Abstract
Background:
The research focused on the effects of pseudorabies virus on gene expression in piglet brain tissue.
Objectives:
The goal was to understand the changes in gene expression in piglet brains due to pseudorabies virus infection.
Design:
The study used a comparative approach with infected and control piglet groups.
Methods:
Twelve 2-month-old piglets were divided into pseudorabies virus-infected and PBS-treated control groups, with brain tissue analyzed after 7 days.
Results:
Infected piglets showed increased oligodendrocyte counts and pseudorabies virus-positive signals. Transcriptomic analysis revealed 269 differentially expressed genes, with 149 up-regulated and 120 down-regulated. Gene ontology and Kyoto Encyclopedia of Genes and Genomes analyses indicated these genes are involved in signal transduction, transmembrane transport, apoptosis, and neuroactive ligand–receptor interaction. Quantitative fluorescent PCR validated these findings, particularly for genes related to neuroactive pathways, ferroptosis, and IL-17 signaling.
Conclusion:
The study provides valuable insights into the molecular alterations caused by pseudorabies virus in piglet brain tissue, enhancing our understanding of pseudorabies virus’s pathogenic mechanisms.
Introduction
Pseudorabies (PR), also known as Aujeszky’s disease, is a disease caused by the Pseudorabies virus (PRV). PRV has caused significant economic losses to the global swine industry. 1 PRV is a complex DNA virus and a member of the Alphaherpesvirinae subfamily of the Herpesviridae family. Pigs are considered the only natural host of PRV. When piglets and adult pigs are infected with highly virulent strains of PRV, sudden death or severe clinical symptoms may occur, including neurological symptoms. 2 In addition to pigs, various animals such as cattle, sheep, cats, dogs, raccoons, minks, skunks, and mice are also susceptible to PRV. 3 Occasionally, there are reports suggesting that humans may be infected with PRV, exhibiting symptoms such as conjunctivitis, dermatitis, fatal encephalitis, and severe central nervous system disorders. 4 These reports suggest that PRV may pose a potential threat to public health. Therefore, it is particularly important to delve into the pathogenic mechanisms of PRV infection.
In recent years, transcriptomics technology has shown broad application potential in exploring the interactions between viruses and hosts. The application of transcriptome sequencing has become a key means to reveal the pathogenic mechanisms of PRV. Xu et al. 5 used mice as subjects of study and infected them with the PRV XJ delgE/gI/TK strain. Transcriptomics results showed that differentially expressed genes were mainly enriched in signaling pathways such as TNF, NOD-like receptor, JAK-STAT, MAPK, IL-17, and apoptosis. Shangguan et al. 6 performed RNA sequencing analysis on PK-15 cells infected with the recombinant strain PRV-Becker-GFP, revealing that the activation of the cytosolic DNA sensing pathway and NOD-like receptor signaling pathway may play a key role in the recognition process of PRV, while the activation of NF-κB and TNF signaling pathways may be involved in the immune response to PRV. These studies not only greatly enrich the understanding of the pathogenic process of PRV but also provide a new basis for the prevention and control strategies of PRV.
Jiangkou radish pigs, as a precious indigenous genetic resource breed in China, are mainly distributed in Tongren City, Guizhou Province. This unique local breed is facing the threat of PRV. However, there are very few reports on the interaction between Jiangkou radish pigs and PRV. This study selected Jiangkou radish pigs as the research subject, used cutting-edge transcriptomics technology, and successfully constructed a brain tissue-specific transcriptome library after PRV infection. Subsequently, we performed transcriptome sequencing to screen for host differentially expressed genes induced by PRV infection, and conducted gene ontology (GO) classification and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis. Further validation of key genes from the transcriptome sequencing was carried out using quantitative fluorescent PCR (qRT-PCR) technology. This study provides supplementary data for the complex in vivo interactions between PRV and its host and lays a solid theoretical foundation for further in-depth study of the pathogenic mechanisms of PRV and other pathogens, providing an important reference for the development of disease prevention, control, and treatment strategies.
Materials and methods
Experimental animals and grouping
Twelve 2-month-old Jiangkou radish piglets, with an average weight of 10.76 ± 0.32 kg, with an equal number of males and females, and all castrated, were randomly divided into a control group and a PRV infection group. After a 1-week adaptation period, the formal experimental phase began. Six piglets in the infection group were inoculated with the PRV-HLJ strain via intramuscular injection, with an infection dose of 1 mL per piglet, and the virus titer was 1 × 104 TCID50/100 μL. Another six piglets were injected with an equal volume of PBS as the control group. On the 7th day postinfection, brain tissue samples from the infected and control piglets were collected after electrical anesthesia, then fixed in 4% paraformaldehyde solution, and the remaining brain tissue was cooled with liquid nitrogen and stored in a −80°C refrigerator for later use.
