Abstract
Background:
The ability of Pseudomonas aeruginosa to produce biofilm has established it as one of the most significant pathogens. The purpose of this study was to evaluate antimicrobial resistance and conduct a molecular investigation of the virulence genes PslB and PelD in Pseudomonas aeruginosa species isolated from patients.
Methodology:
One hundred clinical isolates were collected from patients of different age groups who were hospitalized in Kermanshah and Sonqor hospitals. The isolates were obtained through culture on specific media, biochemical confirmatory tests, and gram staining for confirmation. Biofilm production was assessed using an indirect quantification method with crystal violet. Additionally, antibiotic resistance was determined through the disc various method following Clinical and Laboratory Standards Institute guidelines. Finally, the presence of genes related to PlsB and PelD in resistant strains was examined using The polymerase chain reaction (PCR).
Results:
The results indicate that the highest resistance and lowest sensitivity were related to nitrofurantoin 100 μg, while the lowest resistance and highest sensitivity were related to cefepime 30 mg. Biofilm phenotypes were categorized as weak in 7% (n = 7) of isolates, medium in 13% (n = 13), and high in 80% (n = 80). The PslB and PelD genes were identified in 86% (n = 86) and 38% (n = 38) of isolates, respectively, while 4% (n = 4) did not possess either of these two genes. Additionally, a majority of the isolates exhibited multidrug-resistance (87%) due to their moderate-to-high biofilm formation.
Conclusion:
All isolates were capable of producing biofilm. A significant association were between strains with the high biofilm and multidrug-resistance species (p < 0.05). Multidrug-resistance (78%) isolates included 28% (n = 28) of isolates that were PslB+ PelD+, 45% (n = 45) of isolates that were only PslB+, and 5 (n = 5) isolates that were only PelD+. A significant relationship was found between the presence of the PslB gene multidrug-resistance and high producer (p < 0.05).
Keywords
Introduction
As one of the most common nosocomial pathogens, Pseudomonas aeruginosa (P. aeruginosa) is involved in many severe opportunistic infections, especially in immunocompromised individuals. 1 This poses a pervasive problem as it complicates antibiotic treatments against P. aeruginosa due to the development of multidrug-resistant (MDR) species.2–4 The ability of P. aeruginosa to produce biofilm has placed this bacterium in the category of bacteria with chronic and complex infections. These bacteria protect themselves by producing biofilms, which include consisting of proteins, extracellular DNA, and exopolysaccharides. Biofilm create a microbial complex that naturally poses many problems for antibacterial treatments. 5
P. aeruginosa thrives in various aquatic environments and can cause both immediate and persistent infections in these settings. Bacterial conditions and host defense mechanisms impose different stresses on bacteria, forcing them to adapt in order to survive in diverse environments. The bacterium sustains itself by creating an extracellular polysaccharide enclosure around itself and is capable of causing both acute and chronic infections. Alginate, Psl, and Pel are three membrane polysaccharides secreted by P. aeruginosa that differ in their chemical structure features and biosynthetic mechanisms. 6 Research on alginate, which is responsible for chronic lung infections, has helped researchers identify its production pathway. However, only a limited number of P. aeruginosa strains produce alginate, while most pathogenic strains produce Pel and Psl polysaccharides. The structural and functional similarity of alginate and bridge protein is significant.7,8 Biosynthesis of Psl involves the aggregation of pentameric Psl subunits linked to an isoprenoid lipid carrier, a process also observed in the Escherichia coli (E. coli) extracellular polysaccharides (EPS)/capsular polysaccharides (CPS) capsular biosynthesis pathway.6–9
Psl is classified as a neutral polysaccharide. Its structure consists of 5 units of D-mannose followed by D-mannose-glucose and finally a L-rhamnose. Psl plays a crucial role in interactions between bacterial cells during biofilm formation, making it important in the process of biofilm formation and maintenance. 10 The geneset that produces this membrane polysaccharide is an operon containing 15 genes (pslA to pslO) that are transcribed and then translated as a polycistronic mRNA. An enzyme from the Psl operon is involved in making sugar-nucleotide precursors of Psl monosaccharide subunits, namely, D-mannose, L-rhamnose, and D-glucose. The PslB protein has GMP at the N-terminus and PMI at the C-terminus, giving it two distinct properties and functions. The PslB enzyme performs the first and third steps phosphomannose isomerase (PMI)/GDP-mannose pyrophosphorylase (GMP) guanosine diphosphate mannose (GDP-mannose) of converting fructose-6-phosphate to. 11
The Pel operon is involved in the formation of a pellicle, which is a layer of polymer/cells at the air-liquid interface of P. aeruginosa standing culture. 12 Structurally, exopolysaccharide P consists of 1→4 acetylated glucosamine sugar subunits and glucosamine sugars, and is sensitive to cellulose. 13 The pel operon contains seven genes (pelA to pelG). 14 Considering the potential of biofilm in increasing antibacterial resistance and prolonging infections caused by P. aeruginosa, identifying isolates with such a factor helps us to better understand the pathogenesis of the organism. Colvin et al. 15 showed that pel exopolysaccharide has a structural and protective function in the biofilm matrix. Yang et al.’s 16 findings also showed that psl plays a role in the antibiotic resistance of P. aeruginosa biofilms. Singkham et al. 17 demonstrated the function of biofilms by decrease several genes, including biofilm synthesis genes (algD and PslB) and biofilm regulators (gDsia, U, Ag).
