Abstract
Objective
Early detection of Parkinson's disease (PD) is essential for halting its progression, yet challenges remain in leveraging deep learning for accurate identification. This study aims to overcome these obstacles by introducing a hybrid deep learning approach that enhances PD detection through a combination of autoencoder (AE) and radial basis function neural network (RBFNN).
Methods
The proposed method analyzes the power spectral density (PSD) of preprocessed electroencephalography (EEG) signals, with artifacts removed, to assess energy distribution across EEG sub-bands. AEs are employed to extract features from reconstructed signals, which are subsequently classified by an RBFNN. The approach is validated on UC SanDiego's EEG dataset, consisting of 31 subjects and 93 minutes of recordings.
Results
The hybrid model demonstrates promising performance, achieving a classification accuracy of 99%. The improved accuracy is attributed to advanced feature selection techniques, robust data preprocessing, and the integration of AEs with RBFNN, setting a new benchmark in PD detection frameworks.
Conclusion
This study highlights the efficacy of the hybrid deep learning framework in detecting PD, particularly emphasizing the importance of using multiple EEG channels and advanced preprocessing techniques. The results underscore the potential of this approach for practical clinical applications, offering a reliable solution for early and accurate PD detection.
Introduction
For centuries, human brains, integral to the nervous system, have been extensively studied. Neurological conditions like Alzheimer's, meningitis, stroke, dementia, and Parkinson's disease (PD) have seen a significant rise in incidence over the past quarter century, driving much of this research. 1 PD is a degenerative neurological condition that destroys dopaminergic neurons in the brain's substantia nigra, 2 rising in prevalence each year, now impacting over 1% of those over the age of 50.3,4 Common symptoms in individuals with PD include tremors, muscle rigidity, postural instability, and slow movement (bradykinesia). Voice problems (dysphonia) 5 due to vocal cord deterioration can also occur. Both motor and non-motor symptoms play a role in the diagnosis, and advancements in neurology, such as electroencephalography (EEG) and modern technologies, now contribute to PD diagnosis.
An EEG is a recording of voltage variations in the brain caused by ion currents in individual neurons. The bioelectrical brain activity captured in this EEG recording is rich with important physiological and disease-related data. Changes in the frequency and rhythm of brain activity during seizures have made the EEG signal the gold standard for diagnosing brain diseases.6,7 Diagnosis based on EEG is simple to implement, non-invasive, and offers excellent temporal resolution. PD illness can influence EEG frequency bands. To distinguish PD patients from healthy ones, analyze alpha, beta, and theta frequency bands' power or coherence. Only intrusive electrodes can sample the brain pathways directly related to PD, restricting the availability of human investigations. Nonetheless, there are established animal models of PD in which disease signs may be induced by giving neurotoxins. 8 To better understand the neurophysiological correlates of PD and how they may help reveal disease processes and other non-trivial aspects in brain data, researchers are turning to machine learning methods. This might help with early diagnostics and new treatments on the road.
Machine learning and deep learning have shown great potential in identifying and diagnosing brain disorders such as PD, with various benefits over old human techniques. 9 It can detect early indications of PD and enable proactive management by evaluating a range of biomarkers, such as patterns of movement. Deep learning algorithms are among the many sophisticated models whose decision-making process is mysterious. Although approaches are becoming more accurate, they are still unable to distinguish between early-stage disease and healthy controls (HCs), or between PD and other neurological diseases. 10
There are still a number of gaps in the literature that need to be filled despite the substantial study on Perkinson's disease that has been done utilizing machine learning techniques. For instance, the performance of machine learning models can be adversely affected by the existence of irrelevant features in the domain of features, which can result in overfitting and poor accuracy. 11 The model's performance may be impacted by adding extraneous details and noise in the form of irrelevant features. Feature selection techniques can assist in removing unwanted features and enhance model performance. These drawbacks call for adopting alternate validation strategies and emphasize the necessity for additional research into the problem of improper validation techniques used in earlier studies. To address the aforementioned concerns, we present in this paper, a novel hybrid deep learning model combining autoencoder (AE) and radial basis function neural network (RBFNN), as well as the use of power spectral density (PSD) for preprocessing raw EEG signals to aid in the identification of PD. The amplitude of a signal in the frequency domain is often described using a statistic called the PSD. 12 PSD removes noise-dominated frequency bands from EEG data, increasing signal-to-noise ratio and offering a more accurate depiction of brain activity. Specific brain processes, such as relaxed wakefulness and motor activity, are related to different frequency ranges. The ability to quantify the power within these bands aids in characterizing PD-related variations in brain activity, which can be employed as biomarkers for PD diagnosis monitoring. 13 Time-domain characteristics may overlook information regarding the signal's frequency content, but PSD records it. It is less susceptible to artifacts and noise that could skew time-domain measurements. Although they need more computing power, time-frequency techniques like the continuous wavelet transform and short-time Fourier transform offer superior time-frequency resolution. PSD is frequently adequate when temporal localization is not essential. Since PSD does not require the additional complexity of temporal localization, it provides the power distribution across frequencies immediately, making it easier to understand. It is a recommended option for real-time diagnostic applications and research investigations in the field of EEG-based neurological examinations since it provides a well-balanced approach that is both effective and efficient. This contributes to a better knowledge of the spectrum properties of PD-related brain activity.
