Abstract
Background
Recent investigations suggest an increasing prevalence of Gram-positive and antibiotic-resistant bacteria causing spontaneous bacterial peritonitis (SBP), probably related to changes in antibiotic prescription patterns, in particular more widespread and long-term use of antibiotic prophylaxis with quinolones.
Objective
The primary objective of this study was to assess potential changes in the microbiology of SBP in two patient cohorts studied at a 10-year interval. Further aims were to study prognostic factors and outcome of SBP.
Methods
A retrospective double-cohort study, including all ascitic cultures from patients with cirrhosis obtained 2003–2005 and 2013–2014, was conducted.
Results
In total 312 patients were included, 125 patients in the first and 187 patients in the second cohort. SBP was diagnosed in 132 of 840 analyzed ascitic fluid samples; 62 samples were culture positive. An increase of Gram-positive bacterial isolates was noted from 26% to 46% between cohorts (p = 0.122). The prevalence of multidrug-antibiotic–resistant pathogens increased from 25% to 32% (p = 0.350). Survival after SBP among the two cohorts was comparable.
Conclusion
This single-center study in the Netherlands found a modest but nonsignificant increase in the proportion of patients with SBP caused by Gram-positive bacteria and multidrug-antibiotic–resistant bacteria over a 10-year period. Our findings differ from reported data in other countries and suggest empiric antibiotic prophylaxis and treatment of SBP should be based on national and regional microbiological findings and resistance patterns.
Keywords
Key summary
Spontaneous bacterial peritonitis (SBP) is a common complication in patients with liver cirrhosis with a 30-day mortality of 33%. A modest but non-significant increase of Gram-positive bacteria and multidrug-antibiotic–resistant bacteria causing SBP was noted over a 10-year period in the Netherlands. There are substantial regional differences in bacterial flora and antibiotic susceptibility causing SBP. Recommendations in antibiotic prophylaxis and treatment of SBP should carefully take microbiologic geographical differences into account.
Introduction
Spontaneous bacterial peritonitis (SBP) is a common infection in patients with cirrhosis and ascites. Reportedly, this infection can be diagnosed in up to 30% of cirrhotic patients with ascites who are admitted to the hospital. SBP is associated with significant morbidity and mortality.1–4 Intestinal bacterial translocation, altered immunity and the presence of ascites are key in the development of SBP.5–7
SBP is diagnosed by a polymorphonuclear neutrophil (PMN) count in ascitic fluid equal to or greater than 250/µl. Approximately 40% of SBP episodes are culture positive.3,8,9 Numerous, in particular older, studies reported that Gram-negative enteric bacteria were involved in the majority of SBP episodes. International guidelines recommend third-generation cephalosporin as empirical treatment for SBP and quinolones for secondary prophylaxis.10,11 However, in the last decade Gram-positive bacteria and antibiotic-resistant bacteria have been increasingly found to cause SBP.3,8,9,12–14 This change in microbiology has been attributed to long-term and widespread quinolone use and increased prevalence of hospital and intensive care unit admissions. These findings have raised doubts about the currently recommended antibiotic strategy in SBP.
The prevalence of antibiotic-resistant pathogens substantially differs geographically. 15 Antibiotic consumption has been identified as the main cause for increasing rates of antibiotic resistance. The Netherlands is known for a restrictive antibiotic policy and has had the lowest antibiotic use in Europe for years.15–17 Consequently, microbiological study results in SBP in our country could differ from those observed in other countries—i.e. Spain, Greece, Germany and the United States—over time. This would mean that international guidelines for prophylaxis and treatment of SBP would need to differentiate between countries based on antibiotic resistance rates. Therefore, we investigated causative microorganisms in two patient cohorts who were hospitalized with a 10-year interval in a tertiary referral hospital in the Netherlands. In addition, we aimed to identify the patients most at risk for SBP and to evaluate the associated short- and long-term survival.
Materials and methods
All consecutive ascitic cultures performed in patients with cirrhosis between January 2003 and December 2005 (first cohort) and between January 2013 and December 2014 (second cohort) at Erasmus MC, University Medical Center, Rotterdam, were included. Demographic, clinical, biochemical and survival data from patient hospital records were retrospectively studied to evaluate the prevalence, risk factors, microbiology, and mortality of SBP. The study protocol conforms to the ethical guidelines of the 1975 Declaration of Helsinki as reflected in an approval by the ethical review board of the Erasmus MC in February 2017.
