemble Network integrates Federated Transfer Learning with class-specific synthetic data generation via Auxiliary Classifier Generative Adversarial Networks to enhance training under data scarcity. Segmentation is performed using a Multi Attention U-Net (MAU-Net), while classification employs a dual-stage ensemble of ResNet50V2, NASNetLarge, and MAU-Net, followed by a meta-classifier. Privacy is preserved through Differential Privacy with Gaussian noise injection and Secure Aggregation for interclient model update protection. The model was trained and validated on the Breast Ultrasound Image (BUSI) dataset (780 images: 80% training, 10% validation, 10% testing) and further evaluated on independent test sets from the Breast Ultrasound Classification (BUSC) (407 images) and UDIAT (163 images) datasets. Statistical significance was assessed using paired
Results
On the BUSI test set, FAME achieved 98.70 ± 0.27% accuracy, 96.82 ± 0.53% F1-score, and 0.978 area under the curve (AUC). On UDIAT, it reached 98.14 ± 0.31% accuracy, 94.04 ± 0.75% F1-score, and 0.960 AUC, while on BUSC, it achieved 96.92 ± 0.27% accuracy, 90.32 ± 0.80% F1-score, and 0.950 AUC. For segmentation, Dice Scores were 89.72 ± 0.53% (BUSI), 93.09 ± 0.49% (BUSC), and 87.98 ± 0.57% (UDIAT), consistently surpassing state-of-the-art baselines. Synthetic augmentation improved performance on underrepresented malignant cases and enhanced generalization under non-IID client data distributions.
Conclusion
Federated Attention-guided Multi-task Ensemble offers a scalable, privacy-compliant, and high-performing solution for multi-institutional breast ultrasound analysis. By combining federated learning, synthetic augmentation, and attention mechanisms, it provides a strong foundation for secure, collaborative breast cancer diagnosis.
Keywords
Introduction
Breast cancer remains one of the most prevalent and life-threatening malignancies among women globally. 1 The early detection and accurate classification of breast tumors are critical in improving survival rates and guiding effective treatment decisions. Among various imaging modalities, ultrasound (US) imaging has gained prominence due to its noninvasive nature, cost-effectiveness, and ability to effectively detect abnormalities in dense breast tissue. 2 The automated analysis of breast US (BUS) images introduces several significant challenges that hinder the development of reliable computer-aided diagnosis systems.3–6 Ultrasound images are inherently affected by speckle noise, low contrast, and operator dependency, which complicates accurate segmentation and classification.7,8 These artifacts require manual interpretation by experienced radiologists, making the process subjective and time-consuming.
Training deep learning (DL) models for tumor analysis necessitates large volumes of annotated data, which are difficult to obtain in clinical settings due to the high cost and expertise required for accurate labeling. 9 Medical institutions face stringent data privacy regulations (e.g., HIPAA and GDPR) that prevent the centralization of patient data across institutions. As a result, collaborative learning efforts are severely limited, leading to poor model generalization and limited deployment in real-world clinical scenarios. Datasets collected from different sources often exhibit significant heterogeneity in image acquisition protocols, resolution, and demographic characteristics, making it difficult for conventional centralized models to adapt to domain variations.
To clarify the motivation behind our proposed solution, we outline the key challenges in automated BUS analysis and the specific strategies employed to address them. (i) Data privacy restrictions across healthcare institutions are addressed via a Federated Transfer Learning (FTL) model, which enables decentralized model training without sharing raw data. (ii) Domain heterogeneity, arising from differences in imaging protocols and equipment, is tackled using a dual-stage ensemble classifier combining Multi Attention U-Net (MAU-Net), NASNetLarge, and ResNet50V2, which enhances generalization across non-IID distributions. (iii) Annotated data scarcity is mitigated by incorporating Auxiliary Classifier Generative Adversarial Networks (ACGAN) that generate synthetic, class-specific images locally to improve training diversity. (iv) Ultrasound imaging noise and low contrast, which obscure tumor boundaries, are handled by enhancing the U-Net into the MAU-Net, integrating both channel and spatial attention modules. (v) Security vulnerabilities in model parameter sharing, such as model inversion and membership inference attacks, are countered through Differential Privacy (DP) and Secure Aggregation (SA) techniques. These components collectively define the architecture of our Federated Attention-guided Multi-task Ensemble Network (FAME) model.
To address these challenges, we propose a novel FAME, an ensemble DL model that employs FTL ACGAN to enable privacy-preserving and data-efficient BUS image classification and segmentation. FTL facilitates decentralized model training across multiple institutions without transferring raw patient data, thereby preserving patient confidentiality and complying with regulatory standards. While Federated Learning (FL) allows for collaborative training without data exchange, it is not entirely immune to privacy risks such as model inversion and membership inference attacks through which sensitive patient information may be inferred from shared model updates. 10 To mitigate this, our study emphasizes distributed training as a key privacy-preserving strategy. It further integrates additional safeguards (e.g., SA and DP) to reduce the risk of information leakage during training. Simultaneously, ACGAN is incorporated to generate synthetic, class-specific US images, augmenting the training dataset and alleviating data scarcity. These synthetic images improve model robustness, particularly in underrepresented categories, and help prevent overfitting.
At the core of our design is a dual-stage feature fusion classifier that combines the outputs of three DL models: NASNetLarge, ResNet50V2, and the proposed MAU-Net. The MAU-Net introduces attention-based mechanisms (channel and spatial) within a U-Net backbone to enhance segmentation performance by focusing on the most informative regions of the image. The multimodel ensemble approach enables the system to capture low-level and high-level features, ensuring robust tumor segmentation and accurate classification into normal, benign, and malignant categories. The novelty points of this study are as follows:
We propose a FTL model that enables decentralized collaborative training across distributed medical institutions, ensuring privacy preservation without sacrificing model performance. The model employs SA and DP to mitigate common threats such as model inversion and membership inference. We introduce the use of ACGANs to generate class-specific synthetic US images at the local client level, effectively augmenting limited datasets and improving model generalization in underrepresented categories. We develop a dual-stage ensemble learning architecture that synergistically fuses discriminative features from three advanced backbones, NASNetLarge, ResNet50V2, and an attention-enhanced MAU-Net to achieve improved segmentation and robust three-class classification. We validate the effectiveness of our model through comprehensive experimentation on three benchmark BUS datasets (Breast Ultrasound Image [BUSI], UDIAT, Breast Ultrasound Classification [BUSC]), demonstrating consistent outperformance of existing state-of-the-art (SOTA) models in both segmentation metrics (Dice Score, Intersection over union [IoU]) and classification measures (Accuracy [ACC], F1-score, area under the curve [AUC]).
This integrated solution presents a scalable, privacy-preserving, and clinically reliable approach for breast tumor analysis using US imaging, offering the promising potential for deployment in multi-institutional healthcare environments.
The application of DL to BUS image analysis has gained substantial momentum due to its ability to automatically extract hierarchical features for both segmentation and classification tasks. 11 However, existing models often rely on centralized datasets, face performance degradation in heterogeneous data environments, and lack privacy-preserving mechanisms, limitations that directly motivate the approach proposed in our study. Several prior studies have demonstrated the efficacy of CNNs in BUS image classification. 12 For instance, 13 applied patch-based U-Net, LeNet, and FCN-AlexNet to a small-scale BUS dataset and demonstrated reasonable classification performance. Similarly, the study 9 improved classification ACC using data augmentation and DAGAN, with NASNet achieving 94% ACC when trained on the enhanced dataset. These efforts highlight the significance of augmentation in addressing data limitations. However, they rely on centralized training, limiting their real-world applicability due to data privacy constraints. The study 14 introduced a grayscale-to-RGB mapping technique and fine-tuned VGG19 on 882 US images, outperforming expert radiologists. The study 15 used Mask R-CNN for simultaneous segmentation and classification, but the performance was highly dependent on annotated data availability. The study 16 compared models such as YOLO, VGG16, Fast R-CNN, and ZFNet using 1043 US images, emphasizing the importance of balanced datasets and multiscale features. While these models exhibit promising results, their reliance on manually annotated datasets and centralized learning remains a key limitation.
Transfer learning techniques have also been employed to enhance BUS classification. The study 17 proposed a Multiview InceptionV3-based CNN architecture, while the study 18 used ensemble learning of CNNs with RGB-fused US images to boost diagnostic ACC. Likewise, the study 19 incorporated DenseNet121 with attention to ROI localization, and the study 20 employed ResNet101 with SVM for classification. Although these architectures offer robust feature representations, they are not inherently designed for distributed learning or data privacy preservation. Recent efforts have focused on enhancing spatial and contextual feature capture for segmentation. The study 21 proposed a multistream segmentation network incorporating global and local features, while the study 22 used adaptive spatial fusion across multiple models to improve classification on the BUSI dataset. The study 23 introduced MTL-COSA, a multitask learning architecture with context-aware self-attention, to jointly perform segmentation and classification. Transformer-based models have also emerged, such as SaTransformer 24 and BUViTNet, 25 which leveraged global self-attention for better boundary delineation. However, these models are computationally intensive and lack integration with privacy-aware training protocols. In parallel, the use of generative models to address data scarcity has also been explored. The study 26 applied a radiomics-based pipeline for classification, while GAN-based models 9 were used for synthetic data generation. Yet, these works often fail to generate class-specific US images with diagnostic relevance, as addressed by our use of ACGAN.
Despite growing interest in FL in healthcare, its application in BUS imaging remains underexplored. Recent surveys, such as,10,27 which review privacy leakage threats and mitigation techniques in secure FL, and,10,27 which explores privacy-preserving FL in remote sensing under adversarial conditions, outline the vulnerability of FL to model inversion and membership inference attacks. These concerns necessitate stronger privacy guarantees, such as SA and DP, especially when dealing with sensitive clinical data. Most BUS studies do not account for these risks, highlighting a critical gap in deploying DL models across distributed medical institutions. In summary, while previous studies have addressed individual aspects such as augmentation, segmentation, attention mechanisms, or classification, there is a lack of a unified, privacy-preserving solution that simultaneously tackles data scarcity, segmentation precision, classification robustness, and data privacy. Our study uniquely addresses these gaps by employing FTL for decentralized training without raw data exchange, using ACGAN to generate class-conditioned synthetic US images, introducing a dual-stage ensemble model combining ResNet50V2, NASNetLarge, and the novel MAU-Net with channel and spatial attention modules, and demonstrating consistent improvements across segmentation and classification tasks on three benchmark datasets (BUSI, UDIAT, BUSC). This integrated strategy offers a scalable and secure pathway for real-world deployment of DL-based breast cancer diagnosis tools across multi-institutional settings.
Methodology
This is a retrospective study utilizing three publicly available BUS datasets (BUSI, UDIAT, and BUSC), all of which were previously collected and anonymized by their respective sources. To enhance tumor segmentation in US images, we extend the U-Net; our encoder is constructed using residual blocks, enabling deeper feature extraction while mitigating vanishing gradient issues during training. Each residual block consists of two stacked 1 × 1 convolutional layers, each followed by batch normalization and ReLU activation, with a nonidentity skip connection to enhance feature propagation. This design improves gradient flow and preserves contextual integrity across network depths. Multi Attention U-Net employs dual attention mechanisms to refine the encoded features before propagating them to the decoder. Channel attention modules are used to recalibrate the importance of feature channels by applying global average and max pooling, followed by shared fully connected layers. This allows the model to learn features that are most relevant for identifying tumor regions. Spatial attention modules identify the location of important features in the spatial dimensions using 2D convolution over concatenated pooled feature maps. Both attention types are strategically inserted after the residual blocks and before skip connections, enhancing semantic consistency between encoder and decoder representations.
Federated transfer learning
Federated transfer learning enables decentralized training of DL models across multiple healthcare institutions without requiring the exchange of raw patient data. In this setting, each institution 
To improve convergence and address data heterogeneity, each local model is initialized with pretrained weights from large-scale public datasets (e.g., ImageNet). Transfer learning enables institutions with small or imbalanced datasets to adapt shared representations effectively. During each communication round, local models are updated using gradient descent, as shown in equation (2):

