Abstract
Objective
Cervical cancer stands as a leading cause of mortality among women in developing nations. To ensure the reduction of its adverse consequences, the primary protocols to be adhered to involve early detection and treatment under the guidance of expert medical professionals. An effective approach for identifying this form of malignancy involves the examination of Pap smear images. However, in the context of automating cervical cancer detection, many of the existing datasets frequently exhibit missing data points, a factor that can substantially impact the effectiveness of machine learning models.
Methods
In response to these hurdles, this research introduces an automated system designed to predict cervical cancer with a dual focus: adeptly managing missing data while attaining remarkable accuracy. The system's core is built upon a stacked ensemble voting classifier model, which amalgamates three distinct machine learning models, all harmoniously integrated with the KNN Imputer to address the issue of missing values.
Results
The model put forth attains an accuracy of 99.41%, precision of 97.63%, recall of 95.96%, and an F1 score of 96.76% when incorporating the KNN imputation method. The investigation conducts a comparative analysis, contrasting the performance of this model with seven alternative machine learning algorithms in two scenarios: one where missing values are eliminated, and another employing KNN imputation. This study offers validation of the effectiveness of the proposed model in comparison to current state-of-the-art methodologies.
Conclusions
This research delves into the challenge of handling missing data in the dataset utilized for cervical cancer detection. The findings have the potential to assist healthcare professionals in achieving early detection and enhancing the quality of care provided to individuals affected by cervical cancer.
Keywords
Introduction
For effective disease progression, the EHRs (electronic health records) have the ability to provide valuable insights, which can lead to improved healthcare. 1 As a result, EHRs are getting attention for a large number of practical applications such as disease risk prediction, adverse event detection, and medical decision support. 2 By studying large populations over time and examining the correlation between clinical events and outcomes, mining EHRs has the significant potential to generate new clinical hypotheses. 3 Using HER data, unlike cohort studies that collect data from individuals in a limited time frame, has both pros and cons. One of the top advantages of EHR data utilization is that it is continuously collected and encompasses a more extensive range of information types. Moreover, obtaining the data may require fewer resources. However, since EHRs are not primarily designed for research purposes, the data collected may be sparse and contain high levels of noise, making it difficult to analyze it. 4
Human papillomavirus (HPV) is the leading cause of malignant tumors in the reproductive system of females. 5 The fourth most typical cancer in women to be diagnosed is cervical cancer, which also ranks as one of the main causes of death for females. In the female population, cervical cancer makes for 6.5% of all malignancies. According to estimates, there were around 342,000 cervical cancer deaths and 604,127 new cases of the illness globally in 2020. 6 Cervical cancer is still a significant public health issue, especially in developed nations: In Europe, there are 54,517 newly diagnosed instances of invasive cervical cancer reported each year, and 24,874 women every year pass away as a result of cervical cancers. 7 Regular immunization programs often include vaccinations. A total of 90% of girls in Europe are expected to have received the entire HPV vaccine by the age of 15 by 2030. 8 Additionally, the WHO urges decision-makers in European nations to intensify efforts to eradicate cervical cancer by utilizing the current prevention technologies. 9
The objective of this research is to utilize EHRs as a source of input to detect the presence of cervical cancer, which is the most frequently occurring cancer among women. Each year, over half a million women are diagnosed with cervical cancer, making it the fourth most common disease in women after breast, colon, and lung cancer. The incidence of cervical cancer among women in Europe varies from 12 to 30 cases per 100,000. 10 Detection of cervical cancer at the early stages and preceding alterations has contributed to a continuous decline in the worldwide prevalence of cervical cancer throughout the years. In the past, cervical cancer constituted up to 70% of all genital cancers, but currently, it accounts for approximately 35%–50%, with a tendency for further reduction. The age group most affected by cervical cancer is between 20 and 40 years old, and the incidence decreases among women over 40 years old, where only 1%–2% are diagnosed with cervical cancer. 11
The main cause of cervical cancer is in the lower part of the uterus abnormal changes in the cells of the cervix, which gradually progress from precancerous cells to cancer cells. In the majority of the cases (90%–95%) cervical cancer develops slowly and takes a time of 5 –10 years for pre-stages or dysplasia stages (CIN I, CIN II, and CIN III). On the other hand, 5%–10% of cases, it does not exhibit earlier changes in screening tests. 12 Early detection of cervical cancer through systematic gynecological examination allows for a complete cure of the disease. Severe dysplasia (CIN III) or carcinoma in situ are the direct precursors of cancer and are often preceded by mild (CIN I) and moderate (CIN II) dysplasia changes for an extended period. 13 At the early stages of cervical cancer, several treatment methods have been used such as radiation, chemotherapy, and surgical resection, but the treatment is totally dependent on the stage of the prognosis of cancer. 14