Main reagents
Four percent paraformaldehyde was purchased from Beijing Solaibao Reagent Company; PRV-HLJ strain (MK080279.1) was preserved in this laboratory; DAB staining solution was purchased from Wuhan Boshide Biological Company, China. The porcine IL-1β and TNF-α ELISA kits were purchased from Nanjing Jiancheng Bioengineering Institute, China. Tissue RNA extraction kit (B518621) and primer synthesis were provided by Shanghai Shengong Biological Co., Ltd.. Fluorescence quantitative PCR kit (RR820A) and reverse transcription kit (RR047A) were purchased from TaKaRa Company, China. PRV antibody (ab3534) was purchased from Abcam Company, USA.
Detection of IL-1β and TNF-α in brain tissue
A 10% porcine brain tissue homogenate was prepared using a mechanical grinding method. The levels of IL-1β and TNF-α in the porcine brain tissue were detected according to the instructions of the assay kit.
Distribution of PRV in brain tissue
The expression of PRV in brain tissue samples was detected using immunohistochemical methods. After the brain tissues from both the control and infected groups of piglets were fixed with a 4% paraformaldehyde solution, they were dehydrated and embedded, then sectioned into 5 μm slices. The tissue sections were treated by de-paraffinization, antigen retrieval, and blocking, followed by the addition of PRV primary antibody diluted at 1:300 and incubated overnight. After dilution, biotinylated secondary antibody was evenly applied to the tissue sections and incubated at 37°C for 20 min. The sections were then visualized with DAB staining, counterstained with hematoxylin, blued with ammonia water, and observed under a microscope.
Total RNA extraction
Total RNA from the control and infected groups was extracted according to the instructions and quickly placed in a −80°C refrigerator for later use in constructing the sequencing library. The quality and concentration of the extracted RNA were assessed using a NanoDrop spectrophotometer and agarose gel electrophoresis. The RNA samples with an A260/A280 ratio between 1.8 and 2.0 were considered pure and were used for subsequent experiments. The integrity of the RNA was further confirmed by the presence of clear 28S and 18S ribosomal RNA bands on the gel, indicating that the RNA was suitable for library preparation.
Construction of sequencing library
Three piglet brain tissue samples from each group were sent to Shanghai Baiqu Biomedical Technology Co., Ltd. for transcriptome sequencing. The construction of the RNA-seq library is as follows: RNA meeting the standards was purified to remove rRNA and enrich mRNA. Oligo(dT) magnetic beads were used to specifically enrich eukaryotic mRNA with polyA tails. Next, ultrasonic technology was used to break mRNA into smaller fragments. These fragmented mRNAs were then used as templates to synthesize the first strand of cDNA. Then, RNaseH enzyme was used to degrade the remaining RNA fragments combined with cDNA, and the second strand of cDNA was synthesized. After the double-stranded cDNA was end-repaired, tailed, and connected with sequencing adapters, cDNA fragments of about 200 bp were selected for PCR amplification. The amplified PCR products were purified to ultimately obtain the library for sequencing. The qualified library was then sequenced on an Illumina HiSeq 2500 platform using paired-end 150 bp sequencing strategy. Raw sequencing data were processed and analyzed to ensure the quality of the sequencing results. The obtained clean reads were aligned to the reference genome, and the expression levels of genes were quantified. The subsequent analysis steps included normalization of gene expression data and identification of differentially expressed genes between the experimental groups.
GO and KEGG analysis
DAVID (https://david.ncifcrf.gov/) was used to perform GO enrichment and KEGG pathway analysis on differentially expressed genes. A p-value < 0.05 represents significant enrichment of GO and KEGG pathways.
Differential gene analysis
The transcription level changes of brain tissue samples from the control and infected groups were analyzed using RSEM, FPKM, and TPM tools; at the same time, edgeR, DESeq2, and DESeq software were used to screen and analyze genes with significant transcription differences, and corresponding charts were drawn.
Fluorescence quantitative PCR verification
Total RNA was reverse transcribed into cDNA. β-actin was used as the reference gene, and primers for the corresponding target genes were designed (Table 1), and qRT-PCR analysis was performed. The results of qRT-qPCR were processed using the 2−ΔΔCt method. In this experiment, the calculation method for the ΔΔCt value of a specific gene was (Ct value of the detected gene in the PRV-infected group—Ct value of the reference gene in the infected group)—(Ct value of the detected gene in the control group—Ct value of the reference gene in the control group). Each sample was repeated three times, and statistical analysis and data processing were performed using SPSS and GraphPad Prism 6 software, respectively. The reaction program for qRT-qPCR includes 95°C pre-denaturation for 5 min; entering the cycle at 95°C denaturation for 15 s, 60°C annealing; and extension for 40 s, for a total of 40 cycles. The reaction system includes SYBR Green 5 μL, upstream primer 0.5 μL, downstream primer 0.5 μL, cDNA 1 μL, DEPC water 8 μL.
