Abstract
Introduction
Inflammatory bowel diseases (IBDs) and chronic rheumatic diseases (CRDs) are systemic chronic disorders sharing common genetic, immune and environmental factors. About half of patients with IBD develop rheumatic ailments and microscopic intestinal inflammation is present in up to half of CRD patients. IBD and CRD patients also share a common therapeutic armamentarium. Disequilibrium in the complex realm of microbes (known as dysbiosis) that closely interact with the gut mucosal immune system has been associated with both IBD and CRD (spondyloarthritis and rheumatoid arthritis). Whether dysbiosis represents an epiphenomenon or a prodromal feature remains to be determined.
Methods
In an attempt to further investigate whether specific gut dysbiosis may be the missing link between IBD and CRD in patients developing both diseases, we performed here a systematic literature review focusing on studies looking at bacterial microbiota in CRD and/or IBD patients.
Results
We included 80 studies, with a total of 3799 IBD patients without arthritis, 1084 CRD patients without IBD, 132 IBD patients with arthropathy manifestations and 12 spondyloarthritis patients with IBD history. Overall, this systematic review indicates that an increase in Bifidobacterium, Staphylococcus, Enterococcus, Lactobacillus, Pseudomonas, Klebsiella and Proteus genera, as well as a decrease in Faecalibacterium, Roseburia genera and species belonging to Verrucomicrobia and Fusobacteria phyla are common features in IBD and CRD patients, whereas dozens of bacterial species are specific features of CRD and IBD.
Conclusion
Further work is needed to understand the functions of bacteria and of their metabolites but also to characterize fungi and viruses that are commonly found in these patients.
Introduction
Inflammatory bowel diseases (IBDs) are mainly represented by Crohn's disease and ulcerative colitis, whereas chronic rheumatic diseases (CRDs) encompass rheumatoid arthritis (RA) and spondyloarthritis (SpA). These systemic chronic disorders have relapsing and remitting clinical course arising from an interaction between genetic, immune and environmental factors.
CRD and IBD are intercurrent since articular manifestations are observed in up to 40% of IBD patients and intestinal inflammation is often present in CRD subjects. 1 Co-occurring CRD and IBD can be very disabling and are associated with a more severe disease course in IBD patients. 2
