Abstract
Background
Eosinophilic esophagitis (EoE) is an antigen-driven disease mediated by an abnormal immune Th2 response.
Objective
The objective of this article is to investigate genes associated with regulating immune responses leading to disease susceptibility.
Methods
Twenty-seven tag single nucleotide polymorphisms (tSNPs) selected in five candidate genes (
Results
Five tSNPs located in the
Conclusion
Keywords
Key summary
Eosinophilic esophagitis (EoE) results from an abnormal immune response to food and airborne allergens, which are characterized by a predominantly Th2-type inflammatory response. Thymic stromal lymphopoietin ( Primary esophageal epithelial cells express We have replicated the association of the This study describes the The Our results suggest that there is a non-synergic effect between
Introduction
Eosinophilic esophagitis (EoE) is a chronic inflammatory condition that is clinically characterized by symptoms of esophageal dysfunction and histologically characterized by eosinophil-predominant inflammation.
1
The current annual incidence of EoE is estimated to be 7.2 new cases per 100,000 people, and the prevalence in males is approximately 2.5-fold higher than in females.
2
EoE results from an abnormal immune response to food and airborne allergens, and most patients with EoE also exhibit several atopic diatheses, including allergic rhinitis, bronchial asthma and eczema, which are the most common.
3
EoE patients also have a high rate of immunoglobulin (Ig)E-mediated food allergy, which is estimated to be 10 times higher compared to the general population, which suggests that the pathogenesis of EoE and atopy involve common processes4,5 characterized by a predominantly T helper 2 (Th2)-type inflammatory response.
