Abstract
Background
Eradication rates with triple therapy for Helicobacter pylori infections have currently declined to unacceptable levels worldwide. Newer quadruple therapies are burdened with a high rate of adverse events. Whether multi-strain probiotics can improve eradication rates or diminish adverse events remains uncertain.
Methods
Relevant publications in which patients with H. pylori infections were randomized to a multi-strain probiotic or control were identified in PubMed, Cochrane Databases, and other sources from 1 January 1960–3 June 2015. Primary outcomes included eradication rates, incidence of any adverse event and the incidence of antibiotic-associated diarrhea. As probiotic efficacy is strain-specific, pooled relative risks and 95% confidence intervals were calculated using meta-analysis stratified by similar multi-strain probiotic mixtures.
Results
A total of 19 randomized controlled trials (20 treatment arms, n = 2730) assessing one of six mixtures of strains of probiotics were included. Four multi-strain probiotics significantly improved H. pylori eradication rates, five significantly prevented any adverse reactions and three significantly reduced antibiotic-associated diarrhea. Only two probiotic mixtures (Lactobacillus acidophilus/Bifidobacterium animalis and an eight-strain mixture) had significant efficacy for all three outcomes.
Conclusions
Our meta-analysis found adjunctive use of some multi-strain probiotics may improve H. pylori eradication rates and prevent the development of adverse events and antibiotic-associated diarrhea, but not all mixtures were effective.
Introduction
Helicobacter pylori infection remains of global concern with a prevalence ranging from 70–90% in developing countries and 25–50% in developed countries.1,2 The pathogen induces chronic gastritis and, over time, may result in severe consequences ranging from peptic ulcers, gastric adenocarcinoma and gastric mucosa-associated lymphoid tissue lymphoma.2,3 Early eradication of H. pylori prevents these consequences and is the best strategy to reduce the risk of gastric cancer. 4
Treatment with triple therapy (two antibiotics and a proton-pump inhibitor (PPI)) has lost efficacy (now ranging from 71–81%) due to the development of antibiotic resistance in many parts of the world (ranging from 11–63% for clarithromycin, 17–86% for metronidazole, and 14–24% for levofloxacin).5–8 To overcome the development of antibiotic resistance, newer strategies using sequential or concomitant quadruple regimes (adding either another antibiotic or bismuth-subcitrate) have resulted in improved H. pylori eradication rates (85–92%).9–12 However, some newer quadruple combinations still result in a high frequency of side-effects (23% for sequential therapies and 46% for concomitant therapies).8,11,12 and some studies report low cure rates (62–67%). 13 Poor patient compliance with the eradication therapy due to the development of side-effects may be associated with higher treatment failure rates and may favor the development of antibiotic-resistant strains of H. pylori. 14 To reduce the side effects and possibly lower antibiotic resistance development, alternative treatments are being developed.
Adjunctive use of probiotics may be an effective way to improve the H. pylori eradication rates or reduce the related side effects due to antibiotic use.15–17 Probiotics are defined as “live microorganisms that, when administered in adequate amounts, confer a health benefit on the host (p. 507).” 18 Some probiotics have mechanisms of action directed against H. pylori, including inhibiting H. pylori attachment to mucosal cells,19–21 regulating the immune response to H. pylori, 22 or direct physiologic effects. 23 However, other studies found the use of probiotics did not improve the H. pylori eradication rates,24,25 nor reduce the side-effects of antibiotic use.26,27 The heterogeneity of these study outcomes has led to clinical uncertainty about how to best treat patients with this infection. In addition, pooling the data across different probiotic strains leads to erroneous conclusions, as the efficacy of a probiotic strain (or mixture of strains) has been found to be strain-specific.16,18 Thus, pooling data for meta-analyses for probiotics needs to be done only within similar probiotic strain types. A review of the literature identified over 50 trials using single or multi-strain mixtures of probiotics in patients with H. pylori infections. The efficacy of single strain probiotics for H. pylori infections has been reported elsewhere. 28 The aims of this meta-analysis are to analyze the efficacy of multi-strain probiotic mixtures for the eradication of H. pylori, and the reduction of adverse effects and antibiotic-associated diarrhea commonly linked with eradication therapy.
Methods
This meta-analysis follows Preferred Reporting Items for Systematic reviews and Meta-analysis (PRISMA) guidelines29–31 with clearly delineated parameters, a priori inclusion and exclusion criteria and a standard data extraction form (see Supplementary Material, Table S1).28,31 This study was registered at the systematic review website and assigned the number: PROSPERO CRD42015017157.
Eligibility criteria
We considered all clinical trials from 1980 until 3 June 2015 in which participants were being treated with H. pylori eradication therapy and randomized to either treatment with a multi-strain probiotic mixture or a control group. A multi-strain probiotic is a mixture of two or more different strains of bacteria or fungi used at an adequate dose to improve health or functioning of the host’s body. The type of control group may have included either a placebo (blinded study) or no treatment (open study). Additional inclusion criteria included trials in pediatric or adult populations (inpatient or outpatients), published in peer-reviewed journals or on clinical trial websites, or as meeting abstracts. The type of probiotic intervention included probiotics in any delivery vehicle (e.g. capsule, sachet, tablets, drink, etc.) given in conjunction with the H. pylori eradication therapy. Trials investigating non-specific probiotics or yogurts (e.g. articles not providing the probiotic species used) were excluded. The most recent probiotic strain designations are presented in this study for those strains whose names have changed over time (older articles may have reported a different strain designation). The taxonomy of the probiotic strain type was confirmed by correspondence with authors or the probiotic manufacturing company. For studies reporting intent-to-treat (ITT) and as-per-protocol (APP) results, the ITT data was used whenever provided. All participants were required to have received H. pylori eradication therapy (double, triple, or quadruple therapy) including at least one antibiotic and one proton-pump inhibitor. Non-English language trials were translated and included. Exclusion criteria included pre-clinical studies, safety, kinetic or formulation phase 2 studies, case reports or case series, duplicate reports, trials of unspecified types of probiotics, non-randomized trials, or incomplete or no outcomes reported. Trials which did not assess either H. pylori eradication rates or the incidence of adverse events were excluded. Multi-strain probiotics with only one randomized controlled trial (lacking at least one other confirmatory trial) were also excluded. If studies had multiple types of control groups, we chose the controls most similar to the probiotic group (for example, for a trial comparing a probiotic in yogurt against two types of control groups (placebo yogurt or placebo capsule), the most appropriate control group selected was the placebo yogurt).
Information sources
We searched PubMed, Cochrane Central Register of Controlled Trials (http://www.cochrane.org), ISI Web of Science and two clinical trial registries (MetaRegister of Controlled Trials (http:www.controlled-trials.com/mrct) and National Institutes of Health (http://www.clinicaltrials.gov)) from the earliest date of the database to 3 June 2015. We used bibliographies of all relevant studies to do a recursive search. Additionally, we conducted grey literature searches, including abstracts from infectious disease and gastroenterology meetings, probiotic product websites, experts in the field and communication with published authors on H. pylori infections. Search terms included: probiotic*, H. pylori, randomized controlled trial, probiotic mixtures (for example, VSL#3, BioK+, Lacidofil, Lactogermine, Pb Probinul, Bifido Triple) and specific probiotic strains (for example, L. rhamnosus, L. acidophilus, Bifidobacterium lactis, Enterococcus faecalis).
