Abstract
Background
Patients with primary sclerosing cholangitis associated with inflammatory bowel disease (PSC-IBD) have a very high risk of developing colorectal neoplasia. Alterations in the gut microbiota and/or gut bile acids could account for the increase in this risk. However, no studies have yet investigated the net result of cholestasis and a potentially altered bile acid pool interacting with a dysbiotic gut flora in the inflamed colon of PSC-IBD.
Aim
The aim of this study was to compare the gut microbiota and stool bile acid profiles, as well as and their correlation in patients with PSC-IBD and inflammatory bowel disease alone.
Methods
Thirty patients with extensive colitis (15 with concomitant primary sclerosing cholangitis) were prospectively recruited and fresh stool samples were collected. The microbiota composition in stool was profiled using bacterial 16S rRNA sequencing. Stool bile acids were assessed by high-performance liquid chromatography tandem mass spectrometry.
Results
The total stool bile acid pool was significantly reduced in PSC-IBD. Although no major
differences were observed in the individual bile acid species in stool, their overall
combination allowed a good separation between PSC-IBD and inflammatory bowel disease.
Compared with inflammatory bowel disease alone, PSC-IBD patients demonstrated a
different gut microbiota composition with enrichment in
Conclusions
Patients with PSC-IBD had distinct microbiota and microbiota-stool bile acid correlations as compared with inflammatory bowel disease. Whether these changes are associated with, or may predispose to, an increased risk of colorectal neoplasia needs to be further clarified.
Key summary
Summary of the established knowledge on this subject
Primary sclerosing cholangitis is a chronic cholestatic disease of unknown etiology
and frequently associated with inflammatory bowel disease. Emerging evidence suggests that alterations in the microbiome may be associated
with this special phenotype. No studies have yet investigated the net result of cholestasis and a potentially
altered BA pool interacting with a dysbiotic gut flora in the inflamed colon of
PSC-IBD.
What are the significant and/or new findings of this study?
Patients with PSC-IBD presented demonstrated a different gut microbiota
composition, and specific microbiota-fecal BA correlations. Despite no significant differences in the specific BA in stool, the overall
combination of stool BA was discriminant between PSC-IBD and IBD.
Introduction
Primary sclerosing cholangitis (PSC) is a rare chronic cholestatic liver disorder of unclear aetiology. 1 It is characterized by chronic inflammation of the biliary epithelium, that eventually leads to fibrosis, resulting in multifocal strictures of the intrahepatic and extrahepatic bile ducts. It can lead to cirrhosis, and end-stage liver disease requiring orthotopic liver transplantation (OLT). 1 Furthermore, PSC is also associated with an increased risk of cholangiocarcinoma, gallbladder cancer and colorectal cancer.2,3 The strongest risk factor for having PSC is a history of inflammatory bowel disease (IBD). Around 60–80% of PSC patients will also have IBD, most commonly ulcerative colitis (UC). Ironically, despite having mild or quiescent extensive colitis, patients with PSC-IBD have the highest risk of developing colitis-associated neoplasia, which, in comparison with IBD, tends to be located preferentially in the right side of the colon.2,4 The factors that contribute to the increased risk of colorectal neoplasia in PSC remain unknown.5,6 A potential role for altered luminal concentration and/or composition of secondary bile acids (BAs) has been suggested, but never confirmed. 7 Data from basic and clinical studies have long supported the hypothesis that the intestinal microbiota may have a role in PSC pathogenesis.8–10 Recently, studies using next-generation sequencing have reported a distinct fecal or mucosal microbiota composition in PSC-IBD patients.11–18 There is a close interplay between gut flora and BA metabolism. Besides their role in nutrient absorption and lipid digestion, BAs are important signaling molecules, acting in inflammation and metabolism, through activation of BA receptors such as the G-protein-coupled BA transmembrane receptor TGR5, and the nuclear BA receptor Farnesoid X receptor (FXR). 19 BAs have antimicrobial properties, and through FXR-activation they regulate the expression of host genes whose products promote innate defence against luminal bacteria.20,21 On the other hand, BA metabolism is a property of the gastrointestinal microflora; BAs are transformed from primary BA (cholic acid (CA) and chenodeoxycholic acid (CDCA)) to secondary BA (litocholic acid (LCA) and deoxycholic acid (DCA)) by deconjugation, 7-alpha de-hydroxylation and epimerization (CDCA → ursodeoxycholic acid (UDCA)) by the gut microbiota; therefore the degree of activation of the BA receptors is also largely influenced by the gut microbiota.22–24 Nothing is known about the net result of cholestasis associated with PSC, and a potentially altered BA pool interacting with a dysbiotic gut flora in the inflamed colon of PSC-IBD patients. In this article, we have explored the BA profiles, the gut microbiota, and their correlation in PSC-IBD as compared with IBD patients.