It normally takes several years for normal cervical mucus to evolve into detectable malignancy. Frequent gynecological exams, Pre-service Academic Performance Assessment tests, colposcopies, and HPV typing allow for early detection and treatment of these abnormalities, thus, preventing cervical cancer. There are two main types of cervical cancers: squamous cell cancer, which accounts for 70%–80% of cases, and adenocarcinoma, which develops from glandular cells that produce mucus and line the cervical canal. Although adenocarcinoma is less common than squamous cell carcinoma, its incidence has been increasing in recent years and it represents 10%–15% of all uterine malignancies. 15 Adenocarcinoma is more challenging to detect through screening as it develops inside the cervical canal, rather than on the cervix. 16 Nonetheless, the treatment for both types of cervical cancers is the same. The primary cause of cervical cancer is high-risk strains of HPV. Women infected with HPV have a higher likelihood of developing cervical cancer due to risk factors such as smoking, early sexual activity, multiple sexual partners, genital herpes infection, weakened immune system, lower socioeconomic status, poor genital hygiene, and a higher number of childbirths. 17
Detecting cervical cancer can be challenging because the disease is usually asymptomatic in its early stages. Changes in the cervix are often found during routine annual check-ups. Cervical cancer symptoms vary according to the stage of cancer, with advanced stages exhibiting distinct signs in around 90% of cases, with irregular bleeding being the most common symptom. 18 Postmenopausal bleeding, spotting, bleeding during sexual intercourse or defecation (known as contact bleeding), and brownish discharge are all common signs of this condition. In addition to the symptoms mentioned earlier, another important indication of cervical cancer is the presence of bloody discharge that usually has an unpleasant odor. When the disease reaches an advanced stage, pain in the lower abdomen may develop due to the involvement of nearby organs. 19
Objective of study
Cervical cancer is a serious disease that can be fatal, and many people are not aware of its dangers. However, early detection and treatment can help prevent it. 20 Unfortunately, many countries do not have effective screening methods to detect this type of cancer. Existing datasets are not appropriate for ML (machine learning ) models as they might contain missing or null values that have a huge impact on the efficiency of models. To overcome this issue, this study investigates the use of K nearest neighbor (KNN) imputer. Additionally, an ensemble model is proposed to increase the detection accuracy of cervical cancer detection. We also compare the effectiveness of various ML techniques for cervical cancer detection in comparison to the proposed approach.
Major contributions
The major contributions of this research work are as follows:
This study proposes a novel ensemble model for predicting cervical cancer in patients. The proposed ensemble model is based on the XGB (extreme gradient boosting), RF (random forest), and ETC (extra tree classifier) with a voting mechanism for making the final prediction. For the issue of missing values, experiments are done with the missing values deleted and the KNN imputer to generate missing values. A performance comparison is conducted using various ML models, such as RF, LR (logistic regression), GBM (gradient boosting machine), GNB (Gaussian Naive Bayes), ETC, SVC (support vector classifier), DT (decision tree), and SGD (stochastic gradient descent). Furthermore, the proposed model’s effectiveness is analyzed by comparing its performance to state-of-the-art approaches in terms of accuracy, precision, recall, and F1 score.
The remaining sections of the paper are structured as follows. The “Related work” section presents a review of the relevant literature on cervical cancer detection using ML techniques. The “Materials and methods” section provides an overview of the dataset and the ML models used in the study. It also describes the proposed methodology in detail. The “Experimental results” section presents the results of the study and discusses their implications. Finally, The “Conclusions” section offers concluding remarks and suggests potential areas for future research.
Related work
Over the decades, efforts to cure cancer have taken various approaches. While complete eradication may not be possible, detecting and predicting the disease can help mitigate its impact. Early detection is crucial for successful treatment, but cervical cancer is difficult to detect in its early stages as it typically has no early symptoms. Therefore, regular screening is the only way to predict the presence of cancerous cells. However, screening results can sometimes yield false-positive results or be delayed, leading to potential risks. To address these challenges, AI (artificial intelligence) has been introduced in healthcare. Various algorithms, tools, and techniques are being used to improve the accuracy and speed of predicting cancerous cells while reducing the false-positive rate. Many researchers have used different algorithms to predict cervical cancer. Another interesting work on the topic of cervical cancer detection is done by Dweekat and Lam. 21 In that research work, the key component is dimensionality reduction which is done using principal component analysis (PCA). The authors imply three different types of models for cervical cancer detection that are GA–MLP, PCA–GA–MLP, and PCA–AdaBoost. The combined approach of genetic algorithm and multilayer perceptron (MLP) is proposed using PCA, and the resulting features are then fed into the MLP for model training and achieved robust results for the precise prediction of cervical cancer.
Yaman and Tuncer 22 conducted a thorough study on the categorization of cervical cells in pap smear images using two datasets: SIPaKMeD and Mendeley LBC (liquid-based cytology). The authors utilized NCA (neighborhood component analysis) to select 1000 features and the SVM (support vector machine) technique for classification. To validate the findings, they performed five-fold cross-validation and hold-out validation (80:20). The results demonstrated high accuracy rates of 98.26% and 99.47% for the SIPaKMeD and Mendeley LBC datasets, respectively. Similarly, Huang et al. 23 proposed a method to detect cervical cancer by fine-tuning pre-trained deep network models to extract deep convolutional features, including Inception ResNet, DenseNet-121, VGG19 Net, ResNet-50V2, and InceptionV3. They also used local binary patterns and an oriented gradient histogram to capture typical visual characteristics. The authors found that the combination of deep features extracted from DenseNet-121 and ResNet-50V2 achieved the highest accuracy, with an average classification accuracy of 95.33%, surpassing the results of ResNet-50V2 and DenseNet-121 used separately. Additionally, they observed a 90.89% improvement in the identification ability.