Study selection
Titles and abstracts of papers were screened by two reviewers. One reviewer (LVM) screened all abstracts, extracted and assessed all articles using a pre-constructed and piloted, data extraction form. 28 Each of three other reviewers (YH, LW, PM) independently extracted data from one-third of the articles (each sent different articles). Any disagreements were resolved by a third reviewer. For articles published in abstract form only or for any missing significant data in full articles, further information was sought by contacting authors or by the company manufacturing the probiotic product. Using a standardized data extraction form, we systematically collected items relating to the study design, outcomes, probiotics used and types of controls, study performance and factors relating to study quality.
Data collection process
The data extraction form collected data items on study performance measures and risk of bias: (a) study design (Participants, Intervention, Comparisons, Outcomes, Settings; appropriate targeted study population, interventions well described (dose, strains, duration, formulation), type of comparators, objective outcomes, type of study design), defined inclusion/exclusion criteria, appropriate duration of treatment and follow-up planned and a priori aims); (b) study performance (similar study population characteristics by group, sufficient power achieved, recruitment method described and recruitment rate reported, a priori subgroup analysis done, balanced randomization achieved, CONSORT (Consolidated Standards of Reporting Trials) flow-chart done, premature study discontinuation, limitations and generalizability discussed, comparison of study outcomes with other studies); and (c) domain-based risk of bias across studies (selection bias (random sequence generation and allocation concealment), performance bias (blinding of study staff and participants), detection bias (blinding of outcome), attrition bias (incomplete outcome reporting) and reporting bias (selective outcome reporting) and other (miscellaneous reasons for bias)).32,33
Summary measures
Three primary end-points of this meta-analysis were used to address the following: (a) eradication rates of H. pylori, (b) frequency of adverse events (AEs) and (c) frequency of antibiotic-associated diarrhea (AAD). The outcome for H. pylori eradication was defined by having a positive assay (pre-intervention) and a negative H. pylori assay done after the follow-up period. H. pylori infection was diagnosed using at least one of the following assays: 13 C urea breath test (UBT), histology, serology, rapid urease test, stool test or culture. 10 The outcome for adverse events included any symptoms associated with eradication therapy (nausea, bloating, vomiting, diarrhea, metallic taste) and were grouped as “any AE.” The outcome for antibiotic-associated diarrhea was defined as reported diarrhea or colitis, which developed during the intervention or during the follow-up periods.
Sub-group analysis/meta-regression
Outcomes were also stratified a priori into groups to assess if differences in efficacy are associated with specific variables.32,34 These sub-groups were defined by: daily high or low dose of probiotics (≥5 × 109 colony-forming units (cfu) versus <5 × 109 cfu), type of study population (adult versus pediatric), baseline H. pylori disease state (asymptomatic carrier versus symptomatic), type of eradication therapy (triple versus quadruple).
Statistical analysis
Meta-analysis was conducted pooling the data from individual trials grouped by the type of multi-strain probiotic for each of the three outcomes (eradication for H. pylori, the incidence of AEs and incidence of AAD). For each type of multi-strain probiotic, separate pooled relative risks (RRs) and corresponding 95% confidence interval (CI) were calculated using the DerSimonian Laird method. Heterogeneity was evaluated using Cochran Q statistic (X2 with p-value) based on pooled relative risks by the Mantel-Haenazel method and I2 statistic (which indicates the proportion of total variability attributed to heterogeneity). 32 Statistical analysis was performed using Stata software version 12 (Stata Corporation, College Station, Texas, USA) to calculate pooled relative risks (RRs), number-needed-to-treat (NNT) and publication bias statistics. A meta-regression was done with the type of probiotic mixture as covariates, using tau2 estimates to indicate the proportion of study heterogeneity explained by the subgroup covariates (between study variance). Questionable studies (meeting abstracts or low study quality) were also tested by exclusion sensitivity analysis to assess their impact on pooled outcome estimates. To assess for publication bias, a funnel plot, as well as a weighted regression (Egger's test) and a rank correlation test (Begg's test for small study effects) were conducted.35–37
Results
Study selection
The literature review yielded 317 abstracts relating to probiotics and H. pylori that were screened for inclusion. Of those, 231 were excluded after initial screening according to our exclusion criteria (Figure 1): reviews (n = 146), pre-clinical animal models or phase two studies for pharmacokinetics, formulation or safety (n = 70), no control group (n = 6), not randomized (n = 5) or other miscellaneous reasons (n = 4). Of the 86 full articles or meeting abstracts retrieved, 67 were excluded: 39 were studies using single strain probiotics and have been analyzed separately
28
and 28 studies using multi-strain probiotics were excluded (see Appendix 1) due to: no confirmatory randomized controlled trial (RCT) for mixture (n = 12), developmental studies for safety, kinetics or mechanism-of-action (n = 8), no H. pylori eradication therapy given with probiotics (n = 3), undefined probiotic product with no species identification (n = 3), or non-randomized trials (n = 2).
Flow chart of the systematic review.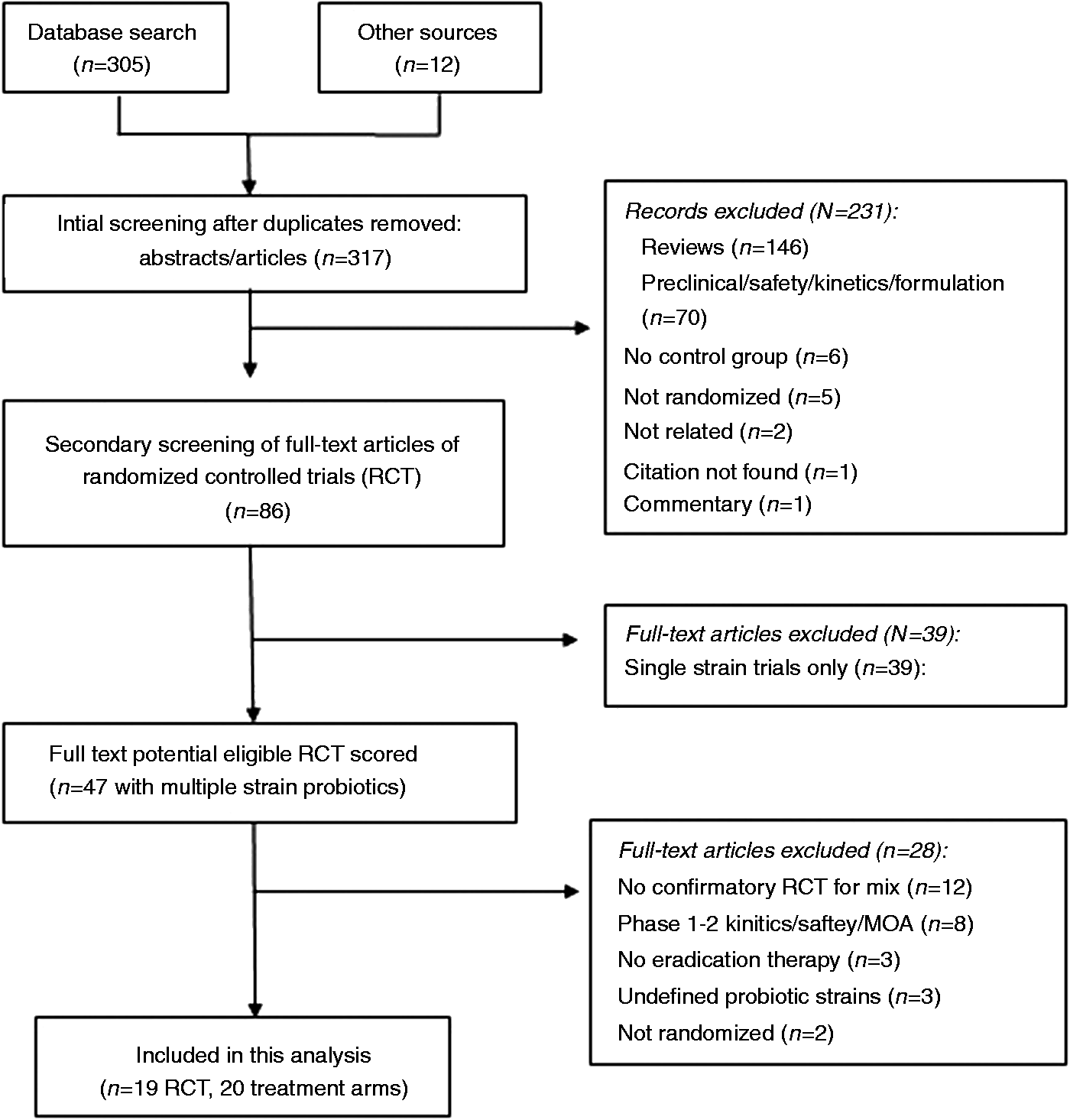
Characteristics of included trials
Included trials
Characteristics of H. pylori eradication and probiotic treatments by type of probiotic mixture.
