Abstract
Objective
Convolutional neural networks (CNNs) have achieved state-of-the-art results in various medical image segmentation tasks. However, CNNs often assume that the source and target dataset follow the same probability distribution and when this assumption is not satisfied their performance degrades significantly. This poses a limitation in medical image analysis, where including information from different imaging modalities can bring large clinical benefits. In this work, we present an unsupervised Structure Aware Cross-modality Domain Adaptation (StAC-DA) framework for medical image segmentation.
Methods
StAC-DA implements an image- and feature-level adaptation in a sequential two-step approach. The first step performs an image-level alignment, where images from the source domain are translated to the target domain in pixel space by implementing a CycleGAN-based model. The latter model includes a structure-aware network that preserves the shape of the anatomical structure during translation. The second step consists of a feature-level alignment. A U-Net network with deep supervision is trained with the transformed source domain images and target domain images in an adversarial manner to produce probable segmentations for the target domain.
Results
The framework is evaluated on bidirectional cardiac substructure segmentation. StAC-DA outperforms leading unsupervised domain adaptation approaches, being ranked first in the segmentation of the ascending aorta when adapting from Magnetic Resonance Imaging (MRI) to Computed Tomography (CT) domain and from CT to MRI domain.
Conclusions
The presented framework overcomes the limitations posed by differing distributions in training and testing datasets. Moreover, the experimental results highlight its potential to improve the accuracy of medical image segmentation across diverse imaging modalities.
Keywords
Introduction
Over the last years, deep convolutional neural networks (CNNs) have been successfully used for a variety of computer vision problems, such as image classification, 1 object recognition, 2 and segmentation. 3 In the medical field, CNNs have been applied to develop intelligent methods that assist medical image diagnosis by segmenting anatomical structures, identifying tumors, studying electronic signals, among many others. Medical image segmentation is a critical step in computer-aided diagnosis, where deep learning models have thrived.4–6 Nevertheless, CNNs often rely on high-quality and large labeled training datasets to perform well. Given that annotating medical data are a time-consuming, tedious, and expensive process, acquiring a large medical dataset can be a challenge. Moreover, many supervised learning models assume that the training dataset (source dataset), and the test set (target dataset), follow the same probability distribution. This assumption is hardly met in medical images, where different acquisition protocols, imaging equipment, imaging modalities, and patient population produce a high variation across datasets. 7 Research has even shown that the performance of a CNN degrades in proportion to the distribution difference between target and source domain.8,9
Several techniques have been proposed to solve the problem of domain shift. The simplest solution is to perform transfer learning from the source to the target domain, and sample images from the target domain to fine-tune the network. 10 Nonetheless, this method needs sufficient labeled samples from both domains, which can be restrictive due to the high cost or complex acquiring process. Domain adaptation approaches (DA) have also been presented, whose objective is to transfer knowledge across domains by learning domain-invariant transformations. In domain adaptation, it is assumed that the source domain dataset is annotated while the target dataset can be fully labeled, partially labeled, or completely unlabeled. 7 The latter, also known as unsupervised domain adaptation (UDA), is especially relevant as the target domain is not required to be annotated and can broaden the applicability to different medical datasets. UDA methods are usually divided into two families, namely feature-level adaptation models 11 and image-level adaptation models. 12 Feature-level adaptation aims to align the feature space distribution of the source and target domain. Meanwhile, image-level adaptation methods reduce the gap between the two domains by aligning the data in pixel space. Recently, a third family of methods has been proposed, which combines image- and feature-level adaptation models to further reduce the domain shift.13,14 This type of method has been shown to provide a better segmentation performance on the target domain as feature and image adaptation are complementary perspectives.
While there has been relevant progress in the development of UDA models, most works focus on the problems of natural image segmentation13,15,16 or medical image classification17–19 Given the complex nature and dimensionality of medical images, segmenting medical data is a more challenging task. The works devoted to medical image segmentation have mostly used a CycleGAN model 20 to map the source domain images to target domain and train the segmentation network with the translated source domain images.21,22 The challenge encountered with this approach is that by only using an adversarial loss to train the generators responsible for the translation, there is no guarantee that the original shape will be conserved during the mapping. Considering that the annotation correctness must be preserved during the transformation to succeed in a segmentation task, this is an issue that requires special attention. In Ref.23,24, a loss function that encourages the preservation of anatomical structures during translation was proposed. Nevertheless, both works present solely an image-level adaptation method that might not be enough when the source and target dataset suffer from a severe distribution shift. Chen et al. 14 presented an unsupervised cross-modality adaptation method that implements an image and feature alignment. However, the whole framework is trained end-to-end in one step, which is computationally and memory-intensive and can prohibit its application in high-resolution imagery or settings were powerful computer systems are not available.
In this work, we present Structure Aware Cross-modality Domain Adaptation (StAC-DA), an unsupervised StAC-DA framework for medical image segmentation. StAC-DA implements an image- and feature-level adaptation in a sequential two-step approach. The first step performs an image-level alignment, where images from the source domain are translated to the target domain in pixel space by implementing a CycleGAN-based model. The latter model includes a structure-aware module composed of two segmentation networks that preserves the shape of the anatomical structure during translation. The second step consists of a feature-level alignment. In this step, a U-Net network with deep supervision is trained with the transformed source domain images and target domain images in an adversarial manner to produce probable segmentations for the target domain. Furthermore, an auxiliary discriminator network that receives the predicted segmentations of the deep supervised layer is added to the model to improve the feature-level alignment. StAC-DA is purposely designed in a sequential manner to reduce the computational requirements during training. Furthermore, to prevent the loss of information between the two steps, transfer learning is applied from the architectures from Step 1 to Step 2. The proposed framework is evaluated on the problem of bidirectional cardiac substructure segmentation from the Multi-Modality Whole Heart Segmentation Challenge dataset. 25 We validate the proposed method in unpaired Magnetic Resonance Imaging (MRI) and Computed Tomography (CT) images by adapting images from MRI to CT domain, and from CT to MRI domain. The experimental results show that StAC-DA outperforms leading unsupervised domain adaptation and is ranked first in the segmentation of the ascending aorta when adapting from MRI to CT domain and from CT to MRI domain. Moreover, StAC-DA is ranked third in the segmentation of the left ventricle blood cavity and left ventricle myocardium in the adaptation from the MRI to CT domain.