Using the Harvel dataset, Alquran et al. 24 combined DL (deep learning) and a cascading SVM classifier to classify cervical cancer into seven classifications, obtaining an accuracy of up to 92%. The research revealed the efficacy of integrating DL with classic ML approaches for accurate cervical cancer classification. According to Alsmariy et al., 25 introduced a ML system to forecast cervical cancer using the UCI cervical cancer risk factor dataset. To address the issue of class imbalance, they utilized SMOTE (synthetic minority oversampling technique) with PCA. The system achieved a remarkable accuracy score of 98.46%. Meanwhile, Lilhore et al. 26 suggested an ensemble model to identify cervical cancer. The proposed approach included a feature selection technique and prediction model based on the Boruta analysis and an SVM classifier. Boruta analysis, an improved version of RF, identifies subsets from the data source that have a significant impact on the classification accuracy. The findings of their research revealed that Boruta with SVM achieved a precision score of 0.912.
To detect the malignant cervical formation, Mehmood et al. 27 proposed an automated system named CervDetect. CervDetect is basically a ML-based system that uses the Pearson correlation between the input and output variables for the data pre-processing. For the significant feature selection, CervDetect uses the RF model. The result of the study shows that the CervDetect achieved a 93.6% accuracy score and 0.07111 MSE (mean square error). An overhead cross-section sampling ML model is proposed by Anandaraj et al. 28 for cervical cancer prediction. To handle the class imbalance problems, the authors used the oversampling technique SMOTE. To check the efficacy of the proposed system, they used the 13 ML models. On the imbalanced dataset, the RF classifier achieved an accuracy score of 96% and the same classifier sustained 98% accuracy using the oversampling technique.
In the study by Alsmariy et al., 29 a ML algorithm was developed for predicting cervical cancer. The study used three machine learning models: LR, DT, and XGBoost. The results of the study demonstrated that ML classifiers achieved the highest performance when using the selected features. Mudawi and Alazeb 30 proposed an astute way of cervical cancer prediction. They used the six ML models and on the selected features the ML models RF, DT, ADA, and GBM give good accuracy scores. Quinlan et al. 31 made a comparison of different ML models for the classification of cervical cancer. The dataset used is an imbalanced dataset which is handled by the resampling technique SMOTE-Tomek in conjunction with a tuned RF. Results show that the RF with S-Tom achieved an accuracy score of 99.69%.
Nithya and Ilango 32 conducted a study on cervical cancer prediction, utilizing optimized feature selection methods and classification techniques. Five ML models, including RF, KNN, SVM, Rpart, and C5.0 were employed. Feature selection methods were applied to identify important features, and an optimized feature selection was developed. The findings of the study indicate that C5.0 outperformed other models in terms of accuracy score. Similarly, Gowri and Saranya 33 proposed a ML system for cervical cancer prediction, aiming to achieve high accuracy. They used DBSCAN and SMOTE-Tomek for outlier detection from the dataset and performed prediction using two scenarios: DBSCAN+SMOTE-Tomek+RF and DBSCAN+SMOTE+RF. The study revealed that DBSCAN+SMOTE+RF attained an accuracy value of 99%.
These studies demonstrate the potential of ML techniques, such as DL and ensemble methods in detecting and predicting cervical cancer. They also highlight the importance of addressing class imbalance and feature selection to improve the accuracy of the models. Overall, these approaches show promise in improving the accuracy and efficiency of cervical cancer detection and prognosis.
Materials and methods
In this section, we provide a brief overview of the dataset used, the applied data preprocessing techniques, employed ML algorithms for cervical cancer detection, and a summary of the class-balancing techniques used in this study.
Dataset for experiments
This study utilized a publicly available dataset by Fernandes et al. 34 , gathered at the Hospital Universitario de Caracas in Venezuela. This dataset is currently the only publicly available that can be utilized for creating a potential cervical cancer screening survey using AI algorithms and questionnaires. The researchers aimed to evaluate the suitability of AI models and class-balancing techniques for developing such a study. Table 1 provides a comprehensive overview of the 35 input variables and an output variable included in the dataset, which is comprised of a total of 858 instances and 36 attributes. Each of the input variables is described in detail in Table 1.
Description of the dataset used in this study.

The dataset consists of the output variable biopsy. Table 1 demonstrates that the dataset exhibits severe class imbalance. Given the difficulties associated with imbalanced data classification, this study opted to implement the KNN imputer as a data balancing technique.
Data preprocessing
Achieving optimal performance from ML models relies heavily on data preprocessing, which involves removing irrelevant or redundant data from the dataset to enhance model efficacy and reduce computational time. During the data preprocessing stage of the research, it was discovered that the dataset contained a considerable number of missing values, which are classified by class in Table 1. Table 1 reveals the presence of a significant amount of missing values in the dataset. Given that the data in the dataset are categorical, there are two viable approaches for dealing with the missing values
Removing the missing values from the dataset entirely. Employing the KNN imputer technique.
