Abstract
This paper reviews the advancements in deep learning for hepatic vascular segmentation and its clinical implications in the holistic management of hepatocellular carcinoma (HCC). The key to the diagnosis and treatment of HCC lies in imaging examinations, with the challenge in liver surgery being the precise assessment of Hepatic vasculature. In this regard, deep learning methods, including convolutional neural networksamong various other approaches, have significantly improved accuracy and speed. The review synthesizes findings from 30 studies, covering aspects such as network architectures, applications, supervision techniques, evaluation metrics, and motivations. Furthermore, we also examine the challenges and future prospects of deep learning technologies in enhancing the comprehensive diagnosis and treatment of HCC, discussing anticipated breakthroughs that could transform patient management. By combining clinical needs with technological advancements, deep learning is expected to make greater breakthroughs in the field of hepatic vascular segmentation, thereby providing stronger support for the diagnosis and treatment of HCC.
Introduction
Hepatocellular carcinoma (HCC) ranks as the sixth most prevalent cancer globally and the third leading cause of cancer-related mortality. 1 Given the intricate vascular network within the liver and the risk of vascular injury and tumor seeding associated with histopathological biopsy, 2 imaging findings serve as crucial data for patient screening, diagnosis, and therapeutic guidance. The treatment modality for liver cancer involves a systematic approach comprising both surgical intervention and medical anti-tumor therapy. 3 The refinement of its treatment regimen necessitates a detailed interpretation and analysis of the patient’s radiological data, encompassing tumor localization, vascular assessment, volume measurement, and watershed analysis, among other steps, with vascular assessment serving as the foundation for completing a series of procedures. The hepatic vasculature, comprising the hepatic veins, portal vein (PV), and hepatic artery, with the latter two typically running in close proximity, determines the hepatic vascular drainage, along with the hepatic veins, which forms the basis of accurate segmentation of modern liver therapy. 4 The spatial relationship between vessels and tumors, the involvement of hepatic vascular territories by tumors, and the remaining healthy vascular territories govern the selection of treatment approaches, scope, and pathways. Meanwhile, during anti-tumor drug therapy, the dynamic changes in the spatial relationship between the tumor and blood vessels are of particular concern to clinicians. However, manual extraction and interpretation of vascular information from multi-dimensional, multi-perspective, multi-temporal, and multi-modal imaging data are challenging, subjective, and lack reproducibility for further analysis.5,6 Hence, computer-based hepatic vascular segmentation has emerged as a solution.
Image segmentation is the process of dividing the entire image into several regions with similar properties, essentially separating objects from the background within the image. 7 Hepatic vascular segmentation specifically involves separating the target vessels from the background. However, abdominal medical images are characterized by high resolution, high dimensionality, complex tissue structures, multimodality, noise, artifacts, and difficulties in standardization, posing challenges for this segmentation task. Traditional image processing, computer vision, and machine learning segmentation methods, including thresholding, 8 region growing, 9 and edge detection, 10 primarily rely on features of vascular pixels and vessel boundaries. However, these traditional methods exhibit limitations when dealing with complex vascular structure data and large datasets. With the rapid advancement of deep learning, deep learning-based image segmentation methods have achieved significant success in the field of hepatic vascular segmentation. 11 Compared to traditional machine learning and computer vision methods, deep learning demonstrates advantages in segmentation accuracy (ACC), speed, multimodal and multi-tasking, and big data processing.
To comprehensively summarize various methodologies, we conducted searches on Google Scholar and Web of Science using the keywords “liver vessel segmentation,” “hepatic vascular segmentation,” and “deep learning” to retrieve the latest literature. The original search yielded 293 studies. We screened 30 studies, excluding non-English articles, studies outside the scope of deep learning-based segmentation, studies not related to liver vessel segmentation, and duplicate articles from the same research. Inclusion criteria encompass recent research work (2015–2024), methods tested and validated, articles written in English, papers specifically addressing liver vessel segmentation, and those employing deep learning methods, ensuring that all results presented are validated. Due to advancements in imaging technology, a large number of early stage and small HCC cases have been detected. In these cases, the liver images are almost indistinguishable from those of healthy individuals, lacking the imaging characteristics of liver cirrhosis or other chronic liver damage, and the lesions occupy an extremely small proportion of the entire background in terms of pixels and voxels. However, there is a lack of publicly and privately available datasets for small and early stage HCCs. In response to this clinical reality, we incorporated data from both HCC patients and healthy individuals in our study on hepatic vascular segmentation. We focused on segmentation performance in both scenarios with liver cirrhosis and chronic liver damage and with minimal background noise, thus enriching our investigation. Unlike existing reviews, this review not only reviews the latest developments, advantages, and limitations of liver vessel segmentation from the perspective of deep learning, compares and summarizes relevant methods, but also provides further insights from a clinical application standpoint. Additionally, it summarizes the challenges and prospects of deep learning methods in liver vessel image segmentation tasks. The structure of this paper is as follows: In the second section, we will provide an overview of deep learning technologies. The third section will review the progress of deep learning in liver vessel segmentation research. In the fourth section, we will discuss the requirements and applicability of deep learning-based liver vessel segmentation methods from the perspective of systematic diagnosis and treatment of HCC. Lastly, We will summarize the full text and put forward the prospect.
Overview of deep learning
Deep learning, as a popular research direction in the field of artificial intelligence, simulates the learning mechanism of the brain to automatically extract features from a large amount of data. It is based on deep neural networks composed of multiple neurons, which process information through complex connections. 12 Deep learning enables the realization of complete image processing workflows, especially as the number of network layers increases. In computer vision, deep learning is widely applied to tasks such as image recognition, restoration, segmentation, etc., demonstrating outstanding performance. Convolutional neural networks (CNNs) are a landmark model in deep learning, combining deep learning and image processing techniques to achieve significant advancements in image analysis. CNNs reduce the number of parameters and improve training efficiency through weight sharing and spatial relationships, making them a supervised learning model. 13 Since the concept of visual receptive fields was proposed by Hubel and Wiesel 14 in 1962, through Fukushima’s 15 introduction of the neural cognitive machine based on receptive fields in 1980, to LeCun et al.’s 16 introduction of LeNet5 using the backpropagation algorithm in 1998, CNNs have undergone a process from theoretical exploration to experimental development. The introduction of AlexNet 17 marked a breakthrough for CNNs in the ImageNet competition, solidifying their core position in computer vision research and continuously driving further research in the field. We summarize the fundamentals in the aspects shown in Figure 1. These are also advantages of deep learning. In the field of complex medical image segmentation, including hepatic vascular segmentation, deep learning is gradually replacing traditional machine learning methods and has become the mainstream of research.

Different components of a typical deep learning pipeline.
Application of deep learning in hepatic vascular segmentation
Assessing hepatic vasculature is fundamental and critical for refining HCC treatment, and hepatic vascular segmentation serves as an efficient approach to acquiring vascular information. Therefore, research on hepatic vessel segmentation has become one of the bridges and links facilitating the translation of deep learning technologies into clinical practice. In this section, we will focus on the problem of hepatic vascular segmentation and analyze it from the following perspectives, summarizing the existing research achievements of deep learning in this field. We summarize the basic information of the 30 included studies in Table 1.
Basic information of hepatic vascular segmentation research based on deep learning.

2D: two-dimensional; 3D: three-dimensional; AFF-Net: adaptive feature fusion network; LSTM: long short-term memory; BCLSTM: bidirectional convolutional LSTM; CNN: convolutional neural network; CT: computed tomography; CTA: CT angiography; GAN-cAED: generative adversarial network-convolutional autoencoder; GAT: graph attention network; HPM-Net: hierarchical progressive multiscale network; IBIMHAV-Net: inductive biased multi-head attention vessel net; US: ultrasound; LUS: laparoscopic US; MRI: magnetic resonance imaging; MRI: magnetic resonance imaging; MTCL: mean-teacher-assisted confident learning; nnU-Net: neural networks for U-Net; PCD: photon-counting detector; PET: positron emission tomograpy.
Motivations for liver vessel segmentation
We summarized the 30 studies included in this paper from a novel perspective. Currently, the motivation for hepatic vascular segmentation primarily focuses on addressing the challenges of low vessel contrast and complex morphology using specialized techniques and algorithms across different modalities, angles, and phases of datasets. We further categorize this motivation into six aspects: improving segmentation ACC, automation, and standardization, addressing challenges in small vessel segmentation, multi-modal and multi-task learning, practicality of new datasets, and real-time or near-real-time analysis. These aspects are summarized in Table 2. It is evident that the current motivation behind hepatic vessel research is primarily focused on addressing graphical and image-related issues through algorithms and technologies, including segmentation ACC and processing efficiency, rather than being driven by clinical problems and demands. The motivation for hepatic vascular segmentation should encompass a broader spectrum of clinical needs, which is also crucial for achieving clinical translation. We will further elaborate on the section titled “Potential clinical translation of hepatic vascular segmentation in HCC.”