Auxiliary classifier GAN-based synthetic image generation
While FTL enables privacy-preserving collaborative learning across decentralized institutions, it remains constrained by the availability of labeled medical data at each site. Medical imaging datasets, particularly in BUS applications, often suffer from class imbalance and limited annotated samples due to the cost and expertise required for labeling. To address this limitation, we incorporate an ACGAN at each local institution to augment the training dataset with high-quality, class-specific synthetic images, thereby improving model generalization and stability without compromising patient privacy. Auxiliary Classifier GAN is an extension of the standard Generative Adversarial Network (GAN) framework, where both the generator
The adversarial training objectives for the ACGAN are defined as follows. The generator loss 
The discriminator loss 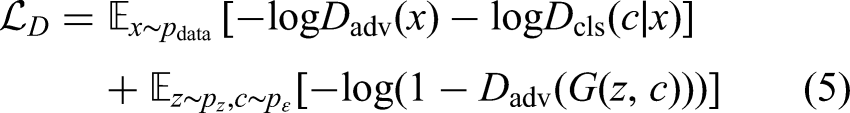
Here,
Image preprocessing
Following the generation of accurate and synthetic US images via ACGAN at each local institution, a standardized preprocessing pipeline is applied to ensure that all images, regardless of source, are normalized and optimized for subsequent segmentation and classification tasks. The preprocessing stage is critical in improving image quality, mitigating modality-specific noise artifacts, and ensuring consistency across decentralized datasets, which is essential in FL environments where data heterogeneity is a key challenge. Let
Resizing
All images are resized to a fixed resolution 
This uniform scaling standardizes input dimensions across datasets (BUSI, BUSC, UDIAT), which originally differ in resolution and aspect ratio (e.g., BUSI images at 500 × 500 px vs. BUSC images at 128 × 128 px). Without resizing, network layers may encounter mismatched feature dimensions, complicating training convergence. Resizing also ensures efficient GPU memory utilization and stable batch processing during federated training.
Speckle noise reduction
Ultrasound imaging is inherently affected by speckle noise, which arises from coherent interference of returning echoes and appears as granular patterns. Such noise obscures lesion boundaries and degrades the performance of feature extractors. To mitigate this, we apply a hybrid denoising strategy using median and Gaussian filtering, as shown in equation (7):
Median filtering is effective at removing salt-and-pepper noise while preserving edges. Gaussian filtering suppresses high-frequency noise and smooths homogeneous regions. The combination is particularly effective for US images, balancing edge preservation and noise suppression. By reducing speckle noise prior to feature extraction, the model receives inputs with enhanced lesion-to-background contrast, improving both segmentation ACC and classifier reliability.
Intensity normalization
Due to differences in acquisition devices (e.g., LOGIQ E9, Siemens ACUSON) and institution-specific imaging protocols, raw US images exhibit significant variation in brightness, contrast, and dynamic range. To mitigate these variations, images were standardized using 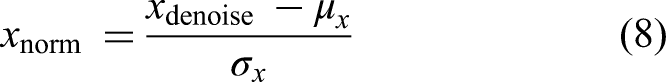
Data augmentation
We apply stochastic data augmentation transformations to enhance generalization further and mitigate overfitting due to limited data, especially in underrepresented classes (e.g., malignant cases). Each image is randomly transformed using a combination of:
Rotation: Horizontal/Vertical Flipping Zooming: scale factor
Mathematically, an augmented image 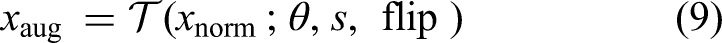
Multi Attention U-Net architecture
The proposed MAU-Net architecture is a multiattention extension of the standard U-Net, tailored specifically for BUS image segmentation, as illustrated in Figure 1. It follows a fully convolutional, symmetric encoder–decoder structure designed to learn hierarchical spatial features while preserving high-resolution details necessary for precise tumor boundary delineation. The input to the network is a preprocessed grayscale US image 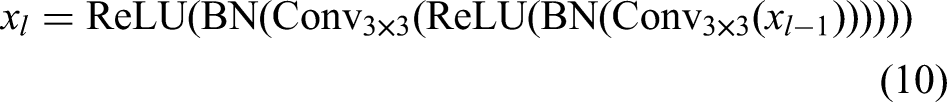