Primer information for qRT-PCR.
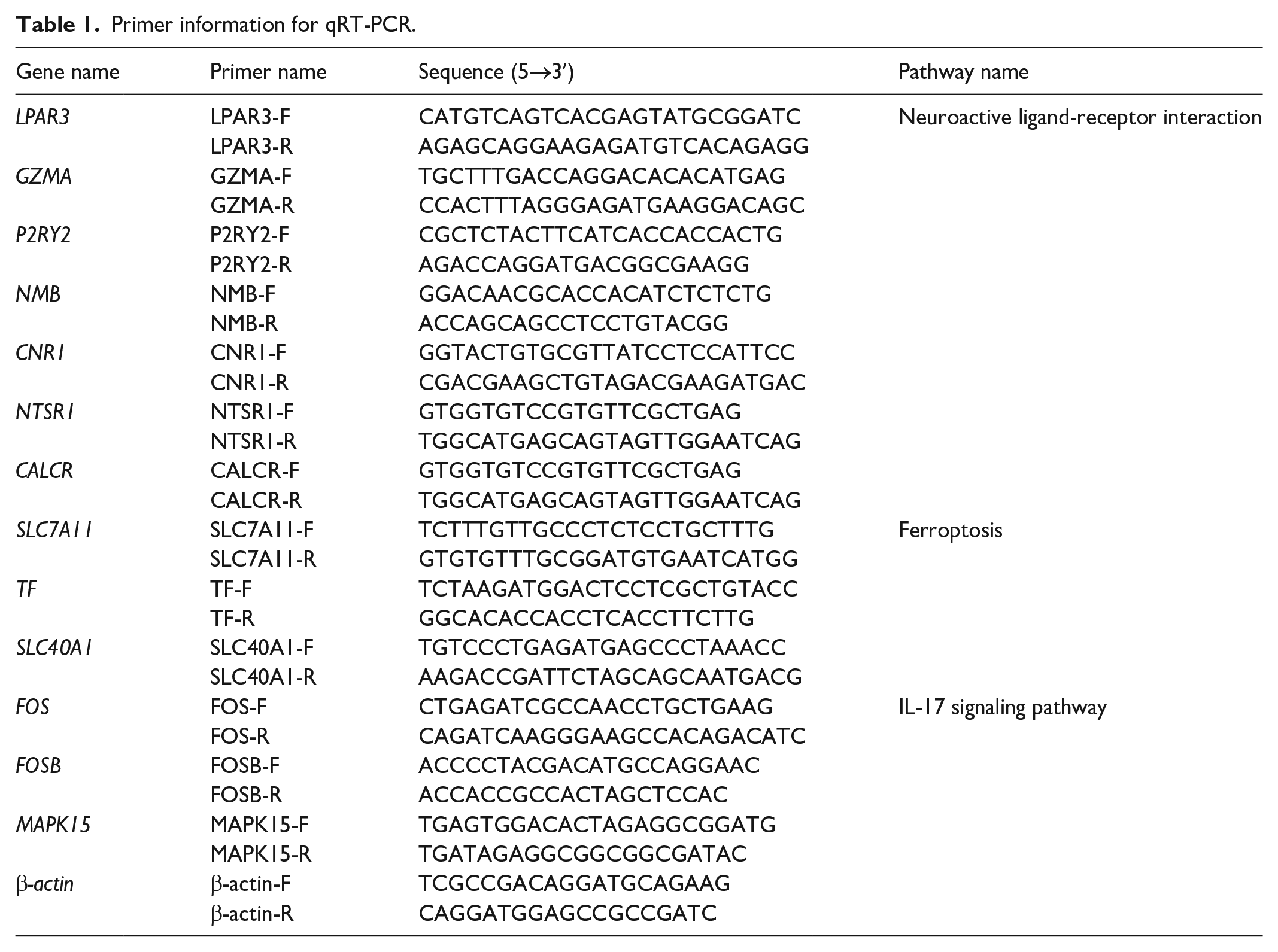
Results
ELISA and immunohistochemical analysis of PRV-infected piglet brain tissue
From the clinical symptom observations, the PRV challenge group exhibited increased body temperature, depression, and decreased appetite. The pigs showed typical neurological symptoms, such as ataxia and sneezing. There were no deaths during the trial, and the injection sites showed redness and swelling, while the control group piglets did not show any clinical symptoms. According to the results of ELISA detection in brain tissue, the concentration of IL-1β and TNF-α was significantly increased compared to that in the control group (Figure 1(a)). In addition, immunohistochemical detection using PRV antibody (Figure 1(b)) revealed that the number of PRV-positive cells (indicated by the red arrow) in the brain tissues of PRV-infected group was significantly higher than that in the control group, indicating that PRV has invaded the brain tissue of piglets.