6
EoE today represents one of the most relevant topics in gastroenterology and allergology, but after 20 years of research on the causes of this disorder, a full explanation of the pathophysiology of EoE has yet to be determined. The familiar associations of cases reveal common genetic predispositions that underlie the increased concordance of EoE in monozygotic (58%) and dizygotic twins (36%) compared to siblings (2.4%).
7
Research on candidate-gene identification and genome-wide association studies (GWAS) have identified multiple genes that are likely contributing to the development of EoE,8–10 among which thymic stromal lymphopoietin (
However, the same studies of a familiar association and others assessing the effects of early life exposures on the risk of EoE also demonstrate the substantial influence that the environment (including exposure to dust mites and microorganisms, foods consumed, antibiotic use and even the esophageal microbiota) 14 has through epigenetic mechanisms. Changes in the esophageal microbiota linked to EoE are arising as a novel but potentially essential causative factor in triggering and maintaining EoE 15 with Toll-like receptors (TLRs) as the interaction point between bacteria and mucosal immunity. In fact, the activation and maturation of antigen-presenting cells (APCs) and regulatory T cells (Tregs) depend on TLR-mediated signaling. TLRs are transmembrane pattern recognition receptors located in intestinal epithelial cells and basal lamina that respond to microbial signals and distinguish different types of pathogens from commensal bacteria. 