Interestingly, IBD and CRD share common pathophysiology, including common molecular and cellular actors and, consequently, common therapeutic armamentarium. Genetic studies have reinforced the importance of genes and pathways contributing to IBD pathogenesis, such as barrier function, the role of T cell subsets and cytokine–cytokine receptor signalling. 3 In addition, recent studies pointed out new genes and pathways, including autophagy or regulation of interleukin (IL)-23 signalling, highlighting the importance of host defence pathways, specifically those involved in the management of mycobacteria. 4 Heredity is also an important feature of CRD, notably in SpA, and several genetic polymorphisms have been shown to influence the disease risk. The most important one is the major histocompatibility complex (MHC) class I allele HLA-B27. 5 Remarkably, a large subset of the IBD and CRD susceptibility identified genes are encoding for proteins involved in immune response, and particularly in the IL-23/Th17 pathway of T cell differentiation, which is primarily implicated in response against extracellular pathogens, including bacteria and yeasts, and/or in microbial sensing.
However, the link between pathological gut and joint inflammation in patients with both IBD and CRD is not fully understood. Taken together, these data suggest that the perturbation of the gut microbiome, also called dysbiosis, represents an attractive target in this context.
In an attempt to further interrogate whether specific gut dysbiosis may be associated with IBD and CRD and promote pathological inflammation within the joint–gut axis, we performed a systematic literature review investigating similarities and differences regarding faecal microbiota in these patients.
Methods
Search strategy and study selection
A systematic literature search was performed according to PRISMA guidelines. 6 The literature review was conducted using PubMed/MEDLINE (from 1950 to December 2018) and Web of Science (from 1958 to December 2018). Abstracts from annual meetings of national and international gastroenterology and rheumatology conferences (United European Gastroenterology Week, Digestive Diseases Week, European Crohn's and Colitis Organization, European League Against Rheumatism, and American College of Rheumatology) were searched manually from 2013 to 2018.
