Abstract
Objective
To achieve an accurate assessment of orthodontic and restorative treatments, tooth segmentation of dental panoramic X-ray images is a critical preliminary step, however, dental panoramic X-ray images suffer from poorly defined interdental boundaries and low root-to-alveolar bone contrast, which pose significant challenges to tooth segmentation. In this article, we propose a multi-feature coordinate position learning-based tooth image segmentation method for tooth segmentation.
Methods
For better analysis, the input image is randomly flipped horizontally and vertically to enhance the data. Our method extracts multi-scale tooth features from the designed residual omni-dimensional dynamic convolution and the designed two-stream coordinate attention module can further complement the tooth boundary features, and finally the two features are fused to enhance the local details of the features and global contextual information, which achieves the enrichment and optimization of the feature information.
Results
The publicly available adult dental datasets Archive and Dataset and Code were used in the study. The experimental results were 87.96% and 92.04% for IoU, 97.79% and 97.32% for ACC, and 86.42% and 95.64% for Dice.
Conclusion
The experimental results show that the proposed network can be used to assist doctors in quickly viewing tooth positions, and we also validate the effectiveness of the proposed two modules in fusing features.
Introduction
Among the many dental imaging modalities, dental panoramic X-ray images are an efficient and cost-effective means of imaging.1,2 The main advantage of this technology is that it provides integrated visualization of the entire oral structure, including the teeth, jaws, and surrounding soft tissues. This information enables physicians to conduct more comprehensive and detailed diagnoses, leading to precise lesion localization and resection extent planning. Compared to other more limited imaging methods, panoramic X-ray imaging holds irreplaceable value in diagnosing a broad spectrum of oral diseases: periodontal disease, dental infections, and maxillofacial tumors. 46 In addition, the technique typically requires a relatively low radiation dose and cost, enhancing its feasibility and availability for clinical applications. The accuracy of image analysis is crucial in the diagnostic and therapeutic processes of dentistry. Manually analyzing panoramic X-ray images is a complex and time-consuming process heavily dependent on the physician's a priori knowledge, even for experienced physicians. 3 Therefore, accurate segmentation of dental panoramic X-ray radiographic images is necessary to help junior practitioners lacking clinical experience to improve the accuracy and efficiency of their diagnosis in the preoperative period, and it plays a significant role in the clinical diagnosis and treatment within dentistry.
Dental panoramic X-ray images contain complete information about the teeth and jaws, from which complete and precise segmentation of tooth morphology information is an important prerequisite for intelligently assisted treatment. The segmentation method based on deep learning currently dominates the field and undergoes extensive research and enhancement. Since the pioneering work of U-Net created a buzz in the field of image segmentation, more accurate and efficient U-Net-based image segmentation methods have been explored from multilayer perceptual machines (MLPs), convolutional neural networks, and attention mechanisms, followed by the emergence of excellent schemes such as UNet++, Attention U-Net, GT U-Net, and UNeXt. Existing methods face the following challenges when segmenting dental panoramic X-ray images: firstly, the characteristics of dental panoramic X-ray images themselves, such as the boundary contours of the teeth are difficult to distinguish obviously, and the pixel values of the teeth are close to the surrounding area, resulting in a lower overall contrast; secondly, the network structure of these algorithms has insufficient ability to extract the target objects of small and large sizes, and the receptive field information in the network model is partially lost during feature propagation, resulting in the edges of the tooth segmentation having a more obvious jagged appearance.
Based on the above analysis, to better adapt and learn dental panoramic X-ray image data, this article proposes a dental panoramic X-ray image segmentation method based on multi-feature coordinate position learning. To ensure that the algorithm can extract detail features and regional features, we design a residual omni-dimensional convolution module, which constructs the main feature extraction branch and the auxiliary feature complementary branch, and effectively learns both large-size regional feature information and small-size detailed feature information. In addition, to alleviate the differences between the feature encoding network and the feature decoding network and at the same time to obtain more accurate tooth region location and tooth contour information, we designed a two-stream coordinate attention module, and this module adds a new maximum pooling processing stream based on the average pooling processing stream, which accurately locates the location of the tooth target region, and at the same time, learns the positional feature information of the tooth edge contour very well.
Literature review
In recent years, tooth segmentation tasks on panoramic X-ray images have attracted increasing attention, and tooth segmentation tasks can be used to solve difficult dental problems. Teeth segmentation methods can be broadly categorized into two research lines: traditional methods based on original image features and deep learning-based methods driven by deep image features.
Traditional segmentation methods use digital image processing techniques to perform tooth segmentation according to the characteristics (e.g. shape, gray value, etc.) inherent in the dental panoramic radiograph. Commonly used traditional segmentation methods include watershed-based segmentation algorithms,4,5 threshold-based segmentation algorithms,6–8 clustering-based segmentation algorithms,9,10 boundary-based segmentation algorithms,11,12 and region-based segmentation algorithms. 13 Watershed-based segmentation algorithm simulates the topographic water flow and fills the local minima to divide the image. The segmentation results of this algorithm are very much related to the number of gray levels and the selection of the threshold value, and the computation is large and prone to over-segmentation. Threshold-based segmentation algorithm separates the background from the target by analyzing the gray level change of the image and setting the threshold value, which is suitable for images with large gray level difference, but has more limitations in the case of small gray level difference. Clustering-based segmentation algorithm divides the pixels in the image into clusters based on certain rules, so that the pixels in the same cluster have similar features and the pixels between different clusters have different features, and finally completes the image segmentation. However, this method is sensitive to noise and is not effective in complex backgrounds, and the stability of the algorithm is also affected by the choice of initial values. Boundary-based segmentation algorithm aims to find the boundary of the image, the pixel at the boundary and the surrounding pixels of the gray value of the difference is very obvious, the boundary pixels will be found in a line to form the boundary of the object contour, to complete the image segmentation. The region growing method is a commonly used region-based segmentation algorithm, which gradually merges the surrounding regions to complete the segmentation by setting rules and initial pixels. However, this method is sensitive to noise, depends on the initial seed selection, and is easily affected by the image quality. In summary, early conventional methods have too many limitations. Therefore, automated methods are important to promote the efficiency and accuracy of tooth segmentation.