A study conducted in 2022 by Divyashree et al. 18 focused on biofilm-related genes of Psl and Pel isolates. The study revealed that ampicillin (88.24%), nalidixic acid (83.82%), and nitrofurantoin (64.71%) had the highest resistance rates. Also, the most dominant gene identified was PslB, with most isolates showing resistance to multiple drugs due to moderate-to-high biofilm formation.
This research aims to evaluate the pattern of antimicrobial resistance and the frequency of PelD and PslB genes in biofilm-producing P. aeruginosa isolated from hospital infectious departments. To the best of our knowledge, there has been little research conducted on the genes PslB and PelD in Iran. This type of study could provide valuable insights for the development and improvement of new and powerful antibiotics that target specific genes or products of combinations of genes involved in the formation of biofilms and the spread of MDR strains.
Methods
Sample collection
This cross-sectional study was conducted in 2024. After obtaining approval from the ethics committee and written informed consent from the legally authorized representative of under 18 years subjects, 100 clinical samples were collected from three different areas of Kermanshah and Sonqor hospitals over a period of 4 months. The samples included urine, bed sores, blood, lung secretions and burn wounds. Diagnostic tests were then performed on the samples. The study included patients of different age groups with at least a positive culture from the target organ, while patients with secondary infections of other organs were excluded from the study.
Phenotypical and biochemical confirmatory tests
All samples were cultured on cetrimide agar medium and then incubated for 24 h at 37°C. The presence of P. aeruginosa in the collected samples was indicated by a green pigment on the cetrimide agar grown on cetrimide agar medium at 42°C. The strains of E. coli ATCC- 29212 (colorless colony in cetrimide agar) and P. aeruginosa ATCC- 9037 were used as negative and positive controls, respectively, for detecting isolates of P. aeruginosa. Biochemical confirmatory tests were then performed on colonies grown on cetrimide agar medium. Isolates that were oxidase positive, hydrogen sulfide (H2S)-negative, triple Sugar Iron (TSI)-positive, catalase-positive, citrate-positive, sulphide indole motility (SIM)- positive, urea-positive, oxidative-fermentative (OF)-positive, able to grow at 42°C, methyl red (MR)-positive and Voges-Proskauer (VP)-negative were selected.
Antibacterial susceptibility tests
Based on the recommendation of the Clinical and Laboratory Standards Institute, specific discs were selected and placement of two Muller Hinton Agar mediums (MERCK., Germany) with a concentration of half McFarland at a distance of at least 2.5 cm apart was made. The antibiotic sensitivity test was conducted by the agar disk diffusion method.
Antibacterial susceptibility was determined by disc diffusion using the following antibiotic discs (MAST, UK): imipenem (IMI 30 mg), gentamicin (GM 10 mg), nitrofurantoin (NIT 100 μg), ciprofloxacin (CIP 5 mg), cefepime (CEF 30 mg) and calactin (CL 10 mg). 3
Biofilm formation assay
The microtitre-plate method 3 along with an indirect quantification method with crystal violet was used to study P. aeruginosa biofilm formation. Initially, trypticase soy agar culture medium and a liquid solution containing 2% glucose were prepared. Bacterial samples were streaked on trypticase soy agar with 2% glucose and then incubated at 37°C for 20 h. The resulting colonies were collected with a loop and transferred to trypticase culture medium with 2% glucose to create a bacterial suspension equivalent to McFarland half turbidity. To prevent testing errors, the turbidity of the samples was measured visually and with a spectrophotometer at a wavelength of 625 nm, and their OD was observed in the numerical range of 0.08–0.1. Next, 200 µL of bacterial suspension were taken and aliquoted into a 96-well microtiter plate, which was then incubated for 24 h at 37°C. After incubation, all the wells of the plate were aspirated and washed with sterile physiological saline to remove any unattached cells. The attached bacteria were fixed with absolute methanol for 10 min, and then the plate was stained with crystal violet (1%). After a few seconds, the excess stain was removed by washing the plate with sterile distilled water. Any remaining stain was washed off with 200 µL of glacial acetic acid (33%v/v), and the OD was read with an enzyme-linked immunosorbent assay (ELISA) reader (Stat Fax 2100, Awareness Tech Inc., USA) at 630 nm (Stat Fax 2100, Awareness Tech Inc., USA). 3 This test was repeated three times for each bacterial sample, with a well containing an empty culture medium that is the same as the trypticase the broth containing 2% glucose was considered as a blank. 19 The standard strain of P. aeruginosa PAO1 ATCC (American Type Culture Collection) 27853 were used as positive control for biofilm production ATCC 15692.
Molecular study
The genomes of all isolates were extracted using the boiling method. Previously described primers were used to confirm the presence of the PslB and PelD genes (Table 1).The primers were manufactured by Cinna gene (Iran). P. aeruginosa strain PAO1 was used as positive control. The PCR assay was performed in a 25 μL volume using the specific primers and the following time and thermal program: initial denaturation at 94°C for 2 min, followed by 35 cycles of denaturation at 94°C for 60 s, annealing at 55°C for 60 s, extension at 72°C for 60 s, and a final extension of 10 min at 72°C. This was carried out in a thermocycler device (Eppendorf Master cycler Gradient, Germany).
