Abstract
Objectives:
The aim of the study was to determine the distribution and antimicrobial susceptibility profile of bacteria recovered from the sewage systems of health institutions found in Hawassa, Sidama Regional State, Ethiopia.
Methods:
A cross-sectional study was conducted from 20 October 2020 to 1 December 2020. A total of 27 sewage samples were collected at two points, namely, before entering the septic tank and from the septic tank of seven health institutions. Samples were inoculated onto Mannitol salt agar, Blood agar, and MacConkey agar, and incubated for 24 h at 37°C. Bacteria were identified using colony morphology, Gram staining, and biochemical tests. An antimicrobial susceptibility test was performed using the Kirby–Bauer disk diffusion method. Data were analyzed by SPSS, version 25, and results were presented in text and tables.
Results:
All sewage samples (n = 27) examined in the current study contained potential pathogenic bacteria. Overall, 129 different types of bacteria were identified. Of isolated bacteria, 14 (10.8%) were Gram positive, while 115 (89.2%) were Gram negative. The most prevalent bacteria were Escherichia coli (n = 27, 20.9%) followed by Shigella species (n = 26, 20.2%), Pseudomonas species (n = 25, 19.4%), Salmonella species (n = 25, 19.4%), Staphylococcus aureus (n = 14, 10.9%), and Klebsiella species (n = 12, 9.3%). All bacteria were susceptible to azithromycin. About 80% of bacteria were resistant to ampicillin, whereas greater than 80% of bacteria were susceptible to norfloxacin, ciprofloxacin, and gentamicin.
Conclusion:
All sewage systems of health institutions included in the current study contained different types of pathogenic bacteria, which are resistant to commonly prescribed antibiotics.
Keywords
Introduction
Sewages from hospitals (hospital wastewater) are a category of wastes that are released from hospitals and health care centers, and are not intended for further use. Wastes from hospitals may arise from biological or non-biological or both sources. Hospital wastes can harm humans when there is direct contact with the waste or when it is released into the environment. Wastewaters from health facilities have a complex nature and serve as a hotspot for the emergence of antibiotic-resistant bacteria (ARB). 1
Activities in hospitals produce fluid wastes that contain harmful chemicals and microbes, which are resistant to antibiotics. 2 Based on the World Health Organization (WHO) information, about 10% to 25% of wastes from health institutions are hazardous, 2 and the burden of infection due to ARB is increasing from time to time. 3 The burden of ARB is not the same across different countries: it ranges from 16% to 75%. 4 ARB will be responsible for 10 million deaths per year in 2050. 5
The most serious problem associated with wastes from health facilities is that they may contain ARB. 6 Hospitals provide a conducive environment for the development of multidrug-resistant (MDR) bacteria, making treatment options limited and expensive. Most importantly, bacteria that reside in wastewater may act as vectors or reservoirs for antibiotic-resistant genes and they may transfer these genes to other bacteria. 7
A previous study conducted on hospital wastewater in Hawassa city reported pathogenic bacteria such as Staphylococcus species (26.6%), Escherichia coli (24.4%), Klebsiella species (20%), and Shigella species (11.1%). 8 The prevalent bacteria detected from hospital wastewater of Gondar were Klebsiella species (26.6%) followed by Pseudomonas species (16.8%) and E. coli (11.5%). 9 On the contrary, the predominant bacteria identified from wastewater of hospitals located in Addis Ababa were E. coli (32%), Salmonella (23%), Klebsiella pneumoniae (15%), and Enterobacter aerogenes (11%). 10 According to a systematic review conducted by Tesfahun et al., 11 all studies from different parts of Ethiopia reported pathogenic bacteria from wastes of health facilities.
Timely and proper management of wastes produced in health facilities has a critical role in reducing the emergence and dissemination of ARB in the community. The purpose of this study was to identify bacteria from the sewage system of health institutions and to determine their antimicrobial susceptibility profile.