In the field of tooth segmentation, compared with traditional segmentation methods, deep learning-based segmentation methods have stronger tooth feature modeling capabilities and can achieve better segmentation results, while eliminating the need for complex and tedious implementation rule definitions. Therefore, deep learning-based segmentation methods are currently the mainstream segmentation methods, which have been widely studied and improved. However, dental panoramic X-ray images are medical image data, which have special features such as unclear boundaries and low contrast compared with general image data, 45 so general image segmentation methods are not suitable for medical image data. After fully convolutional network (FCN), 14 Ronneberger et al. 15 proposed U-Net, whose network structure is similar to the U-shape, and mainly consists of encoder, decoder, and skip connection. The encoder is mainly responsible for the extraction of feature information in the image, the decoder is mainly responsible for restoring the image size to the original size, and the skip connection is mainly responsible for ensuring that more low-level feature information in the image is not lost. Although U-Net achieves good segmentation results, its feature extraction capability is still to be improved due to the backward design of the codec convolutional structure. In general, U-Net has become an important research method in the field of medical image segmentation, and its concise and powerful structural model become a highly influential design idea; some subsequent research teams designed mechanisms such as residual connection, dense connection, multiscale, attention, and so on to better improve the performance of the segmentation algorithm. Therefore, exploring U-Net and related networks is an important reference value for the research of this topic.
The residual connection mechanism originates from ResNet, 16 the core of which lies in the addition of a new data line spanning a number of layers in the network, which effectively solves the problem of gradient information loss caused by the increase of network depth. Inspired by the residual connection mechanism and the dense connection mechanism, Zhou et al. 17 proposed the UNet++ segmentation network model, UNet++ refers to and builds on the basis of the U-shaped network structure, which is different from the skip connection of the U-shaped network, but UNet++ establishes a number of sub-networks in the intermediate part, so that the image feature information extracted by the encoder is processed in the intermediate subnetworks and then handed over to the corresponding encoder. UNet++ builds multiple subnetworks in the middle part, and the image feature information extracted by the encoder is processed by the intermediate subnetworks, and then handed over to the corresponding encoder, which establishes the connection between the low-level detail information and the high-level abstract features, and improves the problem of the large difference of semantic features in the encoding and decoding stages, but the insufficient fusion of the information of the upper and lower layers of UNet++ makes the segmentation results still insufficiently fine. Liu et al. 18 proposed Res-Unet, which introduced the residual unit structure of ResNet into the encoding and decoding structure, increased the number of layers in the network model, and achieved good segmentation results. Rao et al. 19 constructed LeFUNet based on the U-Net by adopting the dense connection and combining improved squeezing and excitation modules to improve the accuracy of X-ray image segmentation.
The multi-scale mechanism functions as a means of sampling the image at different levels, with different levels providing images of varied scales and resolutions, each scale size image contains unique and easy-to-learn semantic features, which provide rich decision-making information for image segmentation. 20 Inspired by the multi-scale mechanism, CE-Net 21 believes that increasing the receptive field is conducive to improving the final image segmentation accuracy, so it incorporates pyramid pooling and atrous convolution into the U-shaped network which is mainly composed of ResNet34 models, and proposes the residual-based multi-kernel pooling (RMP) module and the dense atrous convolution (DAC) module based on dense connectivity. The DAC module captures more semantic feature information by embedding multi-scale atrous convolution into multiple cascade branches; the RMP module combines several pooling operations of different sizes to capture enough background information on the semantic feature information after DAC processing. Through these two modules, CE-Net captures some high-level feature information and reduces the loss of spatial information in the image to get better image segmentation results. Wang et al. 22 proposed ARMS Net and designed adaptive multi-scale feature extraction module (AMFEM), which allows the receptive field to be dynamically adjusted with the feature map size. Wang et al. 23 borrowed the idea of pyramid pooling, designed a multi-scale feature extraction block, and proposed multi-path connected network (MCNet), which obtains larger and more the receptive field information. Chen et al. 24 proposed a multiscale position-aware network, designed a position-aware module for locating the target pixels, and reduced the feature information gap in the multiscale feature branches by an aggregation module to finalize the X-ray image segmentation.
In the recent years of research, researchers widely apply the attention mechanism in image tasks, 25 garnering accolades from many scholars. Inspired by the attention mechanism, Oktay et al. 26 proposed the Attention U-Net, which introduces an attention module based on the U-Net structure. They place this attention module before splicing the encoding path features and the decoding path features at each layer, and it adjusts the feature information extracted from the encoding path, reduces the importance of the irrelevant feature regions in the image, and provides enhanced highlighting of key feature areas in an image. Li et al. 27 proposed GT U-Net, which replaces the encoding and decoding structure with a group transformer with better performance, 28 and also reduces the computation of the network model by grouping and bottlenecking structure to finally complete the X-ray image segmentation. Kaya et al. 29 combined U-Net and OctConv 30 to propose an X-ray image segmentation method with lower memory overhead, and the improved U-Net also achieved better segmentation accuracy. Sheng et al. 31 used SWin-Unet 32 as the network model used in the segmentation algorithm, and the experimental results verified the advantages and future potential of SWin-Unet for X-ray image segmentation. Zhao et al. 33 proposed TSASNet, which divides the X-ray image segmentation task into two phases, with the first phase using the global and local attention modules to coarsely localize the target region, and the second phase using the fully convolutional network to complete the fine segmentation. UNeXt 34 proposed the tokenized multilayer perceptron (tokenized MLP) and introduced it into the lower two layers of the U-shaped network structure, replacing the original convolutional part. The larger size of the downsampling amplitude and the use of fewer convolutional layers make UNeXt lightweight, but these operations lose the receptive field information that is useful for the network model.
In recent years, vision transformers 35 have rapidly become one of the most promising vision-based models as a powerful visual task-aware model. However, the performance of existing transformers in medical image segmentation tasks is still unsatisfactory. Polyper 36 assumes that the localization of the segmented region is accurate; however, in practical applications, false positives or false negatives are often present in medical images, which makes the Polyper perform poorly in dealing with these cases and affects the reliability and accuracy of the segmentation. Swin transformer 37 introduces frequent shifting or reshaping operations, which improve the representativeness of the model, but also introduce significant computational latency, limiting its application to real-time or efficient segmentation tasks. The SegFormer 38 approach combines a hierarchical transformer encoder with no positional coding and a lightweight decoder, and while it can theoretically improve segmentation performance, even the lightest model may still be too heavy to be practical for some edge devices with limited computational resources.
In summary, since the advent of U-Net, deep learning-based methods have demonstrated excellent segmentation performance. However, there are still some deficiencies in the design of the network structure, such as the feature extraction ability of tooth objects with variable sizes needs to be enhanced, the unreasonable use of special convolutions such as atrous convolution leads to serious jaggedness at the edges of the tooth segmentation, and the information of the receptive field of the network model becomes chaotic, which ultimately affects the enhancement of the accuracy of tooth segmentation. Therefore, how to optimize and improve the feature extraction, propagation, and recovery parts of the segmentation network emerge as key issues in dental panoramic X-ray image segmentation (see Table 1).
Summary of the literature.