The contributions of this work are threefold. First, we propose an image- and feature-level adaptation framework for UDA that through its two-step implementation preserves the semantic information and enhances domain alignment. Secondly, we present a structure-aware CycleGAN-based model that performs an image-level alignment to emphasize shape consistency by including a source and target domain segmentation network in the model. Finally, we propose a validation loss function, based on what we have denominated the class area ratio metric, to monitor the performance of the network on the unlabeled target dataset.
The remainder of the paper is organized as follows. Section Related work provides an overview of related work on UDA models for medical image segmentation. Section Method presents the two-step StAC-DA framework and the experimental methodology. Section Results provides ablation studies and benchmark results obtained on the unsupervised cardiac segmentation task. Section Discussion contains the discussion, and Section Conclusion presents the conclusions.
Related work
In this section, we provide a review of works on UDA for medical image segmentation classified by feature-level adaptation methods, image-level adaptation methods, and combined image- and feature-level adaptation methods.
Feature-level adaptation methods
Feature-level adaptation methods transform the source and target domain data from their original feature space to a new shared and aligned feature space. The alignment is usually achieved by minimizing a distance measure such as maximum mean discrepancy, 26 correlation distance, 27 or adversarial discriminator accuracy. 28 In, 11 Dou et al. proposed a plug-and-play adversarial domain adaptation network that aligns the target and source domain in feature space at multiple scales. Using adversarial learning, two discriminators are built to distinguish multilevel features and predicted segmentation masks on the two domains. Degel et al. 29 presented a combined deep-learning-based approach that incorporates shape prior information and a domain discriminator to encourage feature domain-invariance across datasets. Kamnitsas et al. 30 developed a multilevel feature adaptation method to derive domain-invariant features with a multiconnected domain discriminator. Although feature-level adaptation methods are efficient, they can fail to capture low-level appearance variance and do not enforce semantic consistency.
Image-level adaptation methods
In image-level adaptation methods, the images from one domain are transferred to another domain through a pixel-to-pixel transformation. The majority of methods achieve image translation through the application of generative models such as generative adversarial networks (GANs). 31 Zhao et al. 12 proposed a modified U-Net to synthesize MR brain images from CT images using a paired co-registered image dataset. Afterwards, a MALP-EM network 32 is applied to segment the whole brain from the synthetic MR images. Nie et al. 33 proposed a context-aware GAN that generates CT images from MR images. An image-gradient-difference based loss is presented to alleviate the blurriness of the generated CT image. Tomar et al. 24 presented a self-attentive spatial adaptive normalization method that introduces a self-attention module that focuses on the anatomical structures of organs to improve the image translation task. Although these methods have shown promising results, most of them work well mainly in datasets with a limited domain shift and can lose semantic content.
Combined feature-level and image-level adaptation methods
Recently, works have proposed using a hybrid image- and feature-level adaptation method to mitigate severe domain shift. Chen et al. 14 presented a synergistic framework for cardiac image segmentation. The appearance of the source images is translated to the target domain with a cycle-consistent GAN (CycleGAN) 20 while simultaneously a two-stream CNN is trained with a domain discriminator to reduce the domain gap in feature space. Yan et al. 34 applied a CycleGAN with a modified loss, which includes image and feature-level similarity, to transform target images to source domain. Afterward, a U-net network is fully trained in the source domain and used for inference. Cui et al. 35 proposed a GAN-based bidirectional adaptive framework, which applies a CycleGAN-based process to translate the images from the source domain to the target domain. During the image synthesis and semantic prediction, the networks share the same encoder. Furthermore, a self-attention mechanism and spectral normalization are included in the generator, encoder, and discriminator networks to enhance the authenticity of the generated target domain images. In Ref. 36 , we presented a sequential image- and feature-level adaptation method for brain MRI segmentation. In the first step, a CycleGAN model is implemented for image translation between the source and target domain. In the second step, a U-Net network is trained to segment the target domain images in an adversarial manner using information about the shape, texture, and contour of the predicted segmentation. StAC-DA is a significant extension of that work. Specifically, we introduce a new structure-aware CycleGAN-based model in Step 1 to encourage annotation correctness during translation. Furthermore, we enhance the validation by performing experiments on the bidirectional segmentation of a cardiac dataset and provide a more detailed description of the framework.
Method
In this predictive study, we propose the StAC-DA framework. StAC-DA is an unsupervised structure-aware cross-modality domain adaptation method composed of two sequential steps during training, as shown in Figure 1. In Step 1, the images of the source domain are translated to the target domain using a CycleGAN-based model with a structure-aware module to preserve the shape of the anatomical structures during the domain translation. This step converts the source domain images into target style-like images. In Step 2, a feature-level adaptation method is proposed by training a U-Net segmentation network with the translated source domain images and the target domain images using an adversarial training scheme. The objective is to produce probable segmentations for the target domain that follow the same probability distribution as the ground truth segmentations from the source domain. Since the U-Net network is trained with synthesized target domain images, only the trained U-Net architecture is necessary to produce the predictions during inference.