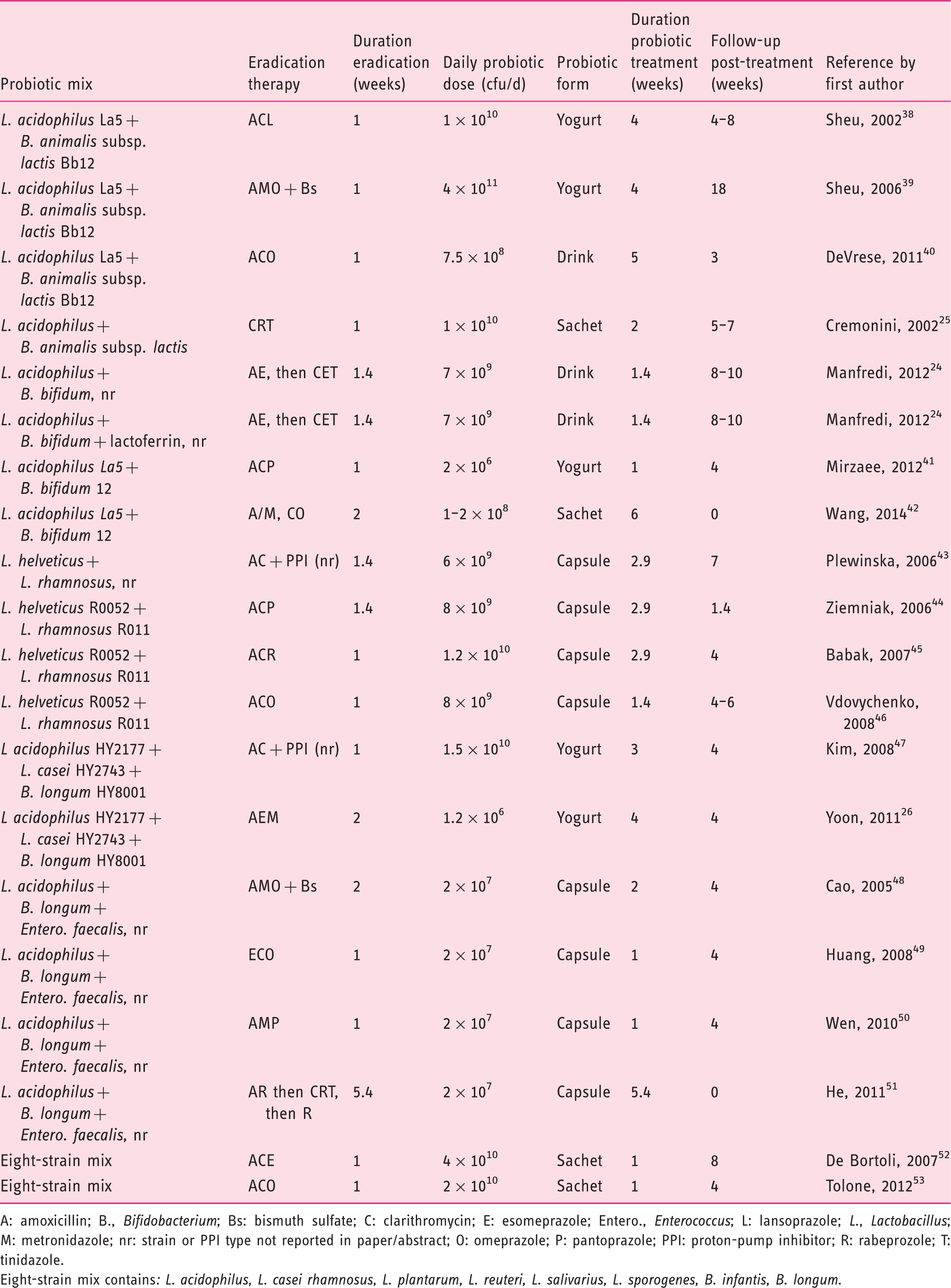
A: amoxicillin; B., Bifidobacterium; Bs: bismuth sulfate; C: clarithromycin; E: esomeprazole; Entero., Enterococcus; L: lansoprazole; L., Lactobacillus; M: metronidazole; nr: strain or PPI type not reported in paper/abstract; O: omeprazole; P: pantoprazole; PPI: proton-pump inhibitor; R: rabeprozole; T: tinidazole.
Eight-strain mix contains: L. acidophilus, L. casei rhamnosus, L. plantarum, L. reuteri, L. salivarius, L. sporogenes, B. infantis, B. longum.
Randomization
Of the 20 treatment arms, all were randomized, but only eight (40%) reported the method of randomization (e.g. computer random number generator, random block design). In one study, randomization was poorly described, 46 and in another study, the equal allocation into different randomized groups did not appear to be successful. 44
Degree of blinding
Of the 20 treatment arms, only three (15%) were double-blinded using placebos and two arms were single blinded (participants assigned to placebo drink, but study staff was aware of treatment assignment. 24 In one study, no placebo was used, but the outcome assessor was blinded as to the treatment given. 39 The placebos used in five control arms were of identical appearance and given in the same delivery vehicle as the probiotic: sachets, 25 or fruit drink, 40 or yogurt, 41 or fermented drink (two arms). 24 Most, 15 (75%) of the treatment arms were open trials (no placebos).
Attrition
Attrition ranged from 0–17% in the 20 treatment arms, usually due to drop-outs due to adverse events or loss to follow-up. Ten (50%) of the treatment arms reported no attrition, seven (35%) had attrition frequencies from 3–9% and only three (15%) reported higher attrition (12–17%).26,41,42 Of the 20 treatment arms, 17 (85%) used ITT analysis and only three (15%) used APP analysis.25,41,42 However, none of the trials with attrition report how the ITT analysis incorporated the missing data in their analysis.