The following keywords were searched in various combinations using the Boolean terms ‘AND’ and ‘OR’ (‘Microbiota’, ‘Microbiome’, ‘Gut’, ‘Gastrointestinal Microbiome’, ‘Microbiology’, ‘Colitis’, ‘Ileitis’, ‘Intestinal’, ‘Enteritis’, ‘Inflammatory Bowel Diseases’, ‘Crohn Disease’, ‘Ulcerative Colitis’, ‘Rheumatoid Arthritis’, ‘Spondyloarthritis’, ‘Arthritis’, ‘Reactive Arthritis’, ‘Psoriatic Arthritis’, ‘Rheumatoid Arthritis’, ‘Infectious Arthritis’, ‘Ankylosing Spondylitis’, ‘Mycobiome’, ‘Fungal Microbiota’, ‘Intestinal Virome’). This strategy was used both as Medical Subject Headings terms if available and as free text. Searching was limited to publications with human subjects. We only selected English language full text papers and abstracts.
Two authors independently reviewed all articles. Inclusion criteria included the presence of IBD and CRD patient samples and 16S rRNA gene sequencing or metagenomic methods to characterize the gut microbiota. Literature reviews did not include meta-analyses, as well as experimental studies based on in vitro findings and animal models.
Study characteristics and outcomes were reported in a Microsoft Excel Office 2016 Professional spread sheets.
Results
Based on defined criteria, 6519 papers were identified (Figure 1). After review of the titles and abstracts 5564 papers were excluded. Amongst the remaining studies, another 881 were excluded because they included reviews, data retrieved from studies using animal models and in vitro findings. Therefore, 80 studies were included: 56 from IBD patients, with one case report
7
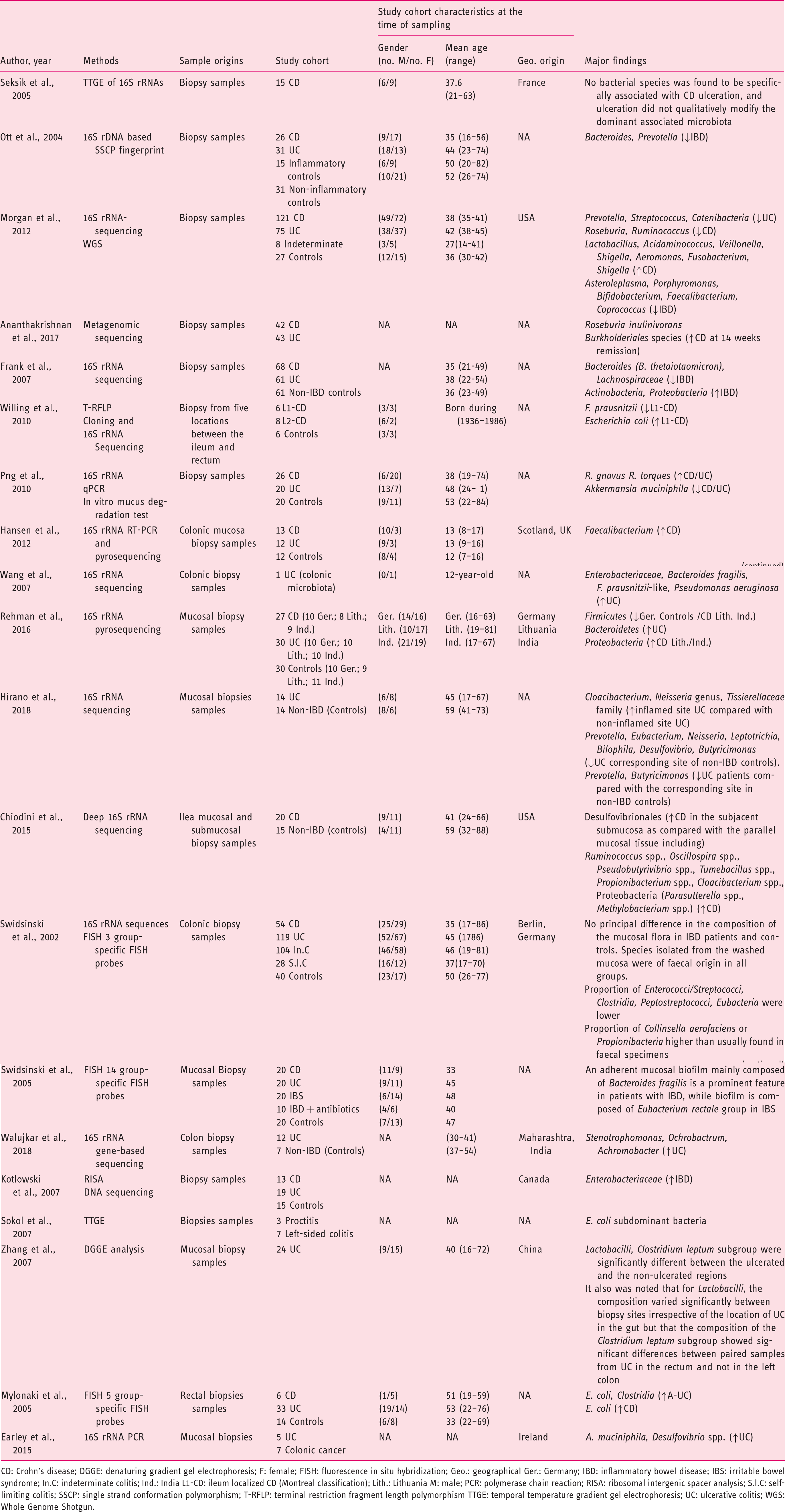
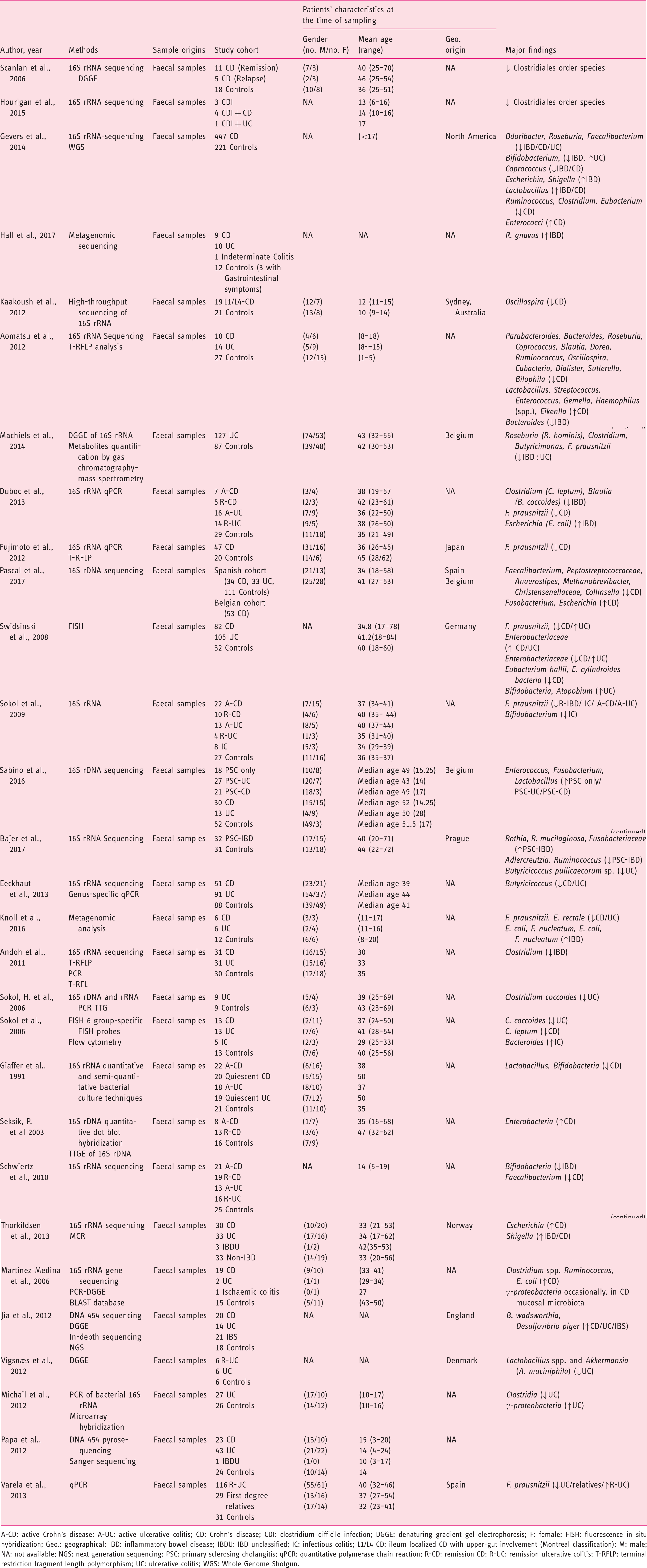
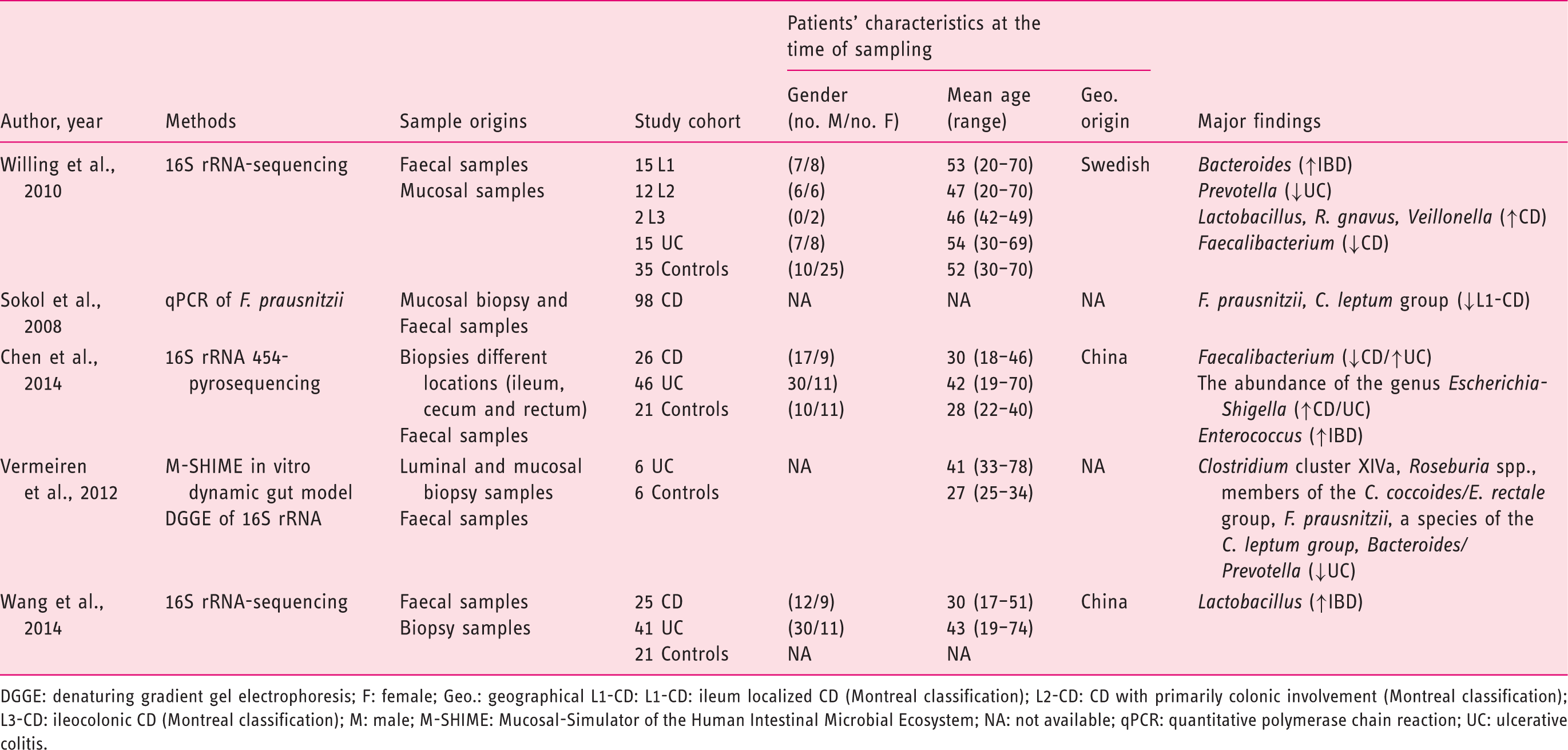
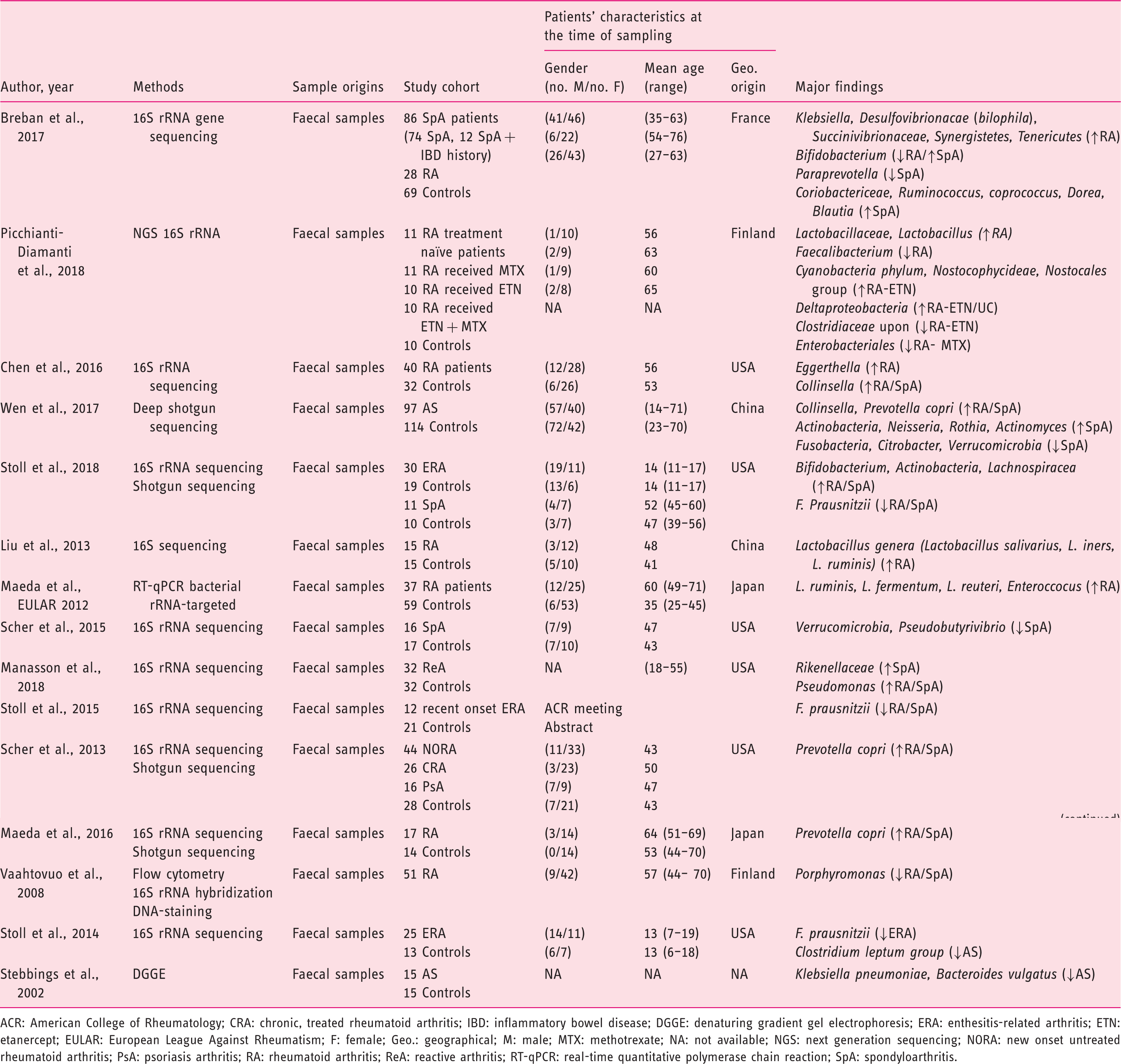
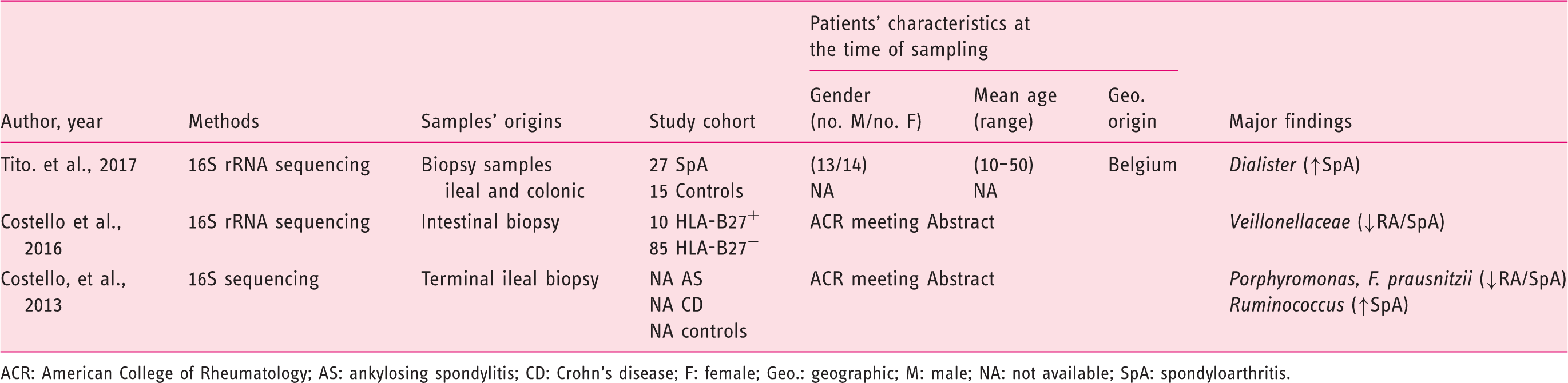
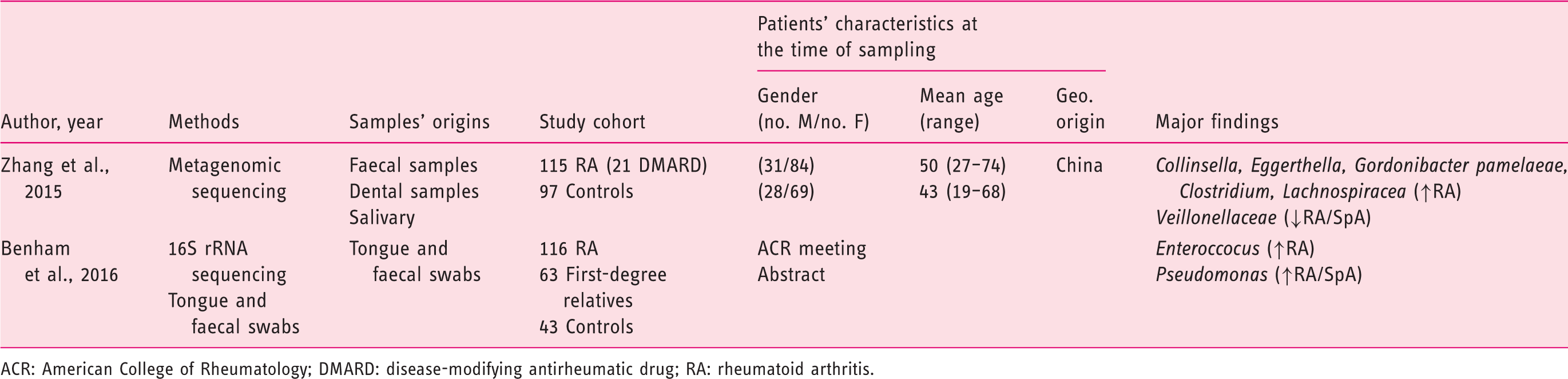
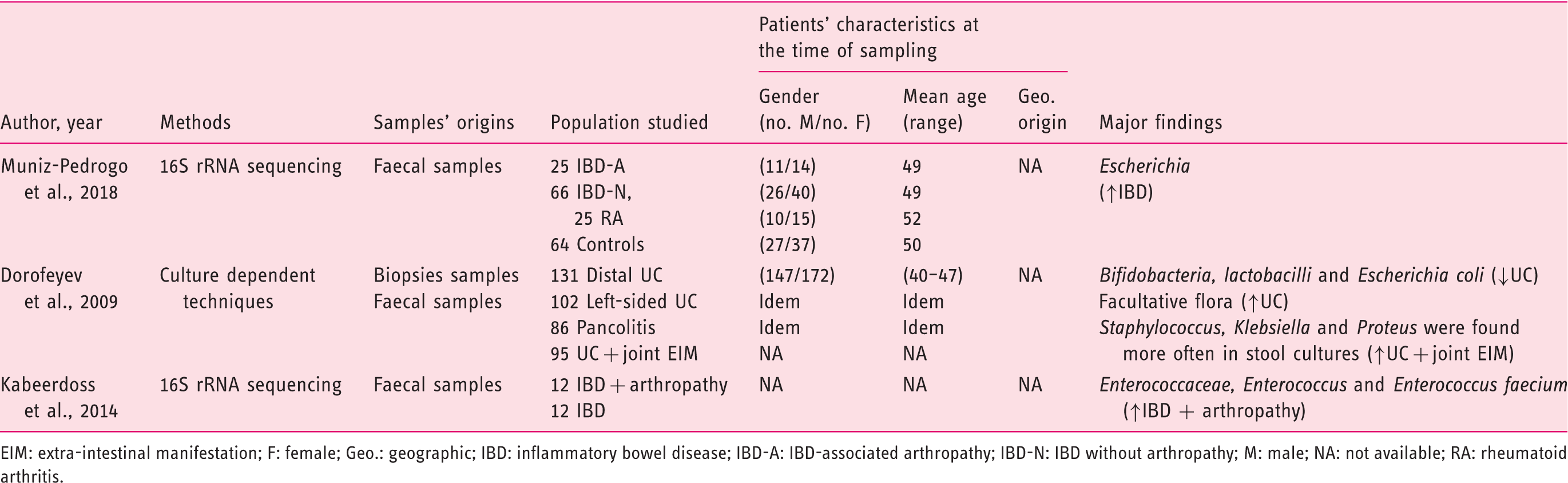
(Tables 1–3), 21 from CRD patients (RA and SpA) including 5 congress abstracts8–12 (Tables 4 to 6). Finally, three publications addressed gut microbiota study in IBD patients developing arthropathy13–15 (Table 7). As a microbiota from one individual is different from one sample location to another, the tables were generated by sample type and are detailed with studied populations characteristics.