The diagnosis of cirrhosis was based on liver histology or a combination of clinical, biochemical, and radiologic findings. 18 SBP was defined as a PMN count equal to or greater than 250/µl in ascites without evidence of an intra-abdominal source of infection.10,11 All ascites samples obtained during the two study periods were studied, implying that, if applicable, multiple samples per patient were taken into account. However, only the first positive culture per SBP episode, and thus one culture per SBP episode, was included in the analysis. Multidrug-antibiotic–resistant organisms (MDR) were defined, according to international guidelines, as an acquired resistance to at least three antibiotic classes. 19 Nosocomial acquisition was defined as SBP diagnosed at least 48 hours after hospital admission. During the study periods, the standard antibiotic prophylaxis for SBP was norfloxacin 400 mg daily.10,11,20 The primary choice for empirical antibiotic treatment for SBP was ceftriaxone 2000 mg daily for five to seven days and the secondary choice was amoxicillin/clavulanic acid 1000/200 mg every eight hours for five to seven days, according to international guidelines.10,11,20
All ascitic fluid samples were routinely analyzed in the central clinical laboratory with automated determination of the white blood cell count with differential. In addition, at least 10 ml of ascitic fluid was inoculated at bedside under sterile conditions in aerobic and anaerobic blood culture bottles (Bactec®) for culture in the central medical microbiology laboratory. For identification of positive cultures, the ascitic fluid was plated on agar and current identification methods were used. Susceptibility was determined with the VITEK® 2 system (VITEK AMS; bioMerieux Vitek Systems Inc, Hazelwood, MO, USA). Cultures collected until 2013 were called resistant using Clinical and Laboratory Standards Institute criteria; later cultures were called resistant using European Committee on Antimicrobial Susceptibility Testing criteria 2013.
Statistical analysis
A mean and standard deviation (SD) was computed for approximately normally distributed variables and compared using the Student’s t test. Non-normally distributed continuous variables were summarized by their median and interquartile range (IQR), and compared using the Mann-Whitney ranks sum test. Categorical variables were expressed with percentages and compared using the Chi-square test. A two-sided p value <0.05 was considered significant. Patients were followed up to a maximum of one year. This time frame was chosen based on the severity of decompensated advanced chronic liver disease. Survival was analyzed using the Kaplan-Meier method. The actual 30-day mortality and one-year mortality was calculated after the first ascitic fluid analysis; both liver transplantation and death were considered as events. The survival rates were compared using log-rank test. Univariable and multivariable Cox’s proportional hazard analyses were carried out to identify independent predictors for 30-day mortality and one-year mortality after SBP. The variables selected for univariable analysis were based on previous studies: gender, age, etiology of liver disease, community- or nosocomial-acquired SBP, positive microbial ascites culture, causative microorganism, antibiotic susceptibility, use of antibiotic prophylaxis, use of immunosuppressant drug, hepatocellular carcinoma (HCC), model for end-stage liver disease (MELD) score, albumin in serum, platelets in serum, and protein in ascites at time of ascites analyses. Variables with a p value of <0.10 in univariate analysis were included in the multivariate Cox’s proportional hazards model.
Results
In the first (2003–2005) cohort of 125 patients, 343 ascitic fluid samples were obtained for analysis. In the second (2013–2014) cohort of 187 patients, 497 samples were obtained. The diagnosis of SBP was established in 132 of the total 840 (16%) ascitic fluid samples in 95 patients (Figure 1).