Operational definition
MDR is a bacterium that is resistant to at least three or more classes of antimicrobial agents.
Sewage is the wastewater released by residents, business organizations, health care facilities, and industries into a community.
Methods
Study design and area
A health-institution-based cross-sectional study was conducted from 20 October 2020 to 1 December 2020 among health institutions located in Hawassa city. There are 7 hospitals (4 private and 3 governmental), 15 health posts, 7 health centers, 47 drug stores, 12 diagnostic laboratories, and 12 pharmacies in Hawassa.
Study population and sampling technique
The source population for this study was all health facilities which provide laboratory diagnosis and treatment services at different levels (n = 14). Health institutions that provide health care service during the study period, those who were voluntary to take part in the study, and those who had a functional sewage system that is suitable for sample collection were included. Initially, we intended to include all health facilities that provide diagnosis and treatment services; however, we managed to include seven health facilities. The remaining seven were excluded from the study: three health facilities did not give us permission to collect samples; sewage systems of four health facilities were not suitable for sample collection.
Of seven health facilities included in the current study, Bete Abraham Primary Hospital (BAPH), Loke Tulo Bushulo Clinic (LTBC), and Alation Primary Hospital (ALPH) belong to private, whereas Hawassa University Comprehensive Specialized Hospital (HUCSH), Adare Primary Hospital (APH), Tula Primary Hospital (TPH), and Alemura Health Center (AHC) were governmental. A total of 27 sewage samples were collected from different source lines of sewage systems of health institutions before entering the septic tank (n = 24) and from the septic tank (n = 3), and transported to Hawassa Public Health Institute of Southern Nations, Nationalities, and People’s Regional State (SNNPRS) for bacteriological analysis and antimicrobial susceptibility testing (Table 1).
Sources, frequency, and numbers of sewage samples collected for analysis from health institutions found in Hawassa city, Sidama Regional State, Ethiopia, from 20 October 2020 to 1 December 2020 (n = 27).

HUCSH: Hawassa University Comprehensive Specialized Hospital; APH: Adare Primary Hospital; BAPH: Bete Abraham Primary Hospital; ALPH: Alation Primary Hospital; AHC: Alemura Health Center; TPH: Tula Primary Hospital; LTBC: Loke Tulo Bushulo Clinic.
Sample collection and isolation of bacteria
A duplicate 250 mL sample was collected in a sterile bottle of 500 mL capacity from each site and transported within the sample box to the Public Health Institute of SNNPRS for bacteriological analysis and antimicrobial susceptibility testing within 30 min. The sample was thoroughly mixed and diluted 1 in 10 before inoculating onto a culture media; 0.5 mL of the diluted samples were inoculated onto MacConkey agar, Mannitol salt agar, and blood agar, and incubated at 37°C for 24 to 48 h. Identification of bacteria was based on colony appearance, Gram reaction, growth on selective media, and biochemical tests. Pure colonies were prepared by subculturing onto blood agar. For Gram-negative bacteria, biochemical tests such as indole production, citrate utilization test, oxidase test, triple sugar iron agar, lysine decarboxylase, urease test, hydrogen sulfide production test, carbohydrate fermentation tests, and Sulfide Indole Motility medium were used. For identification of Gram-positive cocci, catalase and coagulase tests were used.
Antimicrobial susceptibility testing
Kirby–Bauer disk diffusion method was used to determine antimicrobial susceptibility profiles of the isolated bacteria according to the Clinical and Laboratory Standards Institute (CLSI) guidelines. 12 Antibiotics tested include tetracycline (30 µg), penicillin (10 µg), gentamicin (10 µg), erythromycin (15 µg), clindamycin (2 µg), azithromycin (15 µg), ampicillin (10 µg), ceftriaxone (30 µg), chloramphenicol (30 µg), ciprofloxacin (5 µg), kanamycin (30 µg), vancomycin (30 µg), and doxycycline (30 µg).