Primers sequences of PslB and PelD genes, as well as characteristics of their PCR products.

Statistical analysis
By SPSS version19.0 statistical software (IBM® SPSS®) was conducted statistical analysis. Chi-square test (c2) and Fisher’s exact test (in cases where the frequencies were less than or equal to 5) were applied to check the differences in allele frequency and genotype distribution between healthy controls and affected patients. Logistic regression analysis was performed to calculate the odds ratio (OR) with confidence interval (CI) 95% adjusted for age and sex.
Results
Characteristics of clinical isolates of P. aeruginosa
One hundred isolates were obtained from various clinical samples taken from patients of different age groups who were hospitalized in Sonqor and Kermanshah hospitals. These isolates were identified using culture, Gram staining, and diagnostic-differential tests.
Out of 100 samples examined, 48 belonged to men and 52 to women. A statistical analysis using Chi-square and Kramer correlation coefficients and agreement (Cohen’s Kappa (κ = 0)), showed no notable association between P. aeruginosa infection and the gender of the patients. The age distribution of the patients was as follows: three individuals were below 6 years old; 12 individuals were between 6 and 10 years old, and 85 individuals were older than 10 years old (Table 2). A significant relationship was found between P. aeruginosa infection and the age of the patients (p < 0.05). The average age of the patients was 29.88 years. Out of 100 isolates, 30% (n = 30) were from urine, 10% (n = 10) from bed sores, 5% (n = 5) from blood, 32% (n = 32) from lung secretions, and 23% (n = 23) belonged to burn wounds.
Frequency of isolated patients according to age.

Results of antibiogram test
The isolates demonstrated resistance to most of the commonly used antibiotics that were tested. Out of the 100 isolates, 90% of isolates showed maximum resistance to NIT 100 μg, and the lowest resistance was to CEF 30 mg (Figure 1).
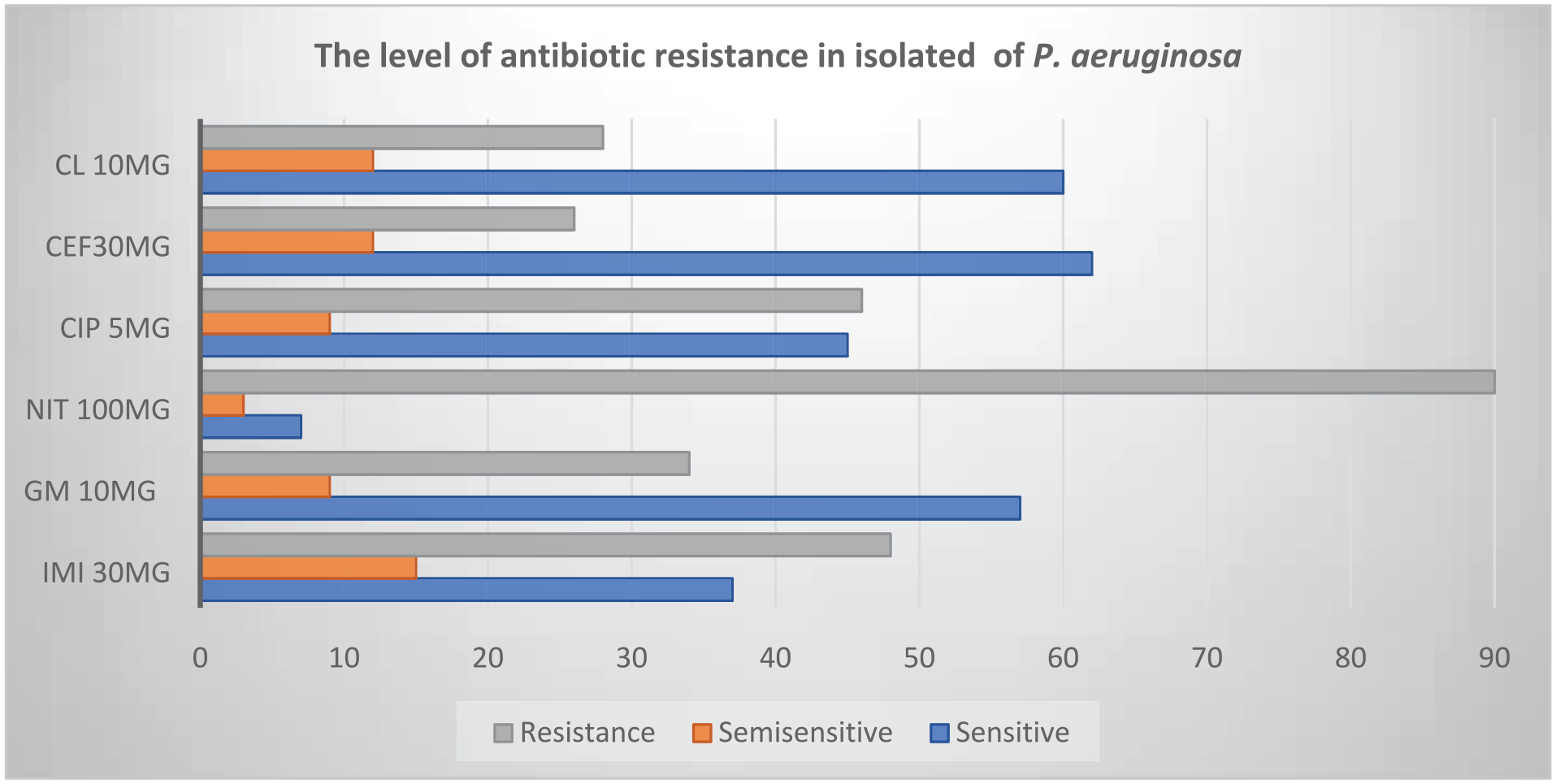
The level of antibiotic resistance in the 100 investigated P. aeruginosa isolates. The results indicate that the highest resistance and lowest sensitivity were related to NIT 100 μg, while the lowest resistance and highest sensitivity were related to CEF 30 mg.