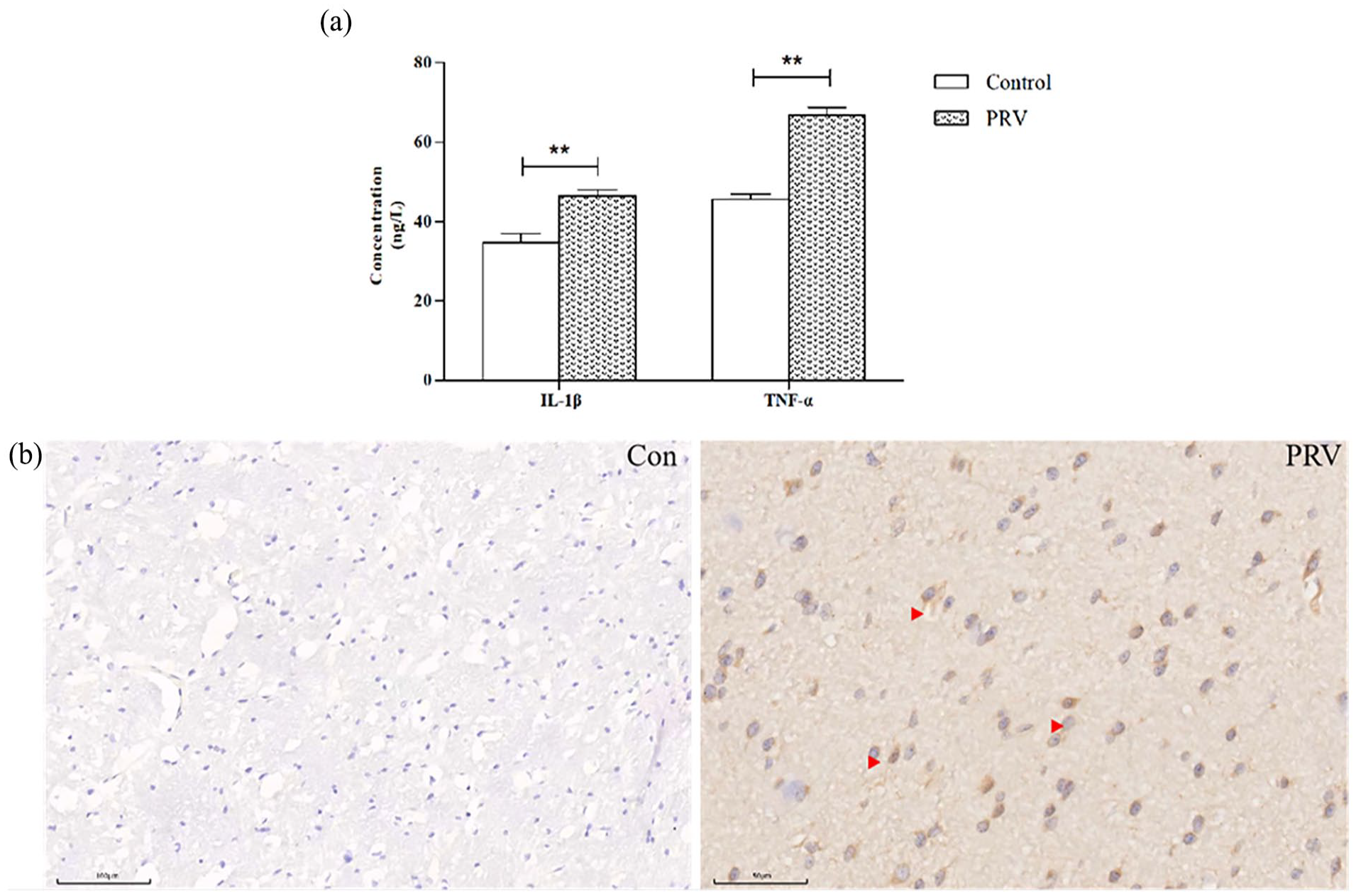
ELISA and PRV expression in brain tissue after PRV infection. (a) The results of the brain tissue IL-1β and TNF-α ELISA detection and (b) Immunohistochemical results of PRV, red arrows indicating PRV-positive cells in the brain tissues (Bar = 100 μm).