Architecture of the proposed MAU-Net segmentation network enhanced with dual attention mechanisms. The network consists of a symmetric encoder–decoder design with skip connections, where each encoding block captures hierarchical spatial features through convolution and max pooling. Channel and spatial attention modules are integrated into the skip pathways to refine the flow of discriminative features. The decoder mirrors the encoder structure with upsampling and concatenation operations, followed by convolutional refinement. The final pixel-wise segmentation is achieved through a classification block using 1 × 1 convolution and softmax activation. Bottom: Detailed view of the channel and spatial attention mechanisms used to recalibrate feature responses along with channel dimensions and spatial locations.
After the double convolution, a 
At each subsequent level, the number of feature channels is doubled to allow the network to capture increasingly complex and abstract semantic features. If the first encoder block outputs 64 channels, the next block will output 128, then 256, and so on, resulting in a channel expansion pattern of
The channel attention mechanism emphasizes informative channels (e.g., tumor texture), while spatial attention focuses on spatially relevant regions (e.g., tumor location), thereby improving feature discrimination in noisy US environments. The decoder path mirrors the encoder structure, with each decoding block consisting of a transpose convolution (also called upconvolution or deconvolution) for upsampling, followed by a concatenation of the corresponding encoder features (skip connection) and two standard convolutional layers. For the decoder level 
The skip connections between the encoder and decoder ensure that fine-grained spatial information lost during pooling is preserved during reconstruction. This fusion of low-level and high-level features is essential for accurately capturing BUS images’ heterogeneous and often indistinct lesion boundaries.
At the final decoder layer, a 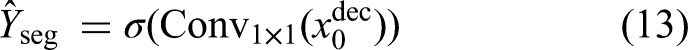
Here, each pixel value in
Channel attention module
The Channel Attention
Given an intermediate feature map

These descriptors are passed through a shared multilayer perceptron (MLP) composed of two fully connected layers with a ReLU activation. Let 

This mechanism effectively boosts channels critical for identifying tumor textures or contrast changes while suppressing irrelevant background or noise channels. The integration of channel attention within the MAU-Net bottleneck and decoder stages improves the model's ability to focus on semantically rich and diagnostically relevant features, enhancing segmentation performance in US data characterized by low contrast and speckle noise.
Spatial attention module
Complementing the channel attention mechanism, the Spatial Attention (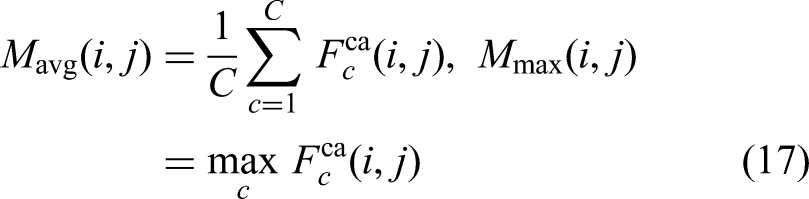


This operation ensures that regions with high attention scores (e.g., tumor boundaries, dense textures) are amplified while background regions or irrelevant structures are suppressed. By integrating spatial attention into the decoder blocks of MAU-Net, the network can better reconstruct accurate segmentation masks, particularly in complex or ambiguous regions of BUS imagery. With channel attention, the spatial attention module forms a powerful dual-attention mechanism that enables MAU-Net to dynamically prioritize critical features and their spatial locations, enhancing segmentation robustness across datasets with varying imaging conditions.
Segmentation output and loss
At the final stage of the MAU-Net architecture, the decoder produces a high-resolution feature map 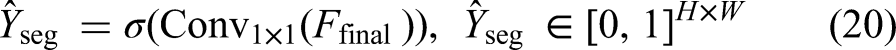
Here, 
To train the segmentation model effectively in the context of class imbalance often present in medical datasets, employ the Dice Loss, which directly optimizes for the Dice Similarity Coefficient, a widely used metric in medical image segmentation. The Dice Loss 
Ensemble strategy with cross-validation
To enhance the robustness and generalization of the segmentation output within each federated institution, we incorporate a three-fold cross-validation ensemble strategy during MAU-Net training. This technique mitigates overfitting, accounts for data heterogeneity across institutions, and ensures more stable lesion boundary prediction in both real and ACGAN-augmented datasets. The ensemble also strengthens local models before global federated aggregation, aligning with the decentralized learning paradigm of our study. During training, the local dataset 

This final segmentation mask
Classification network: dual-stage feature fusion
Following segmentation, the next stage of the pipeline is the classification of BUS images into three diagnostic categories: Normal, Benign, and Malignant. The “Normal” category refers to US images showing no abnormal tissue or lesions, whereas the “Benign” category includes nonmalignant abnormalities such as fibroadenomas or cysts. The “Malignant” category refers to cancerous tumors requiring clinical intervention. To ensure high ACC and robustness, we propose a dual-stage feature fusion classifier, which integrates the strengths of three diverse yet complementary DL backbones-ResNet50V2, NASNetLarge, and MAU-Net-through a two-tiered architectural strategy as shown in Figure 2. Each input image, whether real or ACGAN-generated, is first passed through the three base networks in parallel. Let 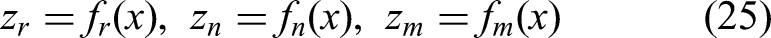
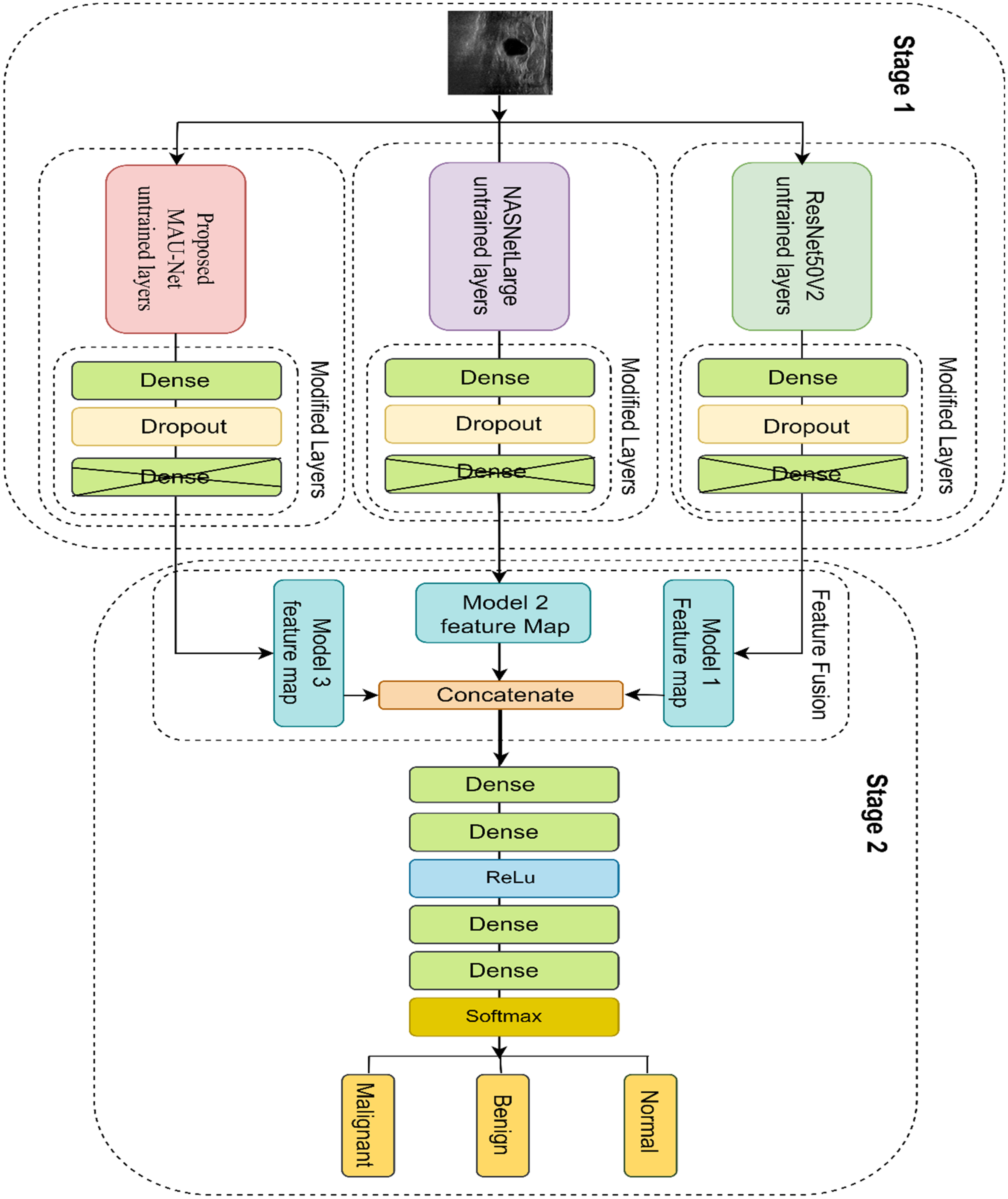
Our developed dual-staged feature fusion-based classifier framework. In Stage 1, feature maps are extracted from the untrained layers of three base models: Proposed MAU-Net, NASNetLarge, and ResNet50V2, each followed by custom dense and dropout layers. In Stage 2, the feature maps from the three models are concatenated and passed through additional dense layers with ReLU activation, leading to a final softmax layer for classifying the input image as Malignant, Benign, or Normal.
Each feature vector
On top of each frozen backbone, we append a customized classification head composed of a fully connected (Dense) layer with 1024 units and ReLU activation to introduce nonlinearity. To reduce the risk of overfitting, a Dropout layer with a rate of 0.3 follows. The final classification output is produced by a fully connected Dense layer with 3 neurons, activated by a Softmax function to yield normalized class probabilities across the three diagnostic categories. Instead of simply averaging the predictions from the three models, we proceed to a second fusion stage that combines the learned representations into a higher-level ensemble. Specifically, the intermediate feature vectors 
The model is trained using the categorical cross-entropy loss function defined as shown in equation (27):