A suspension of bacteria was prepared by mixing a pure colony of bacteria in 4 to 5 mL normal saline and adjusted to the turbidity to 0.5 McFarland standard. The suspension was spread uniformly over the entire surface of the Mueller–Hinton agar using a sterile cotton swab to produce confluent growth. Antibiotic discs were placed on the surface of the inoculated medium and incubated at 37°C for 18 to 24 h. Zones of inhibitions were measured and compared to the standard, and interpreted as Resistant (R), Intermediate (I), and Susceptible (S).
Data quality control
The expiry date of all reagents, culture media, and antibiotic discs was inspected before using them. Pseudomonas aeruginosa (American type culture collection (ATCC) 27853), E. coli (ATCC 25922), and Staphylococcus aureus (ATCC 25923) reference strains were used to check the performance of culture media. The sterility of culture media was checked by incubating 10% of the culture media at 35 to 37°C for 24 h.
Data analysis
Data were entered and analyzed by SPSS, version 25. Descriptive statistics such as frequency and proportion were used to analyze the data. The results were organized, summarized, and presented in text and tables.
Results
Culture positivity rate
A total of 27 sewage samples were collected in the present study. Of these, 24 samples were collected before sewage enters the septic tank and 3 samples were collected from the septic tank. All samples collected in this study were positive for bacteria (Table 2).
Sample collection site and culture positivity rate of sewage samples collected from various health institutions found in Hawassa, Sidama Regional State, Ethiopia, from 20 October 2020 to 1 December 2020 (n = 27).
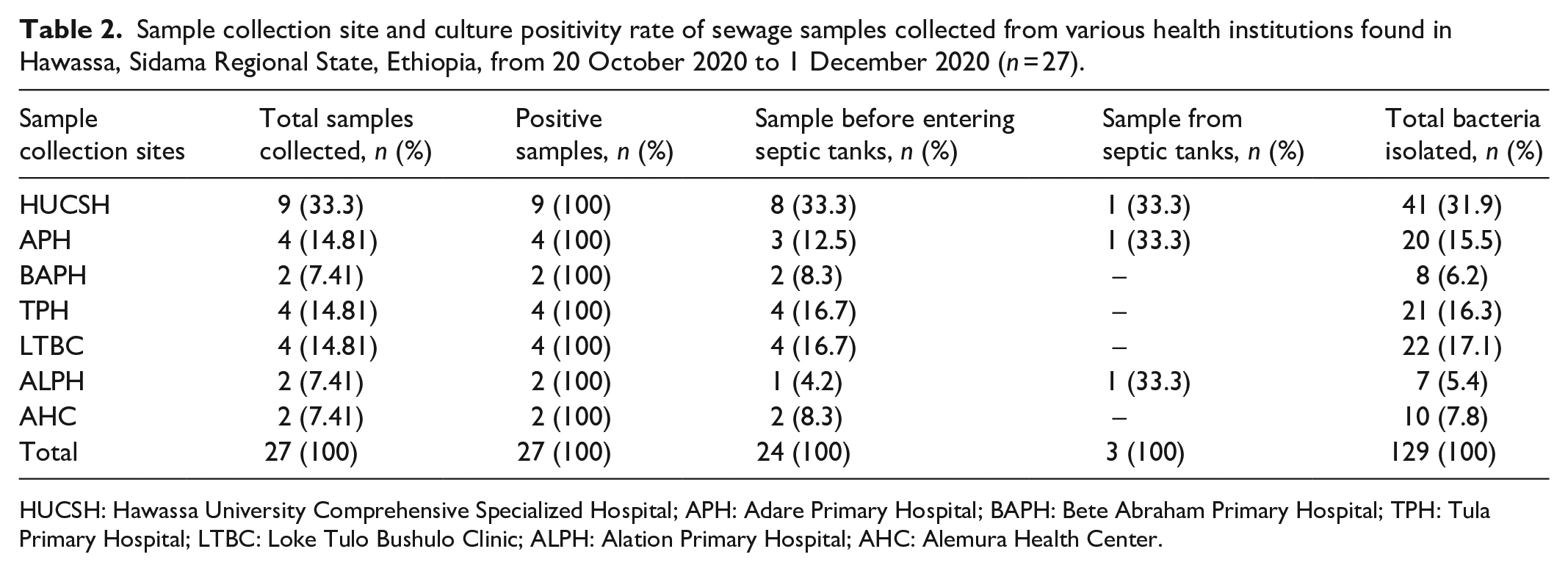
HUCSH: Hawassa University Comprehensive Specialized Hospital; APH: Adare Primary Hospital; BAPH: Bete Abraham Primary Hospital; TPH: Tula Primary Hospital; LTBC: Loke Tulo Bushulo Clinic; ALPH: Alation Primary Hospital; AHC: Alemura Health Center.
