Abstract
Background
Breast cancer is a leading malignant tumor among women globally, with its pathological classification into benign or malignant directly influencing treatment strategies and prognosis. Traditional diagnostic methods, reliant on manual interpretation, are not only time-intensive and subjective but also susceptible to variability based on the pathologist's expertise and workload. Consequently, the development of an efficient, automated, and precise pathological detection method is crucial.
Methods
This study introduces RSDCNet, an enhanced lightweight neural network architecture designed for the automatic detection of benign and malignant breast cancer pathology. Utilizing the BreakHis dataset, which comprises 9109 microscopic images of breast tumors including various differentiation levels of benign and malignant samples, RSDCNet integrates depthwise separable convolution and SCSE modules. This integration aims to reduce model parameters while enhancing key feature extraction capabilities, thereby achieving both lightweight design and high efficiency.
Results
RSDCNet demonstrated superior performance across multiple evaluation metrics in the classification task. The model achieved an accuracy of 0.9903, a recall of 0.9897, an F1 score of 0.9888, and a precision of 0.9879, outperforming established deep learning models such as EfficientNet, RegNet, HRNet, and ViT. Notably, RSDCNet's parameter count stood at just 1,199,662, significantly lower than HRNet's 19,254,102 and ViT's 85,800,194, highlighting its enhanced resource efficiency.
Conclusion
The RSDCNet model presented in this study excels in the efficient and accurate classification of benign and malignant breast cancer pathology. Compared to traditional methods and other leading models, RSDCNet not only reduces computational resource consumption but also offers improved feature extraction and clinical interpretability. This advancement provides substantial technical support for the intelligent diagnosis of breast cancer, paving the way for more effective treatment planning and prognosis assessment.
Introduction
Breast cancer is one of the most common malignant tumors in women worldwide, with more than 2 million new confirmed cases each year, seriously threatening women's lives and health.1–3 Tumor pathology is an important basis for the diagnosis and prognosis of breast cancer, and its results directly affect the formulation of individualized treatment plans. For example, highly differentiated breast cancer usually progresses slowly and is suitable for conservative treatment, while poorly differentiated tumors often have a higher risk of invasiveness and metastasis and require more aggressive treatment strategies. 4 However, traditional pathological evaluation relies on manual microscopic analysis by pathologists, which is not only time-consuming but also easily affected by subjective factors. Efficient and standardized solutions are urgently needed.
Although pathology is the “gold standard” for breast cancer diagnosis, manual analysis has significant limitations. First, the experience level and workload of pathologists may lead to differences in diagnostic consistency5,6; Second, complex pathological features observed in microscopy (such as cell morphology and structural heterogeneity) may lead to missed or misdiagnosis. In addition, in areas with a high incidence of breast cancer, the shortage of pathologists further exacerbates the problem of diagnostic efficiency and quality. 7 These challenges underscore the need for artificial intelligence to automate pathological diagnosis, offering a promising solution to enhance the efficiency and consistency of breast cancer detection.