H. pylori eradication therapy
All trials were required to use an H. pylori eradication therapy, including at least one antibiotic and one PPI for both the probiotic and control group (Table 1). Of the 20 treatment arms, most used triple therapy (n = 15, 75%), which most commonly included two antibiotics (amoxicillin and clarithyromycin) combined with a PPI. Less commonly used were concomitant quadruple therapy (n = 2 arms, 10%) or sequential quadruple therapy (n = 3 arms, 15%). Overall, the duration of eradication therapy ranged from one week (60% of treatment arms), or 10 days (20%), or two weeks (15%) or five weeks (5%). Most trials (n = 18, 90%) followed patients after eradication treatments were completed (40% for four weeks), with follow-up duration ranging from 10 days–18 weeks. Only half of the trials had sufficient follow-up times (>4 weeks) to allow determine long-term eradication or delayed-onset adverse events, and 10% did not have any follow-up post-treatment.
Participants
The characteristics of the enrolled study populations by trial arm are presented in Appendix 2. Most (85%) of the studies were in adults, only three were in children42,43,53 and all studies included both genders. Race or ethnicity was not reported in most clinical trials. All treatment arms enrolled H. pylori positive participants who were either symptomatic (n = 17, 85%), or asymptomatic carriers (n = 2, 10%), or had a mixed population (n = 1). 52
Probiotic treatments
In the 20 treatment arms, six different multi-strain probiotic mixtures were tested (Table 1). Most (12, 60%) of the mixtures consisted of two strains, six (30%) had three strains and two (10%) were mixtures of eight strains. Strains used as fermentation starters for yogurts or fermented drinks (Lactobacillus bulgaricus or Streptococcus thermophilus) are not counted as probiotic strains. Daily doses of probiotic mixtures ranged from 1 × 106 to 4 × 1011/day, with 8/20 (40%) arms using low dose (<1 × 109/day), five (25%) using 1–9 × 109/d and seven (35%) using higher doses (>1 × 1010/d), as shown in Table 1. Most of the probiotic formulations were in capsules (8/20 arms, 40%), while other formulations were less common: yogurt (5, 25%), or sachets (4, 20%) or fermented drinks (3, 15%). A few probiotic formulations contained additional ingredients not given to the control group: additional Vitamin B complexes, 25 or lactoferrin, 52 or prebiotics (galacto-oligosaccharide 24 or inulin).52,53 The length of the probiotic treatment ranged from one week to six weeks, with 10 (50%) arms for 1–2 weeks and 10 (50%) for 3–6 weeks.
Efficacy of probiotics for H. pylori eradication
Pooled efficacy by probiotic mixture type
Helicobacter (H.) pylori eradication rates and prevention of adverse events in 20 treatment arms with adjunct probiotics.
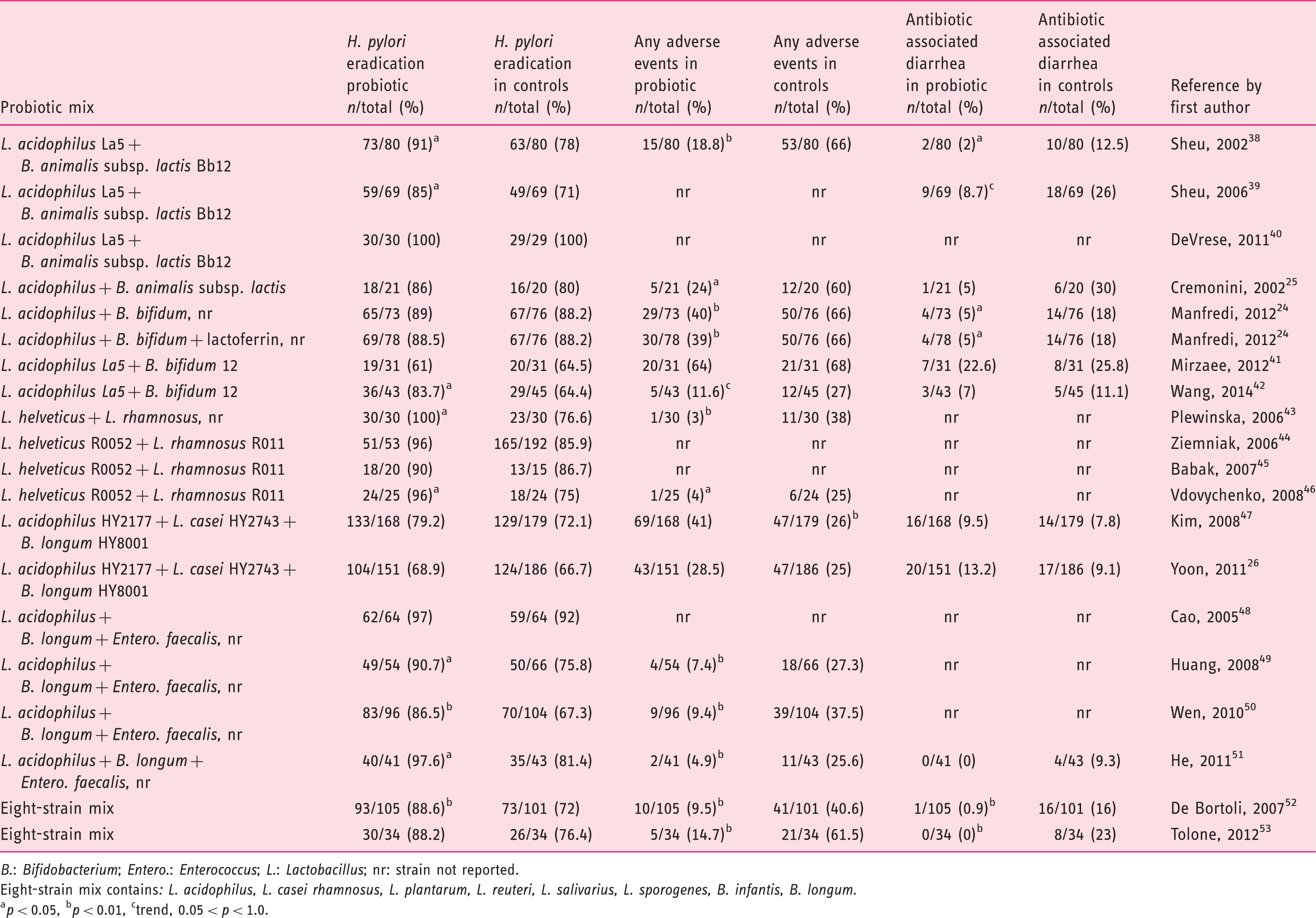
B.: Bifidobacterium; Entero.: Enterococcus; L.: Lactobacillus; nr: strain not reported.
Eight-strain mix contains: L. acidophilus, L. casei rhamnosus, L. plantarum, L. reuteri, L. salivarius, L. sporogenes, B. infantis, B. longum.
p < 0.05, bp < 0.01, ctrend, 0.05 < p < 1.0.