Flow-diagram of identified studies. Bacteria associated with inflammatory bowel disease analysed from biopsy samples. CD: Crohn's disease; DGGE: denaturing gradient gel electrophoresis; F: female; FISH: fluorescence in situ hybridization; Geo.: geographical Ger.: Germany; IBD: inflammatory bowel disease; IBS: irritable bowel syndrome; In.C: indeterminate colitis; Ind.: India L1-CD: ileum localized CD (Montreal classification); Lith.: Lithuania M: male; PCR: polymerase chain reaction; RISA: ribosomal intergenic spacer analysis; S.l.C: self-limiting colitis; SSCP: single strand conformation polymorphism; T-RFLP: terminal restriction fragment length polymorphism TTGE: temporal temperature gradient gel electrophoresis; UC: ulcerative colitis; WGS: Whole Genome Shotgun. Bacteria associated with inflammatory bowel disease analysed from faecal samples. A-CD: active Crohn's disease; A-UC: active ulcerative colitis; CD: Crohn's disease; CDI: clostridium difficile infection; DGGE: denaturing gradient gel electrophoresis; F: female; FISH: fluorescence in situ hybridization; Geo.: geographical; IBD: inflammatory bowel disease; IBDU: IBD unclassified; IC: infectious colitis; L1/L4 CD: ileum localized CD with upper-gut involvement (Montreal classification); M: male; NA: not available; NGS: next generation sequencing; PSC: primary sclerosing cholangitis; qPCR: quantitative polymerase chain reaction; R-CD: remission CD; R-UC: remission ulcerative colitis; T-RFLP: terminal restriction fragment length polymorphism; UC: ulcerative colitis; WGS: Whole Genome Shotgun. Bacteria associated with inflammatory bowel disease analysed from both faecal and biopsy samples. DGGE: denaturing gradient gel electrophoresis; F: female; Geo.: geographical L1-CD: L1-CD: ileum localized CD (Montreal classification); L2-CD: CD with primarily colonic involvement (Montreal classification); L3-CD: ileocolonic CD (Montreal classification); M: male; M-SHIME: Mucosal-Simulator of the Human Intestinal Microbial Ecosystem; NA: not available; qPCR: quantitative polymerase chain reaction; UC: ulcerative colitis. Bacteria associated with chronic rheumatic diseases analysed from faecal samples. ACR: American College of Rheumatology; CRA: chronic, treated rheumatoid arthritis; IBD: inflammatory bowel disease; DGGE: denaturing gradient gel electrophoresis; ERA: enthesitis-related arthritis; ETN: etanercept; EULAR: European League Against Rheumatism; F: female; Geo.: geographical; M: male; MTX: methotrexate; NA: not available; NGS: next generation sequencing; NORA: new onset untreated rheumatoid arthritis; PsA: psoriasis arthritis; RA: rheumatoid arthritis; ReA: reactive arthritis; RT-qPCR: real-time quantitative polymerase chain reaction; SpA: spondyloarthritis. Bacteria associated with chronic rheumatic diseases analysed from biopsy samples. ACR: American College of Rheumatology; AS: ankylosing spondylitis; CD: Crohn's disease; F: female; Geo.: geographic; M: male; NA: not available; SpA: spondyloarthritis. Bacteria associated with chronic rheumatic diseases analysed from faecal and other origin samples. ACR: American College of Rheumatology; DMARD: disease-modifying antirheumatic drug; RA: rheumatoid arthritis. Bacteria associated with inflammatory bowel disease and chronic rheumatic diseases. EIM: extra-intestinal manifestation; F: female; Geo.: geographic; IBD: inflammatory bowel disease; IBD-A: IBD-associated arthropathy; IBD-N: IBD without arthropathy; M: male; NA: not available; RA: rheumatoid arthritis.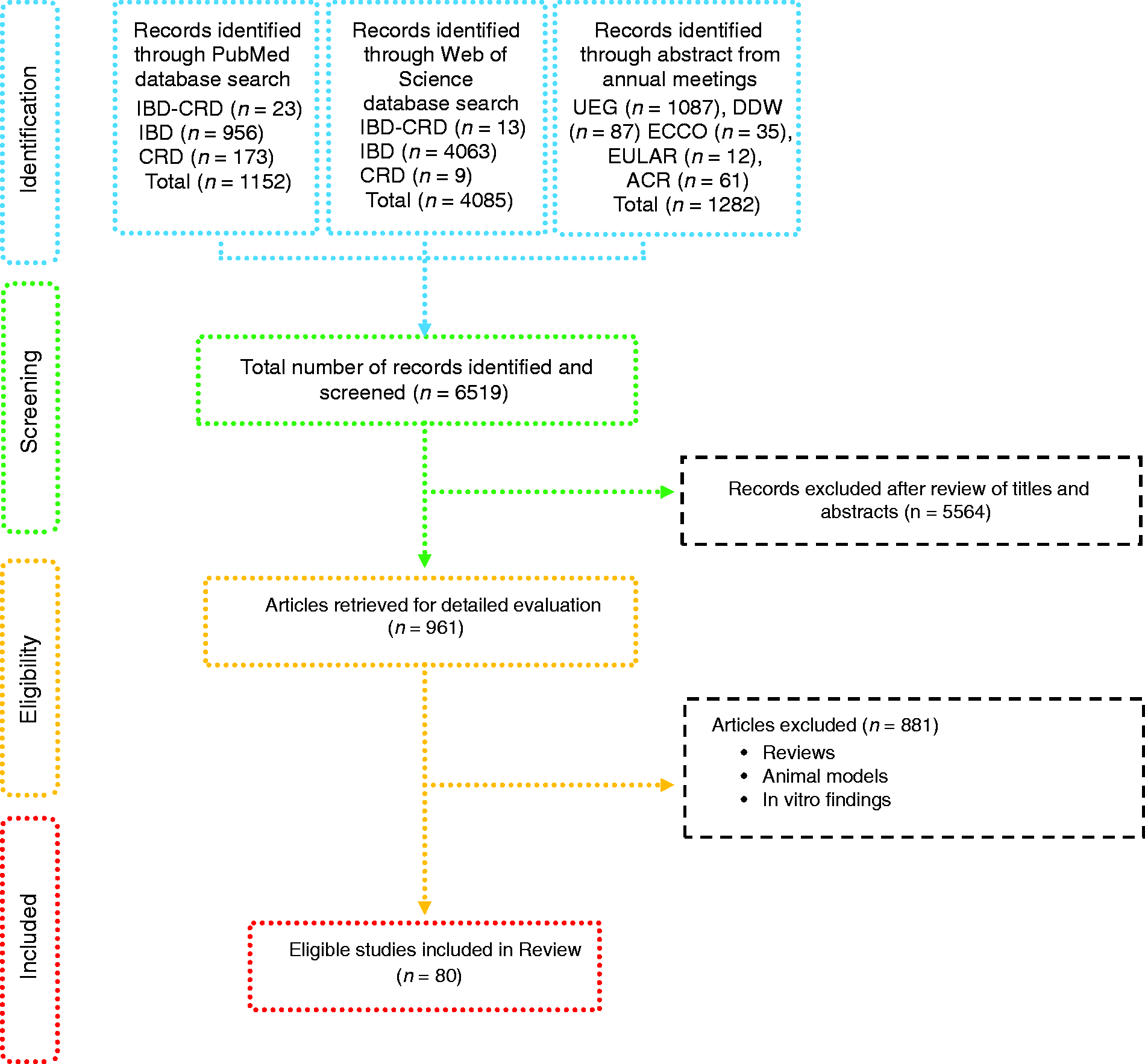
Literature search results: distinct dysbiosis in IBD and CRD
In order to identify bacterial variations specific to IBD and not found in CRD, and vice versa, we adopted two complementary methodologies: we first reviewed bacterial changes reported in studies enrolling IBD patients without information on possible concomitant arthritis, then all studies involving CRD patients without information on possible concomitant IBD. We looked finally at studies comparing gut microbiome in patients with or without IBD-associated CRD.