Methods
Ethical considerations
This study was approved by the Portuguese National Committee for Data Protection and the local ethics committee. All patients signed an informed consent form.
Subjects and samples
Between October 2014–July 2015, 15 patients with PSC-IBD and 15 patients with IBD were prospectively recruited. The inclusion criteria were age greater than 18 years old, confirmed diagnosis of PSC based on histology and/or abnormal cholangiogram (Endoscopic Retrograde Cholangio-Pancreatography or Magnetic resonance cholangiopancreatography), a confirmed diagnosis of IBD by conventional endoscopic and histological criteria, and the presence of extensive colitis. Patients with a personal history of colectomy, a diagnosis of secondary sclerosing cholangitis or a history of OLT were excluded. All patients provided clinical and demographic information, and completed a semi-quantitative food frequency questionnaire (FFQ) validated for the Portuguese population. 25 Clinical activity was scored according to the Mayo score for ulcerative colitis, 26 and the Harvey-Bradshaw index for Crohn’s disease. 27 Endoscopic activity was scored according to the Mayo endoscopic score for UC 26 and the Simple Endoscopic Score for Crohn's Disease (SES-CD) for CD. 28 All study participants collected serum sample, and a stool sample for BA analysis and microbiota analysis. All PSC patients on UDCA therapy were required to stop it two weeks before specimen collection. A minimum interval of three months was required between antibiotic intake or bowel preparation (for colonoscopy) and sample collection. During colonoscopy disease severity was recorded, and biopsies for colorectal neoplasia screening were obtained, according to current guidelines.
Serum BA profiles
A fasting serum sample was obtained from each patient. Individual amidated BAs in serum (1 ml) were determined by high-performance liquid chromatography (HPLC), 29 after solid-phase extraction using Sep-Pak C18 cartridges (Waters Corp., Milford, Massachusetts, USA). 30 Only the conjugate fraction of BAs was measured in serum.
Stool BA profiles
A morning stool sample was obtained and dried to obtain a lyophilized extract. To lyophilized faecal samples weighing 1 g, 80% methanol was added. All samples were sonicated for 30 min, refluxed for two hours, and then cooled and filtered. 31 The residue was re-suspended in chloroform/methanol (1:1, v/v), refluxed for one hour, and filtered. The combined extracts were taken to dryness, and re-suspended in 10 ml methanol (MeOH). An aliquot of 1 ml was added with 2 µl of 1 mg/ml nordeoxycholic acid, and was diluted in 10 ml deionized water and deposed on a 300 mg HLB Oasis column, washed with 10 volumes deionized, 1 volume cyclohexane, and the BAs were then eluted with 5 ml MeOH and were taken to dryness and resuspended in 250 µl MeOH. Four microliters were injected on liquid chromatography coupled with tandem mass spectrometry (LC-MS/MS) as previously described. 32 Results are reported in nmol/g of dried stool for total BAs and in proportion of the median after calibration of the method, with weighted mixtures and normalization relative to the internal standard. 32 The conjugate and non-conjugated species were quantified.
BA analysis
BAs were not normally distributed according to the Shapiro-Wilk test; therefore, their distributions were compared using non-parametric tests. The relative proportion of a given BA corresponds to its concentrations divided by the total of BAs. BA results are presented as the median proportion. For example, the total primary stool BA is the sum of CA and CDCA and their respective glyco-, tauro-, and sulphoderivatives. Linear discriminant analysis (LDA) was conducted to illustrate the classification of disease groups (IBD only and PSC-IBD) using stool BAs. LDA is a dimension reduction statistical technique that looks for a combination of features (continuous variables) that maximize the separation between classes. LDA was performed using the MASS package in R software.