In the examination of 30 urine isolates, it was observed that the highest antibiotic resistance was related to NIT 100 μg, while and the lowest was related to CEF 30 mg (Figure 2).
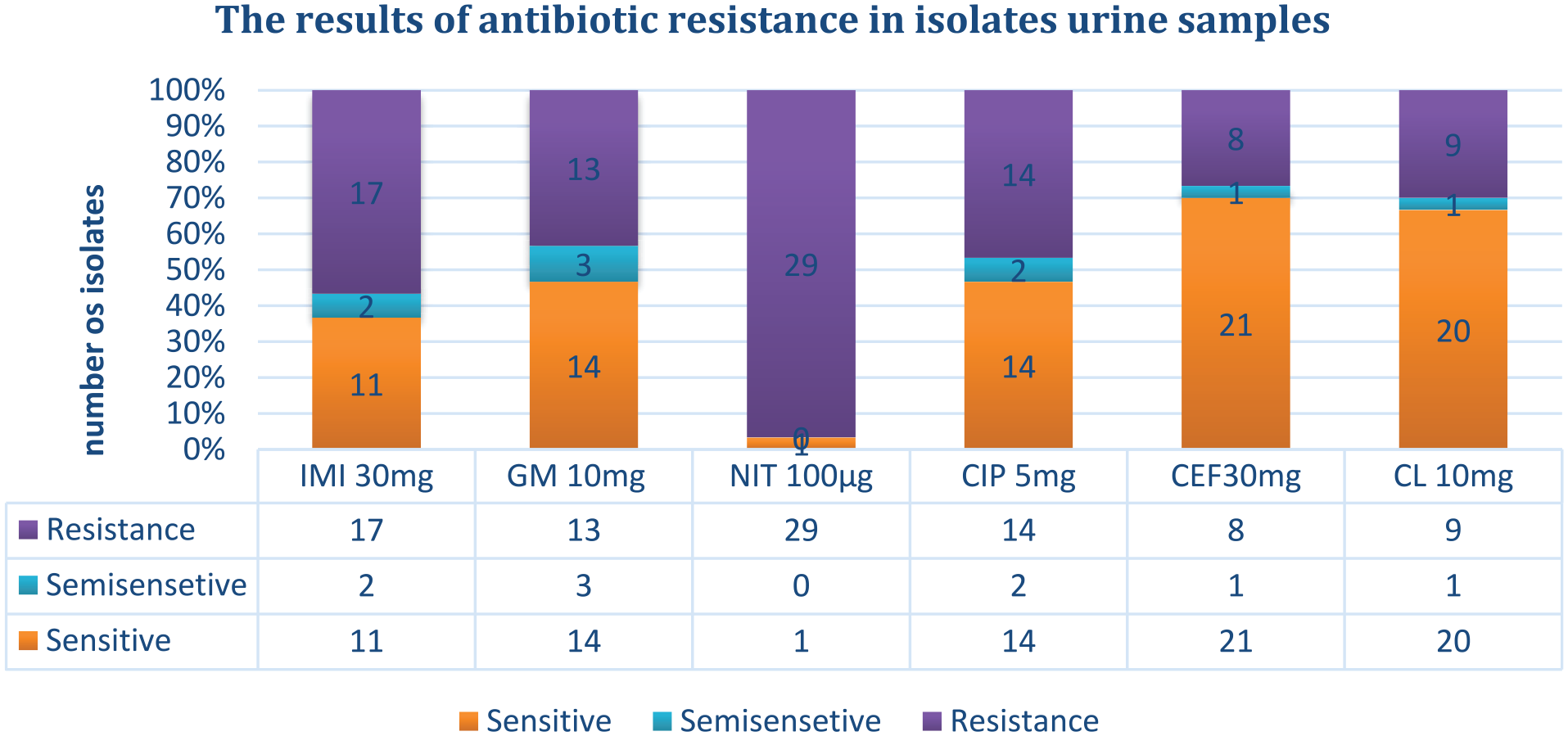
The level of antibiotic resistance of isolates related to urine samples.
In 10 bedsore isolates, it was observed that the highest resistance was related to NIT 100 μg, while the lowest resistance was related to GM 10 mg, CEF 30 mg, and CL 10 mg, respectively. Among these samples, the most sensitive disc was GM 10 mg (Figure 3).