Brain tissue transcriptome library and differential gene analysis
After filtering out reads with adapter sequences, a total of 237,942,304 clear and nonrepeated reads were collected. The average proportion of reads with a quality score greater than 30 (Q30) in the control and infected groups were 98.27% and 98.22%, respectively, indicating the construction of high-quality transcriptome libraries. Genes with |log2FC| > 1 and FDR < 0.05 were considered significantly differential, and a total of 269 differential genes were screened out, including 149 upregulated genes and 120 downregulated genes (as shown in Figure 2(a)).

Differential gene analysis of PRV infection: (a) The volcano plot of differential genes illustrates the differences in gene expression between the PRV-infected group and the control group. The red dots in the plot represent upregulated genes, while the blue dots indicate downregulated genes. The gray dots signify genes with insignificant changes in expression. (b) The heatmap of differentially expressed genes, with columns representing samples from the control group and the infected group, and rows corresponding to genes that exhibit significant differential expression during the comparison process.
The heatmap of differential gene clustering analysis for individual samples is shown in Figure 2(b). The red blocks represent highly expressed genes, while the blue blocks correspond to lowly expressed genes. Through the heatmap, it can be clearly observed that the differential gene expression clustering among samples within the control group is closely similar, and similarly, the samples within the infected group also show a similar pattern of differential gene expression clustering, which is basically consistent with the expected level of the experiment.
GO analysis of differential genes
After mapping the significantly differential genes to the GO database, the results are shown in Figure 3. Significant differences in GO enrichment analysis were found in different treatment groups, including 25 biological processes (BPs), 15 cellular components (CCs), and 10 molecular functions (MFs). The 25 BPs mainly involve signal transduction, transmembrane transport, G protein-coupled receptor signaling pathways, apoptotic processes, and glutathione metabolic processes. The 15 CCs mainly include cellular membrane, cytoplasm, extracellular matrix, and cytoskeleton. The 10 MFs mainly include hydrolase activity and DNA binding.
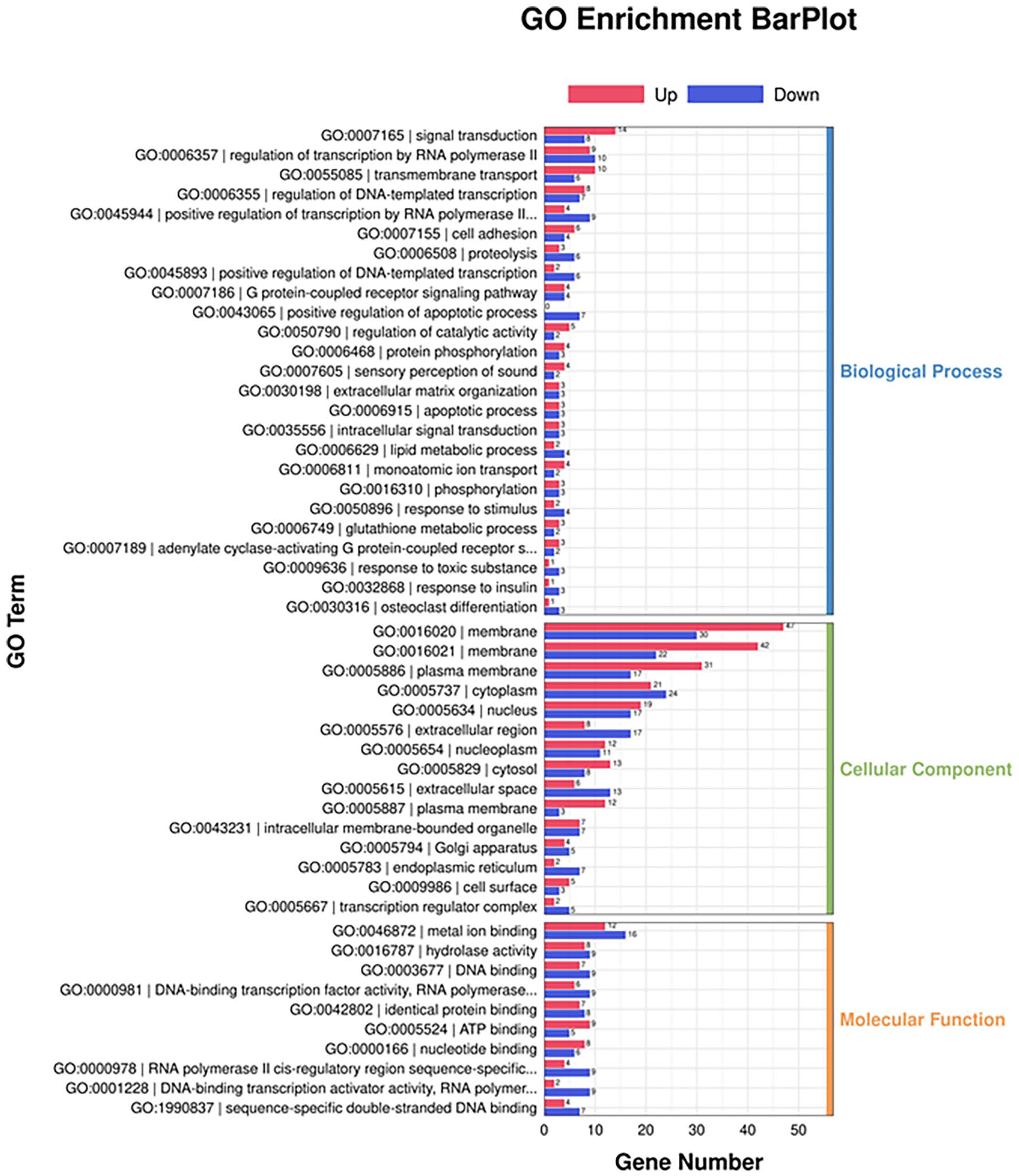
GO enrichment analysis plot of differential genes after PRV infection.