Removing missing values from cervical dataset
One possible approach for handling the missing values in the dataset is to remove any instances that contain missing values. In the first experiment, we opted to explore this approach by removing all fields that contained missing values.
KNN imputer
For the second option to deal with the missing values, the KNN imputer can be employed to address missing values in the data. In today’s world, data is collected from a wide range of sources and used for various purposes, including analysis, valuable insights, and hypothesis validation. However, errors during data extraction or collection can result in missing information, making handling missing values a critical preprocessing step. The choice of imputation method is crucial, as it can significantly impact model performance. The KNN imputer is a widely used technique for imputing missing values, serving as an alternative to traditional imputation methods.
35
It replaces missing values by identifying the nearest neighbors using the Euclidean distance matrix. The Euclidean distance is calculated by ignoring the missing values and giving more weight to the non-missing coordinates. For the distance calculation, the Euclidean distance can be computed as

ML models used for cervical cancer detection
This section covers the ML algorithms that were employed for detecting cervical cancer, utilizing Python programing language to implement all eight of the supervised ML algorithms. These algorithms are frequently employed for solving classification and regression problems. The proposed system’s effectiveness was evaluated using tree-based algorithms, regression-based models, and ensemble models. The classification problem was addressed in this study by utilizing a total of eight distinct ML algorithms, as well as an ensemble learning model.
Random forest
RF classifier is an algorithm based on the decision trees that employ multiple weak learners to produce highly accurate predictions.36,37 To train multiple decision trees using different bootstrap samples, RF utilizes a technique called “bootstrap bagging.” A bootstrap sample is created by randomly subsampling the training dataset, with both the training and test datasets having the same size. As with other ensemble classifiers, RF employs DTs to make predictions. When constructing decision trees, selecting the root node at each stage can be a challenging task. RF can be defined as

Decision tree
DT is a well-known ML algorithm that has broad applications in solving regression and classification problems. 38 At each level of the DT, selecting the root node is a critical aspect of the process, which is commonly referred to as “attribute selection.”
Logistic regression
LR is a commonly employed method for addressing binary classification problems, owing in part to its utilization of the logistic equation (also known as the sigmoid function). This function transforms any given numerical value into a number ranging between 0 and 1 via an S-shaped curve, which is what makes LR so widely used.39,36
Support vector classifier
The SVC is a popular supervised ML algorithm that is widely used for solving classification problems. 40 SVC identifies the optimal line in two dimensions using the radial basis function kernel and can also be used to identify the regression line.
Extreme gradient boosting
In this study, XGBoost is employed as a high-speed supervised learning algorithm for achieving accurate and precise classification of cervical cancer.
41
Its regularized learning features facilitate the smoothing of final weights and prevent overfitting. The XGBoost algorithm involves minimizing a loss function
Extra tree classifier
ETC is a popular ensemble learning algorithm that combines multiple DTs to make accurate predictions. 42 Unlike RFt, ETC randomly selects a subset of the best features to use for each split in the DT. This process results in the creation of de-correlated trees, where each tree is less sensitive to individual features and more robust to noise. ETC uses the Gini index as a criterion for selecting the best feature to split the data. It also ranks the importance of features based on their Gini score.
Gaussian Naive Bayes
GNB is a Bayesian algorithm used for classification. It assumes that the features are normally distributed and independent of each other. 43 The algorithm calculates the likelihood of a feature value for each class and then uses Bayes’ theorem to calculate the probability of each class given the input values.
Stochastic gradient decent
SGD is an algorithm that combines the working principles of LR and SVM. SGD uses the convex loss function of LR and is a robust classifier. 44 It is particularly useful for multiclass classification and combines multiple classifiers using the OvA (one-versus-all) approach.
Proposed approach for cervical cancer detection
The dataset utilized in this study was obtained from Kaggle, a well-known source for publicly available datasets. In order to address the issue of missing values and improve the performance of the learning models, preprocessing was carried out. To handle the missing values, the KNN imputer was employed. The data was then split into a 70:30 ratio, with 70% designated for model training and 30% for testing. The proposed cervical cancer detection system utilized the XGB+RF+ETC ensemble approach. Ensemble models are a powerful technique that involves combining the predictions of multiple models to improve accuracy and robustness. Each of the models in an ensemble has its own strengths and weaknesses, and by combining them, we can achieve a better overall performance. For cervical cancer detection, this study proposes an ensemble learning model that combines three popular algorithms XGB, RF, and ETC. The workflow diagram of the proposed approach is shown in Figure 1.