Stool DNA extraction
Approximately 200 mg of stool were transferred into bead tubes (MO-BIO, Carlsbad, California, USA) and homogenized using the bead-beating method. Homogenized stool samples were further processed using the Qiagen DNeasy Blood and Tissue Kit following the manufacturer’s protocol (Qiagen, Valencia, California, USA). Total DNA concentration was determined with Qubit 2.0 Fluorometer (Life Technologies, Norwalk, Connecticut, USA).
16S ribosomal RNA (rRNA) sequencing
The phylogenetically informative V3–V4 region of 16S ribosomal RNA (rRNA) gene was amplified using universal primer set 347F/803R. 33 The primers were synthesized by IDT (Integrated DNA Technology, Coralville, Iowa, USA). We used a dual-barcoding approach to label the 16S rRNA amplicons from each sample as described previously. 34 The 16S rRNA amplicons were further pooled with equal molarity and submitted for MiSeq 2 × 300 pair-end sequencing at high depth. The paired sequence readings were merged and filtered by size (>400 bp) and quality score (>Q30) using paired-end assembler for DNA sequences (PANDAseq). 35 The processed readings were further split by dual barcodes for each sample and assigned taxonomic classification using the Quantitative Insights Into Microbial Ecology (QIIME) pipeline 1.9.0. 36 Repeated measurements of the same sample were made to assess sequencing reproducibility. After processing, QIIME provided detailed OTU tables containing the microbiota composition and abundance for each individual sample.
Data analysis
First, we measured the diversity of the overall microbiota communities within or across
each sample. The overall species richness within each patient group, so-called
alpha-diversity, was measured using the Chao1 and Shannon Index on rarefied tables at 8000
sequences per sample.
37
Beta-diversity was measured using unweighted and weighted UniFrac