Gut bacterial changes reported in IBD patients
Fifty-six studies enrolling 3270 IBD patients from which gut microbiota was mainly analysed by 16S rRNA gene sequencing or qRNA of DNA extracted from faeces and/or biopsies. A quantitative and qualitative (biodiversity) reduction of the gut microbiome in IBD patients16,17 is generally observed.
Firmicutes phyla
A reduction of Clostridiales order species from the Firmicutes phylum is observed in the faecal microbiota of IBD and Crohn's disease patients.18–20 An enrichment of Ruminococcus gnavus is observed in the IBD patients’ faecal microbiota.21–23 This phylogenetic group includes several butyrate-producing bacteria, notably Faecalibacterium and Ruminococcus, which are among the main members of the Ruminococcaceae genera. 24 Other bacteria that are considered as ‘beneficial’ for the host have been shown to be quantitatively reduced in the faecal microbiota of these patients. A few studies found a lower number of sequences of the bacterial phylum Firmicutes in the mucosal-associated microbiota (MAM) of Crohn's disease and ulcerative colitis patients, especially species from the Lachnospiraceae genera (Roseburia and coprococcus).21,22,25–30 Within this phylum, an increased amount of Streptococcus genera was observed, in contrast to Ruminococcaceae genera (Faecalibacterium), which seems to be particularly deficient in Crohn's disease.24,29,31–33 Furthermore, Rehman et al. demonstrated population-specific disease-related patterns of Firmicutes phyla, by observing a lower abundance in healthy German samples compared with patients’ samples, while Lithuanian and Indian patients with Crohn's disease show the lowest Firmicutes abundances. 34
In a recent study using molecular methods of bacterial identification, 28 it has been shown that Faecalibacterium prausnitzii was one of the most underrepresented species of the Faecalibacterium genera in the MAM of patients with IBD (compared with healthy subjects).21,22,24,28,31,32,35–38 Therefore, similar to the results from faecal microbiota studies, a significant decrease of bacteria from the Firmicutes phylum was demonstrated in the MAM of Crohn's disease patients.24,31,32
A reduction of Ruminococcaceae, Lactobacillaceae, Veillonellaceae and Erysipelotrichiaceae genera (Faecalibacterium, Streptococcus, Veillonella and Catenibacterium respectively),19,22,39–41 along with Dialister genus in Crohn's disease patients,
42
and Roseburia, Clostridium and Butyricimonas genera is observed in IBD patients, particularly those with ulcerative colitis.24,30,40,41,43 A few studies showed an increased number of the Tissierellaceae family, and a decreased number of Eubacterium genera in inflamed colonic mucosa biopsy samples when compared with the non-inflamed sites in ulcerative colitis patients44–46 (Figure 2).
Similarities and differences regarding gut bacteria between inflammatory bowel disease (IBD) and chronic rheumatic disease (CRD) patients.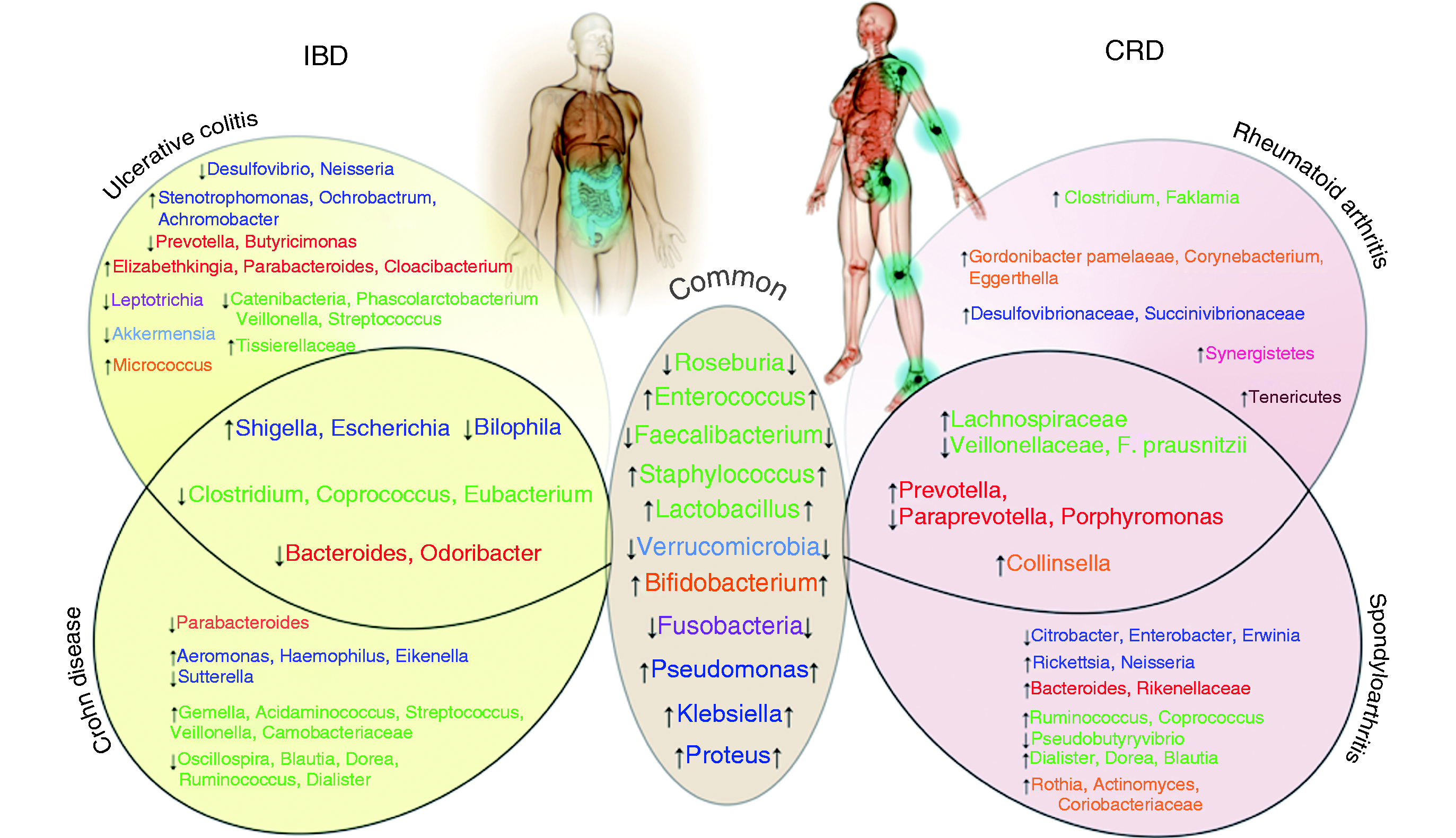
Bacteroidetes phyla
Data concerning the Bacteroidetes phylum are more conflicting. Some studies reported a reduction of the Bacteroides group in IBD patients especially in Crohn's disease patients.18,21,22,29 In contrast, Andoh and colleagues demonstrated an increased amount of this phylum in the context of IBD. 47 To note, one study showed an increase of Bacteroidetes phylum in salivary microbiota in ulcerative colitis patients. 7 Hirano and co-workers showed an enrichment of the Cloacibacterium genus, and decreased abundance of Prevotella (at both inflamed and non-inflamed mucosal site) and Butyricimonas genera at the non-inflamed mucosal site of ulcerative colitis patients compared with the corresponding site in non-IBD controls and in the faecal microbiota of ulcerative colitis patients.21,43,44,48 A greater abundance in these two genera was found in the submucosal tissues of patients with Crohn's disease.21,43,44,48,49 As with Crohn's disease, this strongly suggests a restricted biodiversity in ulcerative colitis and an increased proportion of unusual bacteria.50,51 Bacteroidetes show also interesting age-related patterns and population-independent increase in abundance in the standing and active bacteria among healthy subjects and ulcerative colitis patients. 34 A decreased abundance of Parabacteroides genera and Odoribacteracae family in IBD and Crohn's disease patients respectively has been reported.19,22,24 Similar to the results from faecal microbiota studies, a significant decrease of bacteria from the phylum Firmicutes was demonstrated in the MAM of patients with Crohn's disease.52,53 A recent study by Walujkar and colleagues revealed significant differences in the MAM of patients manifesting acute exacerbations of ulcerative colitis with increased number of Parabacteroides and Elizabethkingia genera as compared to remission stage 54 (Figure 2).