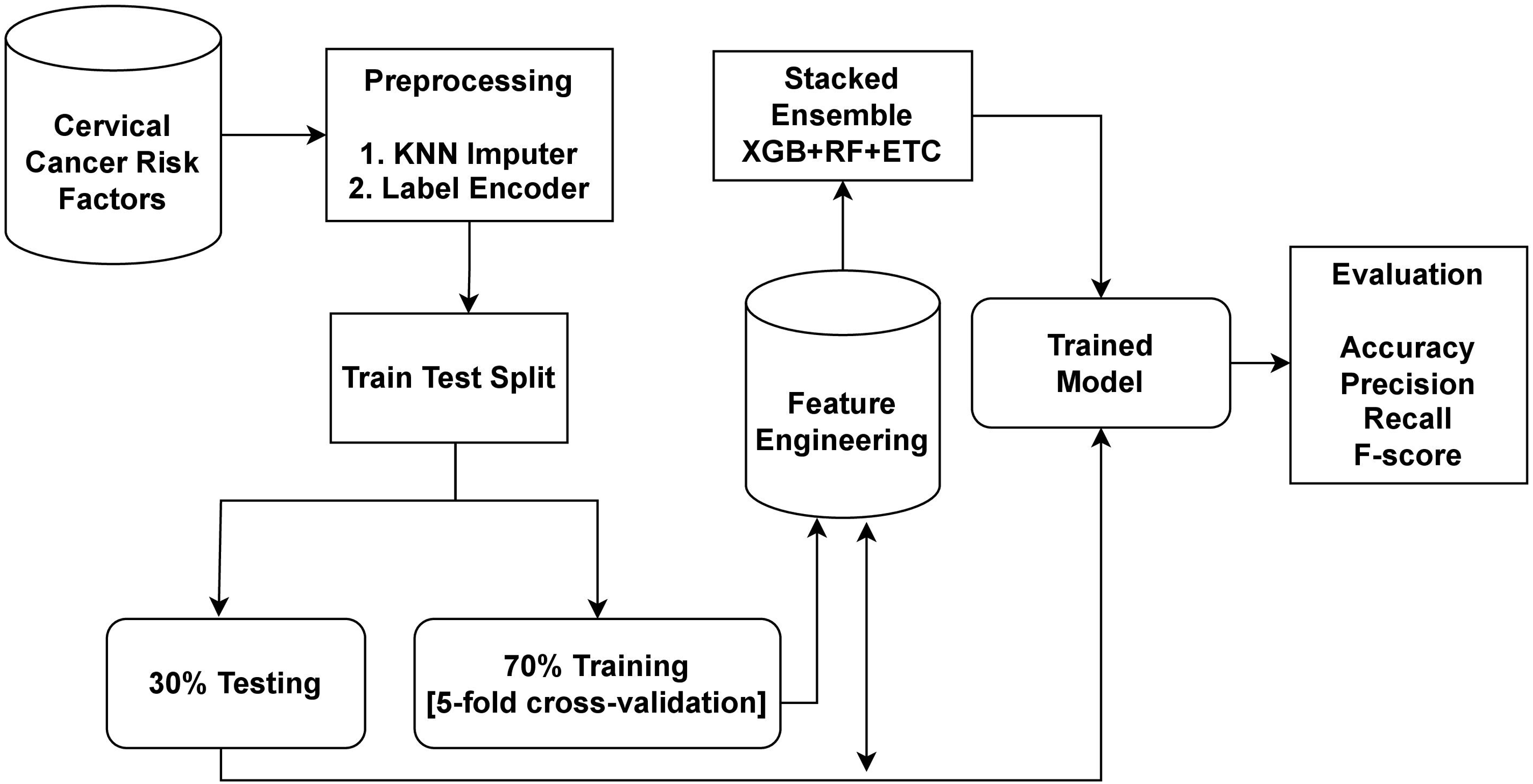
Workflow diagram of the proposed methodology.
The ensemble model works by combining the predictions of three different ML algorithms. The general approach for building an ensemble model is to train multiple models on the same dataset and then combine their predictions. The XGB+RF+ETC ensemble model follows this approach by training XGB, RF, and ETC models on the same dataset separately. Each of these models generates predicted probabilities for each class of the target variable. These predicted probabilities can then be combined to make a final prediction for each observation in the dataset. One common way to combine the predictions is to take a weighted average of the predicted probabilities, where the weights are determined by the performance of each model on a validation set.
The proposed ensemble model works by combining the strengths of three different ML algorithms to produce more accurate and robust predictions. By training multiple models on the cervical cancer dataset and combining their predictions, we can improve the generalization performance of the model and reduce overfitting.
Statistical analysis
The statistical analysis involves comprehending the functionality of the ensemble model’s operations. The ensemble model’s ability to use the prediction strengths drawn from three different ML models can be better understood by looking further into its mathematical representation. Algorithm 1 explains the working of the proposed ensemble model, which can be expressed as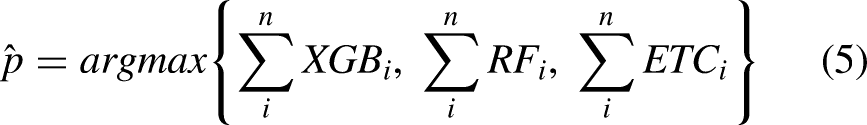

Architecture of the proposed voting classifier.
Algorithm 1 Ensembling of XGB, RF, and ETC.