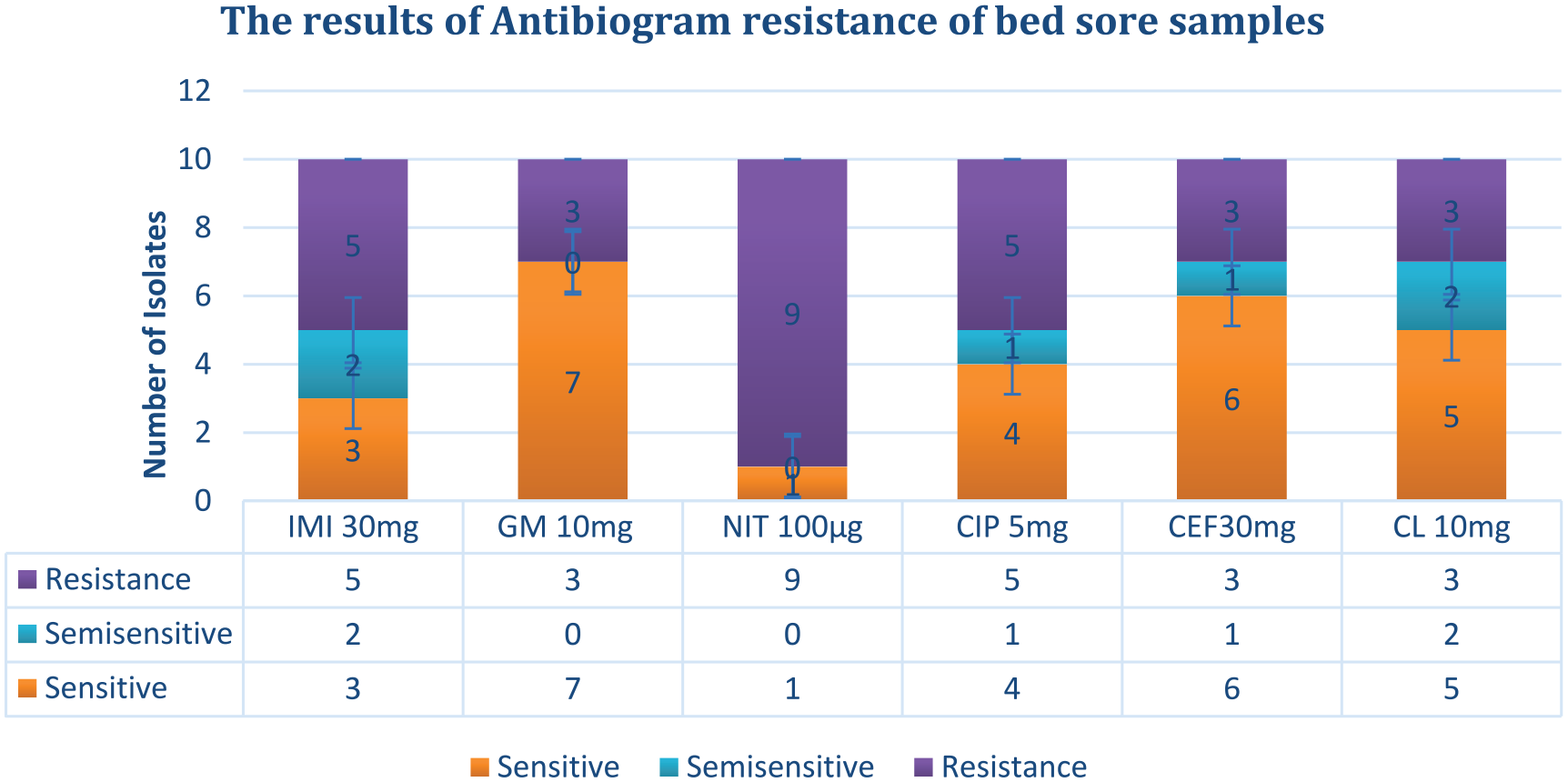
Antibiogram test results of bed sore samples.
In five blood samples, the highest and lowest resistance was related to NIT 100 μg, while all samples were sensitive to GM 10 mg (Figure 4).

Antibiogram test results of blood samples.
The results of the antibiogram test on 32 isolates isolated from pulmonary infections showed that in these samples, the highest resistance was to NIT 100 μg, and the lowest was to CEF 30 mg and CL 10 mg (Figure 5).