Motivation for hepatic vascular segmentation research based on deep learning.

Network type: Platform for addressing hepatic vascular segmentation issues
In deep learning, network structure refers to a series of algorithmic components and their connections that constitute the deep learning model. These components typically include layers (such as convolutional layers, recurrent layers, fully connected layers,

Main network type structure diagram.
U-Net: The foundation of segmentation network architecture
Ronneberger et al. 48 designed a U-Net network for biomedical imaging, which has been widely used in medical image segmentation since its proposal. This method was introduced at the MICCAI conference in 2015 and has been cited over 4000 times. The name “U-Net” originates from its U-shaped network structure, consisting of an encoding path and a symmetric decoding path. The encoding path is similar to traditional convolutional networks, gradually reducing the spatial dimensions of feature maps while increasing the feature depth through successive convolution and pooling layers. The decoding path restores the spatial dimensions of feature maps through upsampling and convolution operations until the output reaches the same resolution as the original image. A key feature of U-Net is the introduction of skip connections in the expansive path, which connect feature maps from the encoding path to their corresponding feature maps in the decoding path. This connection helps preserve and restore detailed information about the image during upsampling, thereby improving segmentation ACC. Convolutional layers in U-Net typically use small kernels and are stacked repeatedly to capture local features. Nonlinear activation functions are typically applied after convolutional layers. In the decoding path, U-Net utilizes transpose convolution for upsampling feature maps to restore their spatial dimensions. Additional convolutional layers can be added during upsampling to adjust and refine feature maps. Similarly, U-Net is one of the most commonly used architectures for hepatic vasculature segmentation. Taking Brown et al.’s 26 study as an example, they applied U-Net to hepatic vascular segmentation research, aiming to address the real-time registration of vessels during laparoscopic liver resection surgery by enhancing the training efficiency of multimodal data and the automation of segmentation. They employed a two-dimensional (2D) U-Net model to automatically segment liver vessels in 2D laparoscopic ultrasound (LUS) images and integrated it into a previously developed untracked LUS-CT registration method to achieve fully automatic initialization, thereby addressing the registration problem of LUS to CT without tracking devices. Batch normalization blocks were added to each layer to accelerate the training process, reduce internal covariate shifts, and help prevent overfitting. A differentiable Dice loss function was utilized to train the network, measuring the similarity between predicted segmentation and ground truth segmentation. By optimizing the loss function, the model could learn more accurate segmentation boundaries. Compared to traditional CNNs, U-Net compensates for the shortcomings of CNNs’ feature extraction ability through its deep network structure, enabling effective integration of global and local information for precise image segmentation, while maintaining good generalization ability and adaptability to small datasets. Furthermore, the structure of U-Net allows researchers to modify and extend it according to specific tasks. This will be further elaborated in the following sections in conjunction with specific research.
Three-dimensional (3D) U-Net and VNet: Enhancement of visualization, interactivity, and efficiency
With the advancement of medical computed tomography (CT) and 3D imaging technologies, the basic unit for hepatic vascular segmentation has shifted from pixels to voxels. From the perspective of images and graphics, voxels provide richer spatial information in 3D space, facilitating the understanding and segmentation of complex vascular structures, while also aiding in capturing the spatial orientation and connectivity of vessels more accurately. Utilizing voxels allows for finer segmentation in 3D space, which is particularly useful for identifying and distinguishing structures such as vessels with small sizes and complex shapes. This differs from the objective of natural image segmentation. From a clinical application standpoint, voxel-level segmentation can provide better visualization effects, with segmentation results being more interpretable, thereby enabling possibilities for treatment planning and disease detection. The 3D U-Net and V-Net are two variants of the U-Net network structure. The 3D U-Net replaces 2D convolution operations with 3D convolutions, directly handling 3D image data. It maintains the symmetric structure of the U-Net and enhances segmentation ACC by combining feature maps of the encoder and decoder through skip connections. The 3D U-Net integrates more contextual information through 3D convolution layers, deepens the network depth to expand the receptive field, captures image features from multiple perspectives, and assigns higher weights to labeled data through a weighted loss function, enabling learning from sparse annotations, thereby reducing manual labeling efforts and improving efficiency. Leveraging this characteristic of the 3D U-Net, Huang et al.
18
proposed a 3D U-Net grid model for hepatic vascular segmentation, achieving automatic extraction of hepatic vascular information under the premise of incomplete hepatic vascular annotation. Similarly, the 3D U-Net can perform multitasking and multimodal data training, inheriting the advantages of the U-Net and extending these advantages from pixels to voxels. Baek et al.
30
introduced a 3D U-Net model utilizing photon-counting detector CT (PCD-CT) for extracting multi-energy information, enabling collaborative training on PCD and CT data to improve the ACC of hepatic vascular segmentation. The V-Net is another variant network structure of the U-Net designed for voxel processing, initially proposed by Milletariet et al.
49
The V-Net combines the characteristics of 3D convolutions with the U-Net architecture. The network structure mainly consists of two parts: the compression path and the decompression path. The compression path gradually reduces the spatial dimensions of the image through consecutive convolution layers and convolutional pooling operations while increasing the number of feature channels to extract deeper features. On the other hand, the decompression path restores the spatial dimensions of the image through upsampling and convolution operations while reducing the number of feature channels to achieve precise pixel-level prediction. Unlike the 3D U-Net, V-Net firstly employs residual connections to facilitate gradient backpropagation, accelerating network convergence while ensuring sufficient representation of foreground voxels even in deep features, preserving vessel features. Su et al.
28
optimized this characteristic by introducing dense connection blocks into V-Net, constructing DV-Net, where each layer’s input includes not only the output of the previous layer but also the outputs of all previous layers. This design enables the network to more effectively reuse features from previous layers, aiding gradient propagation, and improving the network’s performance and ability to fit nonlinear features. Additionally, to better capture liver vessel features, a dual-branch dense connection downsampling strategy is proposed, which adds extra branches before downsampling to reduce the size of feature maps through
U-Net variants: Module modifications
Adapting to various problems and task objectives, and adjusting the components of U-Net is a strategy. Therefore, a series of variant networks derived from U-Net have emerged, combining the advantages of U-Net and specific components. A module refers to a functional unit within a network, which can be a convolutional layer, pooling layer, fully connected layer, normalization layer, activation function, or any other type of neural network layer. Network adjustments can be made by adding or removing modules, changing module types, adjusting module parameters, or modifying module connections. The adjustment of the U-Net module primarily focuses on optimizing the liver vessel segmentation model from two perspectives: training efficiency and speed, and segmentation ACC. From a technical standpoint, the optimization of training speed and efficiency holds significant implications for conserving computational resources. Meanwhile, from a clinical perspective, where patients’ conditions are subject to rapid changes, attaining real-time interactive segmentation results emerges as a critical component for clinical translation. For the former aspect, Thomson et al. 20 reduced the number of filters per layer in the U-Net by one-eighth to avoid bottleneck phenomena, aiming to improve the efficiency of interactive 3D vascular visualization between CT and ultrasound images, providing a possibility for intraoperative navigation. Yu et al. 44 proposed a novel 3D residual U-Net architecture by introducing residual blocks into the 3D U-Net to improve training speed and enhance the representational capacity of discriminative features. For the latter aspect, Tong et al. 42 introduced the multi-axis squeeze-excitation module (MASE) and distribution correction module (DCM) into the U-Net architecture. MASE performs squeeze and excitation operations on coronal and sagittal planes to restrict the activation region of the network only to the dense area (DA) of hepatic veins, reducing background noise interference and enhancing attention to hepatic vein features. DCM calculates and corrects the coordinate distribution of hepatic veins at different segmentation mask scales through a deep supervision strategy to improve segmentation ACC. Alirr et al. 45 incorporated consistency-enhanced diffusion filtering and vascular filtering methods into U-Net to improve vascular contrast and intensity uniformity, while introducing an improved residual block called ResDense block, which passes information between residual blocks using connections rather than summation, helping to retain more information between different layers of the network. Zhou et al. 41 designed a sequence-based context-aware correlation network by adding a slice-level attention module (SAM) to U-Net. SAM bridges the feature differences between the encoder and decoder by fusing high- and low-dimensional features in different spatial and channel dimensions, enhancing the model’s ability to capture details of intrahepatic vessels. Feature maps are fused through element-wise addition, multi-fusion blocks, element-wise multiplication, and concatenation operations to enhance the structural details of intrahepatic vessels in both spatial and channel domains.