Distribution of bacteria
In this study, a total of 129 bacteria were isolated. Out of which, 115 (89.2%) were Gram negative and 14 (10.8%) were Gram positive. Six different types of bacteria were identified, which include E. coli (n = 27, 20.9%), Shigella species (n = 26, 20.2%), Salmonella species (n = 25, 19.4%), Pseudomonas species (n = 25, 19.4%), S. aureus (n = 14, 10.9%), and Klebsiella species (n = 12, 9.3%) (Table 3).
Distribution of bacteria across different sewage sample collection sites of health institution of Hawassa city, Sidama Regional State, Ethiopia, from 20 October 2020 to 1 December 2020 (n = 129).

HUCSH: Hawassa University Comprehensive Specialized Hospital; APH: Adare Primary Hospital; BAPH: Bete Abraham Primary Hospital; TPH: Tula Primary Hospital; LTBC: Loke Tulo Bushulo Clinic; ALPH: Alation Primary Hospital; AHC: Alemura Health Center.
Antimicrobial susceptibility profile
All Gram-negative and Gram-positive bacteria were susceptible to azithromycin. More than 80% of bacteria were resistant to ampicillin. More than 80% of bacteria were susceptible to norfloxacin, ciprofloxacin, and gentamicin (Table 4). Out of 129 bacteria identified, 38 (29.5%) were MDR. Close to 42% of Klebsiella species and 42.9% of S. aureus were MDR (Table 5).
Antimicrobial susceptibility profile of bacteria isolated from sewage system of health institutions found in Hawassa city, Sidama Regional State, Ethiopia, from 20 October 2020 to 1 December 2020 (n = 129).
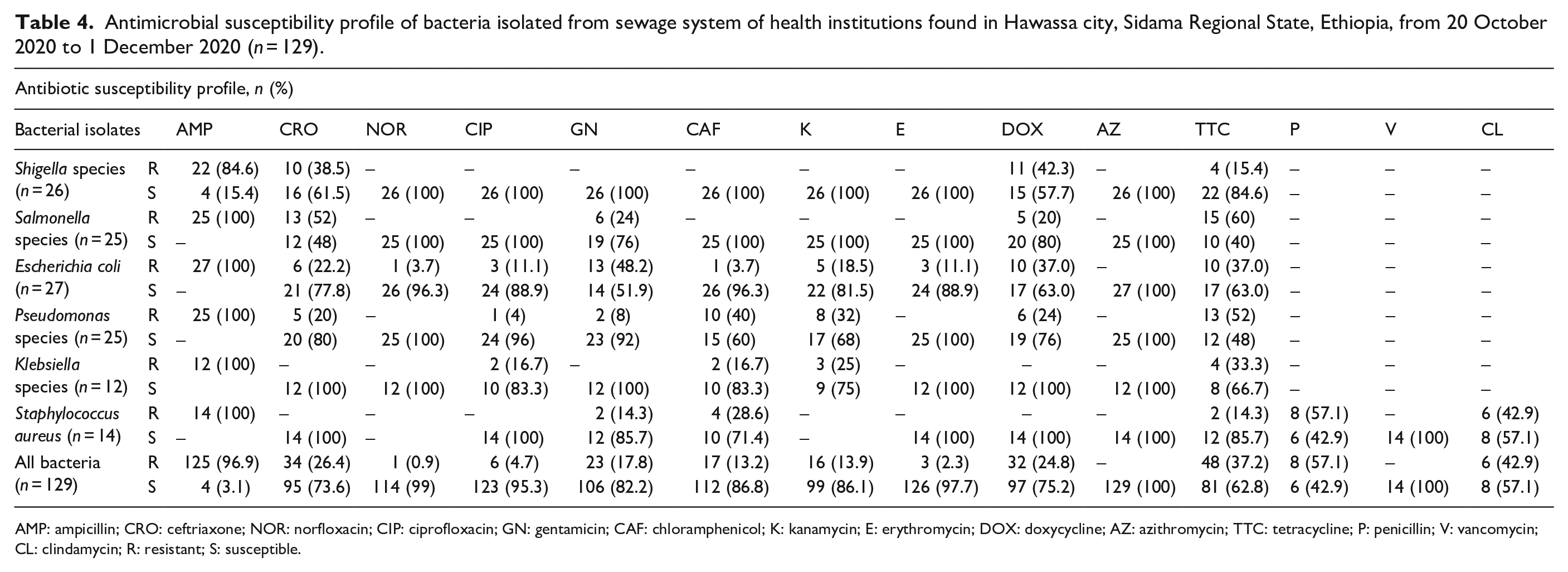
AMP: ampicillin; CRO: ceftriaxone; NOR: norfloxacin; CIP: ciprofloxacin; GN: gentamicin; CAF: chloramphenicol; K: kanamycin; E: erythromycin; DOX: doxycycline; AZ: azithromycin; TTC: tetracycline; P: penicillin; V: vancomycin; CL: clindamycin; R: resistant; S: susceptible.
Multidrug resistance profile of bacteria isolated from sewage system of health institutions found in Hawassa city, Sidama Regional State, Ethiopia, from 20 October 2020 to 1 December 2020 (n = 129).

AMP: ampicillin; CRO: ceftriaxone; DOX: doxycycline; TTC: tetracycline; GN: gentamicin; NOR: norfloxacin; CIP: ciprofloxacin; CAF: chloramphenicol; K: kanamycin; E: erythromycin; P: penicillin; CL: clindamycin.
Discussion