16 Increasing evidence points toward a relevant role for genetic polymorphisms affecting TLR function in inappropriate inflammatory responses. 16 TLR2, 4, 6, 7, 8 and 9 are significantly associated with allergic asthma and atopy,17,18 and TLR4-dependent signals provided by the intestinal commensal flora inhibit the development of allergic responses to food antigens;18,19 TLR3 can signal inflammatory responses in human epithelial cells.20,21 Preliminary results have documented changes in the expression levels of several TLRs that reverse after effective dietary therapy. 22
An impairment of immune homeostasis maintained by CD4+CD25+FOXP3+Tregs, which is a subtype of T cell that expresses the interleukin (IL)-2 receptor alpha chain (CD25) and the transcriptional regulator forkhead box P3 (FOXP3) protein, also arises as potentially involved in EoE. Tregs are important components in the immunoregulatory suppression of T cell proliferation and function, which directly or indirectly suppresses effector cells in allergic inflammation, including eosinophils.23,24 Tregs cells are significantly increased in the esophageal tissue of EoE patients, which suggests that a negative feedback mechanism exists to regulate an inflammatory response triggered by external stimuli or allergen exposure. 25 In this sense, TLR4 is associated with the amplification of the suppressive function of Tregs and may influence allergic responses to food antigens. 18
On the other hand, polymorphisms in epithelium-specific genes have also been associated with EoE susceptibility. A loss of function caused by a single nucleotide polymorphism (SNP) in the filaggrin gene (