Flowchart of study population.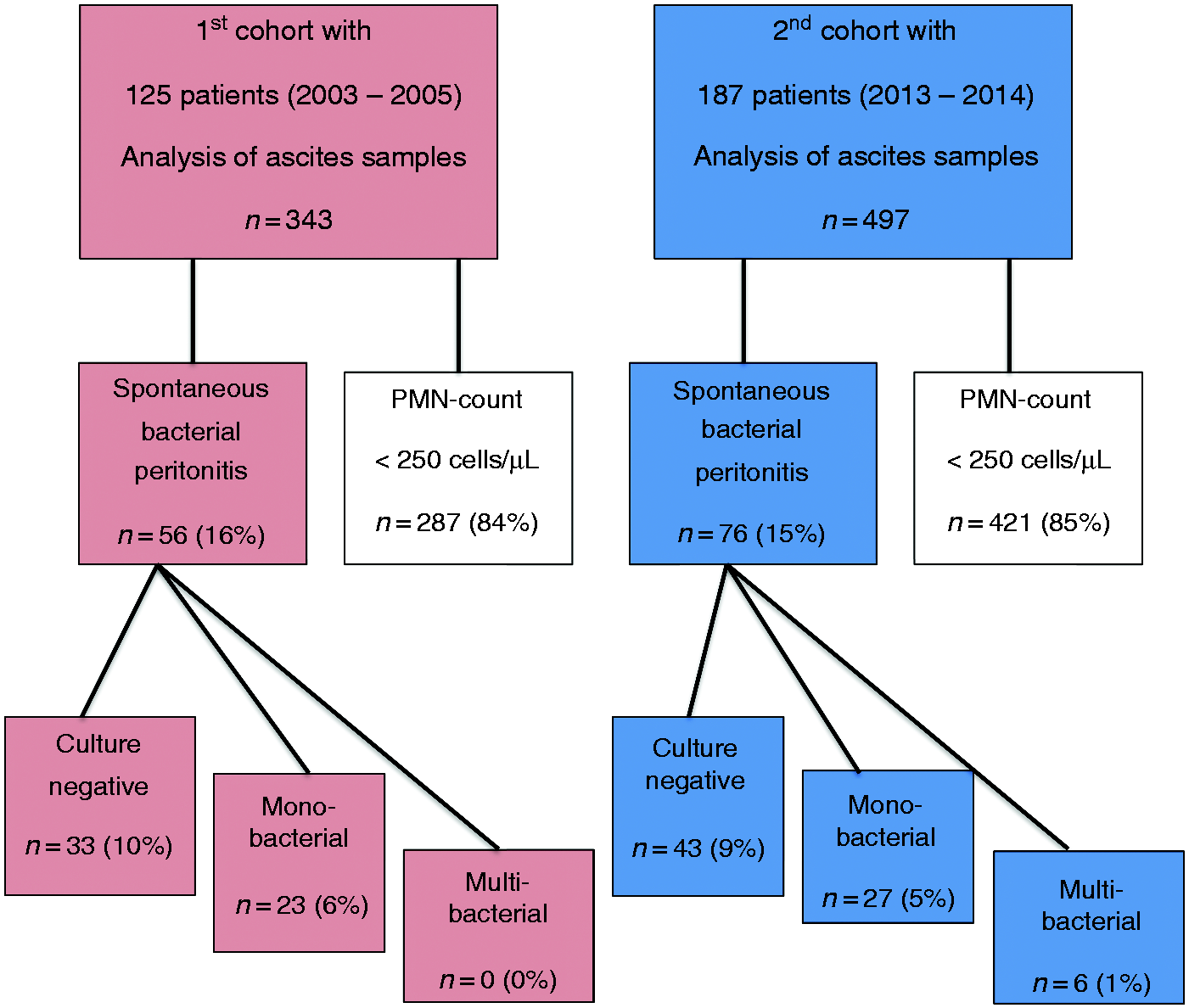
The total study population included 197 men and 115 women with a mean age of 56 years (±12) and a mean MELD score of 19 (±8). Norfloxacin was used by 12% and 10% of patients at the time of paracentesis in the first and second cohort, respectively (p = 0.638).
Clinical characteristics of the study population with and without SBP.