StAC-DA framework is composed of two sequential steps during training. Step 1 performs a structure-aware image-level adaptation by implementing a CycleGAN-based model. Step 2 implements a feature-level adaptation by training a deeply supervised U-Net network with an adversarial training scheme.
In the following subsections, we describe Steps 1 and 2 of the proposed framework, the validation loss function implemented to monitor the performance of the network, the dataset used for the experiments, training details, and the quantitative metrics for evaluation. The computational experiments where performed at Universidad San Francisco de Quito, Ecuador, from August 2021 to August 2022.
Step 1: structure-aware image-level adaptation
In this step, the aim is to learn a structure-aware mapping network that translates the images from the source domain
The CycleGAN-based model consists of two modules, the image synthesis module, and the structure-aware module. The image synthesis module is made up of the source domain generator network 
Furthermore, to avoid model collapse during training and incentivize the mapped images 


Finally, 

Step 2: feature-level adaptation
In Step 2, a feature-level adaptation method in the semantic prediction space is implemented by using an adversarial training scheme. This phase is especially necessary when there is a severe domain gap between the target and source images. After finishing Step 1, the 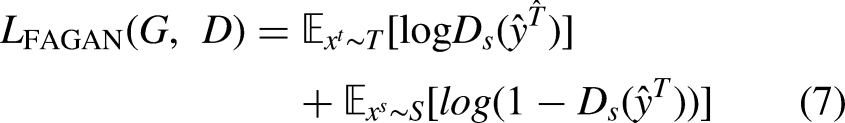