With this background, our research aims to analyze the role of genes associated in regulating immune responses of patients with EoE. First, we will evaluate whether the
Patients and methods
Patients
A total of 218 adult patients with EoE (170 males; mean age at diagnosis 35.4 years) and 376 healthy ethnically matched bone marrow donors were selected for this study. To avoid the confounding effects of the genetic associations investigated with other conditions, EoE patients who had other autoimmune conditions (i.e. celiac disease, diabetes mellitus and thyroid disorders) were excluded. All patients and controls were unrelated Spanish Caucasian individuals who were recruited at two Spanish hospitals: Hospital Universitario Virgen del Rocío (Sevilla) and Hospital General de Tomelloso (Ciudad Real). This study was approved by the Comité de Ética de la Investigación del Hospital Universitario Virgen del Rocío, Sevilla, on April 9, 2012 (Code 2012PI/079). The study protocol conforms to the ethical guidelines of the 1975 Declaration of Helsinki as reflected in a prior approval by the institutions' human research committee. Informed consent for the procedures was obtained from all patients included in the study.
Participants were diagnosed as having EoE if they met the following accepted criteria: 28 (i) symptoms related to esophageal dysfunction, (ii) one or more mucosal biopsies that demonstrated at least 15 eosinophils per high-powered field (hpf), (iii) persistence of eosinophilic infiltration after an eight-week trial with a proton-pump inhibitor drug at double doses, and (iv) exclusion of other potential causes of esophageal eosinophilia.