distance matrices on the rarefied tables. The permutational analysis of variance
(PERMANOVA) test (number of permutations = 999), was performed using QIIME command
compare_categories.py to test the overall microbiota differences between groups by PSC and
IBD status.
38
Secondly, at the taxa level, the LDA effect size (LefSe) analysis was used with default
parameters to select taxa features from phylum to genus level that were associated to PSC
status.
39
Only
features with LDA score >2.0 were kept. A Kruskal-Wallis test on the LefSe selected
differential taxa at the genus level was performed, and corresponding
Correlation networks
We calculated both Pearson’s and Spearman’s correlations between the most abundant (mean
relative abundance >0.1%) 65 genera in the gut microbiota and the stool BA levels in
PSC and non-PSC IBD. To reduce the bias in the correlation analysis due to non-normality,
we removed the variables with more than eight null value results, and removed the
measurements beyond the 5% quantile of the distribution. We computed the raw
probabilities. The
Results
Study population
Demographic and clinical characteristic of patients.
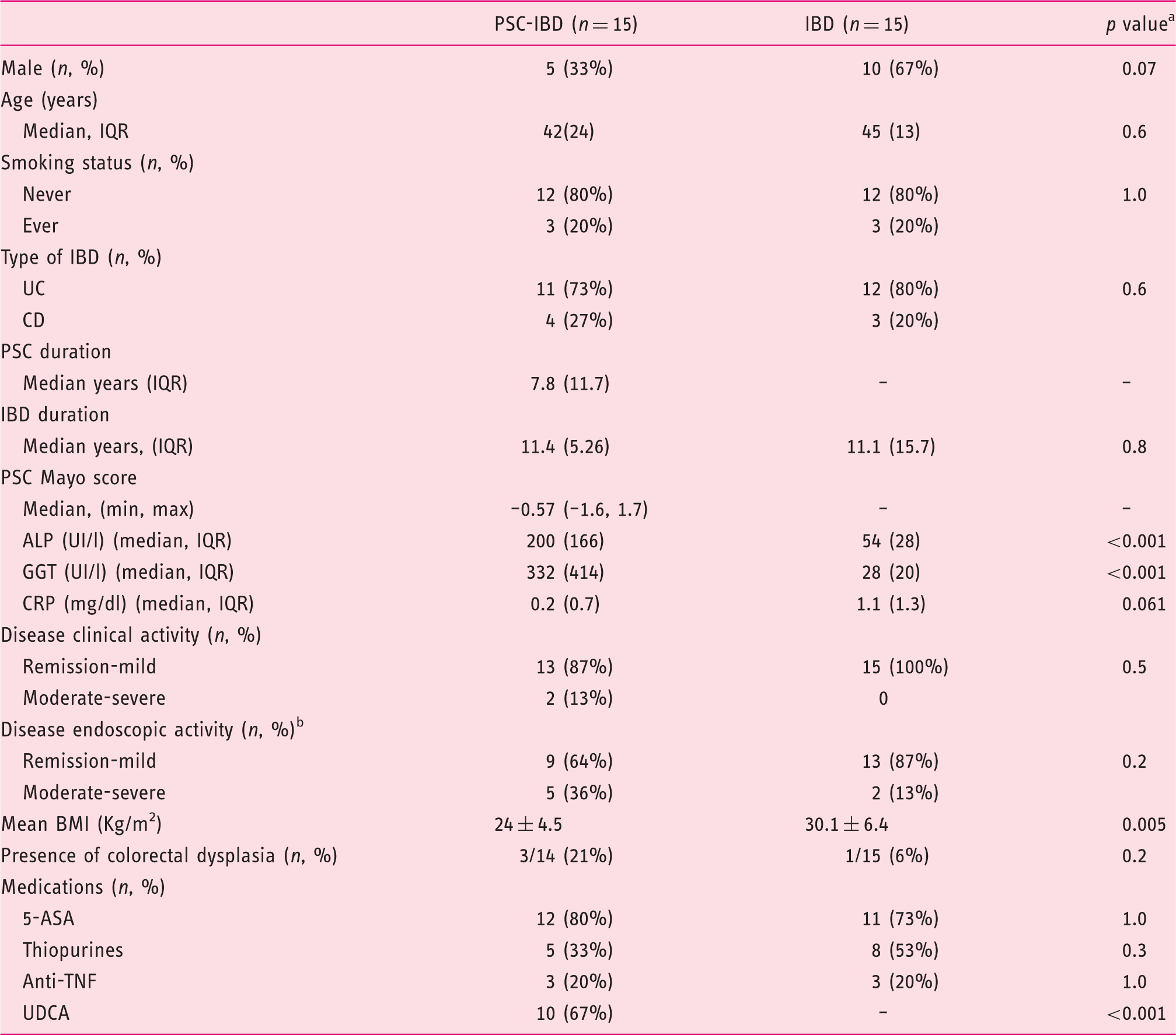
5-ASA: 5-aminosalicilate; ALP: alkaline phosphatase; anti-TNF: anti-tumor necrosis factor; BMI: body mass index; CRP: C-reactive protein; GGT: gammaglutamyl transpeptidase; IQR: interquartile range; PSC: primary sclerosing cholangitis; PSC-IBD: primary sclerosing cholangitis associated with inflammatory bowel disease; UC: ulcerative colitis; UDCA: ursodeoxycholic acid.
Variable distribution was compared using the Student’s
Serum BA
The total BA (µmol/l) pool was significantly expanded in PSC-IBD
(
Stool BA profiles
Stool bile acids (BAs) in primary sclerosing cholangitis associated with inflammatory bowel disease (PSC-IBD) and inflammatory bowel disease (IBD) patients.