Deeply supervised U-Net network trained in Step 2 to segment the unlabeled target domain images. The numbers over the convolutional blocks correspond to the height, width, and number of feature maps.
In addition to the adversarial training, on each iteration the U-Net is also trained in a supervised manner to segment the mapped source domain images 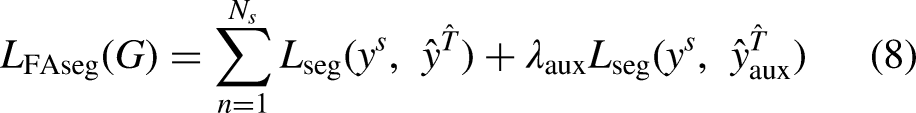
Monitoring validation metric
Since there are no labeled target domain images, it is a challenge to select the best weights for testing. Hence, we propose a pseudo validation loss function based on the segmentation area and dice coefficient to monitor the performance of the network on the target dataset. First, the ground truth segmentations from the source domain are used to calculate the average number of pixels per class 

Dataset and preprocessing
The proposed StAC-DA is evaluated on the task of bidirectional segmentation of cardiac structures on MRI and CT imagining modalities from the Multi-Modality Whole Heart Segmentation
25
(MMWHS) dataset. The MMWHS dataset consists of 20 MRI and 20 CT whole cardiac volumes. The MRI and CT images are unpaired and collected from distinct patient cohorts. For both modalities, the ground truth segmentations for the ascending aorta (AA), left atrium blood cavity (LA-blood), left ventricle blood cavity (LV-blood), and myocardium of the left ventricle (LV-myo) are provided. Following the work of,
24
the images of 16 subjects are used for training, and images from 4 subject for testing. Furthermore, the preprocessing proposed by Ref.
14
is used, where the central heart region of the image is first cropped to size of 256
Training details
We perform two types of experiments. First, we evaluate the effectiveness of the method by performing the adaptation from the MRI to the CT domain (MRI