Allergic sensitization tests
To evaluate allergic comorbidities, skin prick tests (SPTs) were carried out in every EoE patient by using standard methods 29 with a panel of 17 commonly distributed aeroallergens (including mites, molds, pollen and animal dander) and 22 plant- and animal-derived food antigens (milk, egg, shrimp, squid, clam, cod fish, chicken meat, walnut, peanut, pepper, tomato, potato, onion, soy, lentil, carrot, peach, kiwi, melon, banana, wheat and rice). SPTs were performed at the flexor surface of the forearm with reactions read out after 15 to 20 minutes. SPTs were positive if the mean wheal diameter was ≥3 mm over the negative control. Saline solution and histamine solution were respectively used as negative and positive controls. Sensitization to airborne or food allergens was considered as a dichotomous variable and was exclusively based on the positivity or negativity of skin allergy testing.
SNP selection and genotyping
Genomic DNA was extracted from blood leukocytes using the QIAmp DNA Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer's recommendations and stored at –20ºC.
Tag SNPs included in this study.
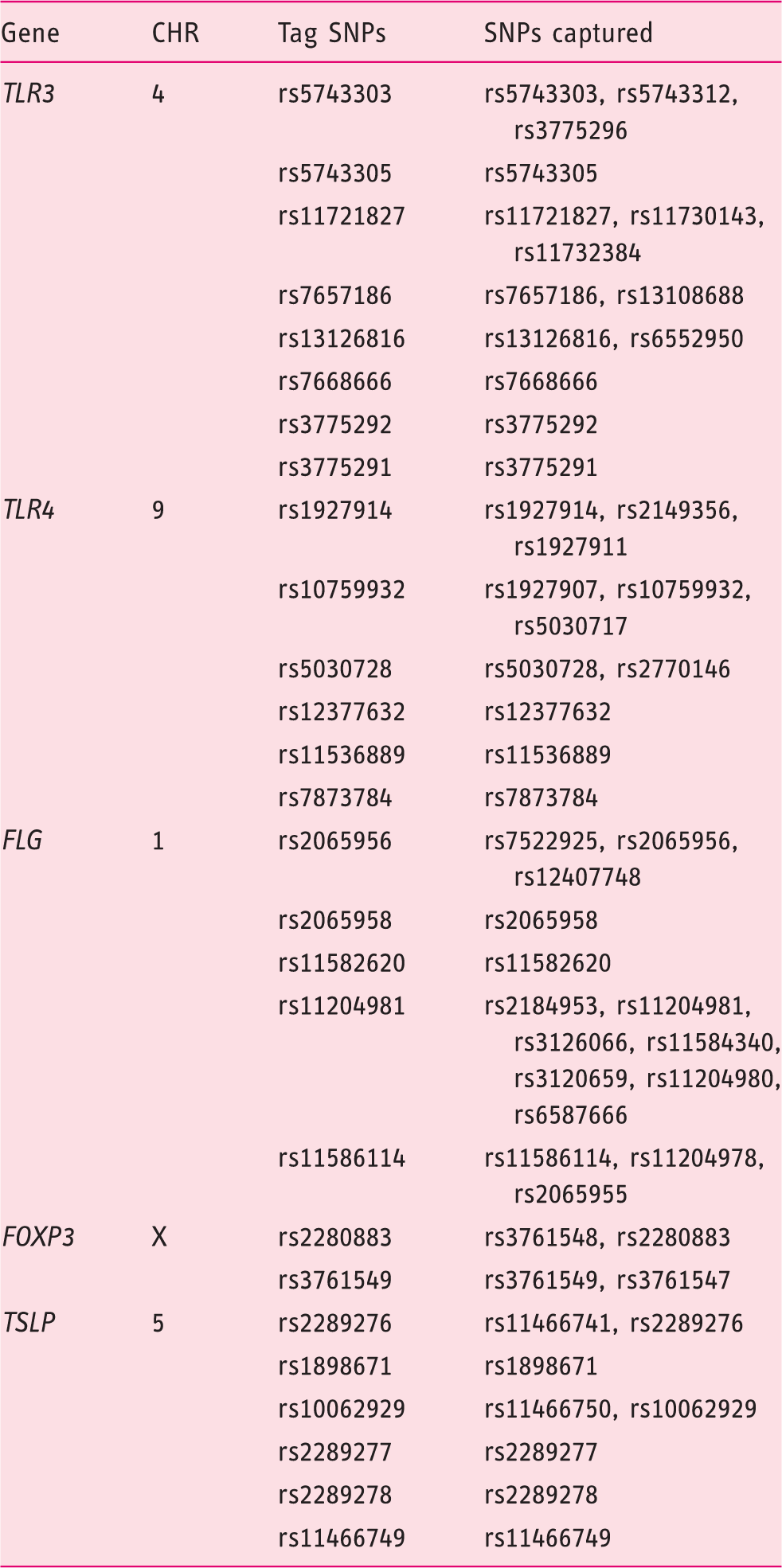
CHR: chromosome; SNP: single-nucleotide polymorphism;
Statistical analyses
Allele frequency distributions were compared using the χ2 test, and a corrected
Gene-gene interactions were evaluated using the non-parametric multifactor dimensionality reduction (MDR) method.31,32 To interpret combination effects, an entropy-based analysis was used. Interaction entropy uses information gain to gauge whether interactions between two (or more) variables that were considered independent exist. 