U-Net variants: Branch network architecture modifications
A branch structure refers to the presence of multiple parallel paths within a network, which can process different features or data streams. Adjustments to the branch structure include methods such as adding or removing branches, adjusting branch depth, merging or splitting branches, and establishing cross-branch connections. The adjustments to the U-Net branch network primarily aim to meet two objectives for liver vessel segmentation models as follows: firstly, to enhance the utilization efficiency of labels and the prediction of vessel segmentation results; secondly, to better adapt to the requirements of multitasking and multiscale liver vessel segmentation. Labels refer to annotations of different regions in images. When training deep learning models for liver vessel segmentation, these annotations inform the model about which pixels belong to specific structures or objects. Labels can guide the model in learning features and optimizing model parameters by comparing predicted results with actual labels. Obtaining high-quality labeled data is often challenging, as it requires clinicians to invest a significant amount of effort and cost to complete pixel-level or voxel-level annotations, which is unrealistic for clinical practice. Therefore, efficient utilization of limited labels is an important direction. Liver vessel medical imaging spans a wide range, with tens or even hundreds of cross-sectional layers, varying vascular features in different enhanced phases of imaging data, and its own complex structure. Training multi-task, multi-scale, and multi-modal segmentation models aligns with this scenario. For the former aspect, Zhao et al. 22 concatenated two additional network models to U-Net, proposing a deep neural network structure combining RetinaNet, U-Net, and long short-term memory (LSTM) networks. This structure first enhances data with histogram equalization for data augmentation and preprocessing, then extracts regions of interest (ROIs) using RetinaNet, utilizes U-Net to extract features of ROIs and performs deconvolution operations to achieve automatic vessel segmentation, and finally uses the LSTM network to predict vessel information in subsequent images. Xu et al. 25 introduced a self-confidence learning framework (mean-teacher-assisted confident learning (MTCL)) into U-Net, where the model utilizes exponential moving averages to update the weights of the teacher model. The teacher model guides the student model by providing consistency loss, enabling the student model to maintain stable predictions in the presence of input perturbations, thereby improving the model’s generalization ability. The model also adapts confident learning technology to identify and handle noisy labels in low-quality annotated data. By estimating the joint distribution between noisy labels and true labels and using the teacher model as a third party to identify label noise, the model can reduce the negative impact of noisy labels on the learning process. The smoothly self-denoising module gradually improves noisy labels through soft correction, transforming noisy labels from a burden into valuable data, and providing beneficial guidance for the model. For the latter aspect, Affane et al. 27 introduced the MultiRes U-Net into liver vasculature segmentation research, which enhances the model’s multi-scale feature learning ability based on U-Net by adding multiple resolution paths, enabling it to handle image information at different scales simultaneously. This multi-scale structure helps the network capture finer image features, thereby improving segmentation ACC. Hao et al. 32 proposed a dual-branch progressive (DBP) 3D U-Net for liver vasculature segmentation, reducing the loss of detailed information during network downsampling by using a DBP downsampling strategy. This network structure achieves feature learning of liver vasculature at different scales through internal and external progressive learning strategies. The internal progressive learning strategy gradually expands the network depth from shallow networks to deep networks to obtain receivers of different sizes for learning different levels of semantic information. The external progressive learning strategy complements and fuses local and global information by transferring and merging feature maps between different stages of training. Similarly, following the hierarchical structure, Li et al. 35 divided the network structure based on the overall vessels and sub-type vessels. This design allows the network to simultaneously complete these two sub-tasks and reduce feature sharing and error accumulation by combining feature maps from encoding layers in the output layer to predict overall vessels and sub-type vessels.
U-Net variants: nnU-Net)
The nnU-Net is an open-source medical image segmentation framework, with “neural networks for U-Net” as its full name.
50
This framework is based on the popular U-Net architecture, which employs a modular design philosophy, allowing users to customize and configure the network according to specific application requirements to provide higher flexibility and automation. The nnU-Net, developed by
Generative adversarial networks (GANs): Dataset capacity and diversity expansion
GANs are a type of deep learning architecture consisting of two competing neural networks, namely the Generator and the Discriminator. The Generator aims to generate data that closely resembles the real data distribution, while the Discriminator’s objective is to differentiate between generated data and real data. These two networks mutually confront each other during training, thereby improving the quality of the generated data. Combining the requirements of semantic segmentation and the characteristics of GANs, Luc et al. 51 trained a convolutional semantic segmentation network and an adversarial network. In the field of liver vessel segmentation, the goal of GANs is to expand the dataset capacity by generating synthetic images through adversarial generation. The synergistic training of high-quality synthetic and real image data enhances the robustness of segmentation models. Cheema et al. 46 applied an improved GAN-convolutional autoencoder (GAN-cAED) model to liver vessel segmentation research. The network structure consists of two core components: the GAN component for synthesizing fused images, and the cAED component for liver vessel segmentation using CTA/synthetic positron emission tomograpy-CT. The training objective of the cAED network is to minimize reconstruction loss, that is, the difference between input images and reconstructed images. To achieve this, the network learns the probability distribution of input data during training and optimizes it through gradient descent and backpropagation algorithms. In this way, cAED effectively utilizes realistic images generated by GANs and the generation capability of the cAED network, thereby achieving finer liver vessel segmentation. Kuang et al. 31 used an improved cycle-consistent adversarial network to achieve segmentation and domain adaptation of arterial and venous phase CT images in liver-enhanced CT. The entire network structure consists of two generators and four discriminators, aimed at achieving global and local style transfer of small vessels through adversarial training. The generators, named GAB and GBA, are responsible for converting arterial phase CT images into venous phase CT images and vice versa while preserving vessel details and achieving image transformation between the two domains. Four discriminators, where DA and DB distinguish real arterial and venous phase CT images, while LDA and LDB focus on local vascular information to ensure effective preservation of generated small vessels during domain transformation. The goal is to achieve segmentation of the entire vascular system on single-phase CT images, ignoring displacement and deformation of vessels in different phases, while reflecting the states of arterial and venous vessels.
GAT+CNN: Optimization of connectivity, spatial features, and edge features
The network architecture combining graph attention networks (GAT) with CNN is an advanced deep learning model. GAT is an important concept in graph neural networks (GNNs), while CNN is the most commonly used architecture in the field of image segmentation. 52 This joint network architecture connects GNN with CNN, combining the advantages of graph attention mechanisms and CNNs to handle both graph-structured and grid-like data. The data processed by GNN consists of nodes and edges. Typically, an adjacency matrix is used to represent the graph structure, capturing the correlations between nodes and edges in the graph. 53 The GAT layer assigns different weights to neighbors of each node through attention mechanisms, allowing the model to focus on the most important node features, effectively capturing complex relationships between nodes. CNN layers are typically used to process grid-like data, such as images. They extract local features through convolution operations, capturing hierarchical structures and local patterns in space. Such a combination allows the network to simultaneously handle graph-structured data and associated grid-like data, enhancing the model’s expressive power and applicability. Vascular data conforms to these characteristics, making the GAT+CNN network architecture a research direction for liver vessel segmentation. From a clinical perspective, the network structure of GAT+CNN offers advantages in handling complex spatial structures of liver vessels. Additionally, it provides a solution for clarifying the connectivity of peripheral vessels and poorly filled vessels, as well as determining the edge features of vessels. This is because the topological configuration of the three sets of hepatic vessels within the liver is precisely the grid data formed by the nodes that GAT excels at handling. In liver vessel segmentation research, Zhang et al. 21 initially generated liver vessel centerline heatmaps using full CNN and initialized vessel graph models through image processing algorithms. The initial tracking graph was transformed into a dual graph, and for each node in the dual graph, features were sampled from specific layers of the CNN. A graph attention layer was used to compute the potential representations for each node. Attention coefficients were calculated by combining node features and weight vectors, allocating an existence probability for each branch segment. Finally, the output layer generated the final output for each node by averaging the outputs of attention heads, namely the existence probability of each branch segment. Based on the calculated probabilities, branch segments with confidence lower than a threshold were pruned from the final reconstructed result to reduce false-positive branches. Li et al. 34 integrated GAT with lightweight 3D U-Net using an efficient mechanism called “plug-in mode.” GAT selectively uses information from neighboring nodes through attention mechanisms to model the connectivity of vessels. GAT, as a multitask branch, is integrated into U-Net and is only used to supervise U-Net with connectivity priors during training, without using GAT during the inference stage, thus not increasing the hardware and time costs compared to U-Net alone, improving model efficiency. The advantages of the GAT+CNN network architecture lie in its flexibility and adaptability, enhancing computational efficiency, and improving performance in multiple tasks, including node classification, image classification, and image-graph matching, as it integrates two powerful feature extraction mechanisms.