KEGG analysis of differential genes
The 269 selected differential genes were classified into 30 KEGG pathways (Figure 4). Specifically, there are five KEGG pathways within each category, which is cellular processes, environmental information processing, genetic information processing, human diseases, metabolism, and organismal systems. The top 20 KEGG enrichment pathways include neuroactive ligand–receptor interaction, ferroptosis, IL-17 signaling pathway, and others.
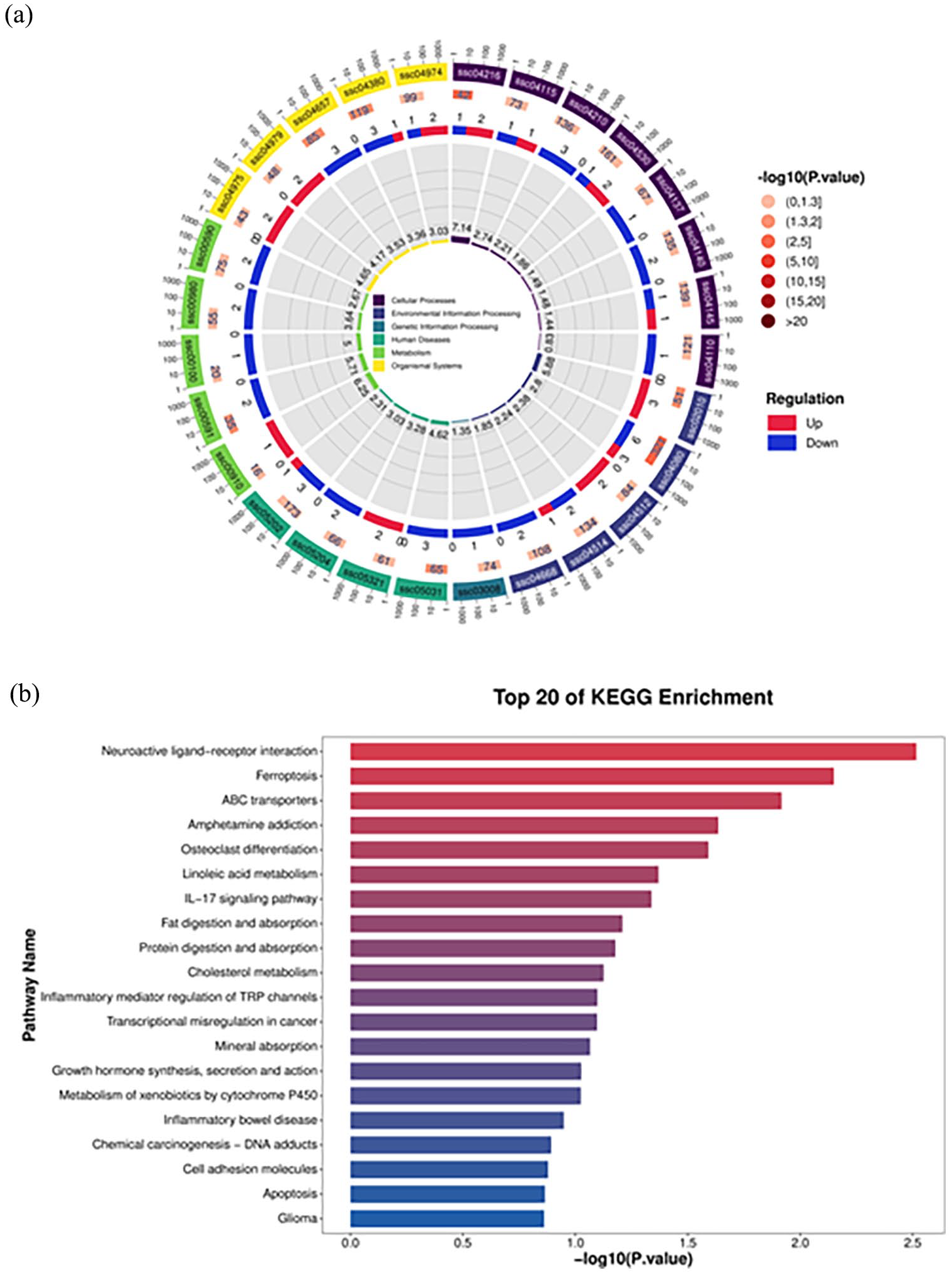
KEGG enrichment analysis plot of differential genes after PRV infection. (a) KEGG analysis loop diagram and (b) Top 20 expressed KEGG analysis plot, the horizontal axis represents the Q-value, and the vertical axis represents the pathway name.