All sewage samples collected from seven health institutions were positive for bacteria. Overall, six different types of bacteria were identified among which 89.2% were Gram negative and 10.8% were Gram positive. The most prevalent bacteria were Shigella species (20.2%) followed by Salmonella species (19.4%), S. aureus (10.9%), E. coli (20.9%), Pseudomonas species (19.4%), and Klebsiella species (9.3%). Similarly, a high proportion of Salmonella species, Shigella species, E. coli, and S. aureus were isolated from the hospital sewage of Hawassa University Referral Hospital. 11 In contrast to this study, the most predominant bacteria from the Al-Shifa hospital sewage system (Gaza) were Pseudomonas species (31.8%) followed by E. coli (21.5%), Klebsiella species (15%), and S. aureus (7.4%). 13
Some species of bacteria were not identified from all health institutions included in this study. Klebsiella species were not detected from BAPH and ALPH, and S. aureus was not identified from BAPH, ALPH, and AHC. Our finding is not in agreement with a report from Mekelle, Ethiopia, in which the most predominant bacteria were Klebsiella species, P. aeruginosa, S. aureus, E. coli, and Salmonella species. 14
About 96% of bacteria isolated in this study were resistant to ampicillin which is not in line with a study conducted in Iran and Nigeria.15,16 The high proportion of ampicillin-resistant bacteria detected in the current study could be due to the wide and misuse of antibiotics in the hospital environment. Greater than 26% of bacteria isolated in the current study were resistant to ceftriaxone. This can challenge the use of ceftriaxone for the treatment of life-threatening diseases. All bacteria isolated in this study were susceptible to azithromycin. A similar finding was reported from Nigeria. 17 More than 85% of bacteria we identified were susceptible to norfloxacin, ciprofloxacin, erythromycin, chloramphenicol, and kanamycin. This finding is in line with studies conducted in different parts of Ethiopia.18,19 In contrast to our study, a high proportion of ciprofloxacin- and aminoglycoside-resistant bacteria were observed among Gram-negative bacteria in Brazil. 20
In this study, all E. coli were resistant to ampicillin. However, according to a meta-analysis done in Ethiopia, a low proportion of E. coli (83.81%) were found to be resistant to ampicillin as compared to our study.17,21 In addition, unlike our study, none of E. coli collected from Addis Ababa, Ethiopia, was resistant to ampicillin. 18 In this study, a relatively low proportion of E. coli (37%) were resistant to tetracycline which is in contrast to a study conducted in Addis Ababa, where greater than 70% of E. coli were resistant to tetracycline. 18 In contrast, most E. coli (54%) recovered from urine in Addis Ababa were resistant to ciprofloxacin. 18