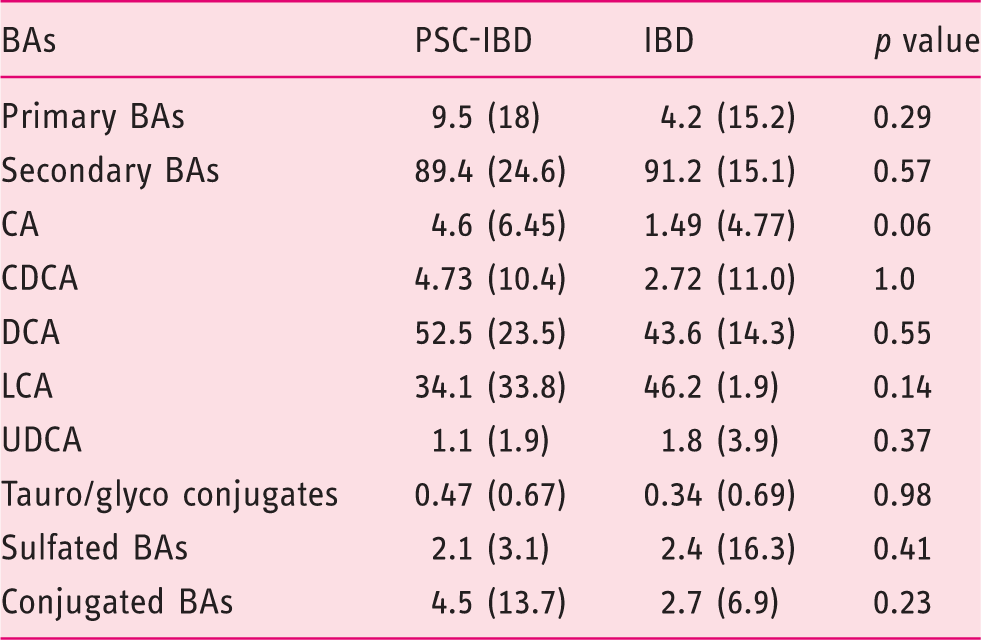
CA: cholic acid; CDCA: chenodeoxycholic acid; DCA: deoxycholic acid; LCA: litocholic acid; UDCA: ursodeoxycholic acid.
BAs are expressed as percentage median (interquartile range) of total BAs. Distributions were compared with non-parametric tests (Mann–Whitney). Due to a small amount of minor BA species in stool (muricholic acid, hycholic acid, hyodeoxycholic acid, or ursodeoxhycolic acid) which are considered by some authors as ‘tertiary BAs’, the sum of primary and secondary BAs is not 100%.
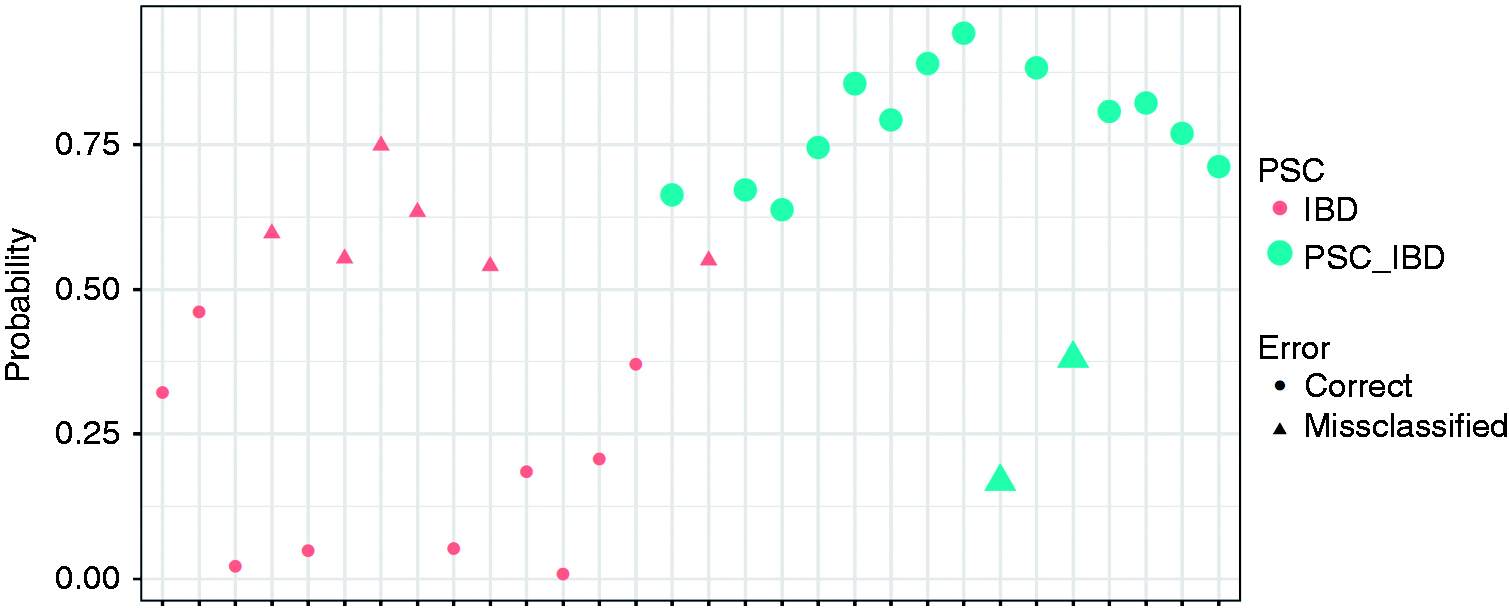
Results of the linear discriminant analysis allowing to see the discrimination of
primary sclerosing cholangitis associated with inflammatory bowel disease (PSC-IBD)
versus inflammatory bowel disease (IBD) alone, based on the combination of the main
bile acids (BAs) present in stool (cholic acid (CA), chenodeoxycholic acid (CDCA),
litocholic acid (LCA), deoxycholic acid (DCA) and ursodeoxycholic acid (UDCA)). On
the
Survey of gut microbiota
Using 16S rRNA sequencing, we surveyed the microbiome composition of 30 stool samples.