In recent years, deep learning technology has made significant progress in medical image analysis, especially in classification, segmentation, and detection tasks.8–10 Algorithms represented by convolutional neural networks (CNNs) have been successfully applied to the imaging diagnosis and pathological grading of breast cancer.11–13 For example, models such as ResNet, EfficientNet, and Vision Transformer (ViT) have been widely used in pathological image analysis and have demonstrated high classification accuracy. However, the high performance of these models usually comes at the cost of huge computing resources and complex architecture, making them difficult to be widely used in resource-constrained clinical environments. 14 Despite their success, existing models still face several challenges. First, the feature complexity of high-resolution pathological images requires a large number of parameters to extract effective information, which leads to high computational complexity and memory consumption. 15 Second, current models often struggle with capturing multiscale pathological features, limiting their ability to identify fine-grained patterns critical for diagnosis. Additionally, model interpretability remains a significant concern in clinical applications, as physicians and patients need clear explanations of AI-based decisions to enhance trust and adoption. These limitations highlight the necessity of developing an improved model that balances accuracy, efficiency, and interpretability. 16
This study proposed an improved lightweight neural network architecture, RSDCNet, for automatic detection of benign and malignant breast cancer pathology. Experimental results show that RSDCNet performs well in multiple evaluation indicators. In terms of classification accuracy, RSDCNet reaches 0.9903, which is significantly higher than RegNet’s 0.9789, HRNet’s 0.9629, and ViT’s 0.8841. In terms of recall rate, RSDCNet is 0.9897, significantly better than HRNet’s 0.9536 and ViT’s 0.8349. In terms of F1 score and precision, RSDCNet is 0.9888 and 0.9879, respectively, far exceeding ResNet50's 0.9310 and ViT’s 0.8917. At the same time, the model parameters of RSDCNet are only 1,199,662, which is much lower than HRNet’s 19,254,102 and ViT’s 85,800,194 . It substantially reduces computational resource consumption while maintaining high performance, demonstrating exceptional practicality and efficiency. The experimental results show that RSDCNet has significant advantages in the classification task of benign and malignant breast cancer pathology. The research workflow is shown in Figure 1.

Workflow of this study.
Method
Patient data collection
This study used the publicly available BreakHis (Breast Cancer Histopathological Database) dataset as the main data source for breast cancer pathology diagnosis tasks. The dataset contains 9109 microscopic images of breast tumor tissue from 82 patients, covering four different microscopic magnifications (40×, 100×, 200×, and 400×) to comprehensively characterize the characteristics of pathological tissues. In terms of image categories, it includes 2480 benign samples and 5429 malignant samples, covering a variety of pathological subtypes of breast adenomas and cancers, respectively. All images have a resolution of 700 × 460 pixels, a three-channel RGB format, an image depth of 8 bits, and are stored in PNG file format. The BreakHis dataset was jointly developed by the P&D Pathology Anatomy and Cytopathology Laboratory of Paraná State, Brazil, and aims to provide a standardized benchmark and evaluation tool for breast cancer pathology image classification tasks.
Data preprocessing
In order to ensure that the images can adapt to the input requirements of deep learning models and improve the generalization ability of the models, this study systematically preprocessed the pathological images in the BreakHis dataset. First, we uniformly resized the images with an original resolution of 700 × 460 pixels to 224 × 224 pixels to adapt to the input format of mainstream deep learning models, while retaining key pathological features and removing irrelevant background information. Second, in order to alleviate the overfitting problem that may be caused by limited data size, a series of data augmentation strategies were adopted, including random horizontal flipping, random rotation (±15°), and color jitter (brightness, contrast, and saturation range is ±0.2). These augmentation operations increase data diversity while enabling the model to better adapt to image variations under different conditions, thereby improving robustness to unseen data. In addition, in order to be consistent with the feature distribution of the pre-trained model, we performed pixel normalization on the images and adjusted the pixel values to a distribution with mean [0.485, 0.456, 0.406] and standard deviation [0.229, 0.224, 0.225] as parameters.
Model construction
RSDCNet architecture
The model is based on RegNet and integrates multiple modules to improve performance and efficiency. In the feature extraction stage, the initial convolution layer and the first two groups of convolution blocks of RegNet are used, where the convolution kernel size is 3 × 3, the stride is 2, and the number of channels is 64 and 128 respectively. Next, the SCSE module (Squeeze-and-Excitation channel attention and Spatial Excitation spatial attention) is introduced, where the channel compression rate is set to 16 to enhance the feature selection ability, meaning that the number of channels is first reduced to 1/16th of its original size before being restored via a gating mechanism. This approach enhances the model's ability to emphasize key features while suppressing irrelevant background information. The spatial excitation mechanism applies a sigmoid activation function to generate attention weights, further improving feature selection by reinforcing important regions in the input image. In deeper feature extraction, Depthwise Separable Convolution and ShuffleUnit modules are used. The ShuffleUnit module is implemented with four grouped convolution layers, where each group contains 256 channels. The depthwise separable convolution operates with a kernel size of 3 × 3 and a stride of 1, significantly reducing computational cost while preserving essential spatial details. Additionally, the ShuffleUnit module employs a channel shuffle operation to enhance cross-group information exchange, improving the model's feature representation capabilities. This design ensures efficient feature extraction while keeping the model lightweight. Finally, the classification task is completed through global average pooling and fully connected layers, and the number of output categories depends on the task objectives.