Gene expression validation by qRT-PCR
The qRT-PCR technique was used to further validate the differentially expressed genes identified in the transcriptome sequencing results. Among these genes, seven differential genes from the neuroactive ligand–receptor interaction pathway were selected (Figure 5(a)), three differential genes from the ferroptosis pathway (Figure 5(b)), and three differential genes from the IL-17 signaling pathway (Figure 5(c)). Overall, the genes lysophosphatidic acid receptor 3 (LPAR3), granzyme A (GZMA), purinergic receptor P2Y2 (P2RY2), and neuromedin B (NMB) in the neuroactive ligand–receptor interaction pathway were downregulated, while the genes cannabinoid receptor 1 (CNR1), calcitonin receptor (CALCR), and neurotensin receptor 1 (NTSR1) were upregulated. In the ferroptosis pathway, the genes solute carrier family 7 member 11 (SLC7A11) and solute carrier family 40 member 1 (SLC40A1) were upregulated, while the transferrin gene (TF) was downregulated. In the IL-17 signaling pathway, the transcription factor subunit (FOS), FOSB, and mitogen-activated protein kinase 15 (MAPK15) genes all showed downregulation. Overall, the expression levels of the differential genes were consistent with the trends observed in the transcriptome analysis (Table 2).

Differential gene expression caused PRV by qRT-PCR: (a) Neuroactive ligand-receptor interaction, (b) Ferroptosis, (c) IL-17 signaling pathway.
Transcriptomics gene regulation situation.

Discussion
PRV causes immunosuppression and is a highly contagious viral disease. Since the emergence of variant PRV strains in China in 2011, it has caused significant economic losses to the swine industry. 7 Research by Li et al. 8 on the impact of PRV infection on various cellular pathways has observed that PRV infection leads to the activation of p65 phosphorylation, stimulation of NF-κB, induction of inflammatory factors, and upregulation of cytokines such as IFN-β, TNF-α, IL-1β, IL-6, and IL-18, thereby triggering a cytokine storm. This is basically consistent with the results of our laboratory’s previous studies using mice as a model with PRV-HLJ strain. 9 Different strains of PRV induce different levels of cytokines in specific tissues, which to some extent reflect the unique pathogenic characteristics of different strains. 10 For example, the LS strain induces high levels of IL-1β and TNF-α in the lungs, indicating that it may cause more severe pulmonary pathological changes compared to other strains. In addition, IL-1β is crucial for activating inflammatory responses, and TNF-α, as a key mediator of inflammation, shows a similar trend to IL-1β. The XJ strain significantly induces IL-1β and IL-6 in the brain, indicating that it may lead to severe neurological damage, which is consistent with observed clinical symptoms such as muscle spasms and mental depression. 11 Brain tissue is one of the important target organs of PRV infection, and encephalitis is the main cause of death in infected animals.12,13 In this study, we found an increase in the number of oligodendrocytes surrounding neurons in the brain tissue of Jiangkou radish piglets, and immunohistochemistry showed that PRV had invaded the brain tissue, implying that the PRV-HLJ strain in this study can also cause brain tissue damage.
By constructing a library from infected brain tissue and performing sequencing and bioinformatics analysis, a total of 269 differential genes were screened out, including 149 upregulated genes and 120 downregulated genes. Further GO enrichment analysis revealed that the main biological processes involved were signal transduction, trans-membrane transport, G protein-coupled receptor signaling pathways, apoptotic processes, and glutathione metabolic processes. Xu et al. 5 established a mouse PRV intracerebral infection model and revealed the differential expression of various inflammation-related signaling pathways and innate immune signaling pathways through transcriptome analysis, mainly including the TNF signaling pathway, Toll-like receptor signaling pathway, NOD-like receptor signaling pathway, MAPK signaling pathway, IL-17 signaling pathway, and JAK-STAT signaling pathway. This experiment found through KEGG enrichment analysis that the differential genes were mainly present in the neuroactive ligand–receptor interaction, ferroptosis, and IL-17 signaling pathways. The differences in transcriptomics may be caused by different strains. The neuroactive ligand–receptor interaction pathway is composed of receptors located on the plasma membrane, and receptors are involved in the signal transduction from the external environment to the inside of the cell. 9 This study found that after PRV infection, the genes LPAR3, GZMA, P2RY2, and NMB in the neuroactive ligand–receptor interaction pathway were downregulated, while CNR1, CALCR, and NTSR1 showed an upregulation trend, indicating that PRV infection can regulate by changing key genes in the neuroactive ligand–receptor interaction pathway.