The ensemble model chooses the final class based on the maximum average probability of a class and combines the projected probabilities of both classifiers. The final prediction will be the one whose probability score is the largest, as
Evaluation metrics
For performance analysis, the evaluation phase is crucial which involves assessing the performance of learning models. Commonly used evaluation parameters such as accuracy, precision, recall, and F1 score are employed to assess the performance of breast cancer detection models. These parameters are calculated based on the values in the confusion matrix, which illustrates the classifier’s performance on the test data. The values of TP (true positive), TN (true negative), FP (false positive), and FN (false negative) are utilized to compute these evaluation parameters, and their values range from 0 (minimum) to 1 (maximum). Accuracy is a measure of the classifier’s ability to predict both positive and negative classes correctly. Mathematically,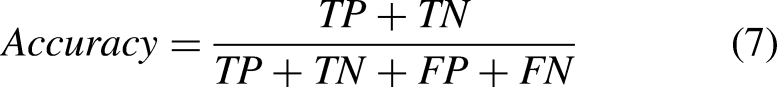



Experimental results
This section provides the outcomes of different classifiers for detecting cervical cancer. The ML models were created using Python 3.8 and a Jupyter Notebook, and the tests were performed on a system equipped with a 7th-generation Core i7 CPU and the Windows 10 operating system. The performance of the learning models was assessed using accuracy, precision, recall, and F1 score. The hardware and software specifications utilized in the experiment are described in detail in Table 2.
Experimental setup for the proposed system.

RAM: Random Access Memory; CPU: central processing unit; GPU: graphics processing unit.
Results of the ML models with deleted values
The first stage of the experiments involved removing missing values from the dataset before implementing machine learning models on the modified data. The performance of the models is summarized in Table 3.
Results of the machine learning models obtained by deleting missing values from the dataset.

LR: logistic regression; DT: decision tree; RF: random forest; SGD: stochastic gradient decent; ETC: extra tree classifier; XGB: extreme gradient boosting; SVC: support vector classifier; GNB: Gaussian Naive Bayes.
The results indicate that the RF, ETC, and XGBoost classifiers achieved the highest accuracy of 71.55%, 72.98%, and 73.41%, respectively. RF also demonstrated a precision value of 79.25%, a recall value of 80.65%, and an F1 score of 80.11%, while ETC exhibited a precision value of 80.25%, a recall value of 80.25%, and an F1 score of 80.25%. Similarly, XGBoost attained a precision value of 79.85%, a recall value of 79.99%, and an F1 score of 79.91%. Conversely, LR had the weakest performance, with an accuracy of 63.47%, a precision value of 76.44%, a recall value of 78.54%, and an F1 score of 77.41%. The proposed VC(XGB+RF+ETC) ensemble model outperformed all other learning models, with an accuracy of 79.93%, a precision value of 83.36%, a recall value of 85.21%, and an F1 score of 84.67%. Overall, the performance of individual ML models using the deleted missing value data was unsatisfactory. Figure 3 illustrates a graphical representation of the ML model results with deleted missing value data, showing that LR has shown the lowest accuracy scores. Non-constant variance in data affects the performance of the model. Results also reveal that, apart from RF, ETC, XGBoost, and VC (XGB+RF+ETC), the performance of other models is average.

Results of the machine learning models obtained by deleting missing values from the dataset.
Results of ML models using KNN imputer
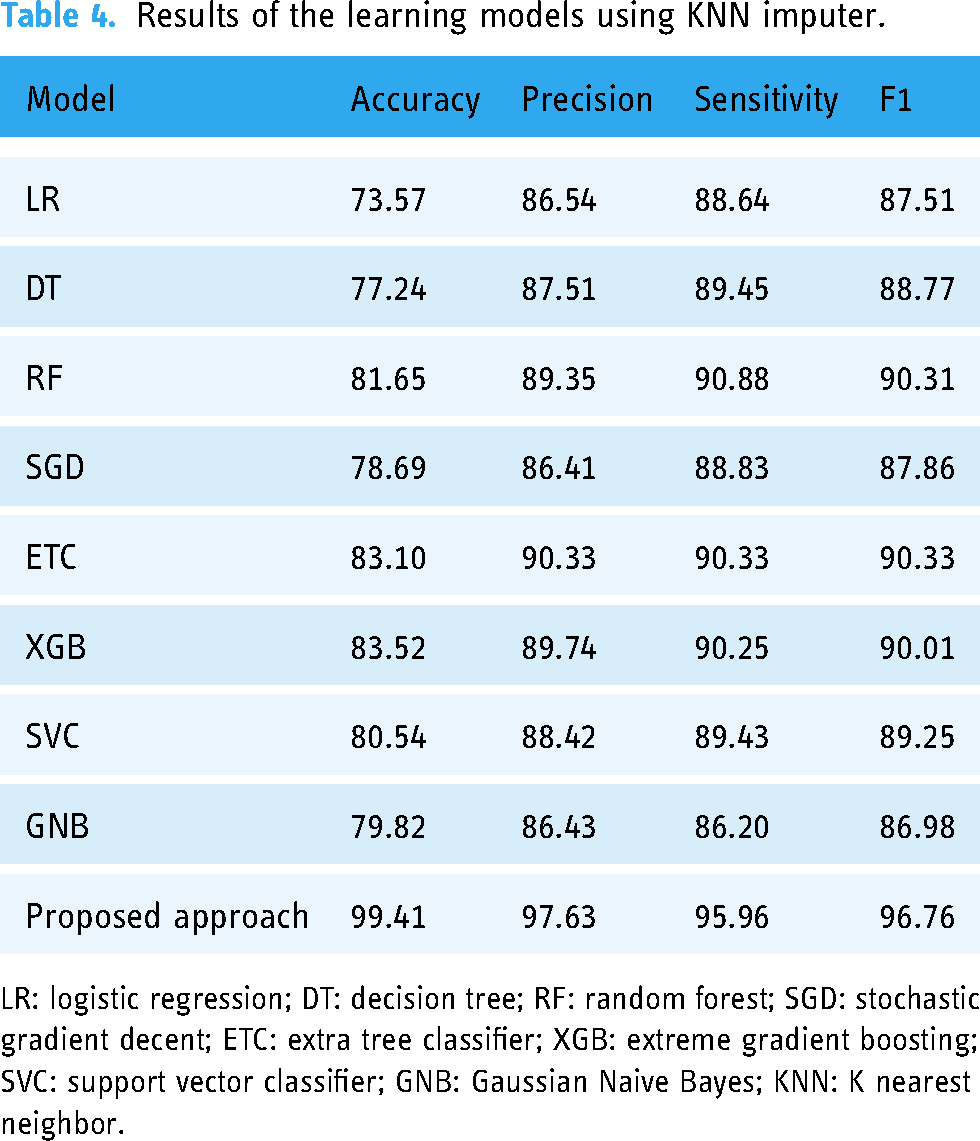
For the second set of experiments, the KNN imputer was employed to address missing values in the dataset. Upon preprocessing the data, it was observed that certain values were absent, prompting the use of the KNN imputer to fill in these gaps. The imputation was conducted using the mean of the given values and the Euclidean distance metric. The resulting dataset was subsequently utilized to train and test several ML models. Table 4 illustrates the performance of various models with the dataset complimented by the KNN imputer. It shows that LR and DT have shown the lowest results with 73.57% and 77.24% accuracy, respectively.