This model combines the design concepts of lightweight and efficient feature extraction. The SCSE module can dynamically adjust feature weights through the combination of channel and spatial attention, so as to capture key information more accurately; the reasonable setting of channel compression rate ensures the efficiency of the attention mechanism. Deep separable convolution can significantly reduce the computational complexity while maintaining high accuracy, making the model more efficient. The ShuffleUnit module further improves the feature expression ability and the lightweight degree of the model through group convolution and cross-channel information exchange. The overall design is based on the excellent characteristics of RegNet and combines a variety of optimization modules to achieve efficient, accurate, and flexible performance, which is very suitable for application in scenarios with limited resources and fast reasoning.
Comparison model
While completing the breast cancer pathology classification task, we tested a variety of deep learning models, including EfficientNet, RegNet, HRNet, ResNet50, and ViT. EfficientNet was proposed by Google. By optimizing the combination of network depth, width, and resolution, it strikes a balance between performance and efficiency, making it an efficient choice for many tasks. 17 RegNet is a regularized network designed by the Facebook AI Research team. It simplifies the network design process and achieves high performance with lower computing resources on datasets such as ImageNet. 18 HRNet performs well in classification, detection, and segmentation tasks by retaining and fusing high-resolution features and realizing multi-resolution information interaction. It is an excellent solution for high-resolution vision tasks. 19 ResNet50 is a deep residual network proposed by Microsoft. It solves the gradient vanishing problem through residual connections, enabling deeper networks to effectively extract complex features and perform well in a variety of vision tasks. 20 ViT was proposed by Google based on the Transformer architecture. It breaks the limitations of traditional CNN, directly processes the unstructured features of images, and demonstrates powerful performance in classification tasks. 21
Model evaluation
This study uses a variety of evaluation indicators, including accuracy, recall, precision, F1-score, and area under the receiver operating characteristic curve (AUC) to comprehensively evaluate the performance of the model in the task of detecting benign and malignant breast cancer pathology. These indicators can measure the performance of the model in the classification task from different dimensions to ensure the comprehensiveness and reliability of the evaluation results.
Accuracy is the ratio of the number of samples predicted correctly by the model to the total number of samples, reflecting the overall classification performance of the model; recall measures the model's ability to capture positive samples and is a reflection of sensitivity; precision indicates the proportion of all samples predicted to be positive that are actually positive, which can evaluate the credibility of the prediction results; the F1 value combines the recall rate and precision, and is suitable for performance measurement of unbalanced data sets; AUC plots the area under the receiver operating characteristic curve (ROC curve) to show the performance of the model at different decision thresholds, providing a global assessment of the model's classification ability.
In order to intuitively display the classification effect of the model, we used a confusion matrix to analyze the classification distribution of the model on positive and negative samples and combined with the Grad-CAM (Gradient-weighted Class Activation Mapping) visualization method to generate a heat map to reveal the key areas of the model's attention. This technology helps us evaluate the interpretability of the model and ensure that the model's decisions are based on the key features of the pathological image rather than background noise. The model evaluation process strictly follows the standardized training and testing division method (the ratio of training set to test set is 7:3) and uses an unseen test set for independent verification to avoid evaluation bias caused by overfitting.