33 Entropy is estimated for each individual attribute (i.e. main effects) and each pair-wise combination of attributes (i.e. interaction effects). Therefore, the main effects of each factor can be compared to the interaction effect to determine whether interactions are additive or non-additive. There is evidence for a synergistic interaction when a positive value is obtained combining two (or more) variables. Conversely, there are redundancy or independent effects when a negative value is obtained by loss of information. 33
Results
Aeroallergen and food sensitization, as estimated by a positive SPT result, were found in 72.22% and 61.46% of the whole series of 218 patients with EoE, respectively. The successful rate of genotyping was >98% for all SNPs included, and genotypes were unequivocally assigned for all the cases except in 11 controls. The study population was found to be in the Hardy-Weinberg equilibrium for all the polymorphisms that were analyzed (
Minor allele frequencies of the SNPs studied in the
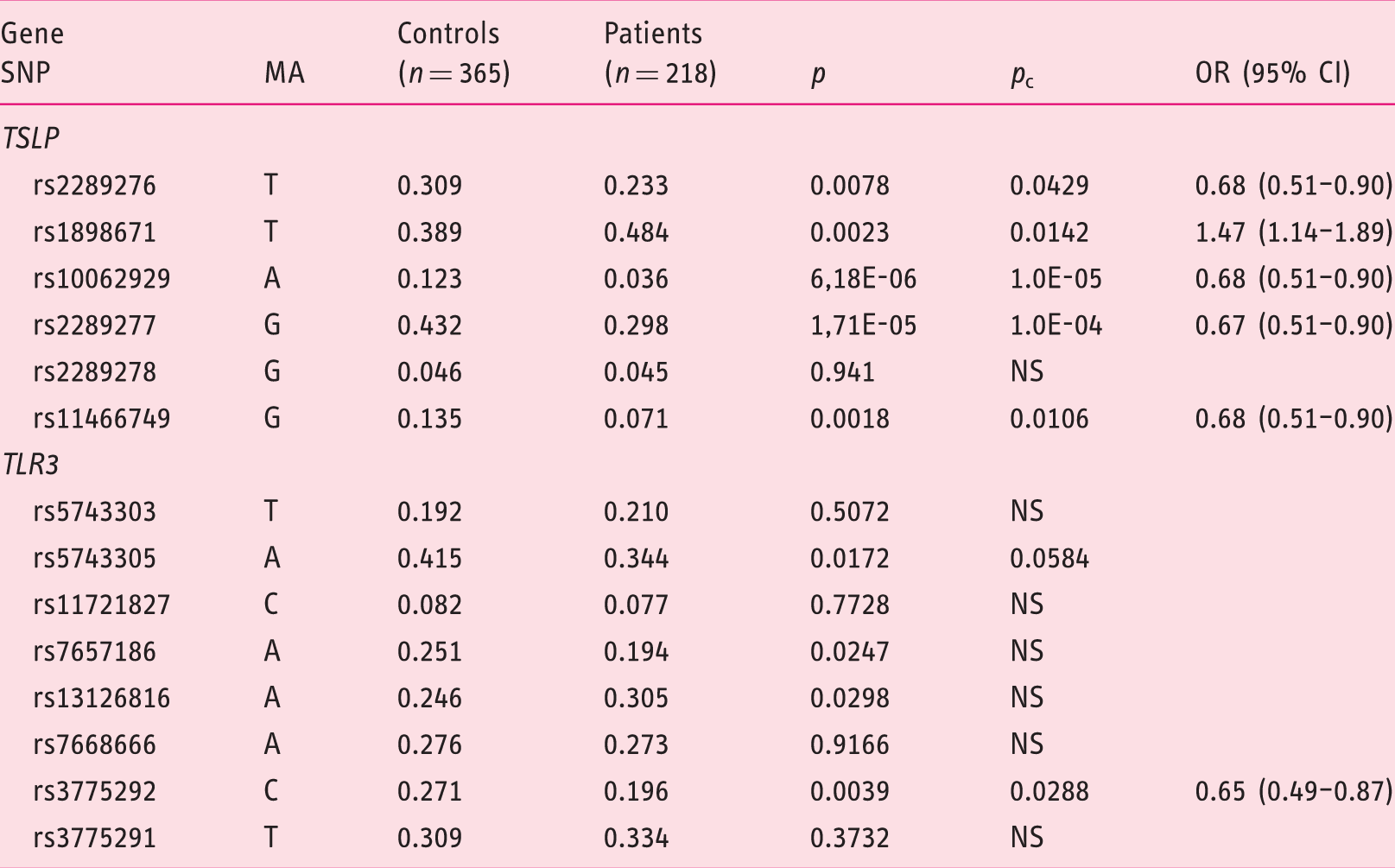
SNP: single-nucleotide polymorphism; MA: minor allele; EoE: eosinophilic esophagitis;
Distribution of allelic frequencies of the SNPs studied in the
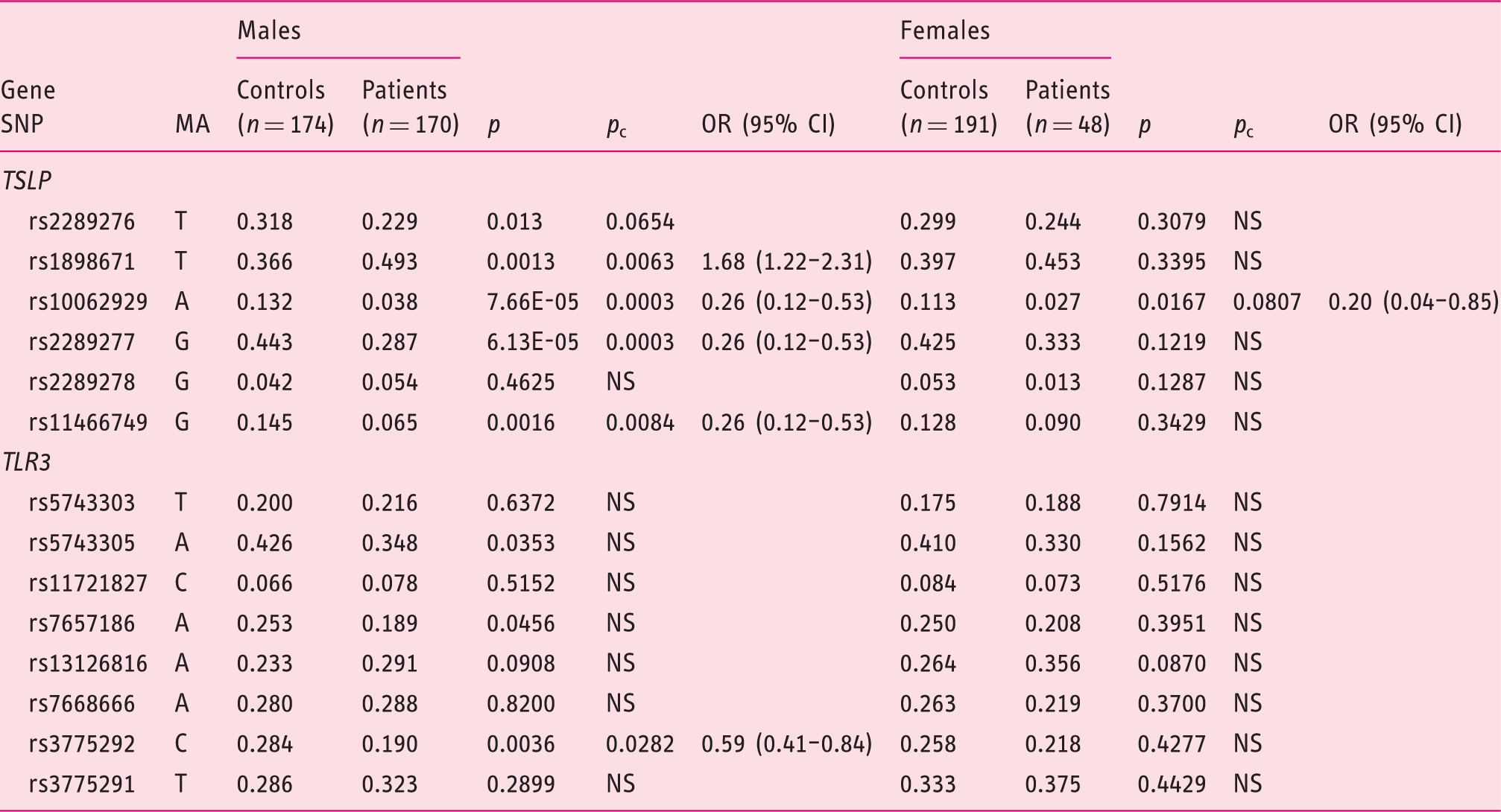
SNP: single-nucleotide polymorphism; MA: minor allele;
Distribution of genotypes of the SNP rs3775292 (
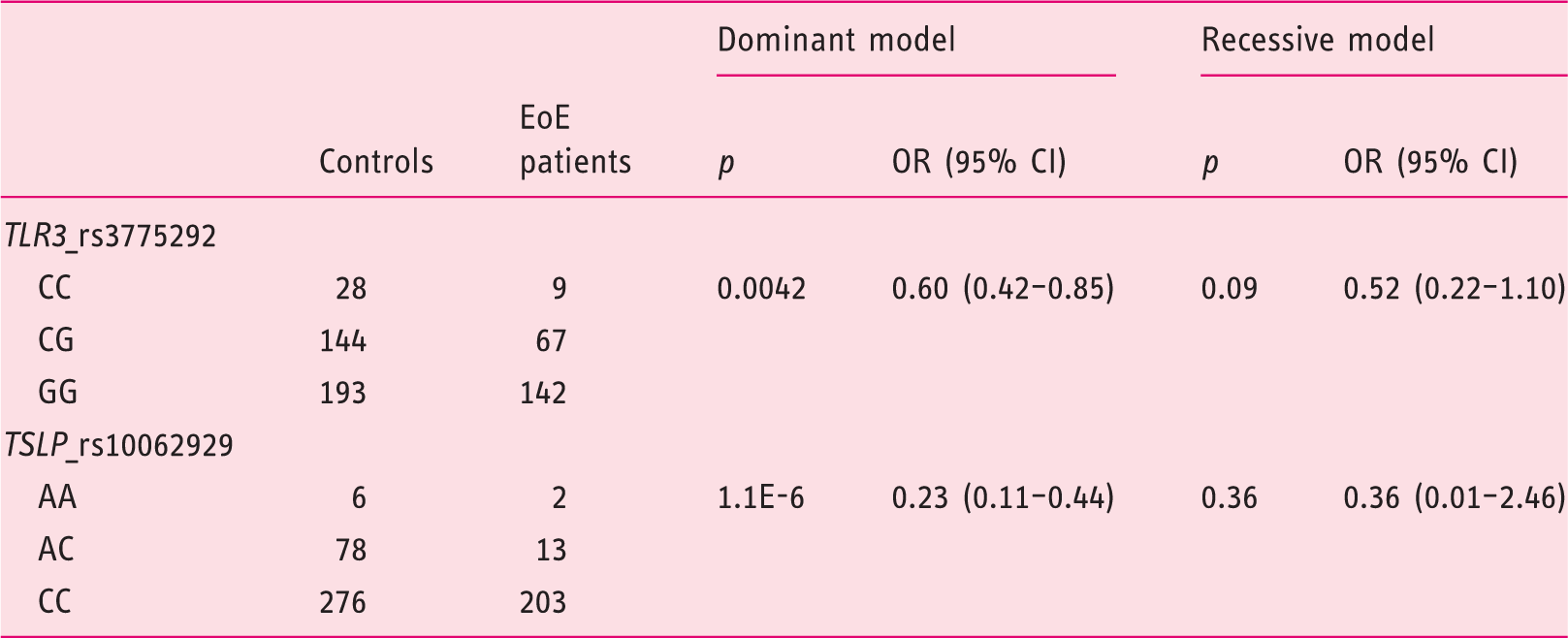
Model dominant: CC+CG vs GG (
Model recessive: CC vs CG+GG (
SNP: single-nucleotide polymorphism; EoE: eosinophilic esophagitis;
Frequencies of combinations of rs3775292 (
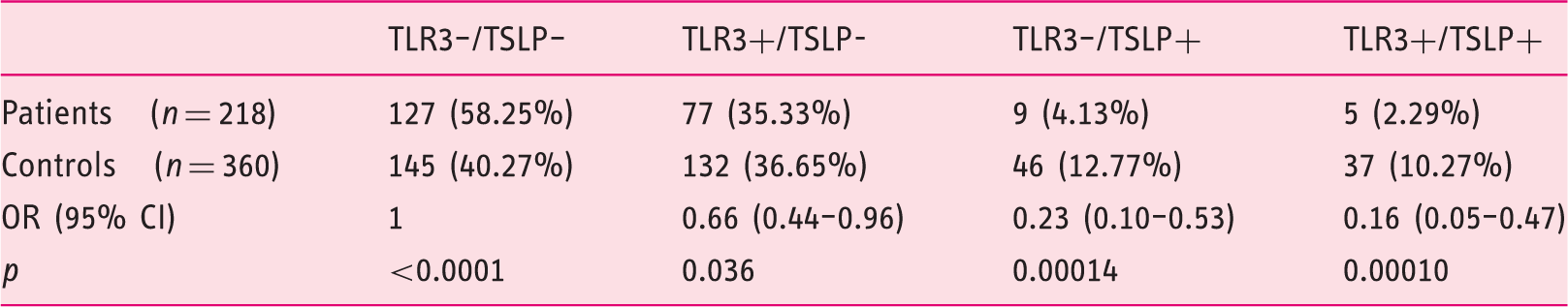
OR (95% CI) and p values were calculated using logistic regression.