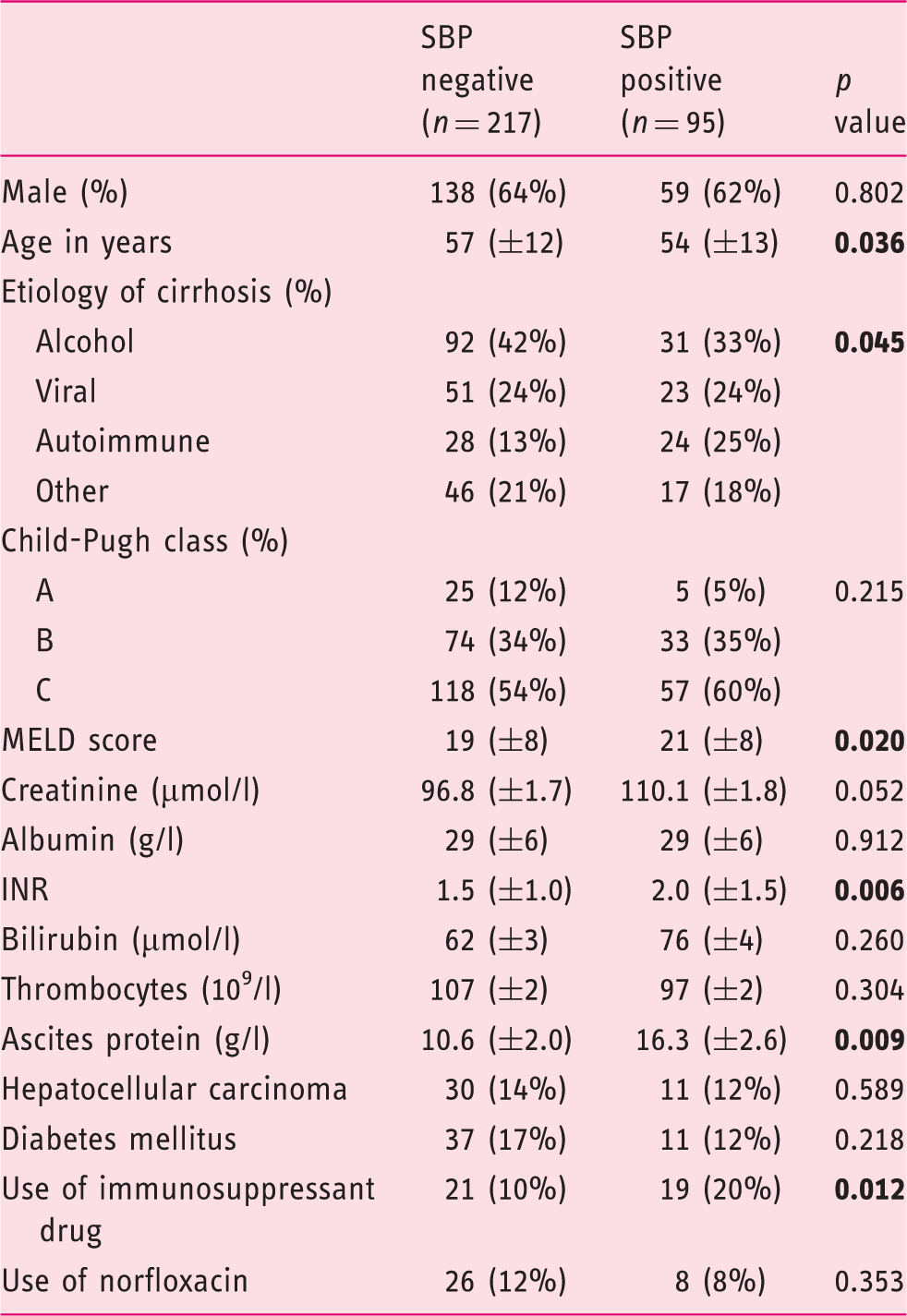
SBP: spontaneous bacterial peritonitis; MELD: model for end-stage liver disease; INR: international normalized ratio. Significant p values (<0.050) are highlighted in boldface.
Microbiology
Microbiological findings in two cohorts of patients; 62 organisms were identified in 56 episodes of culture-positive spontaneous bacterial peritonitis.