The duplicate measurements showed Pearson correlation over 99% at genus level, confirming
the reproducibility of the experimental approach. PSC-IBD presented lower alpha-diversity,
albeit not significantly different (Chao1 899.3 for IBD vs 832.0 for PSC-IBD,
Overall microbiota dissimilarities between samples grouped by primary sclerosing
cholangitis (PSC) and inflammatory bowel disease (IBD) status. (a) Dissimilarities
were measured using UniFrac unweighted distances and visualized using a
multidimensional plot (MDS) plot. The smaller circle represents patients with IBD,
while the larger circle the samples from PSC-IBD patients. (b) The boxplots show the
mean and variance of the richness of the microbial community between different
disease status (Chao1 in the left and Shannon index on the right); no significant
differences are seen (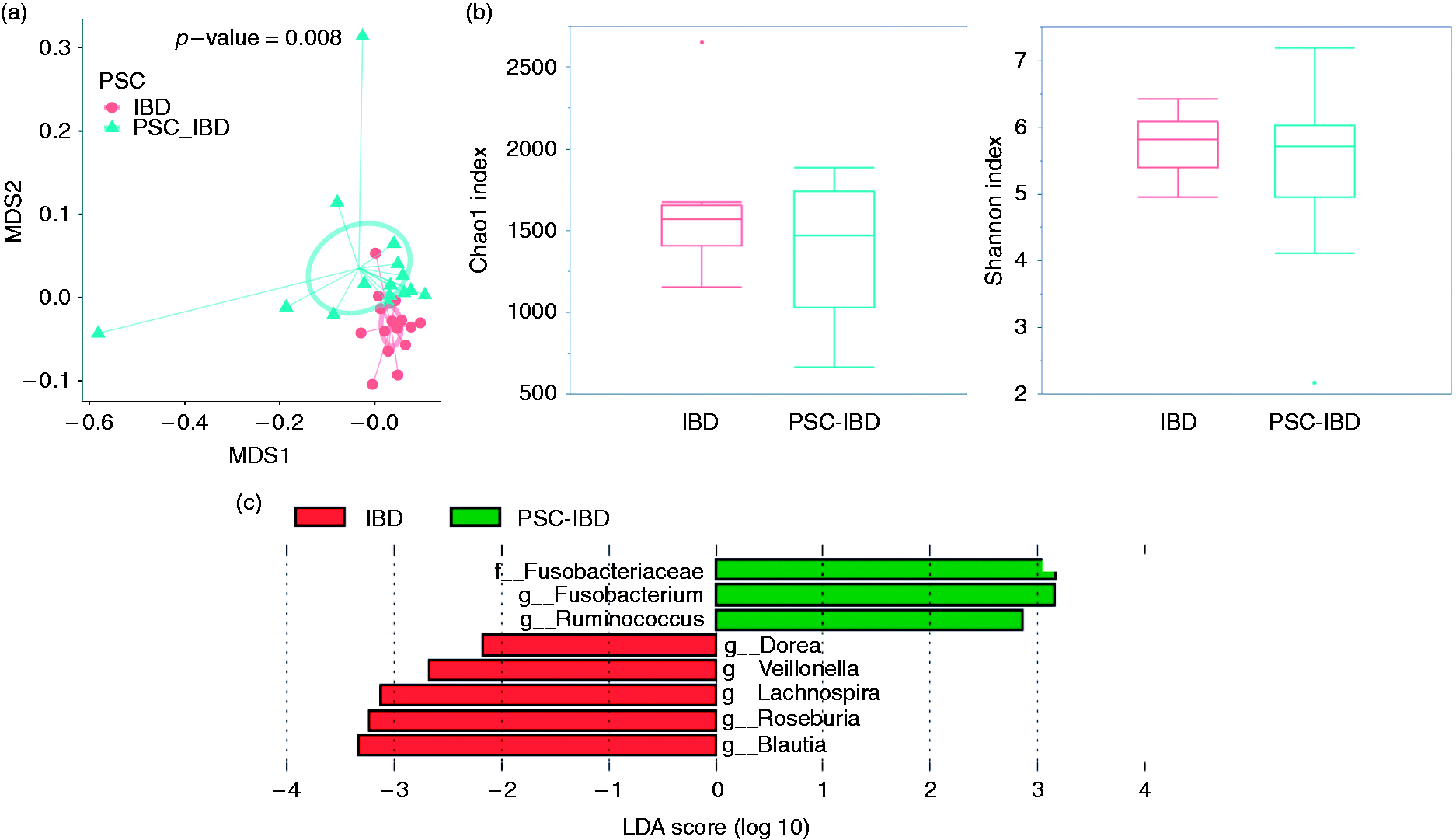
Differential OTUs by PSC status
Based on 97% similarity of the 16S sequencing reads, the open-reference OTU picking using
QIIME pipeline assigned all sequencing reads into individual OTUs. After removing
singletons, we compared 3839 OTUs and selected 143 OTUs which were significantly
(
Correlation between microbiota genera and stool BAs
Correlations between microbiota genus and the stool BA were calculated as described in
our method section to test the interactions between gut microbiota and stool BA. Without
stratifying by PSC status, we found four genera, including
Discussion
Herein, for the first time we have analyzed the stool BA profiles and their correlation