AE research for PD identification is a new area of study that brings together neurology and cutting-edge Machine Learning (ML) methods. AEs are a sort of artificial neural network that has found use in several fields, including biomedical signal processing, for the unsupervised learning of efficient codlings.14,15 Its capacity to run unattended and extract pertinent characteristics makes it a viable tool for feature extraction in PD. 16 It examines the EEG data and extracts hidden, significant aspects that successfully capture the underlying illness signs. This decreases data complexity while keeping critical information, making subsequent classification algorithms more effective at identifying PD patients. AE can disclose which features contribute the most to data reconstruction by evaluating the activations of the hidden layers. This data will be utilized to determine the most important aspects of PD identification.
RBFNs' straightforward topological structure, locally tuned neurons, and fast learning process have made them a favorite for deep learning applications. Connecting fields as diverse as function approximation, regularization, noisy interpolation, pattern recognition, and medicine, serve as a unifying link between these fields. The RBF architecture is a powerful alternate to the multilayer perceptron (MLP) due to its emphasis on a multidimensional space. 17 Classifying data with complicated, non-linear separations, such as those found in EEG signals, is a well-known strength of RBF networks. With such sensitivity, early signs of PD in the brain can be identified. When compared to other types of deep learning networks, it may be easier to train because it needs less training samples to get reliable results. This is especially helpful in healthcare settings where data may be limited or hard to collect. 18
The purpose of this study is to resolve these issues by presenting simple feature extraction and classification methods that maintain high classification accuracy and validating them using a UC SanDiego dataset. Whereas typical methods for PD diagnosis using EEG rely on hand-crafted features and linear classifiers, this method proposes a data-driven strategy that employs PSD, AE, and RBF networks. The literature analysis points out a number of gaps in the current body of research on the use of deep learning techniques with EEG signals to detect PD. Prior research has been hindered by limitations in model architectural design and has had difficulty processing time series brain data. Even though automatic PD identification and EEG data processing have been extensively studied, there is still a significant gap in the use of deep neural networks for PD detection. These drawbacks highlight the need for more reliable and effective methods that can distinguish PD patients from one another and examine the features of PD based on brain activity when the patient is not taking medication. The main contribution of the present work lies in addressing these gaps by proposing a novel hybrid deep-learning architecture for PD detection. By combining an AE and RBFNN, our approach overcomes the limitations of existing techniques and offers a more reliable and timely method for PD detection. Utilizing PSD analysis to derive energy distribution from preprocessed EEG signals, our model extracts features using AE and performs classification using RBFNN.
The study's contributions can be summed up as follows:
The PSD improves the quality of extracted features. By applying this technique, a better feature set for classification is produced, which successfully removes unnecessary and redundant features. The uncontrolled EEG activity of PD patients was used for a multi-pattern study of brain activation and functional connectivity. PD was identified very well by combining AE and RBFNN. An effective and hybrid deep learning-based PD detection model was proposed to identify PD and HC subjects with high performance in the UC SanDiego dataset, where the accuracy, sensitivity, specificity, and AUC (area under the curve) were all >99%.
The remaining content can be broken down into the following sections. In the second section, the relevant literature is reviewed, and the current state of the art in seizure detection is examined from a comparative perspective. The third section demonstrates data processing, the AE, the RBFNN architecture, and the system model presented in the paper. The fourth section explores the prototypical data set. Analysis of results and experiments, experiments on individual patients, cross-validation ablations, and a comparison to other state-of-the-art methods are all described in the fifth section. The final section of the paper is the sixth section.
Related work
In recent years, EEG data's utilization for diagnosing and monitoring PD has grown due to its non-invasive insight into brain activity. Current studies highlight the promising integration of machine learning and deep learning techniques with EEG.
Vanegas et al. 19 proposed three machine-learning frameworks for EEG-based PD biomarker localization. Their first model, analyzing spectral amplitudes in the posterior occipital area during visual stimulation, achieved a remarkable 99.4% AUC in discriminating 29 PD cases and 30 HCs. The logistic regression and decision tree models showed AUCs of 94.9 and 86.2, respectively. Identified key frequency bins in theta, alpha, and beta ranges played a crucial role in distinguishing PD subjects from HCs. Doumari et al. 20 presented a machine learning framework for the early and high-accuracy diagnosis of PD using EEG signal features. This study is motivated by the need for earlier and more objective diagnostic methods compared to traditional clinical examinations, which are subjective and often detect PD only in its later stages. The crow search algorithm and decision tree (CSADT), a novel model for early PD diagnosis, is presented in this work. The suggested model analyzes the critical aspects of each of the four important PD datasets—meander, spiral, voice, and speech-Sakar. Oh et al. 21 built an automated system to detection system that has a 13-layer CNN (convolutional neural network) to categorize participants into PD and HCs using resting-state EEG data collected from 20 PD patients and 20 HCs. This methodology can eliminate the requirement for traditional feature representation steps. The generated model performed well, with 88.25% accuracy, 84.71% sensitivity, and 91.77% specificity. The findings indicated that the classification accuracy achieved through unsupervised feature learning in AE-Convolutional Deep Neural Network (AE-CDNN) was the highest and comparable to those of other research studies. Loh et al. 22 were able to reliably divide patients with or without drugs into HC and PD groups 99.5% of the time using a 2D CNN on the Gabor transform of a resting-state EEG dataset containing 15 people with PD and 16 people without PD. Xinjieet al. 23 introduced