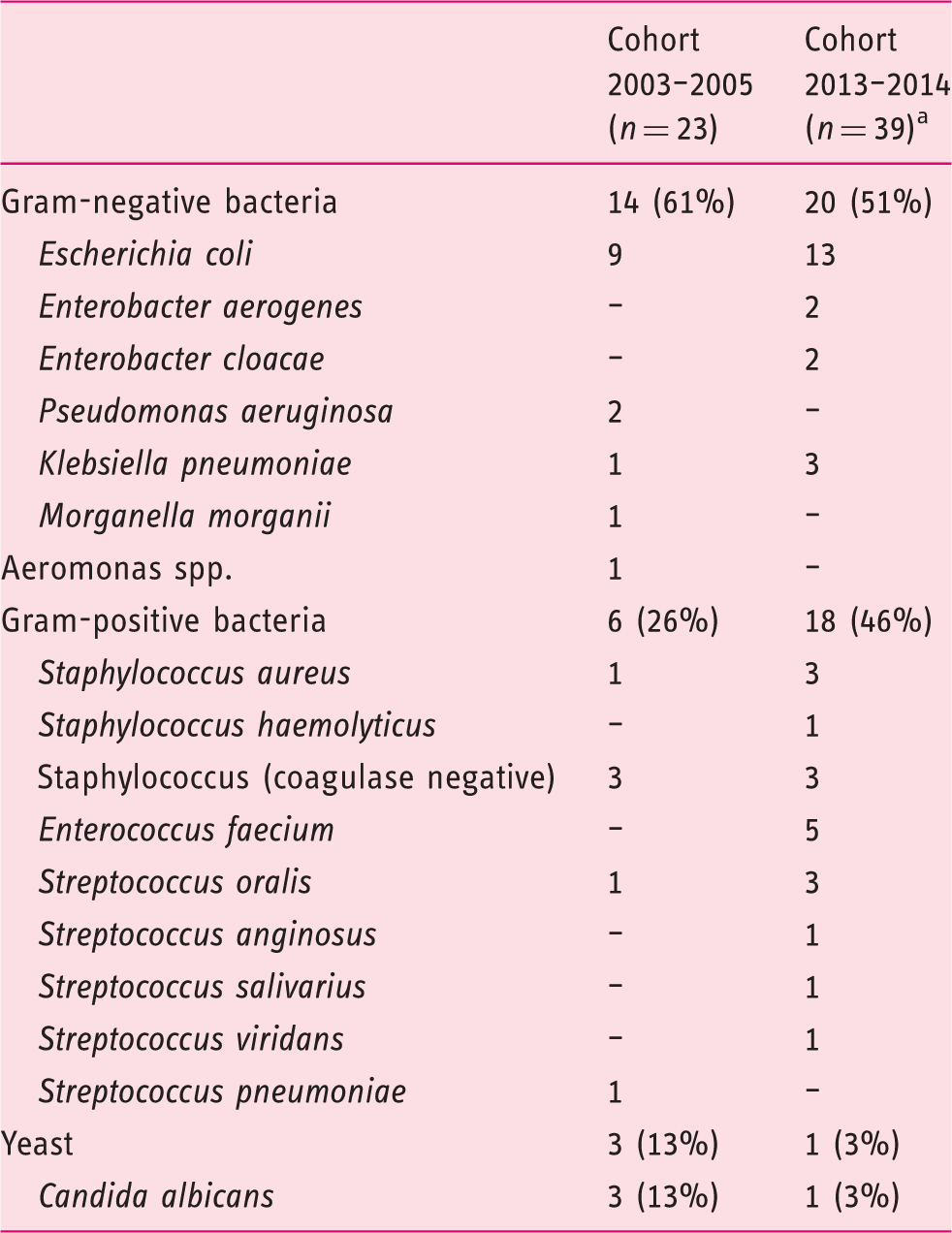
Including six cultures showing two microorganisms.
Antibiotic susceptibility patterns
Antimicrobial susceptibility patterns of bacteria from culture-positive spontaneous bacterial peritonitis (SBP).
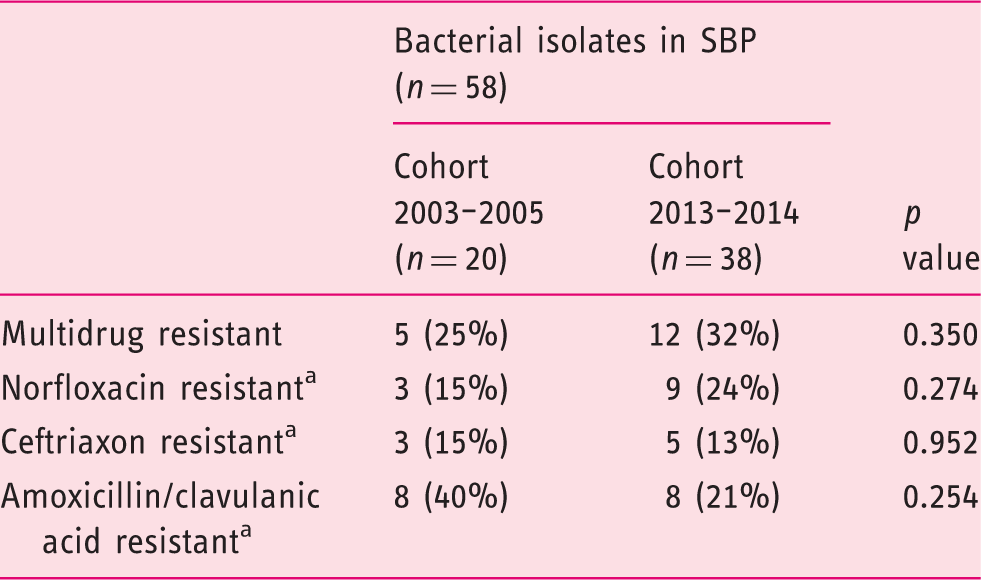
Intrinsically and/or acquired antimicrobial resistance.