Transfomer+CNN: Integration of global and local vascular features
The network architecture combining transformer and CNNs offers complementary advantages in feature extraction. The transformer was initially applied to natural language models and natural image segmentation. However, using the transformer alone for medical image segmentation did not yield satisfactory results. The architecture of the transformer combined with CNNs emerged because researchers aimed to leverage the strengths of both models to enhance the performance of medical image segmentation tasks. Transformers excel at capturing global dependencies and long-range interactions, 54 while CNNs are proficient in capturing local spatial features. CNN’s convolutional operations are effective in capturing spatial patterns and local context in medical images. Transformers are adept at modeling long-range dependencies in sequences, which is beneficial for capturing anatomical relationships and complex structures in liver vessel networks. The combination of transformer and CNN allows for flexible architecture design tailored to the specific requirements of liver vessel segmentation tasks. Researchers can customize the network architecture by adjusting the depth, width, and connectivity of both components to achieve optimal performance. Wu et al. 43 proposed a robust end-to-end vessel segmentation network called inductive biased multi-head attention vessel net (IBIMHAV-Net), which combines 3D swin-transformer with effective convolution and self-attention mechanisms to improve the ACC of liver vessel segmentation. The model adopts a 3D swin-transformer as the backbone network, which can capture the global dependencies of images. By transforming the input 3D volume data into high-dimensional tensors, the model can learn hierarchical object concepts at different scales. Meanwhile, the inductive biased multi-head self-attention (IB-MSA) mechanism is introduced. Different from traditional self-attention mechanisms, IB-MSA improves the recognition ability of liver vessel edges by learning the relative position embedding of inductive bias. This mechanism helps the model better understand and process complex vessel structures. TransFusionNet is another form of a combination of transformer and CNN. 55 Its basic network architecture includes a transformer-based semantic feature extraction module, local spatial feature extraction module, edge extraction module, multi-scale feature fusing module, and multi-task training strategy. These modules work together to extract global semantic features and local spatial details of images while considering edge information to achieve more accurate segmentation results. Meng et al. 29 first introduced TransFusionNet into the field of liver vessel segmentation. In this study, the transformer-based encoder in the transformer semantic feature extraction module captures the global contextual features of the image. The input image is first processed through a feature extraction backbone network and then segmented into multiple small blocks, each of which is transformed into an information matrix through convolutional operations. Learnable position embedding is used to enhance the transformer layer to understand the positional information of each part of the image. The local spatial feature extraction module stacks multiple layers of SEBottleNet to extract local features of the image, such as edges and fine vessels. The squeeze and excitation module enhances the BottleNet residual network, strengthening the interdependence between feature map channels. In the edge extraction module, the edges of CT images extracted by the Canny algorithm are used as input to predict edge information and integrate it into the segmentation network. The predicted class distribution map is output by combining the features extracted by the above three modules. The multi-scale feature fusion (MSFF) module processes different feature maps through convolution and pooling operations to achieve MSFF.
Other CNN network structures: The driving force behind ongoing advancements
There are many variants of CNN-based network architectures for liver vessel segmentation research. Kitrungrotsakul et al.
19
proposed a segmentation method based on CNN called VesselNet. The network explores the 3D structure and improves recognition performance by performing binary classification on training patches extracted from three perspectives (sagittal, coronal, and transverse planes). Yan et al.
23
proposed LVSNet, whose innovative features include the attention-guided concatenation (AGC) module and the MSFF block. The AGC module, located between the encoder and decoder, is used for the effective fusion of features at different levels. The MSFF block enhances the network’s representation of features at different scales by dividing features into multiple groups and exchanging information between different groups to achieve MSFF. Li et al.
33
proposed a new deep learning network called bidirectional convolutional LSTM (BCLSTM), which is a hybrid neural network architecture combining CNN and LSTM. BCLSTM integrates convolutional layers into the traditional LSTM structure to handle sequential data, while utilizing bidirectional LSTM to capture both forward and backward dependencies in sequences. It explores 3D context in segmentation and edge prediction branches. The 3D features from both branches are concatenated to facilitate the learning of segmentation maps. Sobotka et al.
47
introduced the Y-net model to simultaneously predict the segmentation target (vessel labels) and closely related auxiliary imaging modalities (contrast-enhanced imaging) for vessel segmentation in liver MRI without contrast enhancement. The Y-net structure consists of three downsampling blocks and two upsampling blocks, each downsampling block containing two
Supervision: Learning pattern in hepatic vascular segmentation
Supervised learning, unsupervised learning, and semi-supervised learning are the three main learning patterns in machine learning and deep learning, each with different significance and application scenarios in liver vessel segmentation methods based on deep learning. In supervised learning, the model training requires a large amount of annotated data, namely input data (such as CT scan images) and corresponding output labels (the segmentation results of liver vessels). For liver vessel segmentation, supervised learning can provide accurate segmentation results because the model learns directly from known correct segmentations. The drawback of this approach is the need for a large amount of high-quality annotated data, which is often time-consuming and expensive. Additionally, accurate annotation for medical images requires the involvement of radiologists or experts. Unsupervised learning does not rely on annotated data but attempts to discover patterns and structures from the data itself. In liver vessel segmentation, unsupervised learning can be used to identify vessel structures in images. However, due to the complexity of liver vessel structures and the interference of artifacts and noise, and considering the substantial amount of data required for unsupervised training, obtaining liver image data is challenging. Therefore, unsupervised learning is not mainstream in existing deep learning-based liver vessel segmentation research but is generally used for preliminary vessel detection or as a preprocessing step to assist subsequent supervised learning. Kuang et al. 31 applied unsupervised learning during the GAN stage. Semi-supervised learning combines the characteristics of supervised and unsupervised learning, using a small amount of annotated data and a large amount of unlabeled data for training. It can reduce the need for a large amount of annotated data while utilizing unlabeled data to improve the model’s generalization ability. It helps the model learn useful feature representations from unlabeled data and is currently an emerging hotspot in liver vessel research. Li et al., 33 Xu et al., 25 and Sobotka et al. 47 applied semi-supervised learning in their research.
Evaluation metrics and performance
In tasks of liver vessel segmentation based on deep learning, evaluation metrics are crucial indicators used to quantify the segmentation performance and represent an important aspect of liver vessel segmentation tasks. These metrics reflect the algorithm’s performance and the ACC of the segmentation results. This paper will summarize the evaluation metrics currently applied in research and present them along with performance in Table 3.
Performance for liver vessel segmentation research based on deep learning.

DSC: dice similarity coefficient; MAD: mean absolute distance; mAP: mean average precision; VOE: volumetric overlap error; IoU: intersection over union; PV: portal vein; HV: hepatic vein.
Dice similarity coefficient (DSC) and F1 score
The DSC and F1 score are similar statistical tools used to assess the similarity or overlap between two samples, particularly in medical image analysis for evaluating the ACC of image segmentation. The value of DSC ranges from 0 to 1, where a value closer to 1 indicates a higher degree of overlap between two samples and thus a more accurate segmentation result.
The F1 score is used to measure the balance between precision and recall in binary classification models. It is the harmonic mean of precision and recall, serving as a single evaluation metric that comprehensively considers both precision and recall. The F1 score ranges from 0 to 1, with higher values indicating better classification performance of the model. The expression is:
The DSC and F1 scores are equivalent in certain situations. Specifically, when prediction results are converted to binary form, the Dice coefficient and F1 score yield the same results. Some variants of DSC can indeed be derived from the F1 score. For instance, clDice, 56 which focuses on evaluating vascular connectivity, can be defined as the harmonic mean of two measures, and the harmonic mean is essentially the F1 score. However, in some models, such as V-Net, where the evaluation metric such as soft-Dice 49 requires adjustments to enhance the assessment performance for sample similarity, DSC and F1 may differ due to their different emphases.