a hybrid CNN and RNN (recurrent neural network) model for classifying task-state EEG signals in PD diagnosis. Four deep-learning architectures were employed: CNN, RNN, and hybrid models (2D CNN–RNN, 3D CNN–RNN). Results indicate superior performance by hybrid models (3D CNN–RNN 82.89%, 2D CNN–RNN 81.13%) over conventional ones (CNN 80.89%, RNN 76.00%). The hybrids, leveraging RNN for sequential information and CNN for temporal feature extraction, present promising applications in clinical settings. Das et al. 24 utilized a sixth-order Butterworth filter to remove frequencies above 64 Hz. EEG classification has historically centered on feature extraction, which significantly decreases data dimension. There is a strong correlation between the number of features used to characterize EEG data and its classification performance. Time-domain, frequency-domain, time-frequency analysis, and chaotic features are only a few of the many approaches to EEG feature extraction. Aljalal et al. 25 present efficient discrete wavelet transform (DWT)-based methods for detecting PD from HC in two cases: off- and on-medication. EEG signals are preprocessed to remove artifacts and decomposed into several EEG sub-bands using DWT. Features are extracted from wavelet packet-derived signals using different entropy measures, such as log energy entropy, Shannon entropy, threshold entropy, sure entropy, and norm entropy. Machine learning techniques are investigated to classify resulting PD/HC features. The results show the importance of all DWT coefficients and that selecting a suitable number of EEG channels from several brain regions can improve classification accuracy.
Deep learning can autonomously learn features from data, which significantly improves the classification model's performance. Yao et al. 26 demonstrated two AE architectures for learning brief time features from public EEG data. Each test was shown as a two-dimensional picture with pixel intensities that were related to the power of different EEG frequency bands at the location of each scalp electrode and channel-wise, where each EEG electrode was treated as a separate channel. Fully connected layers were fed the learned embeddings to do classification tasks. It led to state-of-the-art performance in cross-subject experiments. CNN-based convolutional AE was proposed by Masci et al. 27 as an unsupervised learning method for learning features. Noh et al. 28 proposed a novel algorithm for semantic segmentation by utilizing a deconvolution network. However, these unsupervised feature learning techniques are predominantly employed in the field of image analysis. In the study, 29 deep neural networks, specifically CNN and convolutional long short-term memory (ConvLSTM) classifiers, were employed to analyze EEG representations in a marmoset monkey model of PD. The neural networks were trained and evaluated using local field potential measurements obtained from both healthy and Parkinsonian subjects. The outcomes demonstrated exceptional accuracies, reaching up to 99.80%. Additionally, a CNN-based AE successfully learned representations associated with PD, showcasing its capability for capturing PD-related features. This research provides further insights into the correlates of PD. Qiu et al. 30 examine PD characteristics based on brain activation and functional connectivity. On the basis of multi-scale convolutional neural network (MCNN), a novel PD detection model was proposed. The analysis of PSD and phase-locked value (PLV) characteristics revealed significant differences in the regions between HCs and PD patients. The combination of PSD and PLV enhanced the performance of PD detection. The proposed MCNN model has the potential to detect PDs automatically. Hongli et al. 31 developed an attention-based sparse graph convolutional neural network (ASGCNN) for PD diagnosis. The model uses a graph structure to represent EEG channel relationships, an attention mechanism to select relevant channels, and the L1 norm to capture channel sparsity. Experiments on the PD auditory oddball dataset showed superior performance, with notable scores in recall, precision, F1 score, accuracy, and Kappa. The study also revealed significant differences in EEG activity between PD patients and HCs, highlighting asymmetry in the frontal lobe. Using EEG channel-based analysis, Siuly et al. 32 have created an AlexNet CNN model based on time-frequency representation (TFR) for the purpose of identifying PD. While the AlexNet CNN finds intricate spatial patterns, the model uses the wavelet scattering transform (WST) to capture temporal and spectral features. Real-time EEG PD datasets were used to demonstrate the efficacy of the model, and the results indicated that frontal and central brain regions—specifically, the electrodes located at AF4 and AFz—play a major role in PD identification. The model outperformed previous techniques with astounding accuracy rates of 99.84% and 95.79%. In previous research, a novel method combining the CSADT was introduced for early diagnosis of PD. This approach was applied to four key PD datasets, including meander, spiral, voice, and speech-Sakar, and demonstrated effectiveness in diagnosing PD by analyzing critical features and extracting valuable outcomes. 33 Additionally, several models were developed, including support vector machines (SVMs), gradient boosting (GB), and random forest (RF) classifiers, to predict membranolytic anticancer activity based on peptide sequences. 34 Another study optimized the Filter Bank Regularized CSP algorithm with Transfer Learning for classifying EEG motor imagery (MI) signals, comparing its performance with the conventional CSP algorithm and presenting the results for both methods. 35 Furthermore, research into MI explored its practical applications and challenges, focusing on improving the classification accuracy of EEG signals during MI tasks. This was achieved by integrating filter bank common spatial pattern (CSP) and Riemannian manifold techniques to extract features while preserving critical signal details.36,37
Proposed method
This section describes the proposed methods for EEG signal processing, including data description, preprocessing, feature extraction, and classification techniques. Figure 1 provides an overview of the components of EEG analysis and classification for PD patients and healthy individuals. Initially, raw EEG signals undergo artifact removal and band-pass filtering to isolate the intended frequency range. Next, the filtered signals are segmented into non-overlapping sections. AEs are then employed for reconstruction, and PD/HC characteristics are extracted. To classify, an RBFNN distinguishes between PD and HC features. The subsequent subsections elaborate on each stage of the block diagram. Figure 1 displays the entire process of this proposed system.