Three of the 20 (15%) and six of the 38 (16%) isolated organisms were norfloxacin resistant, in the first and second cohort, respectively (p = 0.274).
Analysis with respect to ceftriaxone showed that in the first cohort 3/20 (15%) of bacterial isolates were resistant to this agent as compared to 5/38 (13%) in the second cohort. Amoxicillin/clavulanic acid-resistance was slightly more prevalent. In the first cohort 6/20 (30%) organisms were found to be resistant versus 8/38 (21%) in the second cohort. The frequency of ceftriaxone and amoxicillin/clavulanic acid-resistant organisms did not differ significantly between the cohorts (p = 0.952 and p = 0.254, respectively) (Table 3).
Survival
At one year, 153 patients had died (49%) and 48 patients had received a liver transplant (15%). Furthermore, 83 patients (27%) were alive after one year, while 28 patients (9%) were lost to follow-up. The median follow-up time of the patients not reaching the endpoint of death or liver transplantation was 365 days (IQR 12 days). The survival of patients with SBP did not differ significantly between the cohorts (log-rank p = 0.442), nor did survival differ significantly between the cohorts for patients without SBP (log-rank p = 0.216).
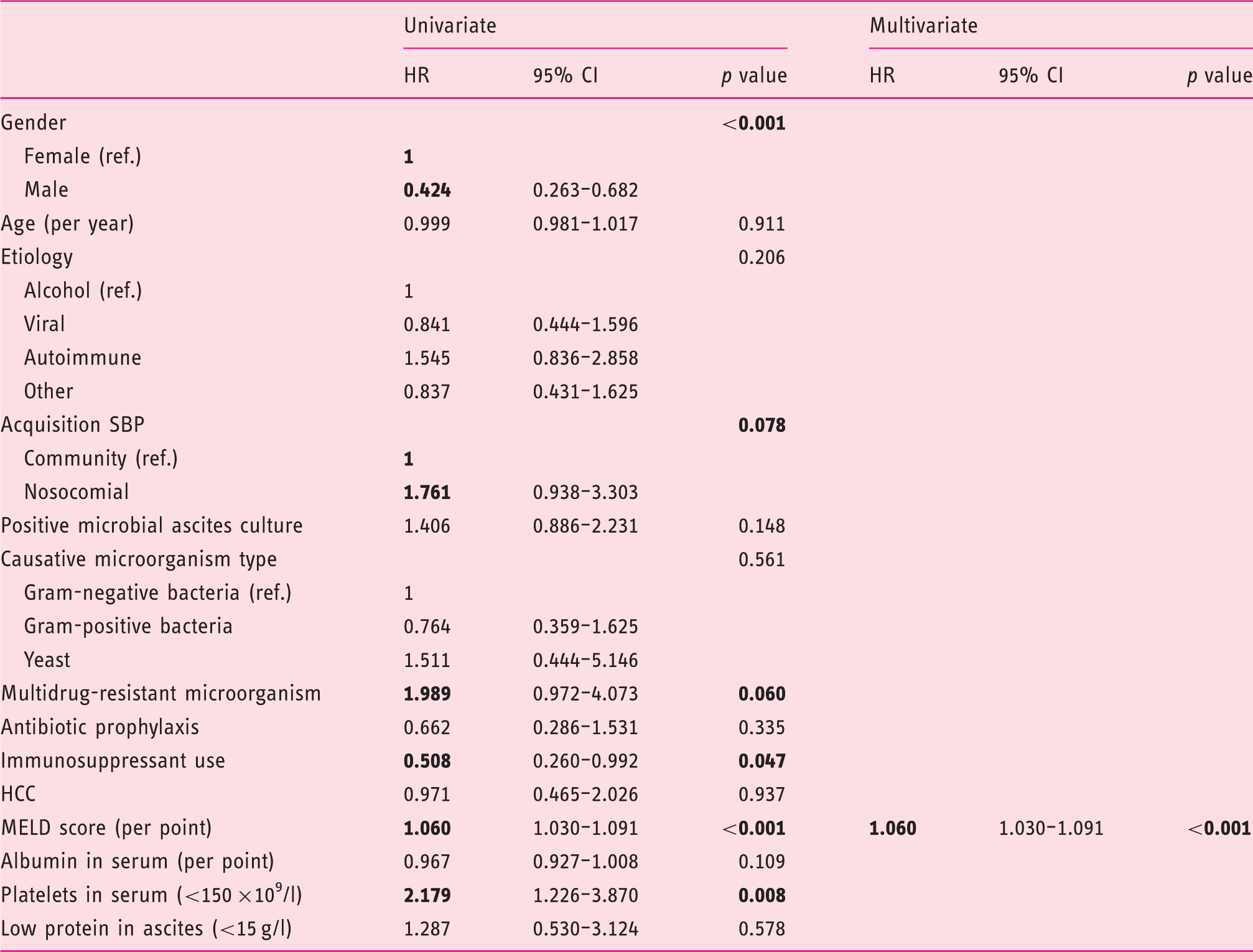
The median survival after the first ascites analyses was 168 days for SBP-negative patients and 77 days for SBP-positive patients (log-rank p = 0.001) (Figure 2). The 30-day mortality rate was 33% (32/95) for patients with SBP compared to 25% (55/217) for patients without