Implementation details
The proposed FAME FL-based architecture for BUS segmentation and classification was implemented and evaluated using a three-client simulated federation, with each client node representing a distinct medical institution. To ensure replicability and efficiency across nodes, we adopted a client–server topology, where the clients perform local training, and a central server coordinates global model aggregation via FedAvg. The overall architecture of our proposed system, combining FTL with ACGAN-based augmentation and personalization via transfer learning, is illustrated in Figure 3. It highlights the interactions between local models, the central server, auxiliary GAN training, and cross-domain deployment for unseen clients. All DL models, MAU-Net for segmentation and ResNet50V2, NASNetLarge, and MAU-Net for classification were implemented using TensorFlow 2.11 and Keras APIs. The training was conducted on NVIDIA RTX 3090 GPUs (24GB VRAM) for centralized simulation and client-side experiments. To simulate realistic federated constraints, each node had access to non-IID subsets of the datasets (BUSI, UDIAT, BUSC), with synthetic data augmentation applied locally using the ACGAN module described previously. During segmentation, ensembling was applied at each client to generate a single binary mask prediction. Classification models were locally fused using our dual-stage feature fusion pipeline. The input images were resized to
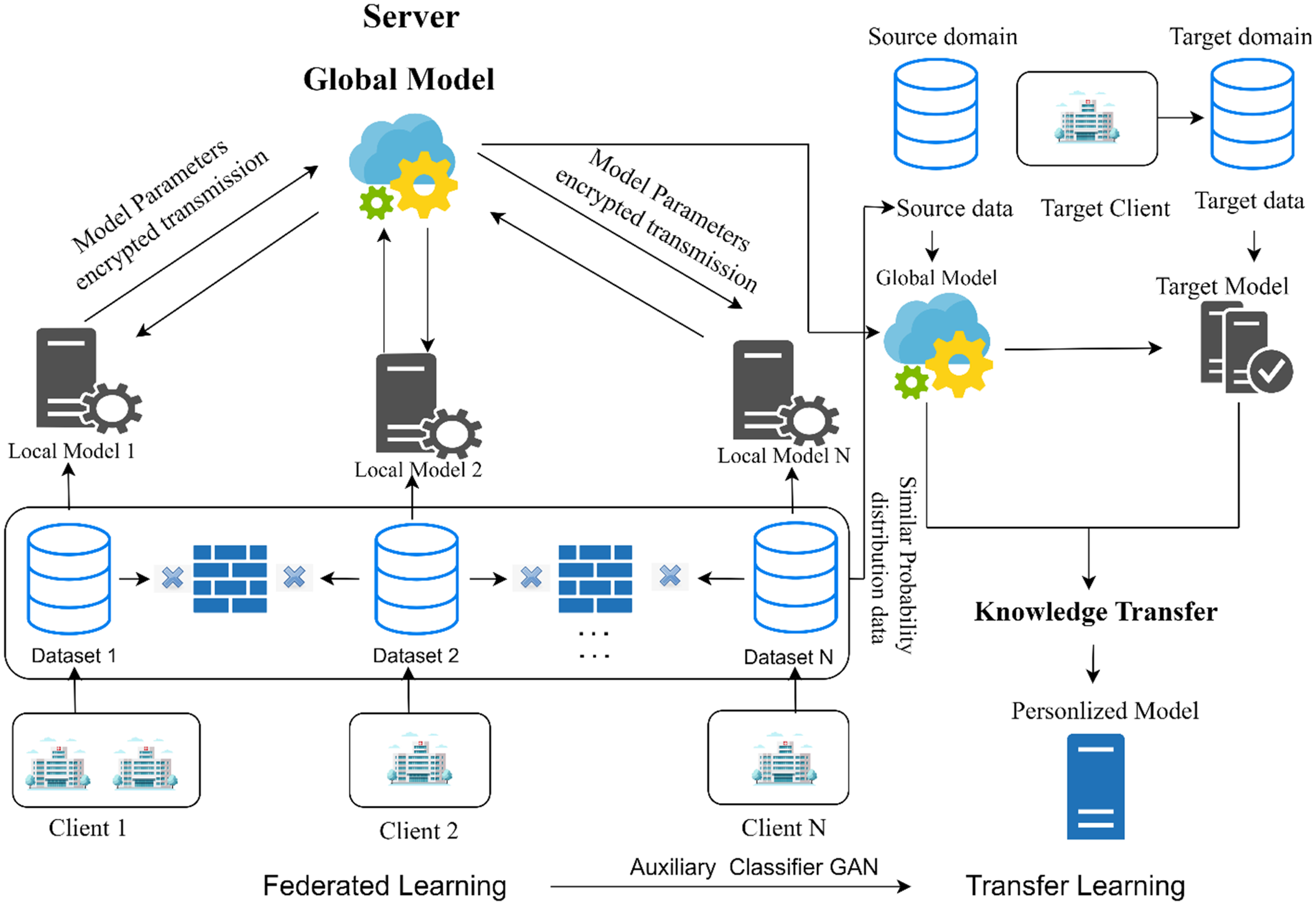
The proposed federated learning framework integrating Auxiliary Classifier GAN (ACGAN) and transfer learning. Multiple clients (e.g., hospitals) train local models on private data without sharing raw images. The server aggregates encrypted model parameters to update a global model, which is distributed back to all clients. Simultaneously, ACGANs generate class-specific synthetic data locally to enhance performance. The global model is transferred to target clients for personalized fine-tuning using transfer learning, ensuring domain adaptation to new clinical environments.
Hyperparameters were tuned through grid search, where batch size (8–16), learning rate (1e-4–1e-5), and dropout rate (0.3–0.5) were varied. The Adam optimizer with an initial learning rate of 1e-4 was selected, with ReduceLROnPlateau (factor 0.1, patience = 5) for scheduling. Early stopping (patience = 10) was employed to prevent overfitting. The final configuration was chosen based on three-fold cross-validation performance across BUSI, BUSC, and UDIAT datasets.
The federated aggregation was performed every five local training rounds, where encrypted model weights were sent to a central aggregator and updated via FedAvg, as shown in equation (28):
Summary of training parameters for segmentation and classification networks.