The system model of the proposed method.
Data preprocessing
In this study, the employed data processing technique was inspired and described by Qiu et al.
30
Each channel's EEG data was first band-pass filtered at 0.5–50 Hz. Next, noise interference such as heartbeat, channel noise, and eye movement artifacts were eliminated using independent component analysis (ICA).
When there are more features than data samples, a classification model may have trouble generalizing. Numerous feature extraction and selection methods have been developed to address this issue, with AE being one such approach. The encoder component can extract features when training an AE. Creating a nonlinear mixture of these factors may help convey all features more clearly. In this study, classifiers are trained on raw and feature-extracted data to compare results.
The AE has a structure of 400, 200, 100, 50, 100, 100, 200, 400, and 753 neurons with a batch normalizer following each layer (Figure 2). The activation function of each neuron in these models is a rectified linear unit (ReLU), and the models are optimized using a root mean square propagation, or RMSprop, with a learning rate of 0.05 and 50 epochs. The model was developed utilizing the Python and the PyTorch implementation.

Electrode map for PSD.
PSD
PSD is a tool used to describe the strength of a signal in time-domain, often used in EEG data to examine brain activity shifts related to attention and mood, capturing natural power disparities.
To filter the EEG signal, high- or low-frequency noise is eliminated, and artifact correction is used if needed. The signal is split into windows, exposed to the Hanning window function with overlap rate of 50% and windowed finite impulse response filter
38
to reduce edge effects:


AE
AE 39 is a specialized neural network structure with input, output, and concealed layers. Because it doesn't require labeled data, this architecture makes the model especially special in that it can find inherent patterns and hierarchical representations. Training is used to adjust the weights of the hidden layer so that the input and output values are as close as feasible. In order to accomplish unsupervised feature extraction, the hidden layer must contain significant original signal features. AE resembles principal component analysis (PCA), which can reduce the dimension of data. 40
Figure 3 depicts the fundamental structure of AE, which consists of two data processing phases: encoding and decoding. The original input signal data during encoding is 

The AE framework consists of an input layer, a hidden layer, and an output layer, with the weights of the hidden layer representing aspects of the input signal. 41


RBFNN
An RBFNN is a type of artificial neural network that uses radial basis functions for activation. Broomhead and Lowe et al. 42 created RBF neural networks. It handles high-dimensional input data, making it appropriate for multi-feature problems. It can use fewer parameters and less overfitting to get a sparse input representation. RBF is a popular alternative to the MLP since it has a simpler structure and faster preparation. TBF is more computationally efficient than other radial basis functions, but Gaussian was chosen since it handles complicated data more accurately and has understandable parameterization. This makes it popular. Symmetrical stabilizers create RBF organizers. Figure 4 illustrates the principal structure of the network.
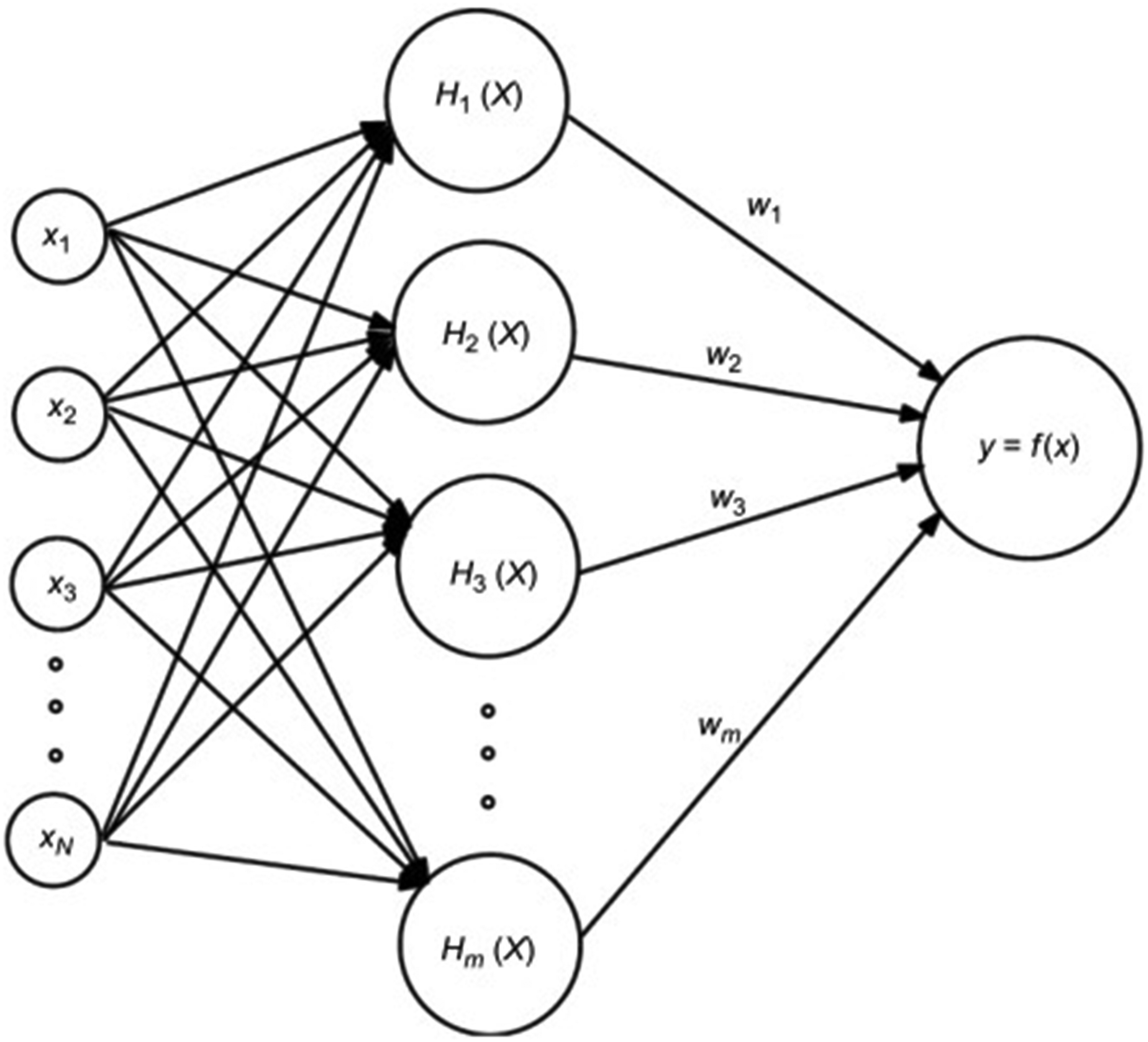
Basic architecture of radial basis function neural network. 43
The three layers that comprise the radial basis function are the input layer, hidden layer, and output layer, 44 as seen in Figure 4. The activation function of the hidden layer is the RBF. The function is real-valued, and its value varies based on the distance from the origin or center.
The RBFNN transfers the input data to an output based on the distance between the input data and the centers of the radial basis functions using radial basis functions. The Euclidean distance is the standard. The Euclidean distance between the input vector 