EoE: eosinophilic esophagitis;
Information gain estimated for each individual locus and combination of both loci.
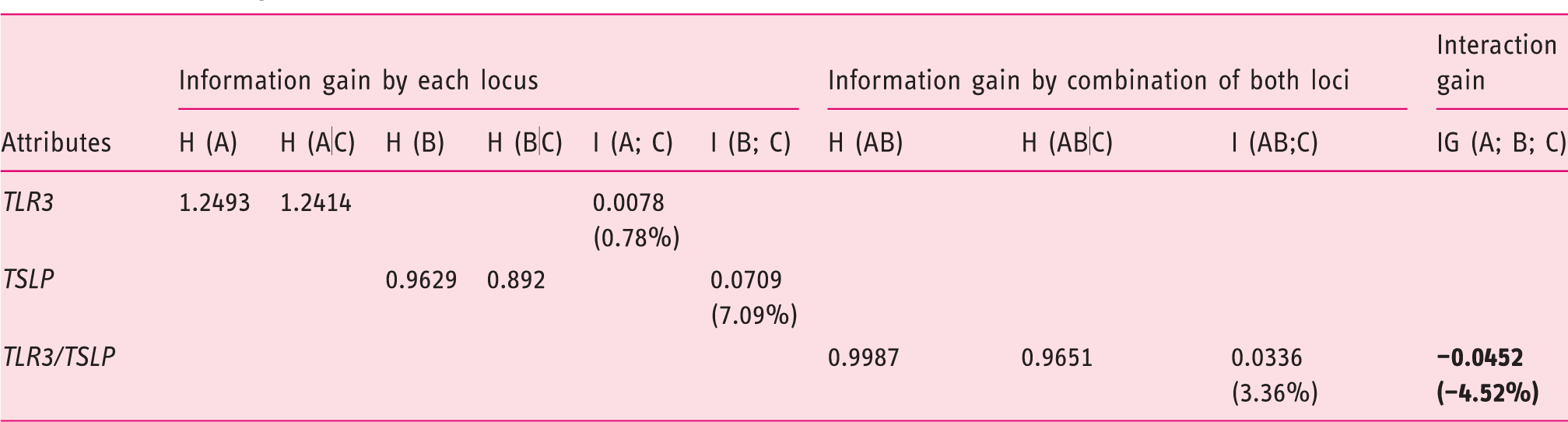

H (A), H (B), H (AB): entropy attributed to
H (A|C), H (B|C), H (AB|C): conditional entropy attributed to
I (A; C) = H (A) – H (A|C): information gain (percentage of entropy removed) by
I (B; C) = H (B) – H (B|C): information gain (percentage of entropy removed) by
I (AB; C) = H (AB) – H (AB|C): information gain (percentage of entropy removed) by the combination of both attributes (interaction effect).
IG (A; B; C) = I (A; C) + I (B; C)–I (AB; C): interaction gain by the combination of both attributes.