Jaccard index and intersection over union
The Jaccard index is a measure of the similarity between two sets, also known as intersection over union (IoU). In fields such as medical image segmentation, the Jaccard coefficient is commonly used as an evaluation metric to quantify the overlap between the segmentation result and the ground truth annotation. The expression is: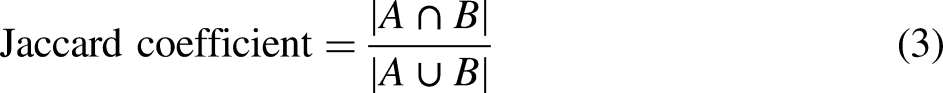
Accuracy (ACC)
ACC is a commonly used evaluation metric in classification tasks, used to measure the correctness of the model’s classification for all samples. It is defined as the proportion of samples correctly classified out of the total number of samples. The expression is:
Sensitivity
Sensitivity, also known as TP rate and recall, is a critical metric for evaluating the performance of classification models, particularly in fields such as medical image segmentation and disease diagnosis. It measures the model’s ability to correctly identify positive samples (such as vascular regions). The expression is:
Specificity
Specificity, also known as TN rate, is a crucial metric for evaluating the performance of classification models, particularly in fields such as medical image segmentation and disease diagnosis. It measures the model’s ability to correctly identify negative samples (such as non-lesion regions). The expression is:
Precision
Precision is an important evaluation metric in classification tasks, used to measure the proportion of true positive samples among all samples predicted as positive by the model. High precision means that among the samples predicted as positive by the model, a higher proportion is truly positive, that is, there are fewer false positives. The expression is:
Mean average precision (mAP)
mAP is a commonly used evaluation metric in the fields of object detection and information retrieval, used to measure the overall performance of the model for multiple classes or queries. mAP is obtained by calculating the average of the area under the curve for each class, which comprehensively considers the precision performance of the model at different recall levels.

Volumetric overlap error (VOE)
VOE is an evaluation metric used to assess the performance of medical image segmentation. VOE measures the error in volume overlap between the segmentation result and the ground truth. It is calculated by computing the ratio of the intersection to the union of the segmentation result and the ground truth. A smaller VOE value indicates higher consistency between the segmentation result and ground truth, indicating better segmentation ACC. The expression is:
Average surface distance
Average surface distance (ASD) is a metric used to evaluate the quality of image segmentation and the performance of models, especially in the field of medical image segmentation. The ASD between the predicted segmentation result and the ground truth is measured. Specifically, it calculates the surface distance between the predicted segmentation boundary and the ground truth boundary and takes the average of all these distances. The expression is:
Hausdorff distance (HD)
HD is used to measure the maximum distance between two sets, here employed to assess the maximum inconsistency between the segmentation boundary and the ground truth boundary. The expression is: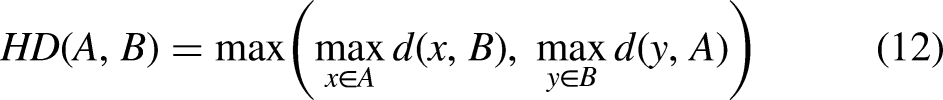
Mean absolute distance (MAD)
MAD calculates the average distance from points on the segmentation boundary to the nearest points on the ground truth boundary. It is used to assess the average inconsistency between the segmentation boundary and the ground truth boundary. The expression is:
Clinically relevant evaluation metrics
Traditional evaluation metrics focus on sample similarity and do not differentiate between pixels that belong to the same hepatic vessel. In different clinical scenarios, the importance of vessel pixels varies by location. For example, in the surgical treatment of HCC, identifying the terminal branches of vessels is more critical than the main vessels. Although there are many branches in hepatic vessels, their pixel proportion is small. Traditional metrics are inadequate for effectively assessing the segmentation performance of liver vascular models on branching vessels. Researchers are increasingly aware of this issue. In current studies related to liver vessel segmentation, some researchers have focused on designing evaluation metrics that are tailored to clinical characteristics. For example, Cheema et al.
46
proposed the branching level identification rate as an evaluation metric, aiming to quantify and evaluate the performance of models in identifying major vessel branches of the liver, serving surgical interventions. The expression is:
Potential clinical translation of hepatic vascular segmentation in HCC
In a broad sense, image segmentation refers to dividing an image into several non-overlapping regions, each possessing similar properties such as grayscale, color, texture, etc. 57 The purpose of image segmentation is to simplify or alter the representation of the image, making the segmented regions easier to analyze and understand. Medical image segmentation is a critical task for clinical diagnosis and research, for dealing with highly imbalanced data remains a significant challenge in this domain. 58 Liver vessel segmentation is a specific application scenario in the field of image segmentation, which focuses on identifying and separating the vascular structures within the liver from medical images. Liver vessel segmentation is a task at the intersection of medicine and computer science. Currently, research methods mostly target computer-related challenges and requirements, emphasizing segmentation ACC and efficiency, while neglecting the analysis and understanding of the segmented regions. Eisenmann et al. 59 conducted a multicenter study incorporating 80 biomedical image segmentation challenges, where they found that the majority of researchers’ efforts primarily focused on fitting past evaluation metrics rather than addressing underlying domain issues. Challenge organizers should pay more attention to ensuring that actual biomedical needs are reflected in their competition designs. Addressing clinical domain issues is crucial to realizing the value of liver vessel segmentation. In this section, we will integrate the comprehensive diagnostic and treatment patterns of HCC (Figure 3) and clinical practice experience to summarize and discuss the demands and challenges of deep learning-based liver vessel segmentation methods in specific clinical scenarios. We aim for this discussion to facilitate the gradual transition of deep learning-based liver vessel segmentation research from the laboratory stage to clinical application, achieving clinical translation.

Systematic clinical procedures for HCC (based on the BCLC staging of HCC). HCC: hepatocellular carcinoma; BCLC: Barcelona clinic liver cancer.
Systematic treament
The systematic treatment of HCC is a multidisciplinary, multimodal treatment regimen aimed at achieving optimal therapeutic outcomes through various treatment modalities. 3 Within this framework, internal medicine treatment for HCC mainly includes supportive treatment and anti-tumor treatment. Supportive treatment aims to maintain the patient’s condition, treat side effects, and improve the quality of life throughout the course of systemic treatment. 60 Anti-tumor treatment encompasses chemotherapy, targeted therapy, and immunotherapy. 1 Sorafenib is the first multi-target tyrosine kinase inhibitor confirmed to improve the survival of patients with advanced liver cancer. 61 Regarding vascular assessment, deep learning-based liver vascular segmentation methods have the potential to be translational throughout the systematic treatment of liver cancer. In terms of diagnosis, achieving visualization and quantification of the spatial relationship between tumors and blood vessels through high-precision vascular segmentation, including whether tumors invade blood vessels, 62 is crucial for evaluating tumor grading and selecting first-line treatments. 63 During the treatment process, liver vascular segmentation technology can be used to monitor changes in tumor blood vessels, 64 evaluate the effectiveness of medical treatments such as targeted drugs or immunotherapy, and adjust treatment plans in a timely manner (Figure 4). Furthermore, research based on vascular segmentation, including predicting PV cavernous transformation65,66 and PV pressure 67 to reflect liver condition, has guiding significance for subsequent treatment selection. It is noteworthy that comprehensive treatment for HCC is not conducted in isolation. Guided by the internal medicine concept, various invasive technologies are combined to form individualized comprehensive treatment plans. Further discussion will continue in the following sections.

In an example of systematic treatment, for advanced HCC patients, the integration and registration of hepatic vascular segmentation and tumor segmentation from CT data during the treatment process allows for the visual assessment of radiological efficacy. HCC: hepatocellular carcinoma; CT: computed tomography
Ablation
Ablation therapy is an invasive local treatment method aimed at directly destroying tumor tissue while minimizing damage to surrounding normal liver tissue. 68 The main categories include radiofrequency ablation (RFA), microwave ablation, cryotherapy, alcohol ablation, and laser ablation.68–72 The challenges and focus of ablation therapy for HCC mainly revolve around the spatial relationship between tumors and blood vessels. Some tumors are located deep within the liver or adjacent to major blood vessels, making localization difficult and increasing the risk of vascular injury. The blood vessels around the tumor may cause energy loss, leading to the heat sink effect, which can affect treatment efficacy. Additionally, the selection of the ablation needle treatment path and the monitoring of treatment efficacy are important aspects. In this clinical scenario, the translation of liver vascular segmentation research is primarily based on multimodal tasks combining ultrasound data with CT or MRI data 73 (Figure 5). Progress has been made in the segmentation of large blood vessels and the basic localization of tumors, which can provide guidance for ablation therapy in some clinical scenarios. However, there is still a need to improve the ACC and registration of small blood vessel segmentation in multimodal datasets, especially the handling of peri-tumoral small blood vessels, which affects the delineation of treatment target areas. This is also one of the reasons why the applicability of ablation therapy is limited.

Xing et al. provided a liver vessel segmentation model for RFA based on the nnU-Net network architecture and multimodal datasets of 3D US and CT/MRI. The above figure shows the schemas of this study, from graph input, vessel segmentation, and model reconstruction to registration. 73 RFA: radiofrequency ablation; CT: computed tomography; MRI: magnetic resonance imaging; nnU-Net: neural networks for U-Net; 3D US: three-dimensional ultrasound.