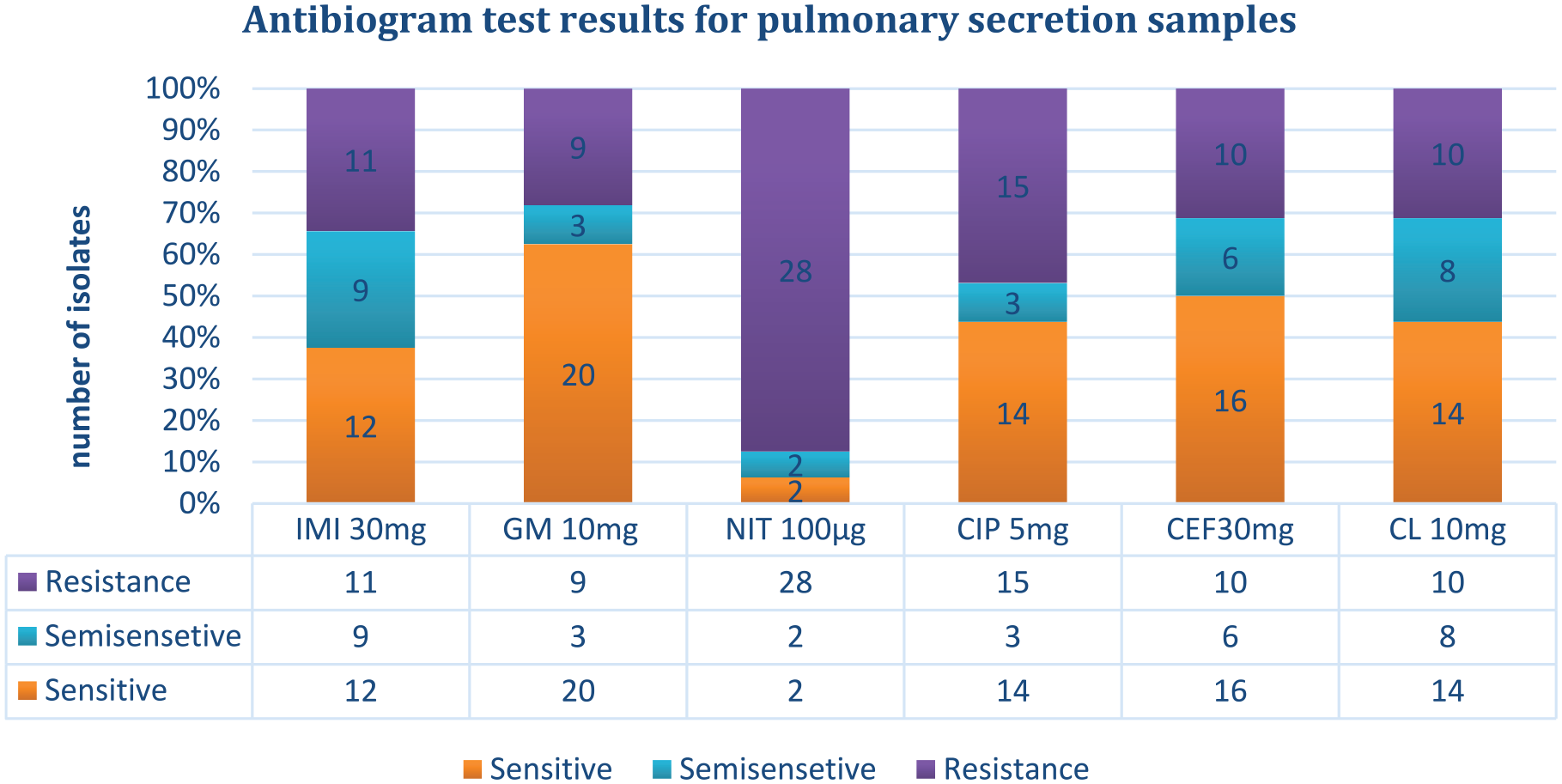
Antibiogram test results for pulmonary secretion samples.
In burn wounds samples, the highest resistance was related to NIT 100 μg, and the lowest resistance was related to CEF 30 mg and CL 10 mg (Figure 6).

Antibiogram test results for burn wounds samples.
Results biofilm
Examining the results of biofilm formation by the collected isolates, it was found that averages optical absorbance of 100 isolates at the wavelength, with three repetitions, was higher than 0.18. This indicates that all isolates were biofilm. Results showed 7% of the strain showed weak biofilm formation (a16, a10, a9, a7, a6, a5, a4), while 13% the strains had medium biofilm formation (a28, a27, a25, a22, a19, a18, a17, a15, a14, a11, a8, a3, a2). The remaining 80% of the strains had strong biofilm formation.
Results of molecular tests
The PslB gene was found to have a frequency of 86% in isolates (Figure 7), while the PelD gene showed a frequency of 38% in isolates (Figure 8).

Electrophoresis of PCR product for PslB gene in the examined isolates. Column 1 on the right: positive control, column 2 of 50 pairs of ladders, column 3 of negative control, the rest of the cells are related to the isolates, all of which have a band of 160 bp.