Minor allele frequencies of the SNPs studied in the
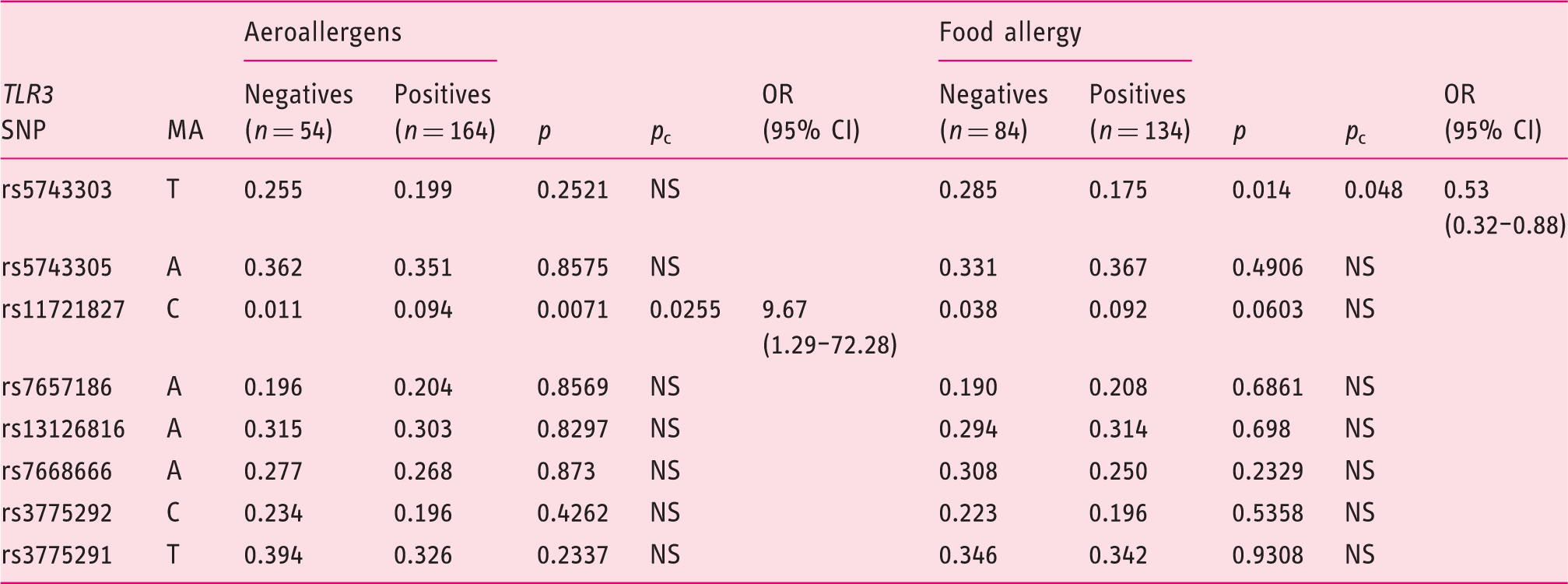
SNPs: single-nucleotide polymorphisms;
Discussion
This study describes the
Since the
The present study also examined the effects of
Taken together, we highlight a critical role for the
The strengths of our research included the analysis of a large series of patients with EoE collected at two referral centers in Spain as well as the selection of a higher number of control individuals, the development of molecular studies to evaluate SNP of the major genetic markers identified as conferring risk to EoE and key factors involved in regulating mucosal immune responses in allergies. The selection of the control group from healthy bone marrow donors is recognized as the most convenient strategy, but it has led to some controversy in the literature since the donors could not completely represent the whole population of the place that they come from. Volunteers who act as donors for organs and tissues are usually healthy people in whom several illnesses have been excluded. For this reason, donors could easily present a better health status than standard populations. Despite those issues, our control group represents the same populations from where the patients with EoE came from, so we are confident that it does not constitute a drawback for our study.
Finally, since no replication cohort was considered in our study, further studies to reproduce our findings in broader populations and to validate the findings of
In conclusion, our research reinforces the role played by
Footnotes
Acknowledgments
The author contributions are as follows: RAC, JRGL and JQ contributed to the conception and design of the research and performed the experiments, analysis and interpretation of the data; SC, AJL and JMB contributed samples, patient information and analysis and interpretation of the data. RAC, JRGL, AJL and JQ drafted the manuscript. All authors read and approved the final version.
Declaration of conflicting interests
None declared.
Funding
This work was supported by Alergosur, Sociedad Andaluza de Alergología e Inmunología Clínica, El Fondo Europeo de Desarrollo Regional (FEDER) and Plan Andaluz de Investigación (PAI CTS-0197).
Ethics approval
This study was approved by the Comité de Ética de la Investigación del Hospital Universitario Virgen del Rocío, Sevilla (Code 2012PI/079).
Informed consent
Informed consent was obtained from all individual participants included in the study. All participants included in the study consented to the publication of the data extracted from the statistical study. No individual patient data are reported.