Transarterial chemoembolization (TACE
TACE is a minimally invasive interventional treatment method used for treating HCC. It involves the direct injection of chemotherapy drugs into the hepatic artery, combined with the use of embolic agents to block tumor blood flow, thereby reducing tumor blood supply and increasing the concentration of chemotherapy drugs in the tumor area. 74 This method enhances the cytotoxic effects of drugs on tumor cells while minimizing damage to normal liver cells. It is often integrated as part of comprehensive therapy for HCC, combined with other treatments such as radiotherapy, targeted therapy, or immunotherapy. Additionally, it serves as a bridge to surgical treatment. It involves the study of multimodal data from CT contrast-enhanced images and digital subtraction angiography (DSA) data, with the task goal being the segmentation of hepatic arterial vessels. As shown in Figure 6, DSA data reveals all arterial vessels filled with contrast agents, while CT data, when adjusted to match the same plane, can only achieve segmentation of arterial data at that level. The reason for this phenomenon lies in the different imaging principles of the two modalities. DSA imaging compresses all 3D imaging information into a 2D plane. The requirements for vascular localization and ACC in TACE are stricter. CT datasets retain spatial information about liver vessels, while DSA datasets have advantages in the ACC segmentation of small vessels and real-time performance. 75 Integrating the advantages of both modalities to achieve hepatic arterial vessel segmentation in this scenario is a future requirement and challenge (Figure 6).

Comparison between the segmentation results of hepatic artery vessels in the TACE scene and the actual DSA interface. Meanwhile, we take paired CT and DSA data as inputs, transfer and reproduce Patel et al.’s 76 method of co-training CT and DSA, and reconstruct the outputs. TACE: transarterial chemoembolization; DSA: digital subtraction angiography; CT: computed tomography.
Radiotherapy
Radiation therapy is a treatment method that utilizes radiation to damage tumor cells’ DNA, inhibiting tumor growth and reproduction. 77 Radiation therapy can be broadly categorized into two main types: external beam radiation therapy (EBRT) and internal radiation therapy (IRT). 78 Stereotactic body radiation therapy (SBRT) in EBRT and selective IRT (SIRT) in IRT play significant roles in the systemic treatment of HCC. Inaccurate radiation therapy can lead to damage to liver tissues. Simultaneously, liver function issues caused by damage to normal liver bile ducts are also known as central liver toxicity. SBRT is used to deliver high doses of radiation in a highly targeted manner and rapidly decrease the dosage away from the center of the radiation zone, preserving much of the adjacent liver parenchyma while delivering therapeutic doses to the tumor. 79 SIRT, on the other hand, aims to locally irradiate tumors by inserting small radioactive beads. Since liver tumors are primarily supplied by arterial blood, these beads are inserted by passing a catheter through the hepatic artery into the tumor-feeding arteries.80,81 In SBRT, PV segmentation is a major task because the planning of the target area relies on the anatomical position of the PV, which represents the location of most intrahepatic ducts. Additionally, PV thrombosis is an indication of SBRT, and precise identification of the PV during treatment is crucial for completing this task (Figure 7). 82 In SIRT, hepatic artery segmentation is key, but it is challenging due to the small proportion of pixels and voxels occupied by the hepatic artery in the entire liver, its small diameter, and rich variability. 83 The translation-related research on liver vessel segmentation also provides possibilities for adaptive planning of radiation therapy target areas and improving ACC. At the same time, it has a guiding significance for radiation dose evaluation and displacement correction during treatment. 84

The figure is taken from the study of hepatic vessel segmentation applied to SBRT by Ibragimov et al. 82 Individual CNNs are applied on each orthogonal cross-section of the CT image in order to generate three PVs enhancement maps. The maps are then averaged to form the resulting PV to preserve as much vascular information as possible from every angle. SBRT: stereotactic body radiation therapy; CNN: convolutional neural network; CT: computed tomography; PV: portal vein.
Minor hepatectomy
Modern minor hepatectomy commonly deals with liver subsegments and cone units, constructing the pre-resection range by piecing together the domains where tumors affect vessels. 85 Hepatic cone units are the smallest anatomical units divided according to the liver’s PV system. 86 Each liver segment consists of six to eight cone units, a division method more aligned with the liver’s vascular supply and anatomical structure, aiding in precise liver resection. 87 In this scenario, directions for clinical translation of hepatic vascular segmentation research lie in interpretable, accurate segmentation, especially in the credible segmentation and combing analysis of PV small vessels. 88 The goal of minor hepatectomy is to minimize normal liver damage while completing anatomical liver resection. 89 Particularly when tumors grow around the liver, surgical approaches involving intrahepatic and extrahepatic anatomy cannot be fully applicable, and liver parenchymal approaches better fit surgical objectives. 90 Achieving liver parenchymal approaches relies on high-precision segmentation and combing analysis of liver small vessels, especially PV vessels. High-precision segmentation of PV small vessels is also a significant motivator in current liver vessel segmentation research, which is not elaborated here. 91 With the optimization of model structures and algorithms, numerous small branches of the PV have been reconstructed. However, for clinical doctors, the efficiency of clinical decision-making is low when facing an uncombed vascular tree. 92 Artificially classifying segmentation results undoubtedly increases the possibility of errors and consumes a considerable amount of manpower. 6 Moreover, if such segmentation results are matched to real-time surgical navigation systems, uncombed and unlabeled results become a disturbance, as surgical teams cannot obtain critical information in real time. Organized PV segmentation results can better assist doctors in completing pre-resection plans and adjusting real-time surgical decisions, which is of great significance (Figure 8). Through the combing analysis of segmentation results, including but not limited to the classification of PV branches and hepatic pedicles, the spatial relationship between tumors and vascular trees, as well as the position of cone units, can be efficiently determined. Additionally, PV vessels of different positions and classifications have different meanings and weights in different scenarios and should not be simply segmented. Therefore, evaluation metrics based on PV model interpretation and classification are crucial for assessing the clinical performance of segmentation models and the applicability of minor hepatectomy. Moreover, visual studies on PV segmentation results after automatic combing analysis are also a direction for development.
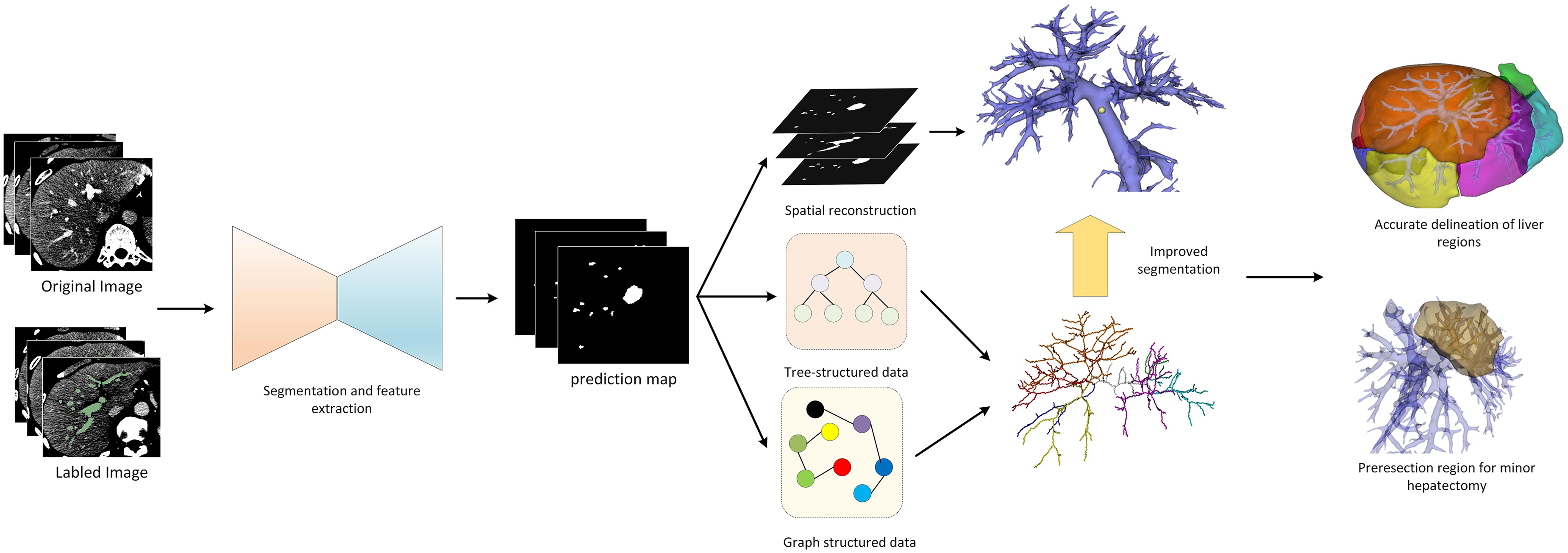
The preoperative planning process for minor hepatectomy includes high-precision hepatic vascular segmentation including small vessel branches, hepatic vascular classification, hepatic segment delineation based on portal vein territory, and selection of the resection margin.