Datasets
This research used the BUSI dataset 28 to evaluate the effectiveness of the proposed architecture. The dataset, sourced in 2018 from Baheya Hospital for Early Detection and Treatment of Women's Cancer in Cairo, Egypt, includes 780 US images accompanied by ground truth segmentation masks. Each image measures 500 × 500 pixels in resolution. The dataset is divided into three categories: 437 benign images, 210 malignant images, and 133 normal images. These scans were collected from 600 female participants, aged 25–75, using the LOGIQ E9 and LOGIQ E9 Agile US devices. Ground truth tumor masks were manually delineated by expert radiologists at Baheya Hospital using MATLAB freehand segmentation, and all annotations were reviewed for ACC and consistency. 28 This dataset provides diverse examples of BUS images, serving as a vital resource for training and validating machine-learning models in breast cancer diagnosis. Each image in the dataset is accompanied by a corresponding binary ground truth mask delineating the tumor region (if present), created by expert radiologists. This dataset presents real-world class imbalance, particularly with fewer malignant cases, which is reflective of screening population distributions. It is frequently used in breast cancer segmentation benchmarks and is particularly valuable for training models in weakly supervised or FL contexts due to its high-quality annotations and balanced image resolution.
The UDIAT dataset 13 was used as the final benchmark to evaluate the performance of the proposed model. Collected from the UDIAT Diagnostic Center at Parc Taulí in Sabadell, Spain, it consists of 163 US images paired with corresponding segmentation masks for ground truth verification. The images have an average resolution of 760 × 570 pixels. The dataset is split into two categories: 109 benign tumor images and 54 malignant tumor images. Captured using the Siemens ACUSON Sequoia C512 system, lesion boundaries were annotated by radiologists at the UDIAT Diagnostic Centre, 29 the UDIAT dataset provides a valuable resource for validating BUS machine-learning models in tumor diagnosis. While the dataset is smaller in size compared to BUSI, it offers high-resolution scans with a moderate degree of visual heterogeneity in lesion texture and shape. Its relatively balanced benign/malignant distribution makes it useful for evaluating generalization and robustness of classification and segmentation models.
The Mendeley BUSC dataset 30 consists of 100 benign and 150 malignant US images. The real resolution of the images is 64 × 64 pixels, which was subsequently transformed to 128 × 128 pixels for this research. As this dataset is primarily intended for classification, it does not include corresponding ground truth segmentation masks. To address this, an experienced radiologist assisted in annotating the benign and malignant tumor images, making them suitable for segmentation tasks. 31 These expert annotations were crucial for adapting the BUSC dataset to support segmentation-based model evaluation. The BUSC dataset was designed primarily for binary classification, reflecting challenging real-world variability in lesion echotexture and boundary clarity. Despite the absence of normal images, its inclusion provides valuable insight into model performance under limited-resolution and class-skewed conditions. The dataset's compact size and lack of metadata (e.g., patient age, acquisition protocol) require careful preprocessing and augmentation to avoid overfitting.
Evaluation metrics
Egmentation metrics
Dice similarity coefficient
The Dice Score is a measure of overlap between the predicted segmentation 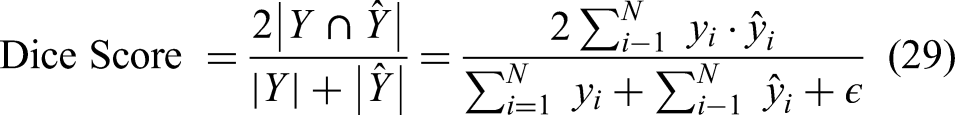
Intersection over union
Also known as the Jaccard Index, IoU measures the overlap between predicted and true regions relative to their union, as shown in equation (30):
These metrics are reported per image and averaged across the test set to assess segmentation consistency and ACC.
Classification metrics
Accuracy
Accuracy represents the proportion of correctly classified samples among all predictions, as shown in equation (31):
F1-score
The F1-score is the harmonic mean of precision and recall, offering a balance between the two, especially important when a class imbalance exists, as shown in equation (32):
Sensitivity (recall)
Sensitivity, or recall, measures the proportion of actual positives correctly identified:
It is particularly important in medical diagnosis tasks, where missing a positive case (e.g., a malignant tumor) is highly undesirable.
Specificity
Specificity measures the proportion of actual negatives correctly identified:
High specificity indicates the model effectively avoids false alarms or misclassification of normal or benign cases.
Area under the curve
Area under the curve evaluates the area under the receiver operating characteristic (ROC) curve, measuring the model's ability to distinguish between classes across varying thresholds. It is defined as shown in equation (33):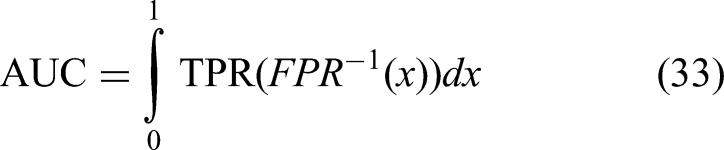
Experimental results
Segmentation
In this study, Dice Score and IoU in equations (29) and (30) were used to evaluate segmentation ACC. Dice quantifies overlap between predicted and ground truth masks, while IoU provides a stricter measure of spatial agreement. The model was trained with an 80:10:10 split on BUSI and tested on BUSI, BUSC, and UDIAT. On BUSI, FAME achieved the highest performance with a Dice of 89.72 ± 0.53%, IoU of 84.81 ± 0.57%, sensitivity of 91.11 ± 0.53%, and specificity of 98.13 ± 0.35%, outperforming TransUNet, D-LinkNet, and ATFE-Net, as shown in Table 2. On BUSC, FAME achieved 93.09 ± 0.49% Dice, 87.13 ± 0.66% IoU, and a sensitivity of 87.41 ± 0.71, along with 99.59 ± 0.27% specificity, surpassing PDF-UNet, MSU-UNet, and DeepLab, as shown in Table 3. On UDIAT, FAME achieved 87.98 ± 0.57% Dice, 78.16 ± 0.66% IoU, 87.41 ± 0.62% sensitivity, and 99.61 ± 0.22% specificity, consistently outperforming ATFE-Net and Axial-DeepLab, as shown in Table 4. Figure 4 presents qualitative segmentation results on BUSI test images using UTNet, LinkNet, TransUNet, D-LinkNet, Axial-DeepLab, ATFE-Net, and FAME. Each row shows different cases with varying tumor sizes, shapes, and textures. Ground truth masks and predicted outputs are displayed for comparison.

Visual comparison of breast lesion segmentation results on the BUSI dataset. From left to right: original ultrasound image, ground truth (GT) mask, predictions from UTNet, LinkNet, TransUNet, D-LinkNet, Axial-DeepLab, ATFE-Net, and our proposed model (FAME), followed by the overlay of FAME prediction on GT. The blue regions represent the predicted lesion masks.
Performance comparison of segmentation methods on the BUSI dataset.

Results are reported as mean ± SD (95% CI).
Performance comparison of segmentation methods on the BUSC dataset.
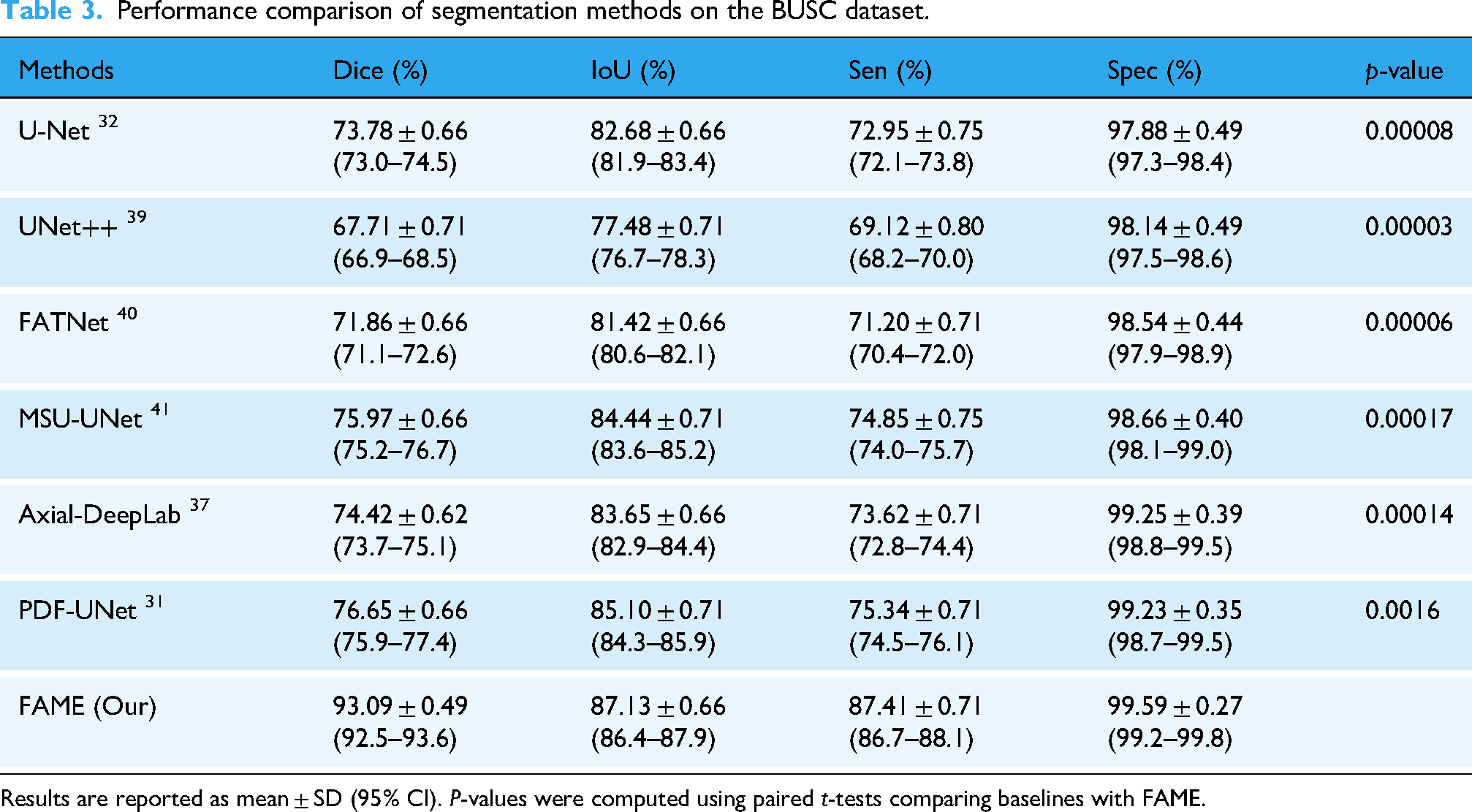
Results are reported as mean ± SD (95% CI).
Performance comparison of segmentation methods on the UDIAT dataset.