In the end, the RBFNN will make something that is a weighted sum of the activation values of the radial basis functions. Using the smallest mean square error (MSE) between the RBFNN's real and predicted outputs, linear regression is used to figure out the output layer weights. We can write the linear transformation equation as:
Dataset
The dataset has EEG data from 31 people, 16 HCs (9 women, mean age 63.5 ± 9.6 years), and 15 PD cases (8 women, mean age 62.6 ± 8.3 years). They were all from the Scripps Clinic in La Jolla, California, USA, and had mild to intermediate PD. The HCs were independent contractors from the area. Each PD patient in this dataset came to the lab twice, once while on medicine and once while not on medication. For each subject, the order of visits was switched around. For the OFF session, at least 12 hours have passed since the last dose of the dopaminergic drug. Patients took their regular morning amount before the ON session. Because of this, the EEG data of all 15 PD patients showed data when they were on medication and when they were not (called PD_ON and PD_OFF in the study, respectively). A 32-channel Biosemi Active Two EEG device was used to record the EEG data of the people who were given the tests while they were at rest at 512 Hz for at least 3 minutes (Figure 5). Rockhill et al. 47 about the 32-channel EEG sensor positions that give more information about these datasets. Table 1 presents the demographic characteristics of the study participants.

The 32 EEG channels utilized in this investigation were shared. 46
Demographics of patients and control participants (mean ± SD).

Experiment and result analysis
This section provides a thorough explanation of our research's findings by outlining our methodology and findings.
Procedure
Using our state-of-the-art Deep Learning (DL) method, PD patients were further classified (including PD_ON and PD_OFF) and HC based on the analyzed features. Raw data was filtered utilizing PSD and extracted valuable features utilizing AE. For that, the glob library function was utilized to identify and compile EEG data files within the specified directory. It helps to extract patient IDs and corresponding file paths from the dataset. In addition, to validate the efficacy of the model, a hybrid classification architecture was constructed by combining an AE and a radial basis function. Prior to being input into the classification model, every feature utilized in the classification process was normalized. All channel data pertaining to subjects that were classified were combined in each classification experiment and subsequently labeled uniquely by group (e.g., PD and HC). For instance, in the UC SanDiego dataset's two-class classifications of PD_ON and HC, HC data was assigned the value 0 and PD_ON data was assigned the value 1. The feature data size for each subject in the
It was discovered, through comparison and analysis of PSD characteristics, that the PD group (including PD_OFF and PD_ON) and the HC group, in addition to PD_OFF and PD_ON, differed significantly in the
Performance evaluation
Several metrics are used to evaluate the performance of the developed models: classification accuracy, sensitivity, specificity,
Accuracy can be calculated in terms of positives and negatives as follows
48
:

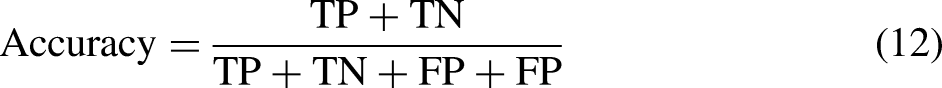
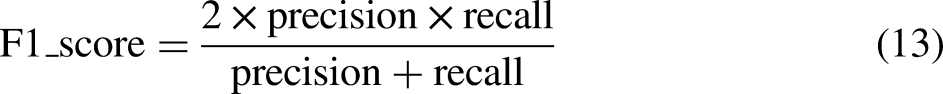

Discussion
The accurate characterization of brain function in PD remains a contentious issue, highlighting the need for more effective and reliable diagnostic methods. In response, a novel PD detection model was proposed to utilize a resting-state EEG data. This model incorporates analysis of both brain activation and functional connectivity patterns to achieve high-precision identification of PD patients. Extensive prior research employing diverse time, frequency, and time-frequency domain features has established that resting-state EEG of PD patients exhibits distinct characteristics compared to HCs. Notably, these studies demonstrate the promising potential of EEG-based spectral and connectivity markers for differentiating between PD and healthy individuals, thereby laying the foundation for our proposed model.
The experimental results are shown in Table 2 and Figure 6 of two classifications based on AE + RBFNN. Figure 7 shows the proposed model's ROC–AUC that were utilized to categorize participants into HC and PD (PD_OFF and PD_ON).

The performance analysis of two-class classifications based on AE + RBFNN.

ROC–AUC score of HC and PD (PD_OFF and PD_ON).
The results of two-class classifications based on AE + RBFNN.

People with PD had a different resting-state EEG than HCs. It was found in studies that used different time, frequency, and time-frequency domain characteristics (Table 3). These studies also show that EEG-based spectral and connectivity markers can be used to tell the difference between HC and PD patients. Most of these studies only look at one pattern (spectral or connectivity) to determine PD and how the brain works in PD patients. This makes it hard to figure out what the brains of PD patients are like when they are functioning. Our findings concur with those of a few earlier research projects. For example, using resting EEG data from PD patients, Gong et al.
53
discovered aberrant phase-amplitude coupling between
Comparison of the proposed method with other state of art methods.