Examples of the translation results using the CycleGAN model and structure-aware model from Step 1. (a) Adaptation from CT to MRI domain. (b) Adaptation from MRI to CT domain. The proposed structure-aware model successfully translates the images between domains while keeping the structure of the anatomical region.
In Step 2, the U-Net model is trained for 100 epochs. The discriminators and U-Net network are optimized with the Adam optimizer and a learning rate of
Evaluation metrics
The Dice similarity coefficient (Dice) and average symmetric surface distance metric (ASSD) are employed to quantitatively evaluate the segmentation performance of the models. The Dice coefficient is an overlap-based metric that measures the intersection between the ground truth segmentation and predicted segmentation. The ASSD distance is a spatial-based metric that calculates the average of all distances between points on the ground truth´s boundary surface to points on the predicted segmentatiońs boundary surface. Let 
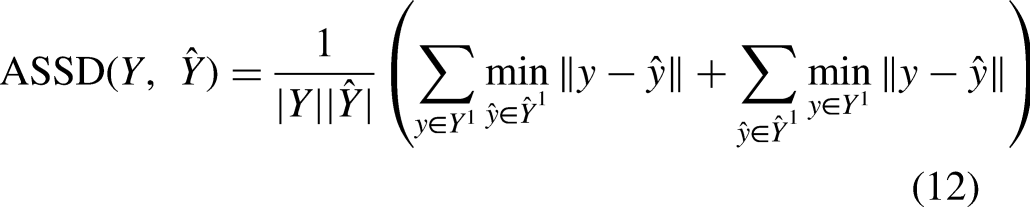
Results
Ablation studies
The proposed StAC-DA model is composed of a structure-aware CycleGAN-based model in Step 1 and a segmentation network with feature-level alignment in Step 2. In this section, we perform ablation studies to evaluate the contribution of each of these steps. For this objective, we train four models for the adaptation from MRI to CT domain. The results are displayed in Table 1. In the first model, we implement the CycleGAN model
20
to translate the images from MRI to CT modality, and train a U-Net network on the translated CT images in a supervised manner. In the second model, we apply the proposed structure-aware CycleGAN-based model for image translation and use the trained
Ablation study of the components of StAC-DA.
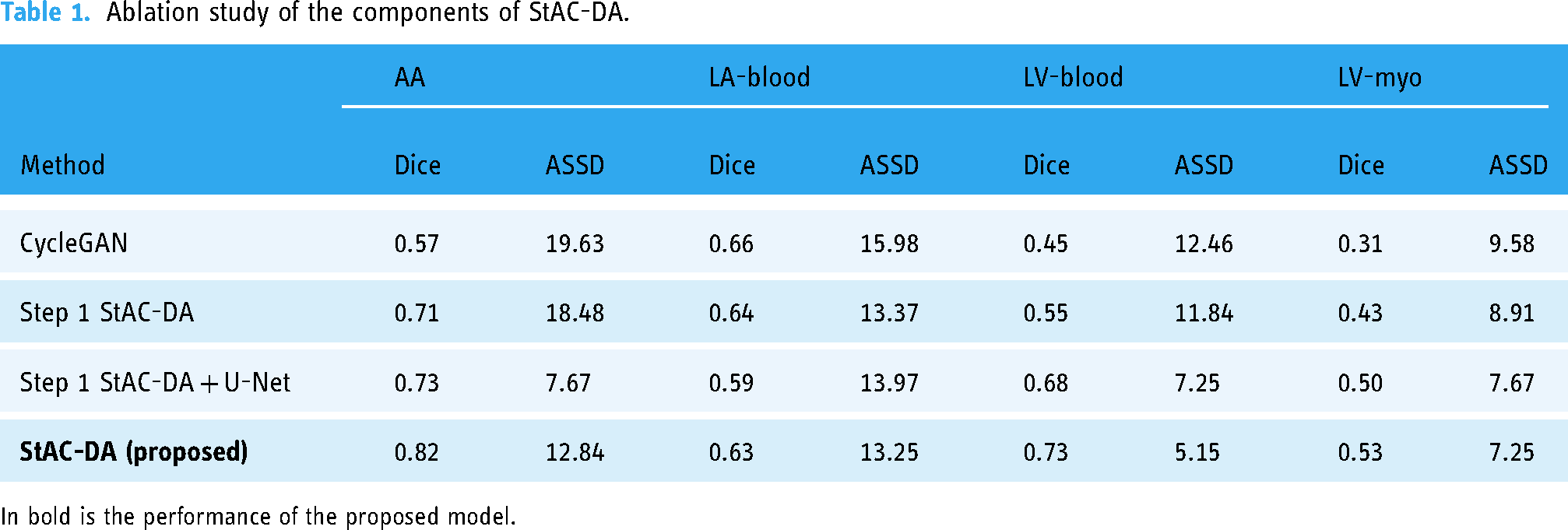
In bold is the performance of the proposed model.
The results demonstrate that including the structure-aware module in the CycleGAN network helps translate the overall shape of the heart better as the segmentation network is able to identify and segment the different cardiac regions with higher accuracy. Particularly, “Step 1 StAC-DA” and “Step 1 StAC-DA + U-Net” have an important improvement over CycleGAN in the segmentation of ascending aorta, left ventricle blood cavity, and myocardium of the left ventricle when considering Dice and ASSD metrics. The experiments also show that training a segmentation network with the synthesized images obtained from the structure-aware CycleGAN-based network increases the performance in comparison to using the
Benchmark results
Our method is compared against six leading unsupervised domain adaptation networks: U-GAT-IT model,
42
PnP-Ada-Net model,
11
SynSeg-Net,
43
AdaOutput,
15
Cycada,
13
and SIFA model.
14
Moreover, we provide a performance upper bound to measure the performance gap by training the U-Net network in a supervised manner on the target domain (denoted as “UpperB U-Net”). To evidence the domain shift, a performance lower bound is also presented (denoted as “LowerB U-Net”) by using the U-Net trained on the source domain to predict the segmentation on the target domain images without any adaptation method. Finally, the results obtained after applying only Step 1 of the proposed framework and using the
Performance comparison of the proposed method (StAC-DA) and leading unsupervised domain adaptation methods for cardiac structure segmentation from MRI to CT domain (MRI

Performance comparison of the proposed method (StAC-DA) and leading unsupervised domain adaptation methods for cardiac structure segmentation from CT to MRI domain (CT