Results are reported as mean ± SD (95% CI).
To benchmark FAME, we compared it against widely used segmentation and classification baselines. For segmentation, U-Net, a canonical encoder–decoder with skip connections, and UNet++, which introduces nested dense skip pathways to reduce semantic gaps, were included. LinkNet was employed as a lightweight residual encoder–decoder optimized for efficiency, while TransUNet combines convolutional encoders with Transformer blocks to capture both local and global dependencies. D-LinkNet leverages dilated convolutions and residual links for multiscale receptive fields, and Axial-DeepLab applies axial attention to efficiently model long-range spatial dependencies. ATFE-Net and FATNet incorporate channel–spatial attention and feature aggregation modules, respectively, to enhance tumor feature extraction, whereas MSU-UNet extends U-Net to address lesions of varying size and morphology. PDF-UNet employs progressive dense feature fusion to improve boundary localization.
Figure 5 shows qualitative results on UDIAT test images with diverse lesion characteristics, including small tumors, blurry boundaries, and variable echotexture. Predictions from LinkNet, UTNet, D-LinkNet, and FAME are compared against ground truth masks. Figure 6 presents qualitative segmentation results on BUSC test images using U-Net, UNet++, FATNet, MSU-UNet, DeepLab, PDF-UNet, and FAME. BUSC samples are shown to highlight noisy, low-contrast conditions, and challenging lesion shapes. Predicted masks and ground truth annotations are displayed for direct comparison.

Qualitative comparison of lesion segmentation results on the UDIAT dataset. Each row shows an original ultrasound image, its ground truth (GT) mask, and the predicted masks from UTNet, LinkNet, TransUNet, D-LinkNet, Axial-DeepLab, ATFE-Net, and our proposed model (FAME), followed by FAME's overlay on the GT. Blue regions indicate predicted tumor areas.

Visual segmentation comparison on the BUSC dataset. Each row displays an original breast ultrasound image, its corresponding ground truth (GT) mask, predictions from U-Net, UNet++, FATNet, MSU-UNet, DeepLab, PDF-UNet, and the proposed model (EDRNet/FAME), followed by an overlay of our model's prediction on the GT. Blue regions indicate predicted lesion masks.
Classification
Classification performance was evaluated using ACC, AUC, and F1-score. Accuracy measures the proportion of correctly predicted samples, while AUC assesses discriminative ability using macro-averaging, and F1-score balances precision and recall. A confusion matrix was also constructed to illustrate prediction distribution across classes. Using the dual-stage ensemble classifier, FAME was tested on the BUSI, UDIAT, and BUSC datasets and compared against baseline architectures. Table 5 summarizes the results. On BUSI, FAME achieved 98.70 ± 0.27% ACC, 96.82 ± 0.53% F1-score, and 0.978 AUC, outperforming GAN + CNN, MTL-COSA, and SaTransformer. On UDIAT, it achieved 98.14 ± 0.31% ACC, 94.04 ± 0.75% F1-score, and 0.960 AUC, surpassing FMRNet, RMTL-Net, and HoVer-Trans. On BUSC, it obtained 96.92 ± 0.27% ACC, 90.32 ± 0.80% F1-score, and 0.950 AUC, outperforming MDA-Net and other approaches. For classification, GAN + CNN combines synthetic augmentation with convolutional classification, while CNN-based image fusion integrates multistream CNN features for improved discriminative power. MTL-COSA adopts a multitask learning strategy with cross-task attention, and FMRNet applies a residual CNN tailored to US to mitigate noise sensitivity. Transformer-based models were also included: BUViTNet adapts vision transformers with patch-based tokenization, SaTransformer leverages self-attention for robust classification, and HoVer-Trans captures both local and global structure. RMTL-Net employs residual multitask optimization, and MDA-Net introduces multiscale discriminative attention for adaptive feature recalibration.
Quantitative comparison of the classification performance of the proposed ensemble DL method with SOTA models on the BUSI, UDIAT, and BUSC datasets.

Results are reported as mean ± SD (95% CI).
Confusion matrices were generated for BUSI, UDIAT, and BUSC to evaluate classification performance in Figures 7, 8, 9. Each matrix summarizes true positives, true negatives, false positives, and false negatives, providing a detailed view of the model's predictions against ground truth labels. Across all datasets, the proposed ensemble classifier showed minimal misclassification, supporting the reported ACC, F1-score, and AUC results. To further assess classification performance, ROC curves were plotted for the BUSI, UDIAT, and BUSC datasets, shown in Figure 10. Across all datasets, the proposed FAME framework achieved consistently higher true positive rates at varying thresholds compared to baseline methods. Specifically, FAME obtained AUC scores of 0.978 on BUSI, 0.960 on UDIAT, and 0.950 on BUSC, outperforming competing approaches such as FMRNet, MTL-COSA, and MDA-Net. These results illustrate FAME's superior ability to distinguish between benign and malignant lesions and confirm the quantitative findings reported in Table 5.

Confusion matrix for the proposed FAME framework on the BUSI dataset, illustrating classification performance between normal, benign, and malignant cases.

Confusion matrix for the proposed FAME framework on the UDIAT dataset, illustrating classification performance between benign and malignant cases.

Confusion matrix for the proposed FAME framework on the BUSC dataset, illustrating classification performance between benign and malignant cases.

Receiver operating characteristic (ROC) curves of FAME and baseline models across three datasets. (A) BUSI dataset, (B) UDIAT dataset, and (C) BUSC dataset.
To mitigate overfitting, several regularization and generalization strategies were employed. These include three-fold cross-validation, data augmentation (e.g., rotation, flipping, zooming), and the use of Dropout layers (rate = 0.3) in classification heads. L2 weight regularization (1e-5) was applied across all models. Early stopping based on validation loss with a patience of 10 epochs was also used to halt training when no improvement was observed. Furthermore, the use of ACGAN-based synthetic data improved class balance and introduced intraclass variation, further reducing the risk of overfitting on minority categories. These techniques collectively ensured that the model remained generalizable across test sets and avoided performance degradation due to overfitting. To further validate the superiority of our FAME model, we conducted paired

All statistical tests were based on classification Accuracy and AUC, computed using 3-fold cross-validation.
Privacy-preserving evaluation: DP and SA
To assess the trade-off between privacy and performance in our federated setup, we evaluated the effect of integrating two privacy-preserving mechanisms: DP and SA. These experiments were conducted on the BUSI dataset under a 3-client federated simulation, using the proposed MAU-Net for segmentation and a dual-stage ensemble classifier for diagnosis.
Differential privacy results
Differential Privacy was applied by adding calibrated Gaussian noise to the local model updates before transmission. The amount of noise was controlled by the privacy budget
Impact of varying differential privacy budgets

Lower values of
Secure aggregation results
We also evaluated SA, where local model updates were masked using random additive noise before server aggregation, ensuring that no individual update was accessible in plaintext. Secure Aggregation was implemented without adding explicit DP noise, preserving model fidelity. Secure Aggregation incurred a negligible impact on performance, demonstrating that communication encryption can be integrated seamlessly without compromising ACC, as shown in Table 8. This supports the deployment of our model in real-world federated hospital networks where privacy regulations require encrypted exchanges of model parameters.
Comparison of model performance with and without secure aggregation (SA) in the federated training process.