The confusion matrices are shown in Figure 8 to examine the suggested approaches' capacity to detect OFF_PD/HC, ON_PD/HC, and ON_PD/OFF_PD. 300 segments were retrieved from off-PD, 297 vectors from on-PD, and 306 from the HC group. Five result values are produced for each assessment parameter (accuracy, sensitivity, specificity, and

Confusion matrices for each classification (HC vs. PD_ON, HC vs. PD_OFF, PD_OFF vs. PD_ON).
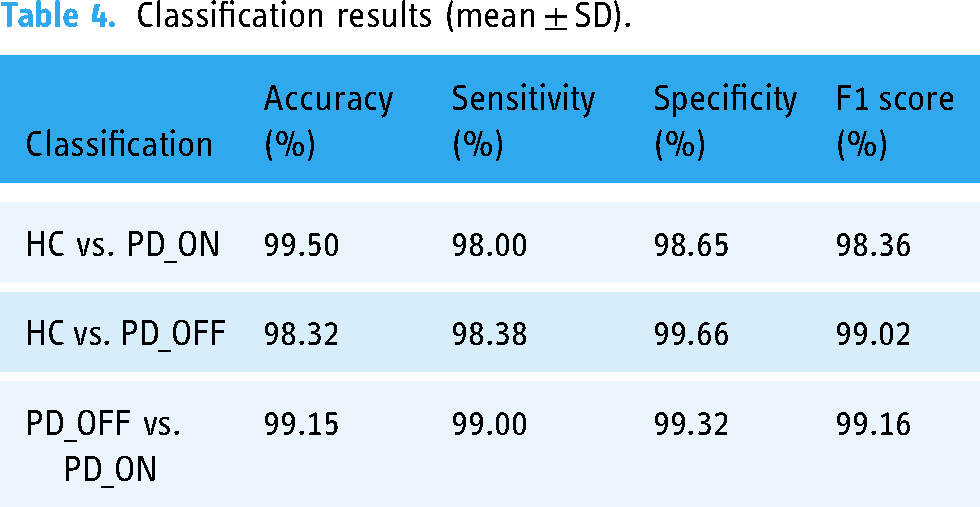
The classification performance is shown in Table 4 with respect to
Classification results (mean ± SD).
In this proposed method, phase-related was integrated into functional connectivity and PSD-based activation patterns into our AE + RBFNN pipeline because we saw the potential of multi-modal research. Compared to relying only on isolated data, this novel fusion enables us to obtain a deeper and more complex picture of brain activity in PD. Functional connectivity patterns in the brain signify information linkages between different regions, while activation patterns capture variations in activity across multiple brain areas. The combination of phase-related connection patterns and power spectral activation patterns from EEG data has demonstrated improved emotion recognition capabilities. 55 Integrating these complementary signals can enhance the accuracy of PD identification, offering more discriminative information compared to single-pattern features. Despite this potential, limited research has explored the joint utilization of these two signals for PD identification. In this study, we combined the techniques of PSD, AE, and radial basis function to detection more precisely. Through the utilization of both activation and connection patterns' complementing capabilities, this synergistic technique represents a substantial advancement above conventional single-feature methodology. This leads to an improved understanding of the intricate neurophysiological causes of PD and enhances the discriminative potential for PD detection. PSD preprocessing, AE-based feature extraction, and RBFNN classification come together to create a multi-step pipeline that makes use of the advantages of several methods. The experimental results (Table 2 and Figure 6) demonstrate that the suggested model is capable of successfully identifying PD, including PD_OFF and PD_ON. Based on the extracted features (γ band), the AE + RBFNN model's classification accuracy, sensitivity, specificity, and AUC for PD (including PD_OFF and PD_ON) and HC, as well as PD_OFF and PD_ON, are all over 95%.
Confidence intervals establish a range where the true model performance is likely to reside. This statistical tool is crucial for evaluating result reliability and consistency. The model's accuracy, sensitivity, and specificity for differentiating HC and PD_ON were assessed. Accuracy yielded a 95% confidence interval of [0.991, 0.999] using a 1000-iteration bootstrap method. Sensitivity and specificity intervals were [0.971, 0.989] and [0.979, 0.994], respectively. These intervals indicate consistently high performance across varied data subsets. The narrow confidence intervals suggest stable and robust model behavior, crucial for dependable clinical applications. While comparable in accuracy to previous studies, the model's narrower confidence interval suggests improved reliability. Our initial findings regarding PD detection offer promising advancements, yet several limitations deserve attention. The study's key limitation is that it depends on a single dataset for both training and evaluation. The limited size and scope of the dataset which was employed in this research may limit how broadly applicable our findings can be, even though it offers insightful information. Larger and more varied datasets may be useful in future research to improve the robustness and generalizability of the suggested methodology in various demographics and contexts. Secondly, while the implemented greedy algorithm provides a readily applicable method for channel selection, it may not yield the optimal configuration for maximizing classification accuracy. Future research will explore more robust optimization techniques, such as genetic algorithms or simulated annealing, to identify the minimal set of channels that achieve the highest performance. This approach not only enhances detection accuracy but also promotes a more practical and user-friendly system with reduced electrode complexity. Thirdly, the proposed model currently lacks comprehensive validation in real-world clinical settings. To address this, we intend to establish collaborations with medical institutions to acquire larger and more diverse datasets and rigorously validate our model using patient-generated EEG data during live examinations. Finally, we are planning to delve deeper into advanced feature fusion techniques that integrate information regarding brain activation and functional connectivity. By leveraging the complementary insights gleaned from both aspects of brain function, we aim to further enhance the model's performance in accurately identifying PD patients.