Experimental setup
The model training in this study was completed on a high-performance computing device equipped with an NVIDIA RTX 3090 GPU (24GB video memory) and an Intel Core i7 13700 K CPU to fully utilize the GPU acceleration capabilities to improve computing efficiency. During the training process, the model optimization used stochastic gradient descent (SGD) and Adam optimization algorithms to achieve efficient and stable parameter updates. The initial learning rate was set to 0.0001, the momentum coefficient was configured to 0.999, and the smoothness of the optimization path was ensured by dynamically adjusting the strategy.
The training was conducted for 40 epochs, and at the end of each epoch, the current learning rate was reduced to 10% of the original value to achieve progressive optimization, thereby avoiding fluctuations caused by excessive learning rates. The number of training samples in each batch (batch size) is 32. In order to enhance the generalization ability of the model, L2 regularization (weight decay coefficient is 0.0005) and Dropout technology (dropout rate is 0.5) were introduced to alleviate the overfitting problem from multiple aspects. In addition, Mixup data enhancement technology is used to expand the diversity of training samples. Specifically, in each training iteration, two images are randomly selected and linearly combined according to the Beta distribution (α=0.4), and the target labels are weighted mixed in the same proportion. This method effectively integrates multiple features and reduces the error rate of boundary sample classification.
The loss function uses the cross-entropy loss function (CrossEntropyLoss) to measure the difference between the predicted distribution and the true distribution. When the validation set loss has not improved for five consecutive epochs, ReduceLROnPlateau is used to dynamically adjust the learning rate and multiply it by 0.1 to jump out of the local optimal solution. In the gradient calculation, the gradient clipping threshold is set to 1.0 to limit the gradient amplitude, thereby preventing the gradient explosion problem during training and further ensuring the stability of model optimization. These optimization strategies jointly build an efficient and stable training process, laying the foundation for the full performance of the model.
Results
Comparison and analysis of model results
In the task of classifying benign and malignant breast cancer pathology, the performance of the model depends not only on the classification accuracy but also on the multi-dimensional indicators such as recall rate, precision rate, F1 value, and model complexity to comprehensively evaluate its practical application value. The RSDCNet model in this study combines a lightweight design with an efficient feature extraction mechanism and performs well in all evaluation indicators while showing strong practicality. Table 1 shows the performance comparison of each model, Figure 2 shows the training process and test results of RSDCNet, and Figure 3 shows the results of each comparison model.