SBP (Figure 2). Univariable and multivariable Cox-regression analyses were carried out to identify risk factors for 30-day mortality and one-year mortality after SBP. MELD score was the only independent predictive factor for 30-day mortality (hazard ratio (HR) 1.106 per point, 95% CI 1.061–1.154, p < 0.001) and one-year mortality (HR 1.060 per point, 95% CI 1.030–1.091, p < 0.001) (Table 4).
One-year mortality after first ascites analysis. Spontaneous bacterial peritonitis (SBP)-negative patients (solid line) have a median survival of 168 days and SBP-positive patients (dotted line) of 77 days (log-rank p = 0.001). Demographic and clinical factors after SBP in 95 patients predicting one-year mortality using Cox-regression analysis. SBP: spontaneous bacterial peritonitis; MELD: model for end-stage liver disease; HCC: hepatocellular carcinoma; HR: hazard ratio; CI: confidence interval. Significant p values in univariable analysis (<0.100) and in multivariable analysis (<0.050) are highlighted in boldface.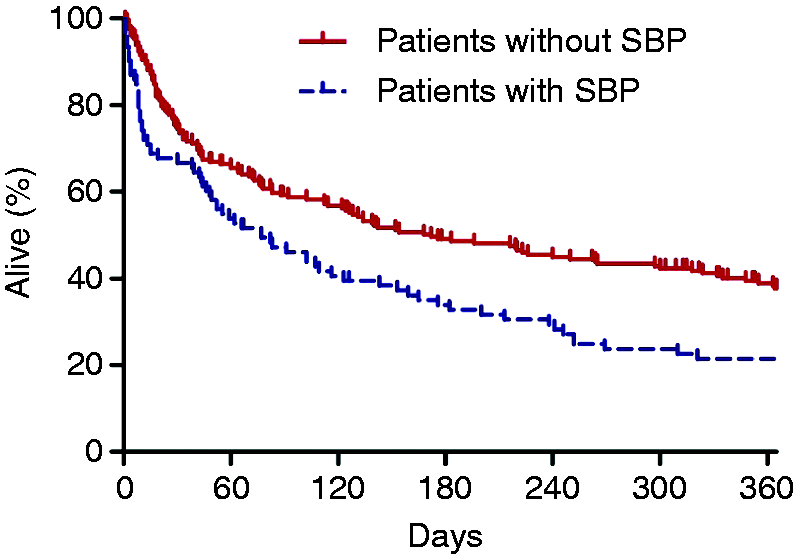
Discussion
In this single-center study in the Netherlands, the microbiological characteristics and antibiotic susceptibility patterns of organisms causing SBP in liver cirrhosis patients were compared between the periods 2003–2005 and 2013–2014. No significant increase in Gram-positive bacteria was observed and Gram-negative bacteria remained the primary cause. Bacteria resistant to empirical treatment with third-generation cephalosporin accounted for 13%–15% of all causative pathogens.