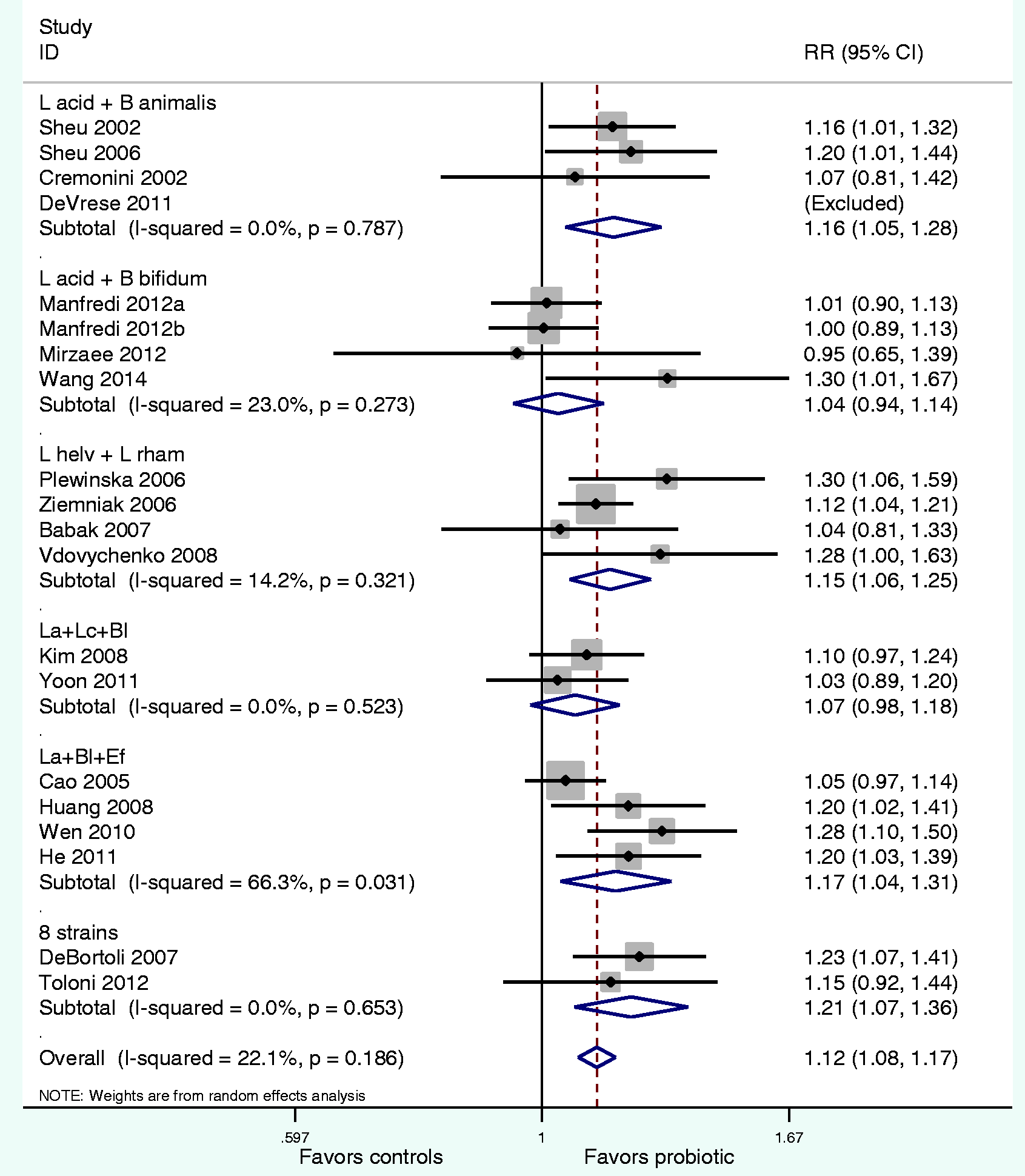
Forest plot of relative risks of Helicobacter pylori eradication by probiotic mixture groups compared to control groups. Eight-strain mix contains: Lactobacillus acidophilus, L. casei rhamnosus, L. plantarum, L. reuteri, L. salivarius, L. sporogenes, B. infantis, B. longum.
Sub-group and meta-regression analyses
Random models were re-run stratifying on subgroup variables for pooled eradication RR. Most subgroup variables (dose, study population, risk of bias) did not result in a significant change in either magnitude or direction of the effect estimate for either eradication. H. pylori eradication rates were significantly better when probiotics were given to symptomatic patients (RR = 1.13, 95% CI 1.09–1.17), but had no significant effect on asymptomatic carriers (RR = 1.07, 95% CI 0.81–1.42, p = 0.63). Results from the meta-regression analyses for the adjunctive use of probiotics found a trend for higher H. pylori eradication rates in adult populations (p = 0.08), but did not find significant differences in associations between daily dose of probiotic (above or below 5 × 109 cfu/d, p = 0.86), or disease state (asymptomatic carriage versus symptoms, p = 0.67). As the majority of the trials used triple therapy (75%) and the eradication rate was low (mean of 74%) in the 15 control groups, we also assessed each type of multi-strain probiotic as to whether they were given with triple or quadruple eradication therapy. Only two multi-strain probiotics achieved high eradication rates when given with triple therapy: L. helveticus/L. rhamnosus (96% cured) and L. acidophilus/B. animalis (92% cured). Of the five treatment arms using quadruple therapy, adjunct use of multi-strain probiotics improved the eradication rate compared to concomitant quadruple therapy controls (91% and 81%, respectively) and in sequential quadruple groups (91% and 87%, respectively). The most effective specific multi-strain probiotic combined with quadruple eradication therapy could not be proved due to the limited number of treatment arms using these regimens.
Efficacy of adjunct probiotics for prevention of any adverse reactions
Of the 20 treatment arms, 15 (75%) planned a priori to document if any adverse events occurred during the intervention and follow-up period, while 5 (25%) did not document total adverse events (Table 2). Significant heterogeneity was found (X2 = 97.1, p < 0.001, I2 = 85.6%) and a random-effects model was used, weighted on study size. The forest plot by the type of multi-strain probiotic mixture (Figure 3) shows five mixtures significantly reduced the incidence of adverse events associated with standard H. pylori eradication therapies: L. acidophilus/B. animalis mix (RR = 0.31, 95% CI 0.20–0.47), L. acidophilus/B. bifidum mix (RR = 0.67, 95% CI 0.50–0.88), L. helveticus/L. rhamnosus mix (RR = 0.12, 95% CI 0.03–0.50), L. acidophilus/B. longum/E. faecalis mix (RR = 0.25, 95% CI 0.15–0.42) and the eight-strain mixture (RR = 0.24, 95% CI 0.14–0.39). Only the one mixture (L. acidophilus/L. casei/B. longum mix) did not significantly reduce the development of adverse events. Subgroup analysis did not find any significant differences by the daily probiotic dose, type of enrolled population, or by symptoms at baseline. Overall, probiotics given with triple therapy decreased adverse events by 14% (mean AEs in probiotics was 22% compared to 36% in controls). Efficacy in reducing AEs by type of multi-strain probiotic was similar to the meta-analyses results above when given with triple therapy. In the three treatment arms using sequential quadruple therapies, multi-strain probiotics reduced AEs by 25% (mean 32% in probiotic groups versus 57% in control groups). The limited number of trials using quadruple therapy and assessing AEs precluded the determination of which multi-strain probiotic was effective when given with sequential therapies. There were no trials assessing AEs using concomitant quadruple therapies.