with the faecal microbiota composition in patients with PSC-IBD as compared with IBD alone.
The serum BA pool was increased and the stool BA pool was significantly reduced in PSC-IBD
as compared with IBD alone. No significant differences in the individual stool BA components
were found, but their overall composition differed from IBD (Figure 1). A significantly different microbiota
composition based on the unweighted UniFrac distances was found between IBD and PSC-IBD,
indicating differences in taxon composition for rare taxa (Figure 2(a)). Specifically,
PSC-IBD patients presented an enrichment in bacteria belonging to the genera
In the past, some authors have hypothesized that the increased risk of right-sided
colorectal neoplasia in PSC-IBD could be linked to an increase in secondary BA, although
this had never been demonstrated.2,7 Normally,
most of the BAs secreted by the liver are efficiently reabsorbed in the terminal ileum,
through the sodium-dependent BA transporter (ASBT), leaving only approximately 5% of the
total BAs to reach the colonic lumen. In the right colon, primary BAs are transformed in to
secondary BAs mostly by bacterial mediated deconjugation, oxidation/reduction,
epimerization, and dehydroxylation.
40
Therefore, faecal BA are mainly deconjugated, secondary BAs. A small
fraction of secondary BA is passively absorbed through the colonic mucosa, whilst the rest
will be extruded with faeces.
41
During obstructive cholestasis, the expression of the apical BA
transporter, which permits intracellular absorption of BAs, is down-regulated, as a
compensatory mechanism.