Actinobacteria phyla
Concerning the Actinobacteria phylum, studies using both culture and recent molecular methods demonstrated an increase of Bifidobacterium genera in the faecal microbiota as well as in the biopsy samples of IBD patients, notably in patients with Crohn's disease.21,22,24,28,55 However, other authors reported that an age-related reduction of bacteria of the Bifidobacterium genera was shown in inflamed sites when compared with non-inflamed ones and salivary microbiota of ulcerative colitis patients.7,21,22,24,40,44,55–57,58 Walujkar and co-workers showed an increase amount of Micrococcus genera in MAM of ulcerative colitis patients when compared with non-IBD subjects 54 (Figure 2).
Proteobacteria phyla
Published studies display a quantitative alteration of Proteobacteria phylum in IBD, especially Escherichia and Shigella from the Enterobacteriaceae family.19,21,22,24,33,38,59 Thus, their increased abundance was reported in the MAM and faecal samples of patients with Crohn's disease, whether using culture33,52 or molecular26,60,61 methods. As with Crohn's disease patients, the MAM of patients with ulcerative colitis contained an abnormally elevated concentration of bacteria, especially anaerobes.52,53 A restriction of the MAM biodiversity similar to that observed in patients with Crohn's disease has been found, such as reduction of Firmicutes and an overrepresentation of Enterobacteriaceae.28,34,53,55,62–64 A decreased abundance of the genera Bilophila and Desulfovibrio was evidenced at the inflamed site of ulcerative colitis patients compared with the corresponding site of non-IBD controls, whereas a decreased amount of Bilophila genera and its species (B. wadsworthia) was detected in the faecal microbiota of Crohn's disease patients.44,65,66 Moreover, an age-related reduction of the Neisseria genera bacteria was reported in inflamed sites when compared with non-inflamed ones and salivary microbiota of ulcerative colitis patients.7,21,22,24,40,44,55–58 Walujkar et al. suggested an increased abundance of Stenotrophomonas, Ochrobactrum and Achromobacter genera in ulcerative colitis patients as compared with the same patients during remission stage. 54 Finally, Proteobacteria phyla displayed also an age-related pattern. 34
Other phyla
Finally, a decreased abundance of Verrucomicrobia (Akkermansia) and Fusobacteria (Leptotrichia) was reported at the inflamed colonic mucosal sites of Crohn's disease and ulcerative colitis patients compared with the corresponding sites of non-IBD controls. However, further investigation concerning an eventual association between Leptotrichia and ulcerative colitis is necessary.21,40,41,44,67–69
In summary, among the 56 available studies on IBD, differential abundance of 40 bacterial species has been reported; 15 were specifically found in Crohn's disease studies while only 16 species were reported in ulcerative colitis studies. These variations mainly concerned Firmicutes, Proteobacteria and Bacteroidetes.
Gut bacterial changes reported in CRD patients
A total of 21 studies, enrolling 993 CRD patients, analysed the gut microbiota by 16S rRNA gene sequencing from faeces. Breban et al. have demonstrated that β-diversity analysis, which evaluates the shared diversity between different microbiomes in terms of various ecological distances, showed a microbiota composition significantly different between the RA, SpA and healthy subjects groups. Both SpA and RA patients differed from healthy subjects as well as SpA from RA patients. This study showed also that α-diversity, which evaluates the species' richness and evenness within the microbiota, assessed by the number of observed species was significantly decreased in both SpA and RA patients, as compared with healthy subjects.70,71 In ankylosing spondylitis (AS) patients, the diversity of the gut microbiome was similar to healthy subjects at the genus level but was significantly higher in the controls at the species level. 72
Firmicutes phyla
Concerning the Firmicutes phylum, several bacteria from the Lachnospiraceae family, including Ruminococcus (R. gnavus sp.), Dorea, Coprococcus and Blautia genera, are overabundant in SpA. 70 Increased amount of several Blautia and Ruminococcus could characterize HLA-B27 + siblings. 70 Likewise, inflamed ileal biopsies of SpA patients revealed an increase in the Dialister genus, which could be a microbial marker of disease activity.73,74 In contrast, SpA patients seemed to display a decreased amount of Roseburia species. 2 Concerning RA patients, fewer Firmicutes of the Ruminococcaceae family but an increase in Lactobacillus species and Faklamia have been observed.70,75 A study by Picchianti-Diamanti et al. characterized the gut microbiota of RA patients under different immunosuppressants treatment strategies (ETN, MTX, or ETN plus MTX) and compared it to that of treatment-naïve patients. This study highlighted a drop in Proteobacteria caused by ETN, which in general are abundant in both intestinal and extra-intestinal inflammatory diseases. 76 Moreover, upon ETN treatment, a decrease in Clostridiaceae was observed, which were previously found enriched in patients with RA and IBD-associated arthropathy. 77 In patients treated with MTX, analysis revealed a significant decrease in Enterobacteriales. 75
Liu et al. reported that RA patients, compared with healthy subjects, exhibited an increased bacterial diversity within the Lactobacillus community with increase in L. salivarius and L. iners,70,78,79 for instance. The analysis of faeces from RA patients has demonstrated the presence of a large cluster including Firmicutes bacteria belonging to the Lachnospiraceae and Clostridiaceae (Clostridium) families, as well as small clusters containing strains from the Lactobacillus and Ruminococcus genera.78–81 In the RA patients' gut, a decrease of bacteria from the Veillonellaceae family was observed.80,82 In contrast to SpA patients, psoriasis arthritis patients showed depletion in Coprococcus, Ruminococcus, Clostridium and Pseudobutyrivibrio compared with healthy subjects.70,82–84 Finally, SpA patients exhibited a decreased faecal abundance of F. prausnitzii compared with healthy subjects. This bacterium may be, at least in part, responsible for the pathogenesis of SpA.8,74,85
Bacteroidetes phyla
There is a significant enrichment of the Prevotellaceae species, and more particularly of Prevotella copri, within the Bacteroidetes phylum, in intestinal microbiota of patients with new-onset RA, compared with chronic RA patients and healthy subjects.9,10,86 This bacterium is relatively scarce in the general population. In addition, Bacteroides genera counts were lower in the same group, while being higher in SpA patients.74,80,86 However, P. copri decreased in the gut of RA patients along with disease chronicity. 9 Breban et al. also demonstrated that SpA and RA patients have decreased populations of Prevotellaceae and Paraprevotellaceae genera compared with healthy subjects. 70 However, in AS patients, Prevotellaceae are more abundant in terminal ileal biopsy samples. 85 Furthermore, a quantitative metagenomics study has shown that the microbial communities in the AS cases were characterized by a higher abundance of Prevotellaceae genera (P. copri) compared with healthy subjects. 72 Other bacteria from the Bacteroidetes phylum, such as Porphyromonas, were shown to be decreased in RA patients while being increased in terminal biopsies of AS patients.85,87
Actinobacteria phyla
Regarding the Actinobacteria phylum, which is a low-abundant one, patients with RA or SpA had a higher amount of bacteria from the Coriobacteriaceae family and especially of the Bifidobacterium genus, including B. bifidum species, than healthy subjects.70,74 However, RA patients are also characterized by an increase of Corynebacterium species. 70 The metagenomic analysis and 16S sequencing have additionally brought to light the presence of the bacteria Gordonibacter pamelaeae, Eggerthella lenta and Collinsella in RA patients.71,72,80 The latter could contribute to the increased permeability of the gut and enhanced production of pro-inflammatory cytokines. 74 In SpA patients, an overabundance of Collinsella, Rothia and Actinomyces genera was reported.71,72,84
Proteobacteria phyla
The Proteobacteria phylum is more abundant in RA patients than in healthy subjects, concerning more specifically the Klebsiella and Bilophila genera from Enterobacteriaceae, Desulfovibrionaceae and Succinivibrionaceae families. 70 In SpA patients there is a decrease of Citrobacter, Enterobacter and Erwinia genera.79,81,84 The last was particularly reduced in the HLA-A24 positive group of patients. In contrast, an overabundance of Neisseria genera was reported in SpA patients. 72
Other phyla
Finally, other phyla, such as Synergistetes, Tenericutes, Fusobacteria and Verrucomicrobia, were also seen to vary significantly in RA and SpA patients21,70,72,83,88 (Figure 2).