Forest plot of relative risks for any adverse events by probiotic mixture groups compared to control groups. Eight-strain mix contains: Lactobacillus acidophilus, L. casei rhamnosus, L. plantarum, L. reuteri, L. salivarius, L. sporogenes, B. infantis, B. longum. B: Bifidobacterium; Bl: B. longum; CI: confidence interval; Ef: Enterococcus faecalis; L: Lactobacillus; La: L. acidophilus; Lc: L. casei; L helv: L. helveticus; L rhamn: L. rhamnosus; RR: relative risk.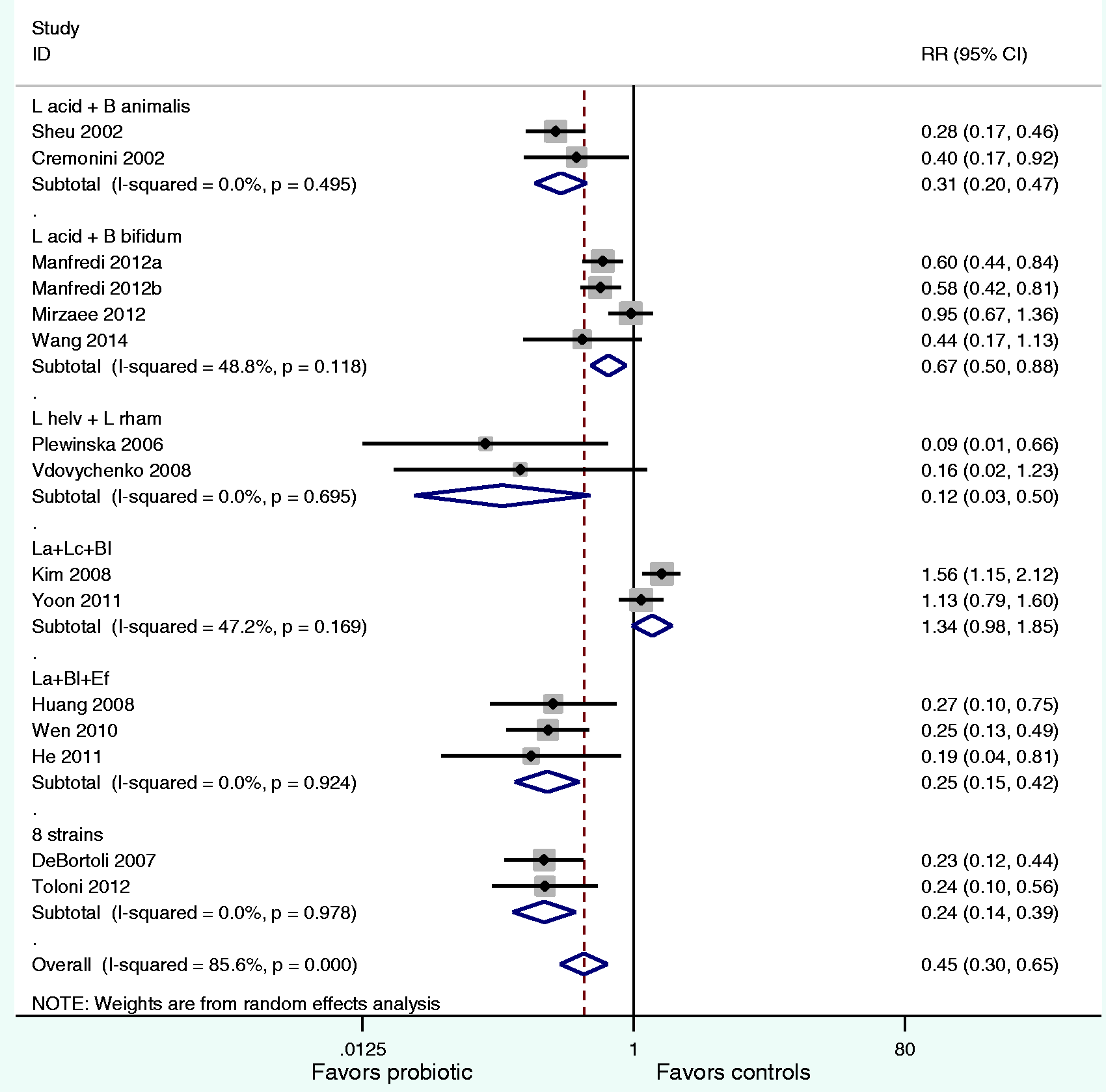
Efficacy of adjunct probiotics for the prevention of antibiotic-associated diarrhea
Of the 20 treatment arms, 12 (60%) planned a priori to document if antibiotic-associated diarrhea developed during the intervention and follow-up period, while eight (40%) did not document AAD outcomes (Table 2). Significant heterogeneity was found (X2 = 30.9, p = 0.001, I2 = 64.4%) and a random-effects model was used, weighted on study size. The forest plot by the type of probiotic mixture (Figure 4) shows only three multi-strain probiotics were effective: L. acidophilus/B. animalis mix (RR = 0.38, 95% CI 0.20, 0.72), L. acidophilus/B. bifidum mix (RR = 0.47, 95% C.I. 0.26, 0.86), the eight-strain mix (RR = 0.06–95% CI 0.01–0.30). Two of the multi-strain mixtures were not effective against AAD: L. acidophilus/L. casei/B. longum mix and L. acidophilus/B. longum/E. faecalis mix. One multi-strain mixture could not be assessed with pooled RRs due to insufficient trials with AAD data. Subgroup analysis did not find any significant differences by the daily probiotic dose, type of enrolled population, or symptoms at baseline. Overall, probiotics given with triple therapy decreased AAD by 4.5% (mean AAD in probiotics was 7.9% compared to 12.4% in controls). Efficacy in reducing AAD by type of multi-strain probiotic was similar to the meta-analyses results above when given with triple therapy. In trials using sequential quadruple therapies, multi-strain probiotics reduced AAD by 12% (mean 4.2% in probiotic groups versus 16% in control groups). There was only one trial assessing AAD using concomitant quadruple therapy (13% in probiotic versus 26% in controls). The limited number of trials using quadruple therapy and assessing AAD precluded the determination of which multi-strain probiotic was effective when given with sequential therapies.