Results of RSDCNet, where a is the loss during the training iteration, b is the change in accuracy of the validation set during the iteration, c is the ROC curve in prediction, and d is the confusion matrix.
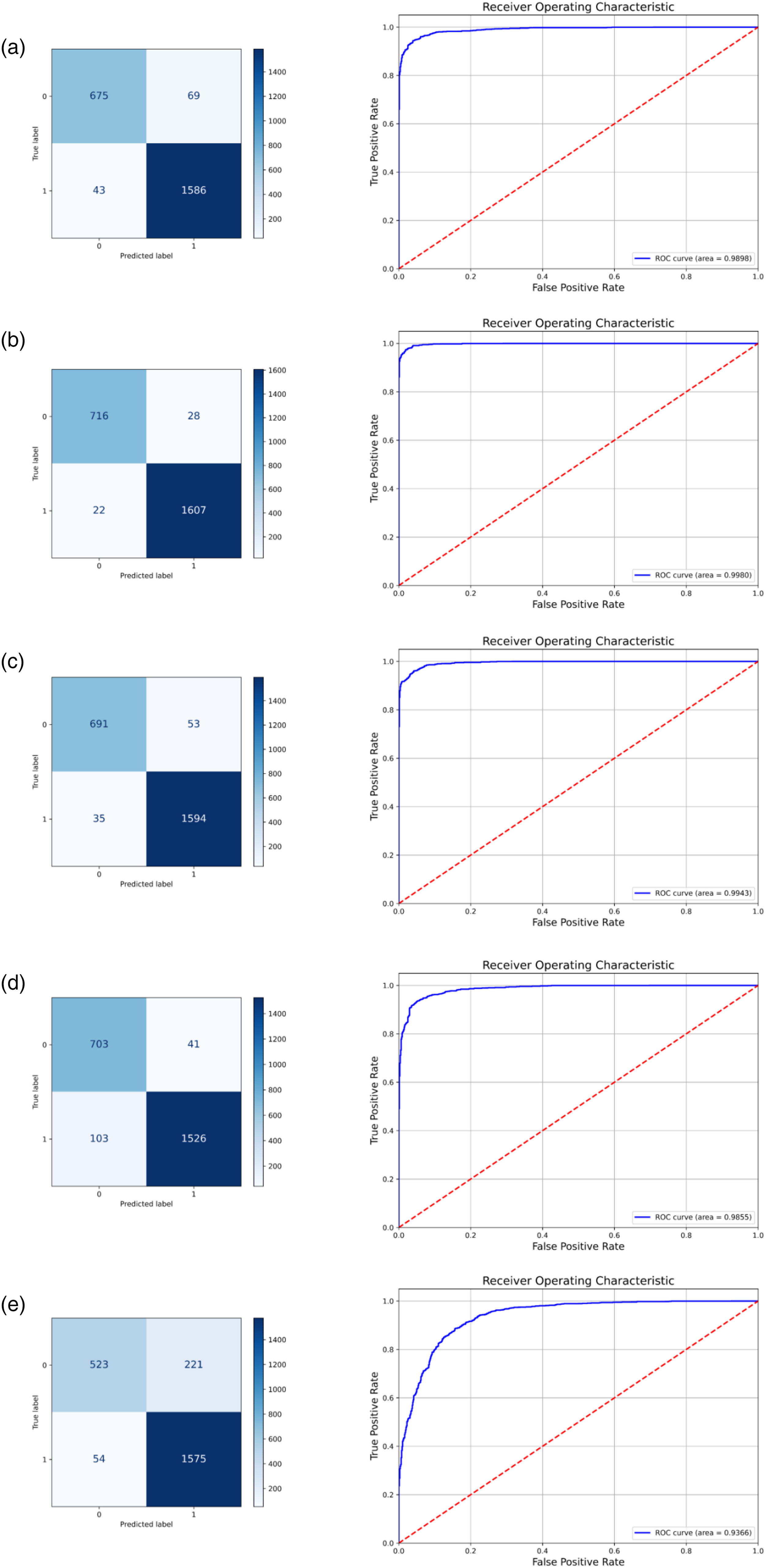
Confusion matrix and ROC curve of each comparison model, where a–e represents the results of EfficientNet, RegNet, HRNet, ResNet50, and ViT, respectively.
Comparison of test results of various deep learning models.

RSDCNet reached 0.9903, indicating that it can reliably distinguish the categories of pathological images in the breast cancer detection task. This high accuracy is due to the synergy of the model's deep separable convolution and SCSE module: the deep separable convolution effectively reduces the computational complexity while retaining key features, while the SCSE module further enhances the model's ability to capture key areas of breast cancer pathology through spatial and channel attention mechanisms. The recall rate is an important indicator of the model's ability to detect positive samples and directly reflects its sensitivity. The recall rate of RSDCNet is 0.9897, which is significantly higher than that of other comparison models. This result shows that RSDCNet can more comprehensively identify positive samples when processing breast cancer pathology images of different differentiation levels, especially in cases with blurred boundaries or unclear features. In contrast, other models such as ViT (0.8349) may not be able to fully exert their advantages in this task due to their large parameter scale and limited ability to capture local features. The F1 value is a comprehensive indicator of recall rate and precision rate, which can better reflect the balanced performance of the model in practical applications. The F1 value of RSDCNet is 0.9888, which means that the model ensures high accuracy of positive class prediction while maintaining high sensitivity. This performance not only relies on data enhancement strategies (such as Mixup) to provide diverse samples during the training phase but also benefits from the dynamic adjustment strategy of the learning rate to make the optimization path more stable, thereby avoiding the problem of overfitting. In terms of accuracy, RSDCNet reached 0.9879, indicating that its reliability in predicting positive samples is extremely high. This is particularly important in pathological diagnosis, as false alarms (false positives) may lead to unnecessary therapeutic interventions. In contrast, although other models such as EfficientNet and HRNet perform well in terms of accuracy, the increase in the number of parameters and the complexity of the architecture limit their application in resource-constrained environments.