Model and methodology
This study is an applied research aimed at improving the tooth segmentation technique for dental panoramic X-ray images by using a multi-feature coordinate position learning approach to improve the accuracy and efficiency of segmentation. This study was initiated in April 2023 and concluded in May 2024. The entire research was conducted at Xi'an University of Science and Technology, Xi'an, Shaanxi Province, China.
General architecture of the network model
The overall network architecture of the proposed method, as shown in Figure 1, comprises a residual omni-dimensional convolution module (ROCM), two-stream coordinate attention (TSCA) module, downsampling layer, upsampling layer, and rectification output layer. The feature encoding network, positioned on the left side of the overall network architecture, actively learns tooth and background features layer by layer from images, taking pre-processed dental panoramic X-ray images as its input. The feature decoding network, located on the right side of the architecture, is responsible for progressively restoring image features layer by layer to yield the final segmentation results.

General network architecture of the proposed method.
Firstly, the designed ROCM is introduced into each layer of the feature encoding network and feature decoding network to construct two branches with different scopes of action and learn to acquire large-size regional features and small-size detailed features at the same time, to achieve better feature learning effect and more comprehensive feature recovery effect. Next, the designed TSCA module is introduced between the feature encoding network and the feature decoding network, which focuses on the target object itself and the junction between the target object and the background, learns these two kinds of closely related feature relationships, and then supplements the original feature information of the feature encoding network by skip connections for the feature decoding network; it is worth noting that the output features of the previous layer of the feature decoding network will be resized by the upsampling layer and finally spliced together with the output results of the TSCA module. In addition, the feature encoding network needs to use the downsampling layer to reduce the size of the feature map, which is achieved by the maximum pooling operation, and similarly, the feature decoding network needs to achieve a gradual expansion of the feature map size to the input image size through the upsampling layer, which is specifically achieved by the bilinear interpolation. Finally, the rectification output layer converts the output of the previous network part into the segmentation result on the image, which is composed of a convolution kernel size of 1 × 1 and step size of 1 and a Sigmoid function.
Residual omni-dimensional convolution module
Common convolution is a very widely used type of convolution, a common convolution has a static convolution kernel, and the weight parameters trained in the convolution kernel apply to all input samples, that is to say, the weight parameters of the common convolution kernel are independent of the input samples. Dynamic convolution is composed differently from the common convolution, and can be regarded as the use of multiple convolution kernels in combination according to a certain rule, which is usually related to the input sample data, and the dynamic convolution can be expressed by equation (1):
Some previous work on dynamic convolution is mainly conditionally parameterized convolutions (CondConv)
39
and “dynamic convolution” (DyConv).
40
Both serve as extensions of equation (3.1), and their attention function implementations are similar, but DyConv uses a Softmax function while CondConv uses a Sigmoid function. Compared to common convolutions, CondConv and DyConv still lack some considerations, although they improve the performance of convolutions through linear combinations. Specifically, for given
Given the above-mentioned, Li et al.
1
proposed omni-dimensional dynamic convolution (ODConv), which, unlike CondConv and DyConv, is a method that simultaneously takes into account the number of input feature channels, the number of output feature channels, the size of the convolution kernel space dimensions, and the number of convolutional kernel number of the new dynamic convolution. Based on the previous equation (1) and further extended, researchers express ODConv with equation (2):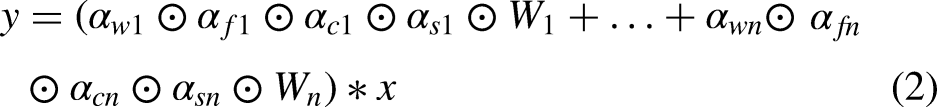

Structure of the omni-dimensional convolutional ODConv.

Positional multiplication operations along the spatial dimension.
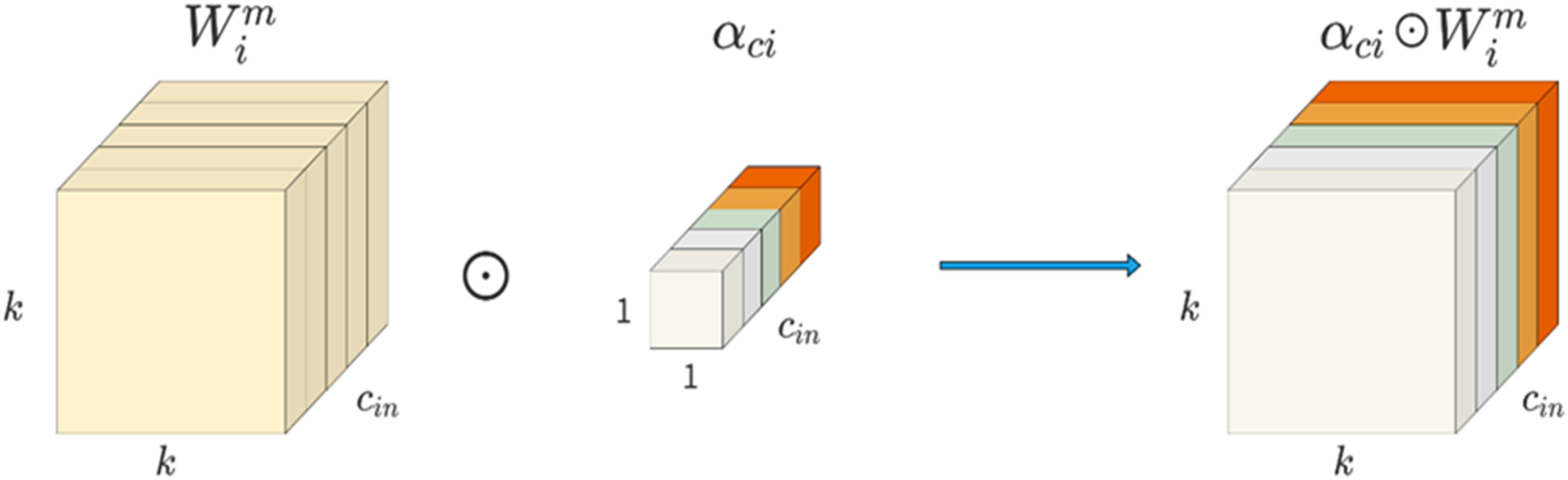
Channel multiplication operations along the input channel dimension.