Forest plot of relative risks for prevention of antibiotic-associated diarrhea by probiotic mixture groups compared to control groups. Eight-strain mix contains: Lactobacillus acidophilus, L. casei rhamnosus, L. plantarum, L. reuteri, L. salivarius, L. sporogenes, B. infantis, B. longum.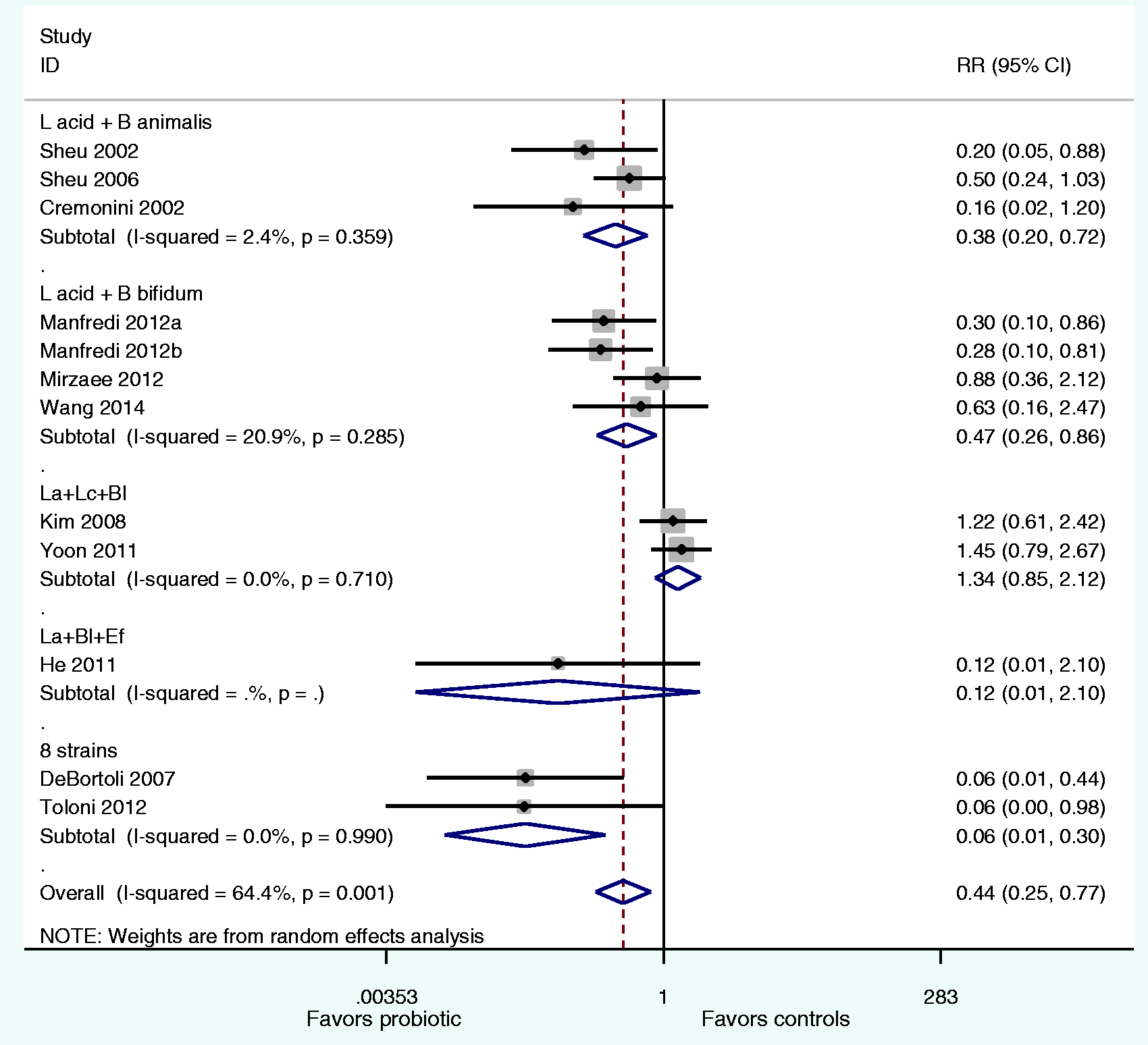
Publication bias
A funnel plot analysis (see Appendix 3) provides no compelling indication of significant publication bias for trials evaluating H. pylori eradication outcomes, showing general symmetry of the funnel for the relationship between risk ratio and standard error. Egger's regression test for small study effects (p = 0.12) and Begg's rank test (p = 0.22) fail to suggest significant publication bias. Potential publication bias is found for trials assessing the prevention of all adverse events (Egger's regression p = 0.04 and studies assessing AAD (Egger's regression p = 0.01), as there are few outliers noted for small study sizes.
Study performance and risk of bias
Study design and available risk of bias data was assessed for all trials. Reviewers reported most of the 33 items assessed using the data extraction form were available (30% had 76–100% of the items, 50% had 51–75%, while only 20% had fewer than 50% of the items). Concordance of the reviewers was excellent (kappa = 0.99, p < 0.001). Any disagreements were resolved after discussion. Study design variables were typically well-described, but only 25% included sample size calculations, and 75% of the trials used open study designs (only 15% were double-blinded and 15% were single blinded). Study performance was also evaluated: all reported their attrition rates and 82% provided the reasons for attrition by the treatment groups, but only 50% described how participants were recruited, 25% failed to provide a comparison of the two treatment groups at baseline and only 21% provided a CONSORT figure describing study flow. In the discussion section of the papers, although 85% compared their results to other studies, only 45% discussed limitations of their trial and 95% failed to discuss the generalizability of their results. Risk of bias was also assessed using the Cochrane Collaboration tool using six key domains. 32 For the 20 RCTs (see Appendix 4), there was a low risk of bias from attrition and reporting bias, but a high risk of performance bias due to the lack of double-blinded study designs (most were open studies). Although all the included trials were required to be randomized, 20% of the studies had high risk of selection bias as the method of randomization was not reported. A lack of reporting how the allocation was concealed (selection bias) and if the outcome assessor was blinded (detection bias) resulted in a high frequency of “unclear” bias in these two domains.
Discussion
Although triple therapy remains a first-line option in many countries, eradication rates rarely exceed 80%.10,55 Newer and more complex treatments consisting of quadruple combinations have targeted a 90% eradication rate.9,11 Our meta-analyses found that only certain types of multi-strain probiotic mixtures are effective for various outcomes of H. pylori infections, while other mixtures are not effective. The pooled data found some probiotic mixtures achieved >90% eradication rates when combined with triple therapy (92% for L. acidophilus and B. animalis mix, 96% for L. helveticus and L. rhamnosus mix).25,38,40,43–46 Two trials combining the mix of L. acidophilus, B. longum and Entero. faecalis with quadruple therapies had a pooled eradication rate of 97%.48,51 In multi-strain probiotics achieving >90% eradication rates, probiotic dose was typically high (67% of these trials used >1010 cfu/day) and 67% used the probiotics for 3–5 weeks.
The search for the most effective combination of multi-strain probiotic and eradication therapy is challenging. Our meta-analysis found 75% of published trials used triple therapy with their adjunct multi-strain probiotic. The low eradication rate in the triple therapy control groups (74% cure) confirms the concern that use of triple therapy may now be considered ineffective and may be unethical to use in clinical trials as a comparator. 56 Dang et al. noted in their meta-analysis of 33 trials of single and multi-strain probiotics that probiotics failed to result in a significant improvement of eradication rates if controls had good efficacy rate (>80%). 57 Unfortunately, this analysis did not report the type of eradication therapy stratified by strain-specific probiotic. We did confirm this observation, in that two of our multi-strain probiotics had similar eradication as triple therapy controls (74% cured), however, four multi-strain probiotic mixtures did significantly improve eradication rates, even when combined with triple therapy. In addition, adjunct probiotics reduced AEs by 14% and AAD by 4.5% when combined with triple therapy. The paucity of probiotic trials using quadruple therapies (only 25%) limited drawing strong conclusions on which combination of multi-strain probiotic and quadruple therapy to use. Eradication rates in controls (81% in concomitant and 87% in sequential quadruple therapy groups) were improved by the addition of multi-strain probiotics (91% and 91%, respectively). And the addition of multi-strain probiotics decreased AEs by 25% and decreased AAD by 12% when combined with quadruple therapies. But clearly, both the type of probiotic and the type of eradication therapy used are two important factors in determining the best clinical recommendations for H. pylori infections.