There was no evidence that mortality was influenced by causative microorganisms, antibiotic susceptibility, use of prophylactic antibiotics, intensive care admission or a nosocomial acquisition of SBP. The main predictors for mortality were age, MELD score, and platelet count.
Our most important finding is that in the era of quinolone prophylaxis for SBP, we cannot confirm observations made elsewhere regarding a significant increase in Gram-positive and MDR organisms causing SBP.3,8,9,12,13 Although in the Netherlands antibiotics are used prudently, a rise in quinolone use in the last decades has been described.21,22
Third-generation cephalosporin may poorly cover the causative pathogens in SBP, with reported antibiotic resistance rates ranging from 57% to 69%.9,23,24 However, our results show a susceptibility rate of 85–87%. We hypothesize this difference can be most likely attributed to different national antibiotic policies. High consumption of antibiotics has been related to higher rates of antibiotic resistance. 15 The Netherlands has always had a restrictive national policy regarding antibiotic prescription and a conservative approach toward the prescription of new broad-spectrum antibiotic agents.15,16,25
The microbiology and susceptibility patterns’ differences can be hospital and region dependent, implying difficulties with recommending antibiotic treatment and prophylaxis in international guidelines. The results of this study in the Netherlands do not support the need to revise guidelines as previously proposed.3,23 Empirical antibiotic treatment should be based on known regional and national differences of antibiotic resistance patterns.
A previous study from our institution on the microbiology of SBP in the period 1987–1991, before the implementation of long-term quinolone prophylaxis in the relevant patient population, reported that causative pathogens were isolated in 25 of 31 SBP episodes. 26 Gram-negative bacteria were detected in 60% of the episodes and Gram-positive bacteria in 40% of the episodes. Despite the small sample size, the proportions of causative pathogens seem comparable to those identified in the cohorts reported here.
Although optimizations in bacterial culture techniques have been implemented, an organism was isolated in a minority of all SBP episodes (40%). This is a stable percentage over the last decade and similar proportions have been documented in other studies.3,8,9 It may be expected that, with technologies arising from bacterial DNA detection and microbiome studies, more causative pathogens can be identified rapidly in the future for targeted antibiotic therapy.
There are a few limitations of the study regarding methodology. This study was designed as a retrospective, double-cohort study, which implied some laboratory and clinical data were missing. For instance, we have no information about short-term antibiotic use prescribed by general practitioners and clinicians outside the hospital, but it is not to be expected that this would differ significantly between the cohorts. Furthermore, multiple tests have been performed that may lead to an increased risk of finding spurious significant results and results should be interpreted while keeping this in mind. Prospective cohort studies comparing multiple regions during a large time frame could provide more insight as to the microbiology of SBP in diverse regions over time.
In this study we show that the microbiology of pathogens causing SBP did not change significantly in our center over the last decade. These findings suggest that guidelines with respect to antibiotic prophylaxis and treatment of SBP should carefully take into account potential national and regional differences in the microorganisms causing SBP and antibiotic resistance patterns.
Footnotes
Acknowledgements
We would like to thank Joyce Kuiper and Dirk Jan Bac for their previous work, in particular the acquisition of data relevant to the present study.
Author contributions include the following: R.C. Oey: study concept and design, acquisition of data, analysis and interpretation of data, critical revision of the manuscript for important intellectual content, and finalizing the article; R.A. de Man: study concept and design, analysis and interpretation of data, critical revision of the manuscript for important intellectual content, and finalizing the article; N.S. Erler: statistical analysis, critical revision of the manuscript for important intellectual content, and approval of the final article; A. Verbon: interpretation of data, critical revision of the manuscript for important intellectual content, and approval of the final article; H.R. van Buuren: study concept and design, analysis and interpretation of data, critical revision of the manuscript for important intellectual content, and finalizing the article.
Ethics approval
The study protocol conforms to the ethical guidelines of the 1975 Declaration of Helsinki as reflected in an approval by the ethical review board of the Erasmus MC in February 2017.
Informed consent
The ethical review board determined that written informed consent was not necessary to obtain from each patient included in this study due to its retrospective design.
Declaration of conflicting interests
None declared.
Funding
This study was sponsored by the Foundation for Liver and Gastrointestinal Research Rotterdam (SLO).