Channel multiplication operation along the output channel dimension.

Kernel-by-kernel multiplication operation along the kernel dimension of the convolutional kernel space.
The feature extraction module in the U-shaped network is closely related to the final segmentation effect, and a good feature extraction module can obtain richer information about image features. The feature extraction module in the U-shaped network is mainly composed of ordinary convolution blocks, which cannot effectively adapt to dental panoramic X-ray images with complex features. To improve this problem and further enhance the feature learning ability of the network model, this article designs the ROCM, as shown in Figure 7, the ROCM has two different branches, one of which consists of a twice repeated convolution kernel of size 3 × 3 ODConv, a batch normalization layer, and the ReLU activation function. This branch is the backbone feature extraction flow in ROCM, which can obtain feature information of a larger-size region or even the global feature information of the whole image; the other branch consists of ODConv with a convolutional kernel of size 1 × 1, batch normalization layer and the ReLU activation function, this branch is the auxiliary feature complementary flow in ROCM, which focuses on learning the detailed feature information of small size, which is beneficial to further improve the segmentation effect of tooth edges. To enable ROCM to simultaneously learn feature information with variable sizes, the output features of the above two branches are then summed to obtain the final fused and complementary output features.

Network structure of the residual omni-dimensional convolution module.
Two-sream coordinate attention module
Skip connection has an important position in the U-shaped network, mainly to ensure that more low-level feature information of the image will not be lost in the feature propagation process, which is conducive to better recovery of detailed feature information in the image. However, limited by the large feature differences between the contraction path and expansion path of the same layer, directly splicing the features of the two sides will still lose part of the intermediate information, and the channel, coordinate, position, direction, and other information in the image are also confused. To solve this problem, an effective approach is to embed the modular structure used to process the image space, channel, and other information into the network, combined with the original skip connection, to make up for the limitations of the previous connection structure, and further improve the performance of the feature transfer process. Such module structures mentioned above mainly include SENet 2 and CBAM. 41 However, SENet only focuses on capturing the channel information of the image, and the spatial location information of the image is neglected by the module structure, although CBAM takes into account the processing of spatial and channel information in the image at the same time, the ability to learn the pixel relationship at a distance is lost. Aiming at the design defects of the above module structure, Hou et al. 42 proposed an advanced and lightweight network module named coordinate attention (CA), CA combines spatial information such as positional orientation with channel information in images more efficiently, further capturing pixel dependencies over long distances on the basis of multi-channel information interactions, which ultimately improves the performance of the network module.
The network structure of coordinate attention is shown in Figure 8, assuming that the size of the input feature map is 


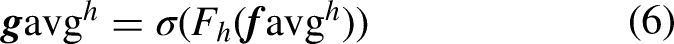
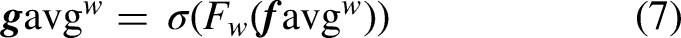