42
This could hypothetically lead to a relative increase in the proportion
of BAs entering the proximal colon in PSC-IBD patients, where they would be converted from
primary into secondary BAs. Interestingly, secondary BAs have been shown to have
anti-inflammatory properties but at the same have been shown to bear carcinogenic
properties.19,32,43–45 Herein, we observed a significant reduction in the total stool BAs in
PSC-IBD as compared with IBD, which was expected taking into consideration the obstructive
cholestatic nature of PSC. However, we did not find an increase in the relative proportion
of the stool secondary BAs in PSC-IBD patients, as previously hypothesized.7,12 No significant differences in individual
proportion of serum or stool BAs were found, which could perhaps be due to the small sample
size. The proportion of DCA, a secondary BA was increased in PSC-IBD, although this did not
reach statistical significance. Furthermore, the proportion of conjugate BAs was also
non-significantly increased in PSC-IBD as compared with IBD, which could indirectly indicate
a decrease in the deconjugation activity of the microbiota. The decrease in
No study had yet looked at the correlations between stool BA and the stool microbiota in
PSC-IBD. However, BA pool size and composition have been shown to be important factors in
regulating the gut microbiota.24,52,53 Herein, despite our
relatively small sample size, and after correcting for multiple comparisons, we were able to
observe unique correlations between stool microbiota and stool BAs in PSC-IBD. Within IBD
alone, this broad BA-microbiota correlation disappeared. In particular, in PSC-IBD, the taxa
that significantly correlated to the stool BA corresponded to ∼12% of the total microbiota,
while in IBD, this was less than 1%. Without any functional data, we may only speculate that
these results suggest that under PSC conditions, the BA changes may have dominant effects on
defining the gut microbiota shifts, potentially towards a more pro-carcinogenic profile.
Interestingly, bacteria from the genus of both
The major limitation of this study is our small sample size, which prevented us from adjusting for potential confounders in the microbiota and BA analysis. To overcome this, we tried to make our cohort as uniform as possible. All patients had pancolitis, and no patient had prior abdominal surgery or history of liver transplantation; all patients had mild to moderate PSC, as measured by the Mayo score, and dietary intake was also similar within groups as assessed by the food frequency questionnaire. Furthermore, all patients stopped UDCA intake for two weeks and had no antibiotics or bowel preparation within at least three months of sample collection, all external factors that could potentially impact microbiota composition. While it may be argued that a two-week interval to stop UDCA may not be enough to remove its effects, we did not observe any differences in the faecal BA composition or in the microbiota composition between those who were medicated with UDCA as compared with those who were not, consistent with what has been previously reported. 13
In summary, in this exploratory study, patients with PSC-IBD had a distinct stool BA and stool microbiota composition, as well as specific microbiota-stool BA correlations when compared with IBD. Whether these changes are associated with or may predispose to the specific PSC-IBD phenotype including the increased risk of colorectal neoplasia needs to be further clarified and warrants further research.
Footnotes
Acknowledgements
The authors wish to thank the OCS genome technology center of New York University Langone Medical Center for the Illumina Miseq molecule library preparation and sequencing service. JT, SHI, MC, JFC, and JH have contributed to the study design and concept. JT, CP, MC, XB, PMS, JPS, CV, and AO have contributed to the acquisition of data. JT, RH, KP, HB, CR, DR, LH and JH analyzed, and interpreted the data. JT and JH drafted the manuscript. All authors have reviewed the final version of the manuscript and provided critical comments. JT and JH obtained funding support.
Declaration of conflicting interests
J Torres has received consulting fees from Takeda. J-F Colombel has served as consultant, advisory board member or speaker for Abbvie, AB Science, Amgen, Bristol Meyers Squibb, Celltrion, Danone, Ferring, Genentech, Giuliani SPA, Given Imaging, Janssen, Immune Pharmaceuticals, Medimmune, Merck & Co., Millenium Pharmaceuticals Inc., Neovacs, Nutrition Science Partners Ltd, Pfizer Inc., Prometheus Laboratories, Protagonist, Receptos, Sanofi, Schering Plough Corporation, Second Genome, Shire, Takeda, Teva Pharmaceuticals, Tigenix, UCB Pharma, Vertex, and Dr August Wolff GmbH & Co. SH Itzkowitz received grant support and fees for serving on a scientific advisory board from Exact Sciences. The remaining authors have no conflicts of interest to declare.
Ethics approval
This study was approved by the Portuguese National Committee for Data Protection and by the local ethics committee at each participating center.
Funding
This study was supported by funds from GEDII (Portuguese Group for the Study of Inflammatory Bowel Disease) (JT) and from the National Institute of Health (NIDDK 1K01DK094986-01(JH) and NIDDK R03 DK106481-01(JH)).
Informed consent
All patients participating in this study signed an informed consent form.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