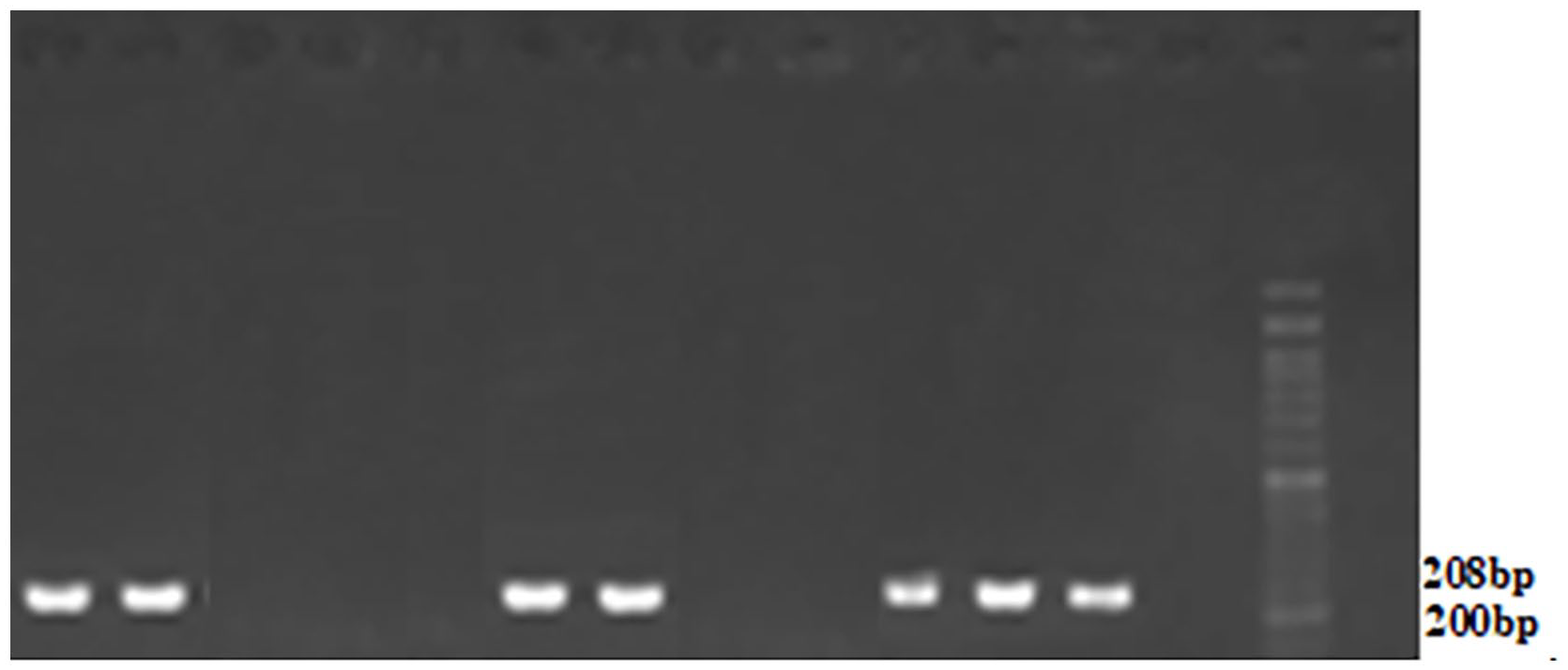
Electrophoresis of pcr product for PelD gene in investigated isolates. Column 1 on the right: positive control, column 2 is 50 pairs of ladders, column 3 is negative control, the rest of the cells are related to the isolates, all of which have a 200 bp band.
Discussion
The presence of P. aeruginosa strains resistant to antimicrobial substances in samples isolated from hospitals, surface water, and other environmental sources poses a serious threat to communities. 17 P. aeruginosa creates conditions for opportunistic pathogens by producing biofilm-forming exopolysaccharides and forming biofilms. 20 Biofilm keeps bacteria together and helps them acquire antibiotic-resistant genes. 21 Resistance of biofilm to antibiotics is a result of frequent exposure to high levels of antibiotics, as well as the repeated close contact of bacteria with these drugs. Additionally, genetic and physical characteristics of the bacterial strain, as well as physiological conditions, play a role in this resistance. Biofilm bacteria are resistant to antibiotic treatment due to internal and mutational resistance against antibiotics. 22
In this research, all P. aeruginosa isolates (100%) were capable of producing biofilms. We found a significant association between strains with the high biofilms and MDR species (p < 0.05). The phenotypic classification of biofilms identified in this study was as follows: weak (7%, n = 7); medium (13%, n = 13); and high (80%, n = 80). Jabalameli et al. 23 reported biofilm formation in more than 96% of the P. aeruginosa isolates collected from burn patients in Tehran. In another study conducted at hospitals in Tabriz, Iran, a figure of 79% was reported for biofilm-forming clinical strains of P. aeruginosa. 24 Previously, Drenkard and Ausubel 25 reported that antibiotic-resistant strains of P. aeruginosa have a high ability to form biofilms.
Out of the 100 clinical strains tested for antibiogram, 48%, 34%, 90%, 46%, 26%, and 28% were resistant to IMI 30 mg, GM 10 mg, NIT 100 μg, CIP 5 mg, CEF 30 mg, and CL 10 mg, respectively.
The results of the antibiogram test on urine (n = 30), bed sores (n = 10), blood (n = 5), pulmonary infections (n = 32), and burn wounds (n = 23) indicated that isolates have the highest resistance to NIT 100 μg. Although the number of isolates differs in each group, it seems that NIT 100 is not appropriate for these isolates. Additionally, in urine, the lowest resistance was related to CEF 30 mg: in bed sores it was related to GM 10 mg, CEF 30 mg, and CL 10 mg, with the most sensitive disc being GM 10 mg (Figure 3). In blood isolates all samples were sensitive to GM 10 mg (Figure 4). In pulmonary infections, the lowest resistance was to CEF 30 mg and CL 10 mg (Figure 5), and in burn wounds, the lowest resistance was related to CEF 30 mg and CL 10 mg (Figure 6). In another study, clinical isolates of P. aeruginosa exhibited high resistance to meropenem, ticarcillin, and other antibiotics. 26
All isolates were examined for the presence of the PslB and PelD genes. Among them, 86% of isolates (n = 86) were positive for the PslB gene and 38% for the PelD gene. Additionally, 28% (n = 28) of isolates had both genes (PslB+ PelD+). All of these were MDR and high producer biofilms. Meanwhile, 4% of isolates were negative for both genes (PslB− PelD−) and were weak biofilm producer. One isolate from the PelD+ group was weak. The high producer biofilm group includes 28 (PslB+ PelD+), 52 PslB+ isolates, and 10 isolates with only PelD+. Among the PelD+ isolates, three were biofilm producers and seven were moderate biofilm producers. Six isolates from the only PslB+ isolates were moderate biofilm producers. Therefore, a significant relationship was found between the PslB gene MDR and high producer (p < 0.05). MDR (78%) isolates included 28% (n = 28) of isolates that were PslB+ PelD+, 45% (n = 45) of isolates that were only PslB+, and 5 (n = 5) isolates that were only PelD+. A significant relationship was found between the presence of the PslB gene MDR and high producer (p < 0.05). Bavashe and Karmoustaji 27 reported that 27.8% of P. aeruginosa clinical isolates were MDR.