It is worth mentioning that the number of parameters of RSDCNet is only 1,199,662, which is much lower than all the comparison models. This lightweight design fully considers the computing resource limitations in actual medical scenarios, making the model more suitable for deployment and application in embedded devices or medical institutions with limited resources. At the same time, the lower number of parameters not only reduces the storage and computing burden of the model but also speeds up the reasoning speed, making real-time diagnosis possible. From an overall analysis, the outstanding performance of RSDCNet is not only limited to numerical advantages but also reflected in the deep fit between its architectural design and task requirements. By balancing performance and efficiency, RSDCNet significantly reduces complexity while meeting high accuracy requirements.
To evaluate the practical efficiency of RSDCNet, we measured its inference time on our platform. RSDCNet achieves an average inference time of 3.2 ms per image, which is 2.5× faster than EfficientNet (8.1 ms) and 3.8× faster than ResNet50 (12.3 ms). These results highlight RSDCNet's advantage in real-time processing, making it highly suitable for integration into AI-assisted diagnostic workflows.
Clinical interpretation analysis
In the task of detecting benign and malignant breast cancer pathology, the clinical interpretability of the model is crucial to verifying its reliability and promoting clinical application. To this end, we combined the Grad-CAM (Gradient-weighted Class Activation Mapping) technology to visualize the decision-making process of the model. As shown in Figure 4, the original images and heat maps of benign (a) and malignant (b) breast tumor samples are shown respectively. In the benign sample (a), the heat map shows that the model mainly focuses on key areas such as the morphological structure of the acinus, the size of the cell nucleus, and the uniformity of the arrangement. This is consistent with the experience of judging benign breast lesions in pathology. For example, the acinus usually retains complete structural boundaries, the cell nucleus is relatively uniform, and the stromal fibrous tissue proliferates significantly but without obvious abnormalities. The model's focus on these features reflects its ability to effectively capture important information in pathological diagnosis and accurately distinguish non-cancerous areas, showing a high degree of interpretability.

Results of the model activation heat map, where a is a benign example and b is a malignant example.
In the malignant sample (b), the heat map highlights the areas where cancer cells are concentrated and the parts with high nuclear heterogeneity. These areas are often accompanied by features such as increased nuclear-to-cytoplasmic ratio, disordered cell arrangement, and increased nuclear division images, which are important pathological bases for judging malignant tumors. At the same time, the model also captures the characteristics of some tumor infiltration areas, which shows that it can identify key pathological features in complex backgrounds and further improve the accuracy of classification. This explanatory analysis enhances the credibility of the model and verifies its applicability in the context of pathology. From the visualization results, it can be seen that the areas of interest of RSDCNet are relatively consistent with the diagnosis of pathologists, which means that the decision-making process of the model is based on real pathological features rather than irrelevant background information. Specifically, the Grad-CAM heatmaps consistently highlighted key pathological features, such as increased nuclear pleomorphism, abnormal mitotic activity, and tissue disorganization in malignant cases, while benign cases were characterized by preserved acinar structures and uniform nuclear morphology.
Discussion
This article proposes a deep learning model RSDCNet for breast cancer differentiation tasks and comprehensively evaluates its performance from multiple dimensions. Experimental results show that the proposed model outperforms mainstream models in terms of accuracy, recall, F1 value, and precision, and also performs well in the ability to locate key feature areas. Through Grad-CAM visualization analysis, the model can accurately focus on the main feature areas of breast cancer, and its heat map results are consistent with the actual lesions. Compared with traditional methods, the proposed model not only achieves higher classification performance but also shows advantages in interpretability and clinical application potential.