Results of the learning models using KNN imputer.
LR: logistic regression; DT: decision tree; RF: random forest; SGD: stochastic gradient decent; ETC: extra tree classifier; XGB: extreme gradient boosting; SVC: support vector classifier; GNB: Gaussian Naive Bayes; KNN: K nearest neighbor.
The results reveal that RF, ETC, and XGBoost attained an accuracy of 81.65%, 83.10%, and 83.52%, respectively. Results support the hypothesis that the use of the KNN imputer to fill the missing values would improve the results as the performance of ML models is elevated compared to their performance with the missing values dataset. The proposed VC (XGB+RF+ETC) ensemble model achieved an accuracy rate of 99.41%, which is the highest among all models. Moreover, the proposed ensemble model demonstrated a precision value of 97.63%, a recall value of 95.96%, and an F1 score of 96.76%. In contrast, the linear model LR had the lowest accuracy value of 73.57%. Figure 4 presents a graphical representation of the ML models’ outcomes using the KNN imputer, highlighting that utilizing the KNN imputer improves the ML models’ performance.

Results of the learning models using K nearest neighbor (KNN) imputer.
Comparison of ML models with and without KNN imputer
We assessed the effectiveness of the KNN imputer by comparing the performance of ML models with and without it. The results showed that when the KNN imputer was used in the second experiment, there was a significant improvement in the performance of the ML models, compared to using data with deleted missing values. Table 5 displays the outcomes of the ML models for both scenarios, making it easier to analyze their performance.
Accuracy comparison of the machine learning models.

LR: logistic regression; DT: decision tree; RF: random forest; SGD: stochastic gradient decent; ETC: extra tree classifier; XGB: extreme gradient boosting; SVC: support vector classifier; GNB: Gaussian Naive Bayes; KNN: K nearest neighbor.
Figure 5 presents a comparison of the ML models’ performance using the deleted missing values and the KNN imputed dataset. The graph illustrates that employing the KNN imputer enhances the individual models’ performance, resulting in a better overall performance from all the ML models.

Performance comparison of the machine learning models regarding the use of K nearest neighbor (KNN) imputer.
Limitations of the current work
This research work has some limitations in terms of the dataset. The current dataset we have used in this research work contains only 858 instances only with 36 attributes. A lot of attributes and less number of instances with missing values in that are the limitations of this work. Well for dealing with this problem, this research work implies KNN imputer. The KNN imputer performs quite well in this case but still, it has some limitations when applies to other types of datasets. The limitations are:
KNN imputer considers all available features to find nearest neighbors. However, if some features are irrelevant or noisy, they may introduce bias into the imputations. Careful feature selection or preprocessing may be necessary to mitigate this issue. KNN imputer is primarily designed for continuous or numeric variables. It may not perform well when dealing with categorical variables or mixed-type data. Additional preprocessing steps, such as converting categorical variables to numerical representations, may be required. KNN imputer can be computationally expensive, especially when dealing with large datasets or high-dimensional data. As the number of samples and features increases, the time required to find the nearest neighbors and compute imputations can become significant. KNN imputation assumes local similarity based on the nearest neighbors. It may not capture more complex patterns or relationships that exist in the data, such as non-linear dependencies or global trends.