In Figures 4 and 5 a qualitative evaluation of the proposed model is shown on the MRI

Segmentation results after the MRI

Segmentation results after the CT
Computational requirements
StAC-DA is designed sequentially to reduce computational requirements during training. In Table 4, we present the GPU and RAM memory used in each step during training, using a batch size of 1, as well as the number of trainable parameters. For comparison, we have also included the computational requirements of the unified model proposed by Chen et al.
14
In terms of GPU and RAM usage, the entire StAC-DA framework has lower requirements. Furthermore, as each step is trained independently, the model can be trained in facilities with less powerful GPUs and RAM memory. Regarding the number of trainable parameters, SIFA is smaller. Nevertheless, since the entire StAC-DA framework is not processed simultaneously, it is unnecessary to save all the models in Steps 1 and 2. Specifically, after training Step 1, only the Generator network is needed in Step 2 to transfer the images from the source to target domain. The Generator network has
Computational requirements of StAC-DA and SIFA during training with a batch size of 1.

Discussion
Deep learning models have been shown to excel in various complex tasks when large amounts of data are available. Nevertheless, when the networks are tested in data that does not follow the training distribution, their performance can significantly degrade. This challenge is of special interest in the medical imagining community, where labeling images is very costly, and due to the different imaging modalities and acquisition protocols, the testing data can differ significantly from the training set. Developing unsupervised cross-modality models is advantageous for the application of AI in medical settings because it decreases the need to obtain costly labeled data while exploiting to the fullest the unlabeled data from the target domain. In this work, we presented StAC-DA, an unsupervised structure-aware domain adaptation framework for cross-modality medical image segmentation. The proposed framework is comprised of a two-step image and feature-level adaptation that importantly reduces the performance degradation when moving from two different imaging modalities.
StAC-DA has been tested on two domain adaptation tasks from a publicly available cross-modality cardiac segmentation challenge. We implemented the proposed framework for domain adaptation from MRI to CT imaging modality (MRI
An interesting observation is that although all competing models seem to reduce the domain shift, there is still a performance gap, particularly in the CT
A limitation of the proposed framework is applying 2D CNNs for image translation and segmentation. Although 2D CNNs are able to capture intra-slice information, they do not fully exploit volumetric information. Hence, some important 3D features that can boost performance might be missed. Since training the 2D structure-aware CycleGAN-based model is already computationally and time intensive, a future direction can be using a 3D segmentation CNN for Step 2 or using 3D patches instead of the whole 3D image for Steps 1 and 2. Furthermore, when utilizing the monitoring validation metric we are assuming the morphology of anatomical structures is consistent across the imaging modalities. However, if the population in the source domain is considerably different from that of the target domain the metric might not be applicable. Lastly, although the proposed framework aims to reduce the computational cost by dividing the training process into two steps, the memory usage can still be limiting with large datasets, high-resolution images or more complex segmentation tasks. To address these issues, future work could explore optimization techniques like model pruning, distributed training strategies, and scalable architectures, potentially broadening the framework’s applicability.
Conclusion
In this work, we present StAC-DA, an unsupervised structure-aware cross-modality domain adaptation framework for medical image segmentation. StAC-DA is composed of two sequential steps. First performs a structure-aware image-level adaptation, where images from the source domain are mapped to the target domain through a CycleGAN-based model. The latter includes a segmentation network that preserves the anatomical structures during translation. In the second step, a feature-level adaptation is applied by training a deeply supervised U-Net architecture in an adversarial manner to produce probable segmentations for the target domain. StAC-DA is evaluated on the task of bidirectional cardiac substructure segmentation from the Multi-Modality Whole Heart Segmentation Challenge dataset. The experiments demonstrate that the proposed model has a very competitive performance, being ranked first in the segmentation of the ascending aorta when adapting from MRI to CT domain and from the CT to MRI domain.
Footnotes
Acknowledgments
This work was supported in part by POLIGRANT No. 17376, Colegio de Ciencias e Ingenierías, USFQ. The authors thank to the Applied Signal Processing and Machine Learning Research Group of USFQ for providing the computing infrastructure (NVidia DGX workstation) to implement and execute the developed source code.
Data availability statement
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article. This work was supported by the Universidad San Francisco de Quito (grant number: 17376).
Guarantor
MB-C