In summary, among the available studies to vary on CRD (N = 21), 33 bacterial species were reported in CRD; among those, 17 were specifically reported in SpA studies while only nine species were reported in RA studies. Variations mainly concerned Firmicutes, Bacteroidetes and Actinobacteria phyla.
Differences between IBD and CRD gut microbiota
In three studies enrolling a total of 554 patients, 356 IBD patients without known arthropathy and a total of 132 IBD with joint extra-intestinal-manifestation (EIM) patients were analysed (Table 7). One study indirectly compared three cohorts of patients, SpA patients without IBD history (n = 74), SpA patients with an IBD history (n = 12) and RA patients (n = 28) compared with healthy controls (n = 69) (Table 7). 70
Firmicutes phyla
Amongst the included studies, some pointed out important differences, including variable amount of several Firmicutes genera. For instance, the overabundance of Veillonella observed in Crohn's disease patients contrasted with its paucity in CRD (RA, SpA) patients. Conversely, the Eubacterium, Clostridium, Ruminococcus and Coprococcus genera, which were increased in CRD (RA, SpA) patients, were decreased in patients with Crohn's disease.9,11,21,22,24,28,80,85–88 Variation of the Ruminococcus genus is the most surprising since a paradoxical overabundance, especially of R. gnavus, has been reported in IBD patients. This increased abundance correlated positively with SpA activity whatever patients' IBD history, even though IBD was inactive at the time of sampling in most of them.21,70 In IBD, R. gnavus was mostly associated with the gut mucosa, which conferred to this mucolytic bacterium a possible role in the triggering or maintenance of inflammation.21,41 Whether its lonely increase could be linked to specific genetic predispositions to SpA warrants more investigation. As for the Dialister genera, belonging to the same bacterial family, an increased number of sequences was observed in SpA groups whereas a decrease was found in Crohn's disease patients. 70 In ulcerative colitis patients with a joint EIM, the Staphylococcus genus was found more frequently in stool cultures. 12
Bacteroidetes phyla
Variations in Bacteroidetes phylum concerned mainly two genera: Bacteroides, which was in increased amounts in SpA patients and in reduced amounts in RA and IBD groups, and Prevotella, which showed a high abundance in CRD (RA and SpA) patients and was lowered in ulcerative colitis patients.9,11,24,29,71,72,74,86,89
Proteobacteria phyla
In the Proteobacteria phylum, the genus Bilophila was overabundant in RA and SpA patients while being found in reduced amounts in Crohn's disease patients.21,59,70,90,91 Dorofeyev et al. showed a significant abundance of Enterobacter, Klebsiella and Proteus genera in stools cultures from ulcerative colitis patients with a joint EIM, compared with healthy subjects and ulcerative colitis patients without EIM. 12 In contrast, in ulcerative colitis a decreased amount of Neisseria was observed.7,21,22,24,40,44,55–58 However, metagenomics studies of gut microbiome in patients with enteropathic arthritis are still lacking. Using quantitative polymerase chain reaction, a relative overabundance of the Enterobacteriaceae family, concomitant to a reduction of the Clostridia group XIVa cluster, was reported in the gut microbiota in IBD patients with joint manifestations. As a whole, the Enterobacteriaceae family seemed to be increased in the gut of IBD patients and this tendency is even more pronounced in those with arthropathy. 92
Actinobacteria phyla
Concerning the Actinobacteria phylum, an overabundance of Gordonibacter pamelaeae, Eggerthella lenta and Collinsella was observed in RA patients.71,72,80,93 However, an increase of Micrococcus genera was also characterized in MAM ulcerative colitis patients. 54 In SpA patients, an overabundance of Collinsella, Rothia and Actinomyces genera was reported.71,72,84
Other phyla
Finally, the Fusobacterium phylum is more abundant in Crohn's disease patients and less abundant in SpA patients. 70 In contrast, amounts of the Tenericutes phylum are increased in SpA patients.19,70,72
Taken together, when considering all available studies (N = 80), 40 bacterial species were reported only in IBD patients, and 33 bacterial species were reported only in CRD subjects (Figure 2). The main variations were mostly observed in the Firmicutes phylum.
Literature search results: similarities regarding bacterial microbiome in IBD and CRD
When comparing studies on IBD patients without known CRD versus studies on CRD patients without known IBD, we first observed that some dysbiotic changes share similarities between chronic IBD and chronic joint diseases, among which are a lower microbial diversity and a diminished abundance of the Firmicutes phylum.
Firmicutes phyla
Amongst the Firmicutes genera, a common decreased amount was described for Faecalibacterium and Roseburia species in both IBD subtypes (Crohn's disease and ulcerative colitis), as well as in SpA and RA patients.11,21,22,24,29,58,70 A few studies using bacterial culture, in addition to recent molecular methods, have demonstrated an increased amount of Lactobacillus and Enterococcus in the faecal microbiota of IBD patients, especially those with Crohn's disease and RA patients, although others demonstrated a reduction of Lactobacillus in Crohn's disease patients.11,16,21,22,24,39,55–57,60,78,80
An overabundance of Staphylococcus was observed in ulcerative colitis patients with arthritis when compared with patients without EIM and a healthy population.
Proteobacteria phyla
In the Proteobacteria phylum, an overabundance of several genera was observed, such as Klebsiela and Proteus in all ulcerative colitis patients with arthritis. These facultative microbiota were significantly higher in these patients than in the healthy subjects and ulcerative colitis patients without EIM.12,54,70,94,95 An increase of Pseudomonas was recently shown by Walujkar et al. in the MAM of ulcerative colitis patients as compared with the same patients during remission stage, 54 as well as shown by Manasson et al. and Benham et al. in patients with SpA or RA.81,84
Actinobacteria phyla
Concerning the Actinobacteria phylum, an overabundance of Bifidobacterium was reported in SpA patients, especially those with enthesitis-related arthritis, and in IBD patients, notably in patients with Crohn's disease.21,22,24,28,55,58,70,72,74,80,87,88
Other phyla
Finally, a common decrease of Verrucomicrobia and Fusobacteria belonging species was reported in both Crohn's disease and ulcerative colitis patients compared with non-IBD controls19,21,40,41,44,67–69 and in RA and SpA patients.21,70,72,83,88
In summary, variations of species belonging to Firmicutes, Proteobacteria, Actinobacteria, Verrucomicrobia and Fusobacteria phyla represent the main common trait between IBD and CRD gut microbiota. A figure depicting similarities and differences observed in bacterial species amounts in biopsy and faeces from IBD and CRD patients is proposed (Figure 2).
Conclusion and perspectives
To our knowledge, this is the first systematic review regarding gut microbiota alterations in IBD and CRD patients. Our analysis highlights the general finding that microbiota favouring proteolytic-fuelled fermentation and lactic acid-producing bacteria are increased in both CRD and IBD inflammatory conditions while those producing butyrate are generally decreased in both diseases. Second, variations of gut microbiota composition in IBD patients mainly concern Firmicutes, Proteobacteria and Bacteroidetes. Within the Firmicutes phylum variations of species such as Roseburia, coprococcus, F. prausnitzii and Streptococcus genera was observed either in the MAM of Crohn's disease patients or ulcerative colitis patients. In terms of the Proteobacteria phylum, published data display a quantitative alteration in IBD Crohn's disease and ulcerative colitis patients compared with control groups, especially of Escherichia, Shigella, Bilophila, Desulfovibrio, Neisseria, Stenotrophomonas, Ochrobactrum and Achromobacter genera. Concerning the Bacteroidetes, variations of Cloacibacterium, Prevotella, Butyricimonas, Parabacteroides, Elizabethkingia genera and Odoribacteracae family in IBD Crohn's disease and ulcerative colitis patients are observed.