Ferroptosis is a new type of nonapoptotic programmed cell death that depends on the formation and accumulation of lipid radicals mediated by iron, 11 characterized by excessive production of lipid reactive oxygen species (ROS) inside cells. When the cell’s antioxidant capacity is reduced, it leads to fatal lipid peroxidation. 12 Therefore, the sensitivity of ferroptosis is finely regulated by biological processes that maintain redox balance, including the generation of ROS and the biosynthesis of the antioxidant glutathione (GSH). Transcriptome analysis indicates that infection with PRV in Neuro-2a cells results in a marked decrease in the expression of 15 genes associated with ferroptosis, such as Slc1a5. On the other hand, the expression of four ferroptosis-related genes, including Alox12, is significantly increased in PRV-infected cells when compared to noninfected cells. 13 In this study, through GO enrichment analysis, differences in the glutathione metabolic process before and after PRV infection were found, and further validation of ferroptosis-related genes showed that SLC7A11 and SLC40A1 were upregulated, while TF was downregulated. Xu et al. 14 found that herpes simplex virus type 1 (HSV-1) enhances Keap1-dependent Nrf2 degradation, thereby promoting ferroptosis. In addition, HSV-1-induced ferroptosis plays an important role in the development of viral encephalitis in mice, and the upregulation of PTGS2 and PGE genes caused by ferroptosis helps the occurrence of encephalitis. Further research confirmed that HSV-1-induced ferroptosis has an important impact on the pathogenesis of viral neuropathy, and ferroptosis inhibitors may be a potential immune therapeutic strategy to combat HSV-1 infection and encephalitis. In addition, Xu et al. 15 found that the PRV XJ gE/gI/TK deletion strain can activate the IL-17 signaling pathway, but did not clarify the location of the differential genes. In this study, we found that after infection with the PRV-HLJ strain, the genes FOS, FOSB, and MAPK15 in the brain tissue of Jiangkou radish pigs all showed a downregulation trend.
This study used transcriptome technology to conduct pathological and immunohistochemical analysis of the brain tissue of Jiangkou radish pigs infected with PRV, and observed a significant increase in the number of oligodendrocytes in the brain tissue and the presence of PRV-positive expression areas. Subsequently, transcriptome analysis showed that PRV infection mainly led to changes in the expression of genes related to neuroactive ligand–receptor interaction, ferroptosis, and IL-17 signaling pathways. While the current research findings offer valuable insights into the pathogenesis of PRV, the precise identification of key genes remains to be confirmed through additional studies.
Ethical statement
This study received approval from the Animal Ethics Committee at Tongren Polytechnic College. The handling and procedures involving animals were strictly adhered to the approved guidelines, aligning with the International Guidelines for the Care and Use of Laboratory Animals, as well as the recommendations set forth by the ARRIVE (Animal Research: Reporting In Vivo Experiments) initiative.
Conclusion
These findings suggest that PRV infection not only disrupts the normal functioning of the central nervous system but also triggers a cascade of immune responses. This could lead to a better understanding of the pathogenesis of PRV and the development of more effective strategies for disease control and prevention.
Footnotes
Acknowledgements
Many thanks to Samuel Kumi Okyere from Wayne University for his advice and guidance in revising this manuscript.
Author contribution
Wei Sun was responsible for drafting the initial manuscript and data analysis; Shan-Shan Liu was in charge of experimental design; Shi-He Long was responsible for analyzing transcriptome data; Yu Fan, Sheng-Qing Deng and Tao-Hua Ren managed animal experiments and collected samples; Fengzhao Zhu and Ling Bai provided supervisory guidance throughout the entire project.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was financially supported by Guizhou Provincial Basic Research Program (Natural Science) (No. ZK[2024]664) and the Program of Cultivating High-Level Innovative Talents in Guizhou Province (No. 2022-(2020)-045).
Ethics approval
Ethical approval for this study was obtained from *Animal Ethics Committee at Tongren Polytechnic College (Approval No. TRZY-2023-06-16)*.
Animal welfare
The present study followed international guidelines for humane animal treatment and complied with relevant legislation.