Given the encouraging results obtained by the suggested technology, it is important to examine its incorporation into clinical decision support systems or telehealth applications. Such integration could improve the accuracy and efficiency of patient monitoring and diagnosis, especially in distant areas where access to healthcare specialists is limited. Future studies could investigate this route by examining the method's performance in different applications and determining its impact on clinical outcomes. Despite the technical and practical hurdles, we face in implementing our algorithm for online detection within a closed-loop system, we firmly believe in its potential to revolutionize real-time monitoring and intervention in PD management. This methodology could be especially useful in telehealth settings, enabling early detection and ongoing monitoring in underserved or remote populations. By implementing this approach in clinical practice, treatment programs could become more individualized, relieving patients of the burden of illness progression and improving their quality of life. With further refinement and validation, our approach holds immense promise to enhance patient care and clinical outcomes. Moving forward, this research will be dedicated to surmounting these challenges and exploring the feasibility of integrating our method into closed-loop systems, ultimately advancing the field and improving the lives of those affected by PD.
Potential additions to the current framework merit additional examination. Expanding the framework to allow for early detection, illness progression monitoring, and individualized treatment techniques could significantly improve its utility in clinical practice. These enhancements would include the creation of more complex analytical tools as well as the incorporation of patient-specific data, paving the way for more personalized and effective healthcare solutions.
Conclusion
In this paper, we present a novel, effective technique for detecting PD from resting state EEG signals using hybrid architecture. EEG signals were processed using PSD. Moreover, our proposed AE + RBFNN model has a high cross-validation accuracy and AUC of 99.0% for sensitivity, specificity, and ROC for automatic PD detection. In EEG data, it reduces artifact signals and improves the signal-to-noise ratio. The features are then extracted using AE architecture which can identify clusters or patterns in the latent space which might correspond to different cognitive states or neurological conditions. The integration of AE architecture enables the identification of latent clusters or patterns in the EEG data's latent space, enhancing our ability to discern different cognitive states or neurological conditions. A critical distinguishing factor is the successful combination of historical data consideration within the RBFNN, making our model uniquely robust in PD detection. This model considers information obtained after the current analysis period and historical data when making decisions. The advantages of both models are used in this method. The SanDiego dataset was used to validate each of these techniques in terms of on and off-medication. Comparing our findings to previous studies in the field, they show a significant improvement in the method for identifying PD. A state-of-the-art technique has been meticulously developed to enhance the pre-processing and interpretation of EEG data, thereby contributing to the body of knowledge. The creation of customized diagnostic models that are based on the traits of each patient and the course of their illness is made possible by the integration of machine learning algorithms. This approach creates patient-specific insights and predictive models by evaluating EEG signals along with clinical data, allowing medical professionals to propose individualized treatment plans and treatments. This individualized strategy maximizes the use of available resources, reduces side effects associated with treatment, and enhances patient satisfaction and care in general.
Footnotes
Acknowledgment
The authors would like to extend their sincere appreciation to the Researchers Supporting Project Number (RSP2024R301), King Saud University, Riyadh, Saudi Arabia.
Data availability statement
Contributorship
FAJ and AT: conceptualization, data curation, methodology, software, resource, visualization, formal analysis, and writing—original draft and review and editing. MAT and MAU: visualization, investigation, writing—review and editing, supervision, and formal analysis. FR and MSU: visualization, investigation, and writing—review and editing. FKA and MK: visualization, validation, investigation, methodology, and writing—review and editing.
Consent to participate
Not applicable.
Consent to Publish
Not applicable.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
Not applicable.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The work was funded by the Researchers Supporting Project, King Saud University (Grant No. RSP2024R301).
Guarantor
Md. Alamin Talukder.