The integration of SA ensures encrypted model communication without degrading segmentation or classification accuracy, demonstrating its effectiveness in privacy-preserving medical imaging applications.
Although FAME achieved strong overall performance, some failure cases were observed. These included lesions with poorly defined or low-contrast margins, very small tumor regions, and cases with heavy acoustic shadowing, where boundaries were either undersegmented or slightly overextended. Representative examples of such cases are shown in Figure 11, providing insight into the limitations of the current framework.
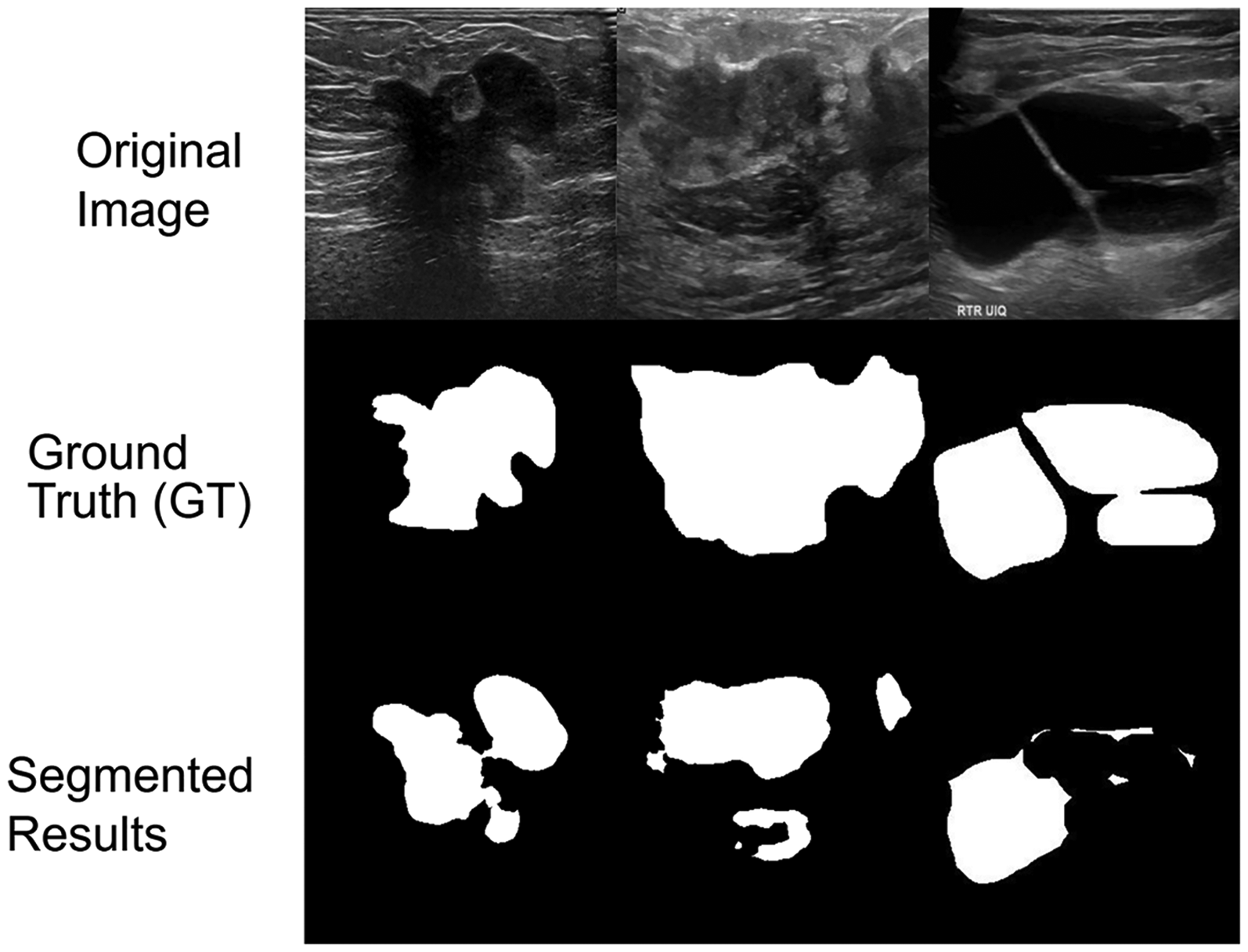
Representative failure cases of the FAME segmentation model on breast ultrasound images.
Ablation study
An ablation study was conducted on BUSI, UDIAT, and BUSC to evaluate the contribution of key components in FAME, including ACGAN-based augmentation, channel attention, spatial attention, the ensemble fusion strategy, and FTL. Components were incrementally removed, and the impact on Dice, IoU, and ACC was measured, as shown in Figure 12. Across all datasets, the complete FAME model achieved the best performance. Removal of ACGAN, attention modules, or the ensemble strategy reduced segmentation ACC, with the largest drop observed when both channel and spatial attention were excluded. Replacing FTL with centralized training also led to decreased Dice and IoU. Figure 13 presents qualitative ablation results, illustrating segmentation differences when key components are omitted.

Ablation study showing the impact of removing key components (Auxiliary Classifier GAN [ACGAN], attention mechanisms, ensemble strategy, and FTL) from the proposed models across three datasets. FAME consistently shows degraded performance when any module is removed, with the full model achieving the highest Dice Score, Accuracy, and IoU on BUSI, UDIAT, and BUSC datasets.

Visual comparison of segmentation performance across ablated versions of the proposed model on the BUSI dataset.
Figure 13 shows qualitative ablation results on BUSI, UDIAT, and BUSC. Excluding ACGAN led to incomplete or fragmented lesion masks. Removing channel or spatial attention individually reduced focus on lesion regions, while omitting both attentions caused notable boundary disruption. Eliminating the ensemble strategy resulted in unstable predictions, and centralized training without FTL produced inconsistent outputs across datasets. In contrast, the complete FAME model generated coherent and precise segmentations aligned with ground truth masks.
Computational efficiency analysis
Computational efficiency was evaluated across ablation settings using training time per epoch, memory usage, inference time per image, and FLOPs, as shown in Table 9. The complete FAME framework, including FTL, ACGAN, MAU-Net, NASNetLarge, and ResNet50V2, required 380 s per epoch, 12.5 GB memory, and 72.5 GFLOPs, while achieving the highest segmentation and classification performance. Removing FTL reduced training time by ∼22% due to the elimination of aggregation. Excluding ACGAN lowered memory and FLOPs but decreased generalization. The absence of MAU-Net improved inference speed but reduced Dice and IoU. Removing NASNetLarge or ResNet50V2 reduced FLOPs but weakened classification ACC. The most efficient configuration (without FTL and ACGAN) achieved the lowest computational cost but also the largest performance decline.
Computational efficiency analysis for different model configurations.