Major hepatectomy
Major hepatectomy is a scenario contrasted with minor hepatectomy. It is typically defined as surgery involving the resection of three or more liver segments. 93 This includes combined resection of liver segments and lobes, hemihepatectomy, and extended hemihepatectomy, among others. In such scenarios, clinicians are most concerned about two issues: the remaining functional liver volume of the patient 94 and the avoidance of important ducts, especially the hepatic veins. Directions for clinical translation of hepatic vascular segmentation research lie in the precise segmentation and analysis of hepatic veins, and in using high-precision segmentation results of important structures in hepatic veins to guide and organize the entire segmentation structure. For major hepatectomy, due to the large incision, the lack of assessment of important ducts in the pre-resection area can result in significant intraoperative bleeding, postoperative bile leaks, and lymphatic leaks, 93 all of which can affect short-term and long-term survival.95,96 Additionally, the volume of the remaining functional liver affects the occurrence of postoperative liver failure in patients. 94 It is important to note that we emphasize functional liver volume rather than actual liver volume. 97 Well-functioning liver tissue includes intact inflow and outflow pathways. 98 Inflow pathways generally refer to the PV and hepatic artery territories. Due to the large resection range, the surgical strategy for the PV and hepatic artery involves graded occlusion starting from the hepatic hilum, whether through an intra-sheath anatomical approach or an extra-sheath structural approach. As most of the occluded PVs are primary or secondary branches, liver vascular segmentation models are mature in recognizing this type of vessel. The outflow pathway is the hepatic vein. Damage to the hepatic vein branches of the remaining liver tissue during surgery can lead to obstruction of the hepatic tissue reflux area, causing congestion, which in turn affects liver function. 98 It also makes the estimation of the remaining functional liver volume inaccurate, increasing the risk of postoperative liver failure. Therefore, the key to assessing the remaining functional liver volume lies in the comprehensive evaluation of the hepatic vein and PV vessels. In plain data and contrast-enhanced CT data, the contrast between the hepatic vein and the background is lower than that of the PV, making high-precision segmentation of hepatic vein small vessels difficult. Unlike the PV, which is a tree-like structure branching out gradually from the first hepatic hilum, the hepatic vein, although originating from the inferior vena cava, has a complex morphology and drainage pattern. It is necessary to use the segmentation results of important structures in the hepatic vein to guide and organize the complex overall segmentation results. The difficulty of completing this task is also greater than that of the PV. This is reflected in two aspects: firstly, the annotation of inherent anatomical points of the hepatic vein, including point B, 99 which indicates the position of the main trunk of the hepatic vein; secondly, the annotation of physiological hepatic fissure veins. Physiological liver fissures are avascular zones of each anatomical unit, generally traversed only by branches of the hepatic vein, 100 making them ideal surgical resection surfaces. Among them, the umbilical fissure and the anterior fissure vein are important boundaries between the right anterior and posterior segments of the liver and segments IVa and IVb of the left medial lobe. 99 Through the spatial positioning relationship analysis and interpretation of the segmentation results of the PV and hepatic vein, accurate segmentation of the fissure veins is achieved, realizing the determination of the position of the physiological liver fissure(Figure 9). Similarly, the detection and effective annotation of the above positions in major hepatectomy are also indicators for assessing the applicability and performance of the model in this scenario.
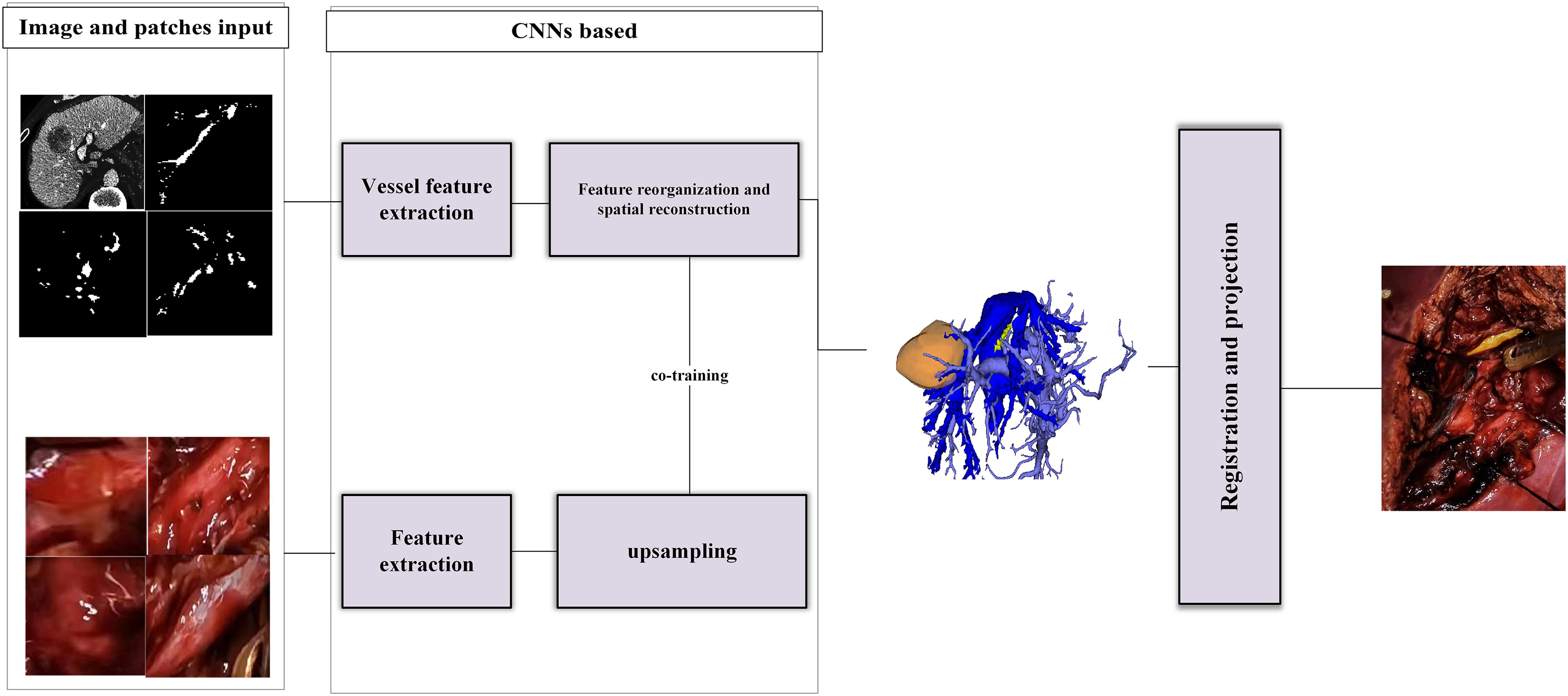
Example of major hepatectomy: precise segmentation of the hepatic vein identifies the middle hepatic vein (yellow-marked area), facilitating surgical planning by organizing and mapping the surgical pathway based on the segmentation results of the middle hepatic vein. Simultaneously during the surgical procedure, the preservation of the functional liver volume of the right anterior segment is achieved, transforming non-resectable hepatocellular carcinoma (HCC) into resectable HCC.