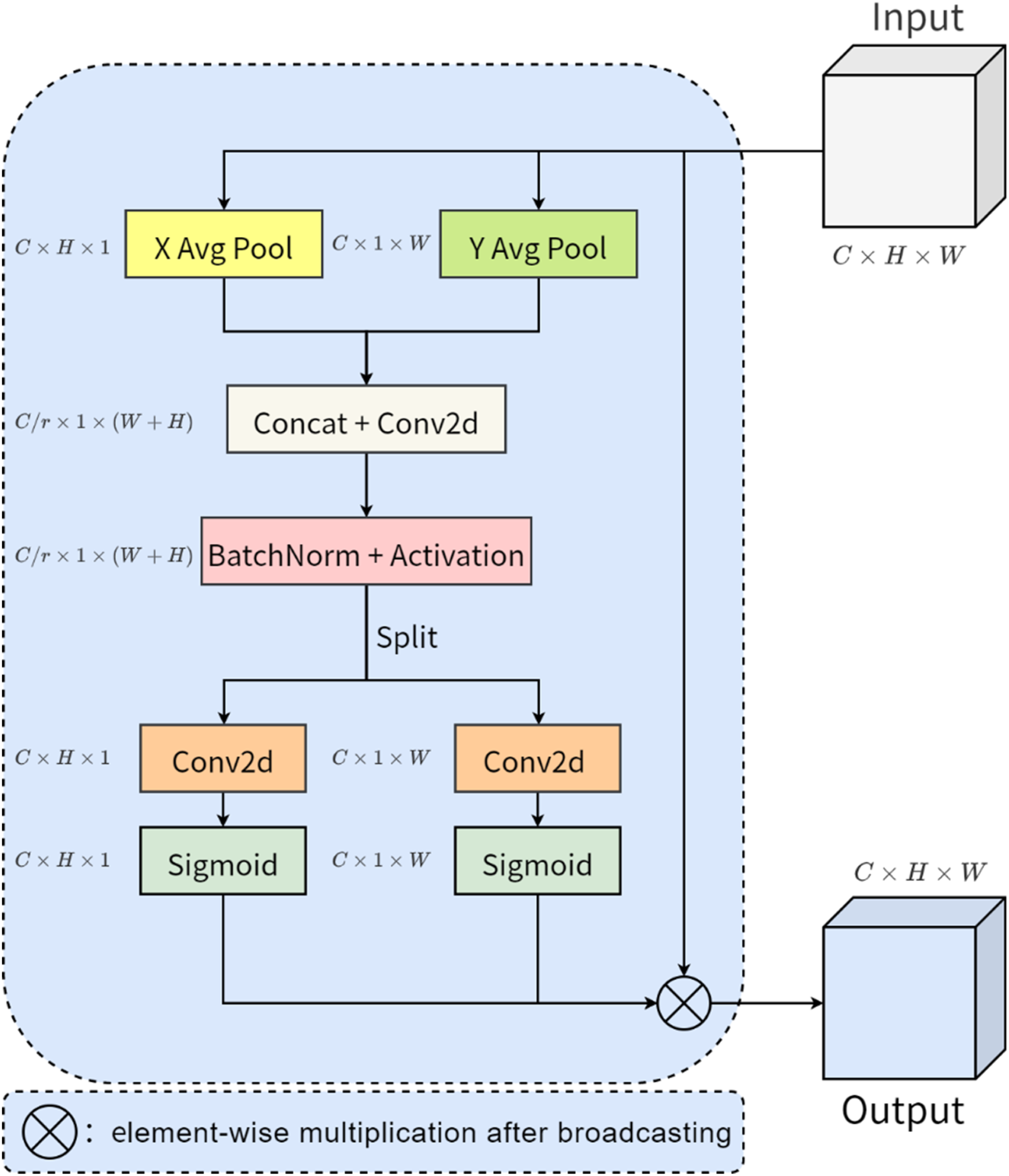
Network structure of coordinate attention.
Coordinate attention is designed with two kinds of average pooling in the horizontal and vertical directions, and after pooling, convolution and other operations successfully abstract the target of interest in the input features into the final weight score, through which the coordinate position of the target object can be more accurately restored. However, it is known that the junction between the target object and the background in the dental panoramic X-ray image is more ambiguous, which means that the average pooling in the coordinate attention only takes into account the tooth object itself, and the differences in the features at the junction as well as in the surrounding area also help to locate the tooth object and optimize the contour information of the tooth object. Therefore, in this article, we design the TSCA module, which retains the position information of the tooth object, and adds a new branch of the network consisting of the maximum pooling operation, which captures the feature information of the junction between the tooth object and the background.
The network structure of TSCA is shown in Figure 9, where the left side of the figure shows the branching structure consisting of average pooling, and the output of the left-branching structure is denoted as 

Network structure of the two-stream coordinate attention module.
Experimental analysis
Experimental setup and data preprocessing
Dataset
We conducted experiments on two datasets, namely Archive 43 and Dataset and Code, 44 to demonstrate the effectiveness of our method. The Archive is a public dataset containing 116 dental panoramic X-ray images and real segmentation labels, with a pixel size of 3104 × 1200 pixels, therefore, it does not require a specific ethical approval number. Additionally, the Dataset and Code contains 1500 dental panoramic X-ray images with real segmentation labels, featuring a pixel size of 1991 × 1127 pixels. The use of this dataset has been approved by the National Research Ethics Committee (CONEP) and the Research Ethics Committee (CEP) under report number 646.050 with an approval date of May 13, 2014. These images have significant aspect inequality, and there are two difficulties in directly using such original dental panoramic X-ray images as network inputs, one is that training very large-size images is a challenging task for devices with limited hardware resources, which is prone to video memory and RAM overflow problems; the other is that, for the input images with unequal lengths and widths, it is sometimes necessary to slice the images beforehand and then feed them into the network sequentially, but the slice will destroy some of the important pixel relationship structures in the original images. Combining the above considerations, in this article, the size of all images in the dataset is uniformly adjusted to 512 × 512, the Archive randomly divides the data of which 60 images are used as the training set and the other 56 images are used as the test set, and the Dataset and Code randomly divides the data of which 1350 images are used as the training set and the other 150 images are used as the test set, and randomly flips the input data horizontally, vertically, horizontally and then vertically during the training.
Evaluation of indicators
To evaluate the effectiveness of the proposed algorithm, four commonly used evaluation metrics, namely Intersection over Union (IoU), Precision, Accuracy, and Dice Coefficient (Dice) are used as the basis for the analysis. The mathematical definitions of these four indicators are shown below:



The hardware environment for the experiment is as follows, the CPU is AMD Ryzen 5 3600, the GPU is NVIDIA GeForce RTX 3060Ti, and the RAM size is 32 GB. The software environment is as follows, Windows 10 64-bit version, Python 3.7, Pytorch 1.7.0+ cu110, CUDA 11.5 Some settings for network training are as follows: the loss function used is BCEWithLogitsLoss, the number of training rounds is set to 200, the optimizer used to train the neural network is Adam, the batch size is set to 1, and the initial learning rate is set to 0.001. The change in the loss value on the training set is shown in Figure 10, and when iterated up to 180 times, the change in the loss curve on the training set tends to converge.
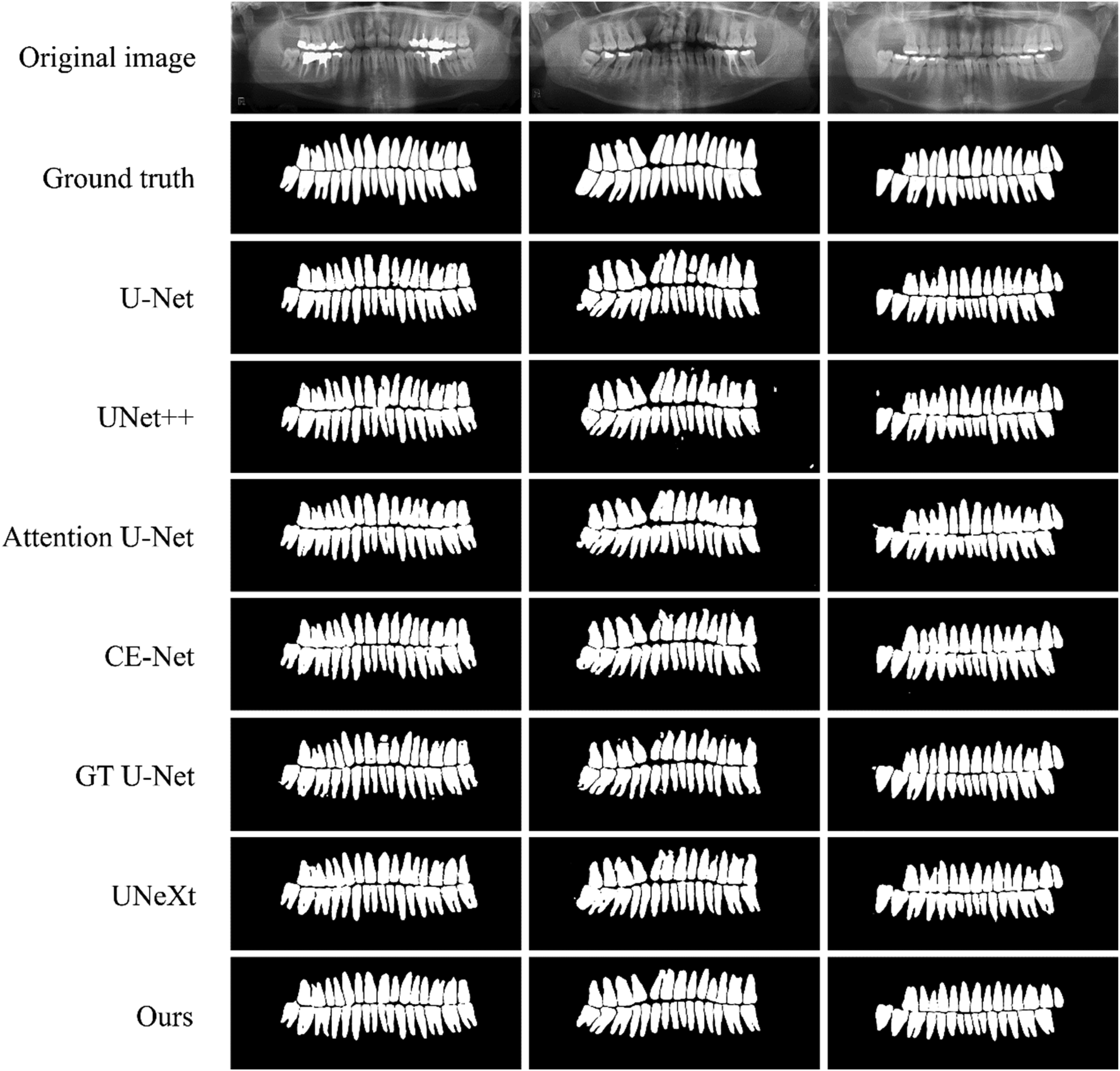
Segmentation results of different methods on Archive dataset.
Comparative analysis of related algorithms
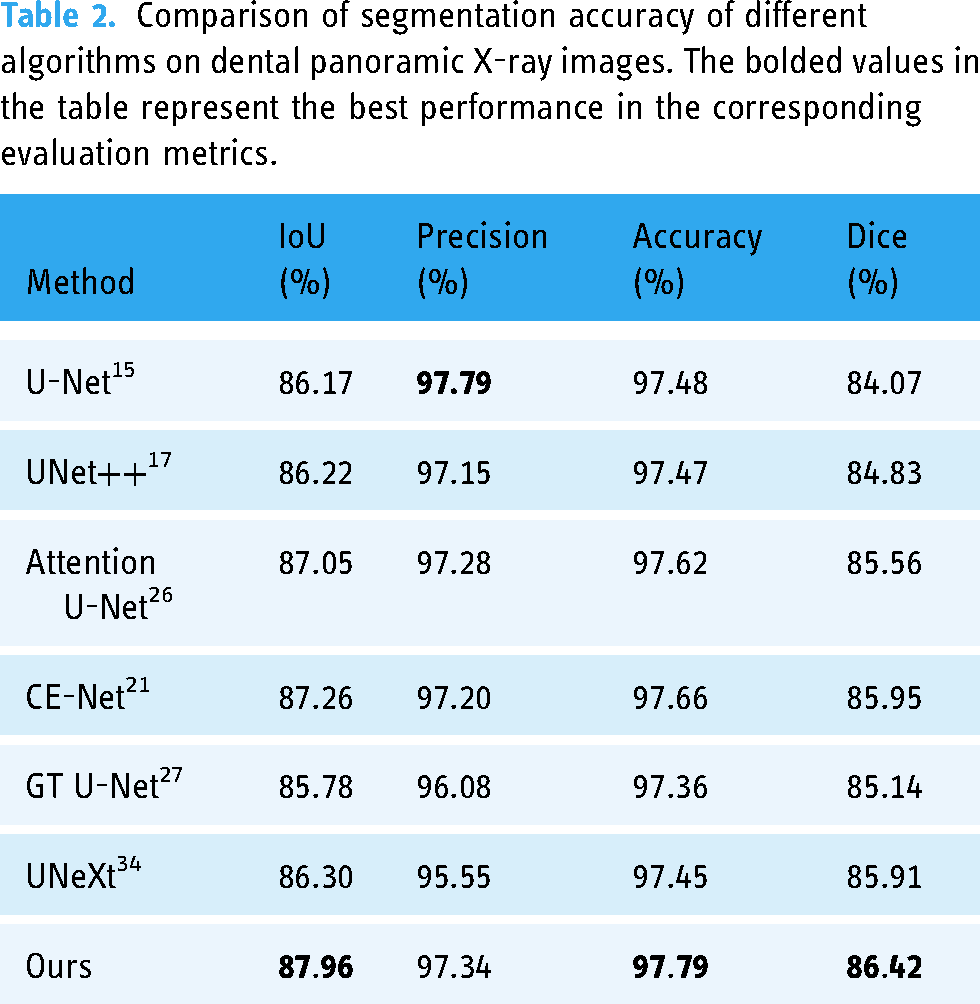
Since there are few deep learning-based tooth segmentation methods with publicly available source code, to better demonstrate the advantages and effectiveness of the proposed algorithm, this article compares U-Net, UNet++, Attention U-Net, CE-Net, GT U-Net, and UNeXt, which are representative segmentation algorithms in recent years. Table 2 shows the comparison of the segmentation accuracy of different algorithms on the Archive, compared with the experimental results of UNet and UNet++, the proposed algorithm achieves the optimum in all three evaluation metrics of IoU, Accuracy, and Dice, which are improved by 1.74%, 0.31%, and 1.59%, respectively, and also achieves a very good result in Precision evaluation metric. Compared with the experimental results of Attention U-Net, CE-Net, GT U-Net and UNeXt, the algorithm in this article has improved the four evaluation metrics of IoU, Precision, Accuracy, and Dice by 0.70%, 0.06%, 0.13%, and 0.47%, respectively. The experimental results prove that the proposed algorithm achieves better-integrated segmentation performance and can distinguish the tooth target region and background region more effectively.
Comparison of segmentation accuracy of different algorithms on dental panoramic X-ray images. The bolded values in the table represent the best performance in the corresponding evaluation metrics.
To reflect the segmentation effect of different algorithms more intuitively, the segmentation results of different algorithms are visualized on a number of images in the test set, Figure 11 is the overall segmentation result visualization of different algorithms, and Figure 12 is the detailed segmentation result visualization of different algorithms. From the figure, it can be seen that U-Net suffers from large semantic feature information differences between the same layers, and is also limited by the lack of feature extraction capability, which results in breaks when segmenting complete teeth; UNet++ has more obvious isolated segmentation error points in the visualized segmentation results, which is mainly affected by the lack of multiscale information extraction; Attention U-Net and CE-Net have obvious sawtooth phenomenon at the edge of tooth segmentation, and some over-segmentation occurs; GT U-Net has some tooth adhesion and splitting phenomenon in the visual segmentation results, accompanied by some isolated segmentation error points; UNeXt's visual segmentation results have more curved tooth morphology, and the edge of the tooth is not smooth enough, which is due to the network model losing part of the important receptive field. Compared with the other algorithms, the visual segmentation results of the proposed algorithm are close to the real labels, the tooth edges are smooth, the jaggedness is obviously reduced, and the control of over-segmentation and isolated segmentation error points is also improved.