Recent advances in deep learning have significantly improved the analysis of histopathological images. Various strategies, such as self-supervised learning, few-shot learning, and reinforcement learning, have been proposed to enhance model robustness and generalizability. Quan et al. 22 introduced Global Contrast-Masked Autoencoders, demonstrating their effectiveness in pathological representation learning. Wang et al. 23 proposed a pyramid-based self-supervised learning approach for histopathological classification, improving feature extraction across different scales. Additionally, Xie et al. 24 developed SD-MIL, a multiple-instance learning framework that effectively integrates scale and distance perception for whole-slide image classification. Few-shot learning techniques have also gained attention in histopathology. Quan et al. 25 proposed a dual-channel prototype network, enabling accurate pathology classification with limited labeled data. In the domain of reinforcement learning, Zheng et al. 26 introduced a deep reinforcement learning framework for detecting melanoma in whole-slide images, enhancing model decision-making through self-optimization. Furthermore, CNN-based models have been refined with improved feature extraction techniques, as seen in recent studies on deep learning for medical imaging.27–29 Compared to these approaches, RSDCNet emphasizes lightweight design and computational efficiency, making it more suitable for deployment in real-world clinical settings. While advanced learning paradigms such as self-supervised learning and reinforcement learning enhance model adaptability, they often require extensive training resources and large-scale datasets. Our model balances performance, efficiency, and interpretability, offering a practical alternative for AI-assisted pathology diagnosis. Future work may incorporate self-supervised pretraining techniques to further enhance RSDCNet's adaptability to diverse histopathological datasets.
Our model performs well in breast cancer recognition tasks while achieving lightweight design. This advantage first stems from the optimization of the model architecture. We adopt a modular design, combined with multi-scale feature extraction and attention mechanism, to achieve an effective balance between global and local feature modeling. By streamlining the distribution and parameter configuration of the convolutional layer, the model has 1,199,662 parameters, which is significantly lower than mainstream models such as EfficientNet and ResNet. In addition, combined with the learning rate adjustment strategy and data enhancement technology, we further improved the training efficiency and generalization ability of the model, providing strong support for the combination of lightweight and high performance.
Given the complexity of distinguishing benign from malignant breast cancer, effective local feature extraction is crucial. Our model leverages an attention mechanism to precisely highlight key regions while integrating multi-scale feature extraction, enabling it to capture lesion morphology and fine details, even in challenging conditions with complex backgrounds and indistinct boundaries. In contrast, other models such as EfficientNet and ViT may perform slightly worse in certain situations due to insufficient local feature capture capabilities or large parameter sizes.
Early diagnosis and correct classification (benign and malignant) of breast cancer are crucial for developing treatment strategies and improving patient prognosis. However, traditional pathological detection methods rely on microscopic observation by pathologists, which is not only time-consuming but also susceptible to subjective factors, which may lead to misdiagnosis or missed diagnosis, especially in the complex pathological environment of malignant tumors. The RSDCNet proposed in this study achieves automatic classification of benign and malignant lesions through deep learning, with an accuracy rate of up to 99.03%, significantly higher than the existing mainstream models. At the same time, the lightweight design of the model (only 1,199,662 parameters) greatly reduces the hardware requirements, making it suitable for deployment in resource-constrained primary medical institutions or embedded devices. Through Grad-CAM heat map visualization, the model can accurately focus on the key areas of the lesion, and its focus is consistent with the pathological diagnosis criteria, further enhancing the credibility of clinical applications. This efficient and reliable automated pathology detection system can not only assist pathologists to improve diagnostic efficiency and alleviate the shortage of pathological resources but also reduce the risk of missed diagnosis and misdiagnosis, providing patients with more accurate diagnostic services. The integration of RSDCNet into existing pathology workflows has the potential to significantly enhance diagnostic efficiency. By incorporating the model into digital pathology platforms, RSDCNet can serve as a pre-screening tool to highlight suspicious areas, reducing the workload of pathologists and enabling more focused manual review. Furthermore, the model's lightweight nature ensures that it can be deployed on edge devices or cloud-based diagnostic platforms, making AI-assisted pathology more accessible, particularly in resource-limited settings where experienced pathologists may not be readily available.