As our meta-analysis shows a distinct strain specificity for both the efficacy of eradicating H. pylori and the prevention of adverse events (including AAD), future studies need to be aware that pooling similar probiotics by species is no longer appropriate and their effects need to analyzed by the same type of probiotic mixture.16,28 The evidence from prior meta-analyses for the eradication of H. pylori in the literature have concluded probiotics, in general, may be efficacious for these types of infections, but two issues limit their conclusions: pooling different types of probiotics together or not performing a sub-group analysis by varying doses and duration of probiotic therapies.57–62 Pooling different strains of probiotics in meta-analysis is not recommended.16,63 Zou et al. pooled eight trials for six different Lactobacilli-containing mixtures and did conduct sub-group analyses by age, symptoms, eradication therapy and sub-species of Lactobacilli, but inappropriately grouped different species within the same group. 59 Wang et al. pooled 10 trials using Lactobacilli or Bifidobacteria containing probiotic mixtures and reported improved H. pylori eradication rates and reduced adverse events, but did not identify which mixtures were more effective than others. 60 Dang et al. reported sub-group analysis by the type of probiotic, but included sub-groups with only one randomized trial in their pooled analysis. 57 Two other meta-analyses reported a pooled protective effect of probiotics for H. pylori infections. Lv et al. pooled 21 RCTs, which included seven types of probiotic mixtures, 64 and Zheng et al. pooled nine trials, of which five were mixtures, 65 but both these meta-analyses failed to report efficacy separately by probiotic mixture type. Zhang et al. also pooled 45 trials with 11 different genera types of probiotics together for one estimate of efficacy, but their sub-group analysis inappropriately reported results by seven sub-groups of probiotic on the genus level and not species or strain level. 66 No species or strains for the probiotics were reported in this paper, which ignores the species specific efficacy of probiotics.
There have been a few attempts to test whether eradication can be achieved by probiotics alone. Only three trials were found using just probiotic mixtures without eradication therapy and two showed only a mild decrease in breath urea levels,67,68 while one mixture of eight strains (VSL#3) significantly improved the H. pylori eradication rates (32% versus 0% in placebo controls), but the effect on adverse events was not reported. 69
Another proposed improvement to eradication therapy is the addition of non-specific molecules with anti-infectious and anti-inflammatory properties. Manfredi et al. did not see any difference in eradication rates or AE or AAD when lactoferrin was added to the mixture of L. acidophilus and B. bifidum compared to the probiotic mixture alone. 24 Two trials added lactoferrin and inulin 52 or just inulin 53 to their probiotic mixtures, but did not compare the effect against probiotic mixtures alone.
This systematic review has several strengths. We collected a large number of published papers on probiotic mixtures and applied rigorous methodology for the systematic review and meta-analysis. By separating the different types of mixtures and assessing efficacy separately by mixture, clinicians can be guided as to which probiotic mixtures are most effective for patients being treated for H. pylori. We used methods to reduce bias: specific outcomes selected a priori, comprehensive literature search including any relevant trials irrespective of language or publication status (i.e. we included published data from meeting abstracts, obtained specific data from authors, and translated seven non-English trials) and had independent duplicate review of the papers. Additional strengths of the review include the evaluation of each of the subgroups (i.e. probiotic dose, type of enrolled population, disease state, and type of eradication therapy). The results of this meta-analysis may be generalizable to a wide population, as we include a range of ages, countries and settings (inpatients and outpatients, adults and children). It should be noted however, that ethnicity and race data were not reported, nor were immunocompromised patients included in most of the trials, so the applicability of our results to these types of these populations is not known.
There were some limitations for this review. Most of the published studies used triple therapy, which has a low cure rate and trials using the current recommended quadruple therapies were scarce. While we did a comprehensive search of the literature, we did not search all conference proceedings or dissertation abstracts. Probiotic strain is the key indicator of efficacy, but the limited number of trials on the same mixture of strains limits our ability draw robust conclusions on many of the strains used for all cited studies. We excluded 12 studies that only had one randomized controlled trial, and as a consequence, not all probiotic strains were included in this analysis. It is also difficult to recommend the best daily dose and duration of a probiotic mixture from this review, but in those trials with >90% cure rate the daily dose was ≥1010 cfu/day for 3-5 weeks. Our subgroup analysis did not show a significant effect of daily probiotic dose using meta-analysis, and doses used in trials with the same strain often had similar daily doses. Several types of probiotic mixtures having evidence-based efficacy for other types of diseases (such as “BioK+” for prevention of AAD or “VSL#3” for treatment of ulcerative colitis and pouchitis) could not be assessed here, as there are no clinical trials using these mixtures as adjuncts with H. pylori eradication therapy.70,71 In addition, most studies did not document the prevalence of antibiotic-resistant strains of H. pylori in their trials, which may also influence H. pylori eradication rates. Another limitation to the trials is that compliance to the eradication therapy was only documented in six (30%) of the studies and compliance to the study treatment was only done in one study. 25 A higher compliance rate was noted in one study (68% in probiotic group versus 44% in controls, p < 0.05), 38 but compliance was equivalent for probiotic and control groups in five other studies, ranging from 55–99%.26,39,47,52,53 It would be of interest to know if the addition of probiotics to the recommended eradication therapies result in better compliance rates, but these data must first be consistently reported.
This review has important clinical implications for current treatment strategies of patients with H. pylori infections and some suggestions on how to improve the scientific literature of probiotic trials in the future. Reliance on meta-analyses for clinical guidance can be useful, if interpreted correctly. Global pooling of efficacy over different types of treatments does not provide assistance in recommending which type of probiotic is clinically efficacious for H. pylori infections or which type of eradication therapy probiotics should best be combined. In this review, we have stratified the data and provided pooled results grouped by the type of probiotic mixture, which provides guidance as to which probiotic mixture are useful and which ones are not. Future studies should strive to collect data on important variables that may impact the efficacy of H. pylori treatments (including the antibiotic-resistance pattern of participant's H. pylori isolate and compliance rates to the treatments) if we are to determine if the observed efficacy of a trial is due to the probiotic, or distribution of antibiotic-resistance to the type of eradication therapy given, or if the probiotic simply improves compliance rates. 56 Future trials should also use quadruple therapy as their eradication therapy whenever possible. Improvements in study design include use of treatment concealment (double blinding), calculating sample size a priori to power a large enough study to detect significant results, use of intent-to-treat analysis to account for patient attrition effects, the collection of adverse event data and having sufficient follow-up time after the treatments are discontinued. In addition, multiple randomized, controlled trials on the same probiotic mixtures are needed, allowing confirmation of single clinical trial results. Last, but not least, the effect of probiotics on the intestinal microbiota in the context of H. pylori eradication should not be neglected in future trials.
In conclusion, our meta-analyses found four multi-strain probiotic mixtures are beneficial and safe for the eradication of H. pylori when combined with standard eradication therapy, five of the probiotic mixtures decreased the adverse events of eradication therapy, while only three reduced antibiotic-associated diarrhea.
Footnotes
Funding
This research received no specific grant from any funding agency in the public, commercial or non-for-profit sectors.
Conflict of interest
Lynne V. McFarland has received fees as a speaker (Biocodex, France and Lallemand, France) and is on the scientific advisory board of BioK+, Canada. Peter Malfertheiner has received speaker fees from Biocodex and is on the scientific advisory board of Danone. Ying Huang and Lin Wang have no conflicts of interest. No authors are employed by, or have stock or equity in, any company manufacturing probiotics.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