Visualization of the results of the detail segmentation for different algorithms.

Segmentation results of different methods on Dataset and Code dataset.
In fact, the shape of teeth captured by different capture devices is different, and to meet the challenge of differences in tooth images and differences in the quality of annotation under different devices, we newly added Polyer and Swim-T, which are segmentation algorithms combined with vision transformer, to the Dataset and Code for comparison. Table 3 shows the comparison of segmentation accuracy of different algorithms on the Dataset and Code, compared with the optimal experimental results of other algorithms, the proposed algorithm achieves optimality in three evaluation metrics, IoU, Accuracy, and Dice, which are improved by 0.25%, 0.10%, and 0.73%, respectively, and also achieves very good results in the Precision evaluation metric.
Comparison of segmentation accuracy of different algorithms on dental panoramic X-ray images. The bolded values in the table represent the best performance in the corresponding evaluation metrics.

Visualizing the segmentation results of different algorithms on some images (adult teeth, implants, children's teeth, missing teeth, etc.) in the test set, Figure 13 visualizes the overall segmentation results of different algorithms, and visualizes the detailed segmentation results of different algorithms. From the figure, it can be seen that U-Net has obvious shortcomings in the tooth segmentation task, especially in the processing of complex structures, which tends to lose important information. UNet++ is prone to errors in detail processing with insufficient multi-scale information extraction, especially in the complex background or near the boundary, where the error points are more significant. The results of the Attention U-Net show jagged tooth edges and partial over-segmentation problems. The increased design complexity and number of parameters in the potential boundary extraction and boundary-sensitive refinement modules of the Polyer network leads to the problem of high computational overhead and insufficient generalization ability of the model during training and inference, which in turn affects the segmentation accuracy. Although the vision transformer introduced by Swin-T can theoretically capture global information better, the problems of insufficient processing of fine edges and omission of segmentation of part of the region due to insufficient training data still exist in practical applications. Compared with other algorithms, our proposed algorithm shows obvious advantages in the segmentation results, which can accurately maintain the boundaries and the complete structure of the tooth, with clear root boundaries and no obvious adhesion between the roots.
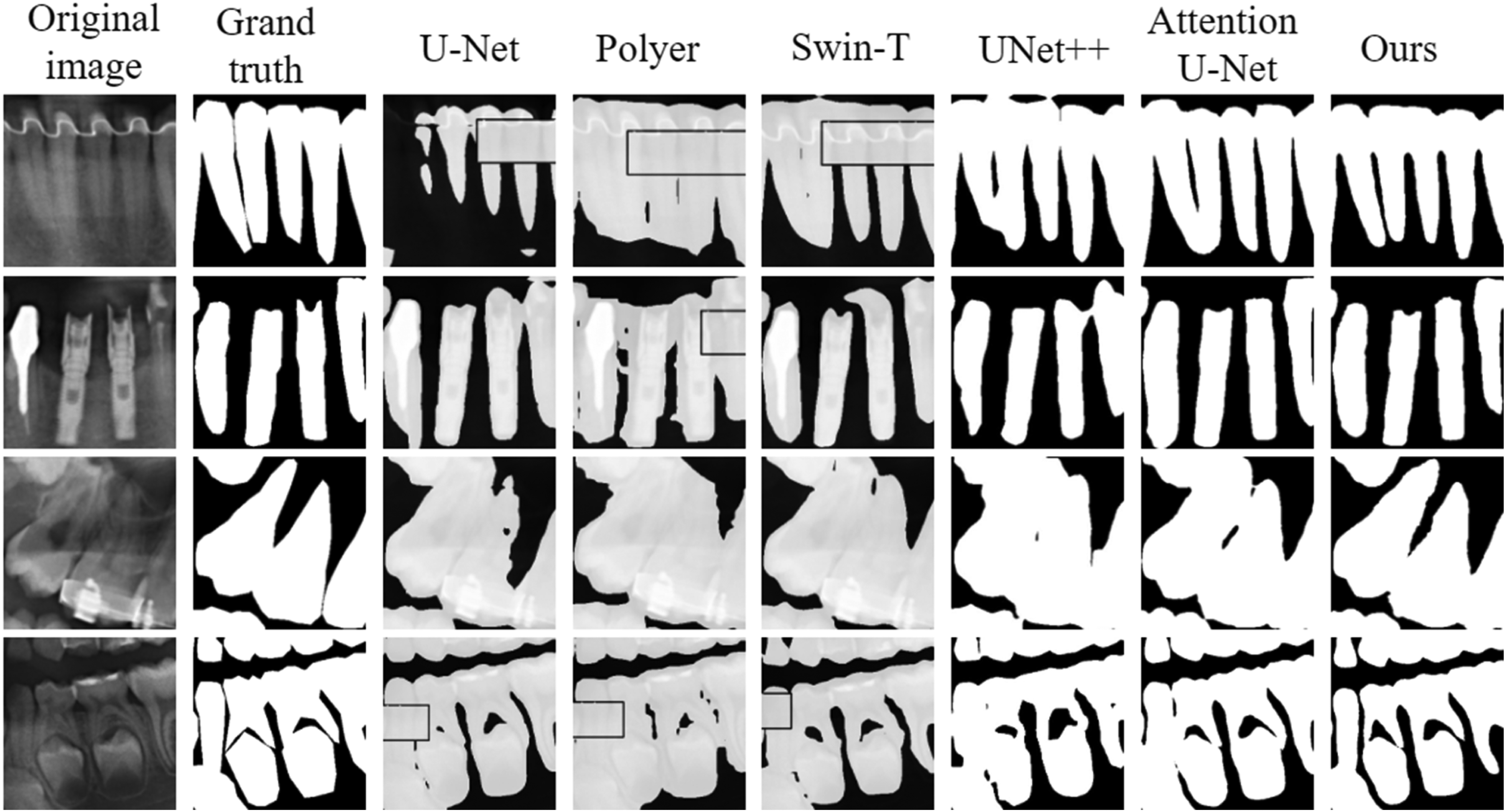