The main exopolysaccharides secreted by bacteria that are important in biofilm formation are Psl, Alg, and Pel.28,29 In nonmucoid bacterial strains of P. aeruginosa, exopolysaccharides Psl and Pel are essential for maintaining biofilm structure. Exopolysaccharide Pel contains a high amount of glucose, necessary for pellicle formation, and plays a structural and protective role in the biofilm matrix. 15 On the other hand, Psl is involved in biofilm formation in both nonmucoid and mucoid strains of P. aeruginosa.30,31 Divyashree et al. 17 found that PslB was the most dominant gene, and that most of the isolates were resistant to several drugs due to moderate and high biofilm formation.
In 2019, Whitfield et al.26,32 identified stable PelDEFG gene clusters over 500 diverse proteobacterial species. Using P. aeruginosa, as a model, they demonstrated that PelD, PelE, PelF, and PelG form a complex in the inner membrane and suggested that this complex represents a previously unidentified Pel polysaccharide synthase, responsible for the polymerization and translocation of Pel to the cytoplasmic membrane. In 2018, Li et al. 33 conducted a study on the effects of diallyl disulfide (DADS) on the growth, virulence factor production (elastase, pyocyanin, biofilm, and swarming motility), and essential gene expression of P. aeruginosa PAO1, specifically in relation to QS and virulence. The lasB gene, which encodes lasB elastase, is regulated by the las, rhl, and pqs systems, so the down-regulation of genes in these systems enhances the down-regulation of lasB. Additionally, phzM (encoding pyocyanin), pslB (responsible for biofilm matrix polysaccharide production), and chiC (encoding chitinase) were positively activated by LasR. Downregulation of lasR transcription further downregulated phzMC and phzM transcription. A study conducted by Kamali et al. 34 in 2020 showed that isolates exhibited resistance to the antibiotic’s amikacin, piperacillin/tazobactam, and levofloxacin. All three genes related to biofilm were present simultaneously in 87.5% of isolates, while13.5% of isolates did not have any of the tested genes. From the results of this study, combination therapy including an anti-pseudomonad beta-lactam (piperacillin/tazobactam or ceftazidime) and an aminoglycoside or carbapenem (imipenem, meropenem) with fluoroquinolones along with an aminoglycoside can be used against Pseudomonas.
One of the shortcomings of this study was the small number of samples. Additionally, it is recommended to collect samples from hospital fluids simultaneously. Another limitation of the study was the small number of antibiotics that were examined, due to limited funding. We did not use a specific formula for calculating the sample size, which was a limitation of this study. Sampling was based on convenience due to the cost of experimentation, limited availability of samples, and time constraints. Additionally, we should expand our research by investigating other biofilm genes in the negative biofilm samples for these two genes. Next, we should investigate the positive samples of the two genes in this assay for other biofilm genes. Another objective is to select different antibiotics and test their effectiveness.
Conclusion
The public health implications of P. aeruginosa should not be underestimated as it remains a major cause of healthcare-associated infections in Asia and has limited treatment options. This study revealed that isolates of P. aeruginosa exhibit a high percentage of resistance to commonly used drugs against this microorganism. In this research, all P. aeruginosa isolates (100%) were capable of producing biofilm. We found a significant association between strains with high biofilm and MDR species (p < 0.05). Out of the 100 clinical strains tested for antibiogram, 90% of them were resistant to NIT 100 μg and 26% were sensitive to CEF 30 mg. All isolates were examined for the presence of the PslB and PelD genes. Among them, 86% of isolates (n = 86) were positive for the PslB gene and 38% for the PelD gene. Additionally, MDR (78%) isolates included 28% (n = 28) of isolates that were PslB+ PelD+, 45% (n = 45) of isolates that were only PslB+, and 5 (n = 5) isolates that were only PelD+. A significant relationship was found between the presence of the PslB gene MDR and high producer (p < 0.05). This study highlights the role of biofilms as a mechanism for bacterial survival.
Footnotes
Acknowledgements
Thank you to everyone who assisted us in conducting this research.
Authors’ contributions
FK and NM developed the concept and design of the study, collected, and analyzed the data. The first draft and revised of the article was prepared by all authors; FK the manuscript; All authors reviewed and approved the article.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethics approval
This research was ethically approved by the Ethics Committee of the Sanandaj Branch, Islamic Azad University, Sanandaj, Iran (IR.IAU.SDJ.REC.1403.071). Also, all protocols involving human subjects comply with the requirements of the Declaration of Helsinki of the Iranian Ministry of Health and Medical Education.
Informed consent
Written informed consent was obtained from the Legally Authorized Representative of under 18 years subjects.
Trial registration
Not applicable.