In CRD patients, variations of gut microbiota are mainly observed in Firmicutes, Bacteroidetes and Actinobacteria phyla. Alterations of gut microbiota observed in the Firmicutes phyla included Ruminococcus (R. gnavus sp.), Dorea, Coprococcus, Blautia and Dialister genus in RA and SpA patients. In addition alterations of Roseburia, Lactobacillus, Faklamia, Staphylococcus, Clostridium, Pseudobutyrivibrio, F. prausnitzii species and Veillonellaceae family was observed in patients compared with healthy subjects. There is a significant variation of species within the Bacteriodetes phylum, particularly of Bacteroides, Prevotellaceae (P. copri), Paraprevotellaceae and Porphyromonas genera in RA and SpA patients compared with healthy subjects. Regarding the Actinobacteria phylum, which is a low-abundant one, in patients with RA or SpA variations of the Bifidobacterium genus, including among others B. bifidum species, Gordonibacter pamelaeae, Eggerthella lenta, Collinsella, Rothia and Actinomyces genera, were reported compared with control groups.
Another major finding of this study is the reduction of bacterial diversity observed in both CRD and IBD and the presence of common bacterial phyla changes. We can mention an increased abundance in Lactobacillus, Enteroccocus, Staphylococcus, Bifidobacterium, Klebsiella, Pseudomonas and Proteus genera in both CRD and IBD, whereas Faecalibacterium and Roseburia genera and Verrucomicrobia and Fusobacteria phyla are decreased in both diseases.
Interestingly, experimental studies have confirmed the role of Faecalibacterium in immune controlled in both type of affections. First, Hablot and colleagues suggested that experimental dextran sulphate sodium (DSS)-induced colitis could altered the gut microbiota of mice with arthritis compared with mice with colitis alone and thus could delay the appearance of ‘pro-arthritogenic’ bacteria. 96 This delay is associated with a difference of microbiota composition between mice with arthritis and colitis and mice with colitis only. Members of the Firmicutes phylum are mainly affected; Lactobacillus genus and Clostridiales order are more present in mice with arthritis and colitis compared with mice with only colitis. Several studies showed that species from Lactobacillus are beneficial in DSS-induced colitis.13,97 Thereby, a Lactobacillus sp. increase in arthritis + colitis group might play a role in the subclinical improvement as observed by the decrease in faecal lipocalin-2 level. A difference of the faecal microbiota composition is also observed between arthritis and arthritis + colitis groups. At arthritis and colitis onset, Lactobacillaceae, and notably Lactobacillus R. gnavus, and S24_7 species belonging to Bacteroidales are more present in mice with arthritis and colitis compared with an arthritis group. Interestingly, these groups of bacteria had been shown to be more present in mice with higher susceptibility to arthritis development.14,96
Viladomiu and colleagues recently identified an enrichment of IgA-coated Escherichia coli in Crohn's disease–SpA with an adherent–invasive E. coli (AIEC) pathotype. Experimental models highlight two features of the host–pathogen interaction that must be considered to understand the specificity of pathogenetic mechanisms, namely, host susceptibility and strain variability. 15 Crohn's disease–SpA-derived AIEC protects against acute injury and death from DSS-induced colitis in WT mice. Resident microbiota, including AIEC, induce colonic RORγt/Foxp3+ CD4+ T cells, which play an important role in restraining inflammatory colitis. 98 Consistently, a higher Enterobacteriaceae in six-month-old infants correlated with better nutritional status. 99 Thus, in situations of nutritional sufficiency or immunocompetence, the response to Enterobacteriaceae may have coevolved to protect the host; however, persistent nutritional deficiency 99 or genetic susceptibility (modelled in IL-10-deficient and K/BxN mice) evokes maladaptive responses, which, in turn, promote more severe inflammatory Th17 disease. Likewise, these data link the shared genetic susceptibility in the IL23R locus in both Crohn's disease and SpA 100 with increased systemic E. coli sero-reactivity and Th17 inflammatory cytokines. These results highlight the functional implication of IgA-coated E. coli enriched in Crohn's disease–associated-arthritis and identify a Th17 immunophenotype characteristic of this EIM. This mechanistic link between intestinal microbiota and systemic inflammation may underlie the clinical efficacy of sulfasalazine in peripheral joint symptoms. 101 While anti-TNFα therapy improves axial symptoms in patients with active Crohn's disease, 102 these data also highlight the overactivation of the IL-23/IL-17 pathway in Crohn's disease patients with peripheral symptoms.
This review displays several methodological and theoretical limitations. First, heterogeneity of studied populations (in terms of age, gender and origins) and microbiota-analysing methodology deeply impact the gut microbiota picture. The purpose of our study, that is, to identify similarities and differences between gut microbiome in IBD and in CRD patients, is challenging considering also the relatively small number of studies in CRD compared with IBD.
Indeed, the first studies analysing gut microbiota in IBD were published in 2005, whereas gut microbiota in CRD has been explored a decade later. Since the first studies, more than 4000 IBD patients have been analysed whereas only 300 have been for CRD.
Second, inconsistencies may exist among the findings from available studies due to the heterogeneity in sample size, biopsy location, local inflammation and types of samples (biopsy vs. stool), which may influence the microbiota composition. Furthermore, complexity of the microbiota must be put into perspective along with current technological limitations (analysing DNA encoding 16S RNA gene still provides only an incomplete picture of bacterial populations and some studies presented here used culture dependent determination methodology).
Despite these considerations and in an effort to synthetize already published data we provide detailed tables by clinical condition and sample type as well as a figure providing an overview of the data available (Figure 2).
Finally, information on the possible concomitant arthritis and IBD was not provided in some of the 80 included studies involving IBD and CRD patients. It is thus impossible to rule out the presence of subclinical joint–gut inflammation in these patients.
We can mention also the absence of healthy control groups in certain studies or the incomplete description of clinical situation of patients (for instance patients with IBD history without information on disease activity or medication or faeces consistency score at time of sampling) that could influence gut microbiota. 103
Bacteria are not the only component of gut microbiota, fungi and virus may have a role in both diseases' initiation or severity. Bacteria and fungi could compete for the same subtracts or produce synergistically metabolites that could affect host immunity and metabolism. Only a few studies on intestinal fungal microbiota and its relationship with IBD have been conducted. Much evidence has shown that fungi and their communities may be involved in the pathogenesis of IBD, especially Crohn's disease. 104 To date fungal microbiota implication in CRD has not been explored.
The enteric virome is known to be altered in patients with IBD, with specific changes assessed between ulcerative colitis and Crohn's disease. Enormous numbers of candidate viruses have been thought to be the triggering factor of arthritis, particularly of RA, but most of the evidence implicating viruses in the pathogenesis of CRD is circumstantial and inconclusive. Tantalizing observations have often been based on in vitro or animal studies, case reports, or studies with small sample sizes, cross-sectional designs or without control groups.
The description of the viral, fungal, bacterial metagenomes in patients suffering from IBD and/or CRD shall provide a better understanding of the interactions between the microbiome and host immunity within the joint–gut axis. The identification of specific species in well-defined categories of patients can provide valuable information, which can be translated into prognostic, diagnostic or therapeutic tools that are critically lacking for these diseases. Furthermore, such studies hold great promise for the development of future strategies aiming at early detection of relapse and at controlling/manipulating the microbiome to reduce the burden of these ailments.
In conclusion, a total of 80 studies investigated the bacterial microbiome in patients with IBD and/or CRD. These studies showed that some bacterial taxons seem specifically imbalanced in IBD (n = 40) and CRD (n = 33), while showing increased abundance in Firmicutes genera Lactobacillus and Staphylococcus, Actinobacteria Bifidobacterium, and Proteobacteria genera such as Pseudomonas, Klebsiella and Proteus, whereas Firmicutes phyla Faecalibacterium, Roseburia genera and Verrucomicrobia phylum are decreased in both CRD and IBD. Large and well-designed prospective studies are eagerly awaited to further elucidate the role of gut microbiome in promoting pathological inflammation within the joint–gut axis.
Footnotes
Acknowledgements
LPB and DM contributed equally to this work. FS and NK: study concept and design; acquisition, analysis, review and interpretation of data; manuscript preparation and critical revisions. NK: study concept and design; review and interpretation of data; manuscript preparation and critical revision. JRM, PN, AL, TK, SD: review and interpretation of data; manuscript preparation and critical revisions. JYJ, LPB DM: study concept and design; review and interpretation of data; manuscript preparation and critical revisions; study supervision.
Declaration of conflicting interests
The authors declare that there is no conflict of interest.
Funding
This work was supported by the French PIA project « Lorraine Université d'Excellence », reference ANR-15-IDEX-04-LUE.