Results of the K-fold cross-validation
To ensure the reliability of the models, K-fold cross-validation was employed. Table 6 presents the results of five-fold cross-validation, which demonstrates that the proposed approach outperforms other models in terms of accuracy, precision, recall, and F1 score, with a small standard deviation.
Five-fold-cross-validation results for the proposed approach.

Discussion
Voting classifiers of tree-based ensembles perform well on a wide range of datasets, particularly when the dataset exhibits certain characteristics
48
:
High-dimensional data: Tree-based ensemble methods, such as RF and gradient boosting, can handle high-dimensional datasets effectively. They can capture complex interactions and patterns between features, making them suitable for datasets with a large number of features. Non-linear relationships: Tree-based ensembles are capable of capturing non-linear relationships between features and the target variable. If the dataset contains non-linear relationships or interactions between variables, tree-based ensembles can often model them more accurately than linear models. Categorical or mixed-type data: Tree-based ensembles especially extra tree classifiers can handle categorical variables directly without requiring explicit encoding or transformation. They can also handle a mixture of categorical and numerical variables, making them suitable for datasets with diverse data types. Robustness to noise or outliers: Tree-based ensembles are relatively robust to noisy or outlier data points. The ensemble’s aggregation of multiple trees can reduce the impact of individual noisy observations, leading to more robust predictions. Imbalanced datasets: Tree-based ensembles can handle class imbalance reasonably well like in this scenario. By adjusting class weights or using specific techniques like balanced subsampling or boosting, tree-based ensembles can mitigate the impact of class imbalance and produce more balanced predictions. Non-parametric nature: Tree-based ensembles do not assume any specific distribution or functional form in the data, making them flexible for a wide range of datasets and data distributions. They can capture complex and non-linear relationships without imposing strong assumptions on the underlying data.
It’s important to note that the performance of the voting classifier of tree-based ensembles can still depend on the specific dataset, the quality of the data, and the appropriate tuning of hyperparameters.
Performance comparison with existing studies
In order to show the performance of the proposed model over previous state-of-the-art models, results are compared with existing models. For this purpose, this research work selects the three most related research works having state-of-the-art models designed for improving accuracy. For instance, Abdoh et al. 45 used the SMOTE features with the ML model RF for cervical cancer detection and achieved an accuracy score of 96.06%. The study by Ijaz et al. 46 used the DBSCAN with SMOTE-Tomek and RF as a ML model to achieve the highest accuracy score of 97.72%. Similarly, RFE (recursive feature elimination) is used with SMOTE-Tomek by Tanimu et al. 47 thereby reporting an accuracy score of 98.81%. Table 7 shows the performance comparison between the proposed and existing studies. Results exhibit a better performance of the proposed model.
Performance comparison with state-of-the-art studies.

SMOTE: synthetic minority oversampling technique; DT: decision tree; RF: random forest; RFE: recursive feature elimination; ETC: extra tree classifier; XGB: extreme gradient boosting; PCA: principal component analysis; XGB: extreme gradient boosting; KNN: K nearest neighbor.
Conclusions
In recent years, cervical cancer is considered the leading cause of premature mortality among women. The developing countries cover the major portion (almost 85%) of this deadliest disease according to WHO report. An early diagnosis and timely treatment could greatly help to reduce the fatality rate of cervical cancer. In this regard, the use of ML approaches is found to provide higher detection accuracy. This research work proposed a framework that consists of two portions for accurately diagnosing cervical cancer in patients. The first step is to normalize the dataset by using the KNN imputer technique and the second part consists of the usage of the stacked ensemble voting classifier (XGB+RF+ETC) model. The results with high accuracy of 99.41% reveal that the use of ensemble models can provide a reliable solution for the early detection of cervical cancer. The comparison with other state-of-the-art models also shows the superiority of the proposed model. The future work of this research work is to make a stacked ensembling of ML and DL models to further enhance the performance of the model on higher dimension datasets and provide generalized and robust results.
Supplemental Material
sj-docx-1-dhj-10.1177_20552076231203802 - Supplemental material for Cervical cancer detection using K nearest neighbor imputer and stacked ensemble learningmodel
Supplemental material, sj-docx-1-dhj-10.1177_20552076231203802 for Cervical cancer detection using K nearest neighbor imputer and stacked ensemble learningmodel by Xiaoyuan Chen, Turki Aljrees, Muhammad Umer, Oumaima Saidani, Latifah Almuqren, Olfa Mzoughi, Abid Ishaq and Imran Ashraf in DIGITAL HEALTH
Footnotes
Acknowledgement
This study is supported via funding from Prince Sattam bin Abdulaziz University project number (PSAU/2023/R/1445), and Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2023R349), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.
Contributorship
Not applicable.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Ethical approval
The dataset used in this study is made publicly available by the Hospital Universitario de Caracas in Venezuela.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study is supported via funding from Prince Sattam bin Abdulaziz University project number (PSAU/2023/R/1445), and Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2023R349), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.
Guarantor
Not applicable.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