Discussion
This study presents a comprehensive and privacy-preserving federated DL model for BUS image segmentation and classification. The framework was evaluated across three benchmark datasets, BUSI, UDIAT, and BUSC, which vary in resolution, imaging devices, noise levels, and lesion complexity. The inclusion of FTL enables collaborative learning across institutions while preserving data locality and complying with privacy regulations.10,27 To further enhance privacy, the model integrates DP, which injects Gaussian noise into shared updates, and SA, which prevents interclient data exposure during aggregation. Robustness to US image variability was evaluated by training and testing across these diverse datasets, each exhibiting domain shifts and acquisition artifacts. The performance gains achieved by FAME reflect several architectural and strategic advances over existing methods in US image analysis. Unlike conventional U-Net variants or attention-guided networks that often struggle with low-contrast and artifact-heavy US data, 32 the proposed MAU-Net, with embedded channel and spatial attention, improved robustness by selectively emphasizing lesion-relevant features and suppressing background noise. The use of a local ensemble strategy at each federated client, unlike global ensemble aggregation seen in prior FL approaches, helped mitigate performance variance due to non-IID data distributions, a challenge highlighted in earlier federated studies but rarely addressed through node-specific architectural enhancements.
Prior works on BUS segmentation have primarily relied on the U-Net and its derivatives. U-Net and UNet++ have shown reasonable performance in delineating lesions but often struggle with irregular boundaries and noisy textures, leading to Dice scores in the 70–80% range.32,39 Attention-based models such as TransUNet and ATFE-Net improved boundary localization by emphasizing lesion-relevant regions, yet their performance decreases on heterogeneous datasets such as BUSC and UDIAT. In contrast, FAME integrates dual attention (channel and spatial) with ensemble learning to achieve significantly higher Dice up to 93.09 ± 0.49% on BUSC and 87.98 ± 0.57% on UDIAT and IoU scores. This demonstrates that our design more effectively handles low-contrast, artifact-heavy US data compared to prior single-model or attention-only approaches. Breast US classification has been addressed by CNN-based models such as GAN + CNN, MTL-COSA, and SaTransformer, which achieved accuracies in the range of 90–95% but often suffered from class imbalance, leading to reduced sensitivity for malignant cases. 42 Recent transformer-based methods, such as HoVer-Trans, have enhanced feature representation but remain dependent on large, balanced training datasets. Our FAME classifier, by contrast, incorporates ACGAN-based augmentation within a dual-stage ensemble pipeline, addressing class imbalance locally at each federated client. This yields consistent improvements, achieving 98.70 ± 0.27% ACC and 98.70 ± 0.27%, F1-score of 96.82 ± 0.53% F1-score on BUSI, and 98.14 ± 0.31% ACC on UDIAT, surpassing all compared baselines. These results highlight the benefit of combining feature diversity with synthetic augmentation for more reliable malignant case detection.
Federated learning has been applied to medical imaging with strategies such as FedAvg and FedProx, which average local models across institutions to preserve privacy. While effective for reducing data-sharing risks, these approaches often degrade under non-IID conditions and do not incorporate data augmentation or attention-based enhancements.10,27 Recent works have begun exploring multimodal federated pipelines, but rarely consider class imbalance or computational feasibility. Our FAME framework advances the field by integrating FTL, dual-attention segmentation, and ACGAN-based augmentation into a single privacy-preserving model. Unlike standard FL methods, FAME achieves SOTA performance across three diverse US datasets while maintaining strong privacy guarantees. This positions FAME as a practical step forward in bridging the gap between FL research and real-world clinical deployment.
Ablation studies (Figures 12–13) confirmed the contribution of each architectural component. The exclusion of ACGAN reduced Dice and IoU, emphasizing the role of synthetic augmentation under limited data conditions. Removing channel or spatial attention impaired focus on lesion boundaries, and the largest degradation occurred when both attentions were removed. Eliminating the ensemble strategy weakened segmentation stability, particularly on heterogeneous datasets such as BUSC. Substituting centralized training for FTL reduced cross-domain consistency and weakened generalization, underscoring the necessity of privacy-preserving distributed optimization. These findings reinforce that each module's attention, augmentation, ensembling, and FTL contribute critically to overall performance. Computational efficiency analysis (Table 9) provided further insight into deployment trade-offs. While the complete FAME model required higher training time, memory usage, and FLOPs, it consistently achieved superior ACC and generalization. Configurations that removed FTL or ACGAN were more efficient but suffered performance drops, demonstrating that modest computational overhead is justified by significant diagnostic gains. These results suggest that FAME is feasible for real-world use, as inference can be executed on midrange GPU-enabled hospital servers or standard workstations, making it suitable for both high-resource and resource-constrained clinical environments.
From a deployment perspective, the federated design reduces the need for centralized data transfer by transmitting only encrypted model updates, lowering both bandwidth and storage requirements. Because FAME is implemented in widely used DL frameworks (TensorFlow/PyTorch), it can be integrated into existing PACS/RIS infrastructure with minimal modification. These properties increase its potential for seamless adoption in diverse healthcare systems. While the results across BUSI, BUSC, and UDIAT datasets confirm the technical promise of FAME, real-world adoption will require validation in larger, prospective, multi-institutional clinical trials. As such, the present findings should be interpreted as preliminary evidence of feasibility and robustness rather than direct clinical readiness.
Limitations and future work
This study has several limitations that warrant consideration. First, the computational overhead of the ensemble-based classification pipeline may constrain deployment in low-resource clinical environments. Although inference can be run on standard GPU-enabled workstations, future work should focus on lightweight model compression and knowledge distillation techniques to facilitate broader adoption in constrained settings. Second, while ACGAN-based augmentation improved class balance and data diversity, expert validation of synthetic image realism remains essential to strengthen clinical interpretability and trustworthiness. Moreover, representative failure cases revealed challenges in segmenting lesions with diffuse boundaries, heterogeneous textures, or severe acoustic artifacts. Addressing these limitations will require integration of additional modalities such as elastography and Doppler US, as well as the development of uncertainty-aware training strategies to better handle ambiguous or low-quality inputs.
Future directions will also emphasize large-scale validation across geographically distributed clinical environments. Deploying FAME using asynchronous federated protocols will allow adaptation to real-world conditions, including communication delays and variable client availability. To further improve personalization, we plan to explore institution-specific model adaptation strategies that account for differences in imaging protocols, scanner hardware, and patient demographics. Another important direction will be extending the framework to support DICOM-formatted US data, enabling seamless integration with hospital PACS systems. This will require metadata-aware preprocessing and the incorporation of acquisition-specific information into the model pipeline. Finally, improvements in preprocessing through adaptive contrast enhancement methods, such as CLAHE, may further enhance lesion visibility and feature extraction, thereby improving segmentation robustness in challenging, low-contrast US cases.
Conclusion
This study presents a comprehensive federated DL model tailored for privacy-preserving and data-efficient diagnosis of BUS images. By strategically combining FTL with dual attention-guided segmentation, class-aware synthetic data generation, and a two-tier classification ensemble, the proposed approach enables high-fidelity diagnostic modeling in decentralized clinical settings. The proposed method not only addresses practical constraints such as data scarcity, privacy regulation, and institutional variability but also advances technical capabilities in multi-source learning through model fusion, generative augmentation, and DP safeguards. The integration of SA and DP establishes a strong foundation for deploying collaborative AI solutions within regulated healthcare infrastructures. This work lays the groundwork for scalable, trustworthy, and interpretable AI systems in medical imaging and opens future directions for real-world deployment across distributed hospital networks, personalized model refinement, and cross-modality generalization. While certain limitations remain, such as the reliance on simulated federated environments and the absence of DICOM support, these are acknowledged and discussed in detail. Looking forward, this work lays the foundation for real-world deployment by enabling collaborative learning across distributed clinical sites, supporting future extensions toward asynchronous protocols, personalized model refinement, and clinical-grade integration with metadata-rich imaging modalities.
Footnotes
Acknowledgements
The author would like to thank Prince Sultan University for their valuable support. Beijing Natural Science Foundation (4232018), the National Key Research and Development Program of China (2022YFB3103104), Major Research Plan of National Natural Science Foundation of China (92167102), the National Natural Science Foundation of China (62271456), and the R&D Program of Beijing Municipal Education Commission (KM202210005026). It was also supported by the Engineering Research Centre of Intelligent Perception and Autonomous Control, Ministry of Education, China.
ORCID iDs
Ethical considerations
We thank the reviewer for pointing this out. As the study exclusively utilized publicly available, fully anonymized datasets (BUSI, UDIAT, and BUSC), no experiments involving human or animal subjects were conducted. Therefore, institutional review board (IRB) approval was not required. We have now added a dedicated Ethical Considerations statement at the end of the manuscript to clarify this point.
Contributorship
Abdul Raheem: Conceptualization, writing—original draft, methodology, and writing—review & editing. Zhen Yang and Ala Saleh Alluhaidan: Supervision, project administration, writing—review & editing, and funding acquisition. Malik Abdul Manan: Methodology, validation, writing—original draft, and formal analysis. Shahzad Ahmed: Software, conceptualization, and visualization. Sadique Ahmad: Writing—review & editing, methodology, and formal analysis. Fahad Sabah: Investigation, methodology, and visualization.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2025R234), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