BreakHis dataset used in this study is a public standardized pathology dataset, its sample size is relatively limited (9109 images) and the source is single, covering only breast tumor pathology slide images from 82 patients. This may lead to a certain lack of generalization ability of the model when processing larger-scale, cross-institutional, or ethnic pathology data. Despite its widespread use, the BreakHis dataset has inherent limitations, including a relatively small sample size, a single institutional source, and an imbalanced distribution of benign and malignant cases. These factors may impact the model's generalizability to real-world clinical settings. To address this, future studies should validate RSDCNet on larger, multi-institutional datasets such as TCGA or Camelyon, which provide more diverse histopathological images. Expanding the dataset scope will ensure the model's robustness across different populations and imaging conditions, ultimately enhancing its clinical applicability. In addition, although there is a certain imbalance in the ratio of benign and malignant samples in the dataset (2480 benign vs. 5429 malignant), the number of benign cases in actual clinical scenarios may be higher, and this distribution difference may affect the performance of the model in real-world applications. Secondly, this study was trained and evaluated only based on image-level labels, without using more sophisticated pixel-level or region-level annotations. This may limit the model's ability to expand in tumor boundary recognition and subtype grading tasks. At the same time, although Grad-CAM provides a certain degree of model interpretability, it can only roughly reflect the model's focus area and cannot deeply reveal the model's weight or mechanism for each feature in a specific decision-making process, which may affect doctors’ complete trust in the model results in clinical practice. Finally, although the design of RSDCNet focuses on lightweight, it may still face the need to optimize inference speed and memory consumption in environments with extremely limited hardware resources (such as low-end embedded devices). While RSDCNet achieves high accuracy on the BreakHis dataset, its generalization to real-world clinical applications requires further validation. Histopathological images vary significantly across different medical institutions due to differences in staining protocols, slide preparation techniques, and imaging equipment, which can affect model performance. To ensure robustness, future work will explore domain adaptation techniques to mitigate dataset shifts and conduct cross-institutional validation on diverse pathology datasets. These steps will help assess the model's adaptability and reliability in varied clinical environments, ultimately enhancing its practical applicability.
Conclusion
This study presents RSDCNet, a lightweight deep-learning model for benign and malignant breast cancer pathology detection. By integrating deep separable convolution and the SCSE module, RSDCNet achieves 99.03% accuracy on the BreakHis dataset while significantly reducing parameter size, outperforming mainstream models in recall, precision, and F1-score. Grad-CAM analysis confirms strong alignment with pathological diagnostic criteria, ensuring high clinical interpretability. Its lightweight design makes it ideal for resource-limited settings, offering an efficient and reliable diagnostic tool. Future work will enhance interpretability with explainable AI, explore multi-modal data integration (histopathology, genomics, proteomics, radiomics), and deploy the model on embedded AI systems for real-time applications. Large-scale validation across diverse populations will assess robustness and generalizability and support regulatory approvals for clinical deployment.
Footnotes
Acknowledgments
The authors thank all the participating patients and doctors.
Guarantor
HL.
Ethical considerations
Ethical approval and patient consent are not required for scoping review because the data are collected from publicly available publications.
Author contributions/CRediT
YL: conceptualization, writing—original draft. HL: data curation, writing—original draft. ZZ: formal analysis, supervision, writing—review editing. CC: writing—review editing. XZ: investigation, writing - review editing. GJ: methodology, writing—review editing. HL: project administration, writing—review editing.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