Boundary detail comparison of different methods.
Analysis of ablation experiments
To validate the effectiveness of the proposed module, this study designed the following partial ablation experiments on the Archive. This series of experiments aims to deeply explore the specific impact of each component on the overall performance by gradually removing or modifying key components of the model. Through this approach, we are able to clarify the function and importance of each module, thus confirming the design rationality and effectiveness advantages of the model.
Effectiveness of the residual omni-dimensional convolution module Effectiveness of the residual omni-dimensional convolution module.

Convolutional kernel size analysis in trunk feature extraction streams
Convolutional kernel size analysis in the backbone feature extraction stream.

Validity of the two-stream coordinate attention module
Validity of the dual-stream coordinate attention module.

Analysis of input image size
Analysis of input image size.

Analysis of different upsampling methods
The upsampling is located before the convolution operation of the feature decoding network, and the output of the upsampling together with the output of the TSCA module constitutes the new input features of each layer of the feature decoding network. To investigate the impact of different upsampling methods on the segmentation results, we designed a series of experimental comparisons in this article. Specifically, four different upsampling methods are compared, including (a) transposed convolution, (b) nearest-neighbor interpolation, (c) single linear interpolation, and (d) bilinear interpolation, and the experimental results are shown in Table 8. Since the bilinear interpolation method can interpolate non-integer pixel positions, while the transposed convolution method can only interpolate integer pixel positions, and the bilinear interpolation method can use the values of the four surrounding pixels to calculate the value of the target pixel, more accurate results than the nearest-neighbor interpolation and single linear interpolation methods are obtained. Therefore, the experiment achieved the best results when the bilinear interpolation method was chosen as the upsampling method.
Analysis of different upsampling methods.

Conclusion
For dental panoramic X-ray image data, a dental panoramic X-ray image segmentation method based on multi-feature coordinate position learning is presented, and a deep learning-based dental segmentation network model is implemented. In this article, two multi-feature coordinate position learning modules are designed to enable the neural network to extract more discriminating multi-scale feature information at different coordinate positions. Multi-feature extraction is performed layer by layer using the ROCM when acquiring larger-size regional feature information and small-size detailed features that affect the trajectory of tooth contours. To reduce the information difference between the feature encoding network and the feature decoding network, we use the dual-stream coordinate attention module to accomplish global information capture. Experimental results show that the proposed method in this article achieves excellent results in terms of accuracy and efficiency of tooth segmentation, and the segmented tooth morphology in the visualization results is close to the real annotation results. However, due to the design of the ROCM and the TSCA module in this article, the number of parameters of the network model has risen. This change increases the computational requirements and runtime of the model, which may affect the utility of the model especially in resource-constrained environments. Therefore, our future research efforts should focus on the lightweight aspect of the model to optimize model efficiency. In addition, annotation of dental data is a time-consuming and costly task, limiting the feasibility of large-scale data processing. Therefore, semi-supervised segmentation methods are particularly important, which can effectively reduce the dependence on labeled data while maintaining segmentation performance by using incompletely annotated dental image data. We plan to further investigate, for example, self-training and pseudo-labeling techniques to improve the accuracy of tooth segmentation under the semi-supervised learning framework.
Footnotes
Acknowledgments
Not applicable.
Contributorship
Conceptualization, T.M. and Z.D.; methodology, T.M. and Z.D.; software, Z.D.; validation, Z.D.; formal analysis, Z.D. and Y.Y. (Yizhou Yang); resources, Z.D. and Y.Y. (Yizhou Yang); data curation, Z.D.; writing—original draft preparation, Z.D.; writing—review and editing, J.Y.; visualization, Z.D.; supervision, T.M., J.Y.; funding acquisition, T.M., J.Y. All authors have read and agreed to the published version of the article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
Not applicable.
Funding
This work was supported by the Shaanxi Natural Science Fundamental Research Program Project (No. 2022JM-508), the Youth Innovation Team of Shaanxi Universities and in part by the National Natural Science Foundation of China (Grant No. 62101432).
Guarantor
Z.D. (Zhenrui Dang).
Informed consent statement
Not applicable.
Institutional review board statement
Not applicable.