Liver transplantation
Liver transplantation is an effective treatment for end-stage liver disease and liver failure; at the same time, it is also the most complex embodiment of internal medicine treatment and surgical technology, and the most representative of the mode of systematic treatment. Common methods of liver transplantation include whole liver orthotopic liver transplantation, living donor liver transplantation, and split liver transplantation. 101 Assessment and reconstruction of hepatic vasculature are crucial steps in the process of liver transplantation. Vascular complications associated with liver transplantation are significant contributors to adverse outcomes. Evaluation involves assessing anatomical and pathological variations of the vessels in both the donor and recipient, including variations in arterial, portal venous, and hepatic venous vasculature, and ensuring compatibility of vascular structures to minimize risks such as intra-vascular thrombosis, stenosis, and rupture. 102 Dynamic monitoring of the anastomotic sites is also essential to detect and manage any adverse events promptly. Additionally, assessing PV pressure in the donor and recipient liver is a critical task 103 (Figure 10). These efforts represent the directions of clinical translation in hepatic vascular segmentation research. Currently, research is underway to establish predictive models for liver transplantation outcomes using deep learning techniques, incorporating results from hepatic vascular segmentation and related studies into these models represents a promising direction for future development. 104
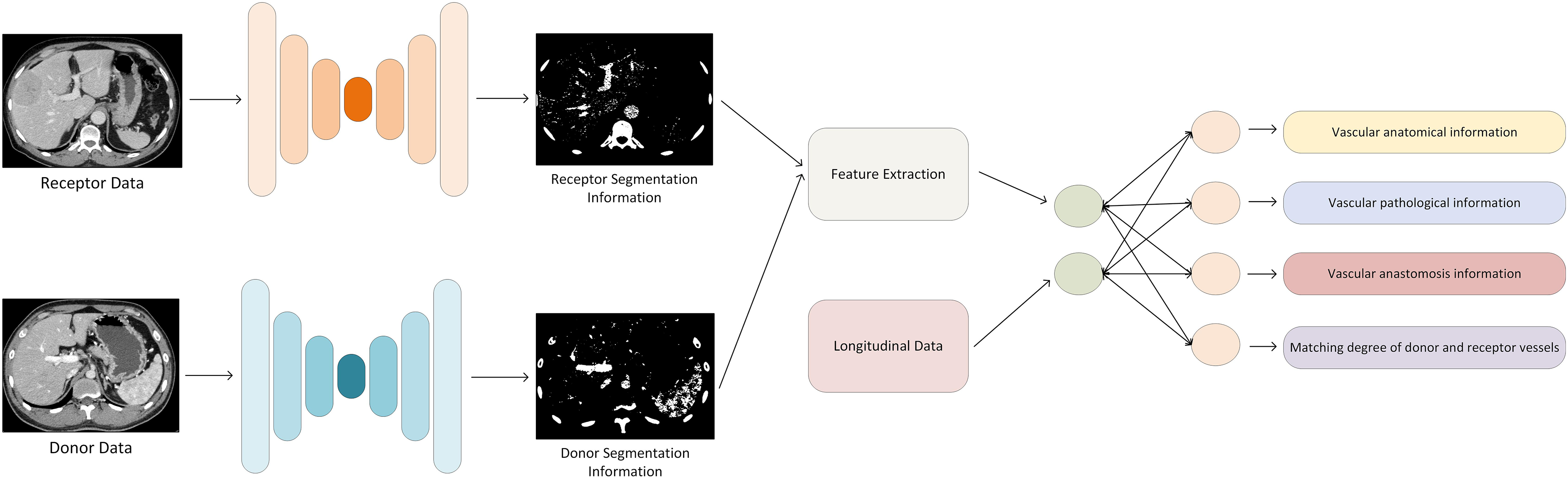
Based on deep learning methods, combined with segmentation information of both donor and receptor hepatic vessels and longitudinal clinical patient data, we achieve the reconstruction of vascular anatomical structures in liver transplant regions, as well as monitoring and prediction of vessel compatibility and anastomosis conditions.
Currently, most clinically used liver vessel segmentation methods are semi-automatic, requiring significant manual interaction, layer by layer, patient by patient, to complete annotation. 5 This process should be automated. Liver vessel segmentation research based on deep learning should aim to reduce the burden on clinicians, be compatible with clinical workflows, provide intuitive and clear interactive interfaces, adhere to clinical guidelines and standards, and ensure that its outputs meet the requirements of clinical practice. By summarizing closely related clinical scenarios, and analyzing and summarizing the needs and challenges close to clinical practice, continuous clinical validation and feedback are also necessary, leading to updates and changes. This helps the model continually adapt to changes in clinical needs and improves its practicality.
Technical challenges and prospects
In this section, we will summarize the limitations of the current deep learning-based hepatic vascular segmentation research from four perspectives, as well as the directions and prospects for solving such problems.
Difficulty in data acquisition and annotation
Medical image data typically involves sensitive patient information and is subject to strict privacy protection and ethical regulations. 105 This restricts the availability of data, making it difficult for researchers to access large and diverse medical image datasets, especially in cross-institutional or international collaborations. Additionally, the lack of a unified data-sharing platform and annotation standards may hinder collaboration between different research teams and the effective utilization of data resources. Accurate liver vessel annotation requires radiologists with professional knowledge, which is not only time-consuming but also costly. Moreover, there may be issues of annotation consistency among different doctors, even if the annotations are performed by experienced radiologists, as different doctors may have different interpretations of vessel boundaries. 106 This inconsistency may affect the learning and generalization ability of models. Due to the diversity of liver diseases and patient physiological conditions, obtaining a comprehensive dataset that represents all possible scenarios is challenging. Biases in the dataset, such as the overrepresentation of specific populations or disease types, may lead to a decrease in the performance of models in actual clinical applications. Therefore, to overcome these limitations, researchers need to explore more efficient data acquisition and annotation strategies, such as using semi-supervised learning, weakly supervised learning, or unsupervised learning methods to reduce the dependence on large amounts of annotated data, 107 as well as developing automated annotation tools to improve annotation efficiency and consistency. 106 At the same time, promoting data sharing and establishing standardized annotation workflows are also crucial for driving development in this field.
Interpretability issues
Deep learning models, especially complex neural networks, are often perceived as “black box” models because their internal workings and decision-making processes are opaque to humans. 108 This means that doctors and researchers find it difficult to understand why the model makes specific segmentation decisions and what the basis for these decisions is. In the medical field, the interpretability of models is crucial for establishing trust between doctors and patients. If doctors cannot understand the predictive results of the model, they may be skeptical about its ACC and reliability, which could hinder the model’s application in clinical practice. Lack of interpretability makes error analysis and model improvement difficult when segmentation results are incorrect. Understanding where and why the model encounters problems is essential for optimizing model performance and reducing future misdiagnoses. In a medical environment, any diagnostic or treatment decision may involve legal liability and ethical considerations. If the model’s decision-making process cannot be explained and verified, determining responsibility and taking appropriate legal action in case of disputes will become complex. The interpretability of deep learning models is also important for medical education and training. Medical students and young doctors need to understand how the model works to better utilize these tools for diagnosis and treatment. To address this limitation, researchers are exploring various methods to improve the interpretability of deep learning models, including: Visualization techniques: Making the model’s decision-making process more intuitive through visualizing activation maps, feature maps, and attention mechanisms. Model simplification: Using simpler or more transparent model structures, such as decision trees or linear models, for easier understanding and interpretation. 109 Interpretability frameworks: Developing dedicated interpretability frameworks, such as local interpretable model-agnostic explanations 110 and Shapley additive explanations , 111 to explain model predictions. Interpretable feature selection: Selecting input features that have clear clinical significance for model predictions, facilitating understanding for doctors and researchers.
Resource requirements
Deep learning models, especially CNNs, typically demand substantial computational resources for training. These resources include high-performance graphical processing units (GPUs), central process units, large amounts of memory, and storage space. For research institutions and hospitals, acquiring and maintaining these hardware devices can incur significant costs. The training process of deep learning models can be extremely time-consuming, especially when using large networks and massive datasets. Prolonged training times may slow down research progress, affecting the rapid iteration and optimization of new models. In clinical environments, particularly in surgical navigation or real-time monitoring, models need to respond quickly and provide accurate segmentation results. Limitations in computational resources may prevent models from running efficiently in real-time or near-real-time environments. To deploy deep learning models on devices with limited computational resources, it may be necessary to compress and optimize the models, using techniques such as knowledge distillation, network pruning, and quantization. These optimizations may reduce model performance or increase the complexity of model design. To overcome this limitation, researchers and developers are exploring various solutions, including: Cloud computing and GPU sharing, 112 model optimization techniques, and developing lightweight models and optimized algorithms to reduce the model’s demand for computational resources. 113
Conclusion
This article systematically reviews the research on hepatic vascular segmentation based on deep learning, from the basic principles of deep learning technology to the current research progress, elucidating aspects such as network architecture, evaluation metrics, motivation, and supervision. Additionally, a summary analysis was conducted on the various clinical scenarios and limitations of the comprehensive diagnosis and treatment of HCC, outlining the challenges and requirements faced by liver vascular segmentation research based on deep learning techniques. By combining clinical needs with technological advancements, deep learning is expected to make greater breakthroughs in the field of liver vessel segmentation, thereby providing stronger support for the diagnosis and treatment of HCC.
Footnotes
Contributorship
TZ contributed to conceptualization, methodology, investigation, and writing–original draft. FY contributed to formal analysis and writing—review and editing. PZ contributed to supervision and project administration.
Declaration of conflicting interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics approval
Not applicable.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Guarantor
Not applicable.