All S. aureus isolated were resistant to ampicillin, which disagrees with the finding from Ethiopia in which 75% of S. aureus were resistant to ampicillin. 19 On the contrary, all S. aureus we isolated were susceptible to ciprofloxacin and vancomycin. However, a meta-analysis from Ethiopia reported S. aureus strains that are resistant to vancomycin (11%) and ciprofloxacin (19%). 19 In contrast to our study, where 57.1% and 14.3% of S. aureus were resistant to penicillin and tetracycline, respectively, a high proportion of penicillin-resistant S. aureus (76%) and tetracycline-resistant S. aureus (62%) were reported from Ethiopia. 19
All types of bacteria we isolated except Shigella species were resistant to ampicillin; this finding is not in line with a report from other parts of Ethiopia.22–24 Most E. coli and all types of bacteria recovered in this study were susceptible to norfloxacin, which is inconsistent with a study conducted in Dessie, Ethiopia, where all Klebsiella species recovered from the wound were resistant to norfloxacin. 19 Unlike the report by Hussen et al., 25 all Shigella species isolated in this study were susceptible to erythromycin.
About 50% of Salmonella species, 38.5% of Shigella species, 22.2% of E. coli, and 20% of Pseudomonas species isolated in this study were resistant to ceftriaxone. Unlike our study, a high proportion of bacteria from different parts of Ethiopia were found to be resistant to ceftriaxone.19,22,26 Furthermore, 100% and 60% of Salmonella species we isolated were resistant to ampicillin and tetracycline, respectively. On the contrary, a low pooled prevalence of ampicillin-resistant (80.6%), ciprofloxacin-resistant (8.7%), and ceftriaxone-resistant (12.2%) Salmonella species were reported from Ethiopia. 27 Like our study, a high proportion of Salmonella species (66.9%) isolated in Iran were resistant to tetracycline. 28 Greater than 23% of Pseudomonas species detected in this study showed resistance to ampicillin, tetracycline, chloramphenicol, kanamycin, and doxycycline. This finding is consistent with a report from India. 29 According to our study, 100%, 33.3%, and 25% of Klebsiella species were resistant to ampicillin, tetracycline, and kanamycin, respectively. The finding is in agreement with a report from China. 30
Out of 129 bacteria identified, 29.5% were MDR which is lower than MDR reported from Addis Ababa, Ethiopia (57.1%), 23 and Hawassa, Ethiopia (57.1%). 24 The proportion of MDR Klebsiella species (41.7%) and MDR S. aureus (42.9%) is lower than a report from Addis Ababa in which 100% of S. aureus and 75% of Klebsiella species were MDR. 23 MDR E. coli (37.3%) we detected in this study is low compared to a report from Hawassa, Ethiopia (57%). 24 Overall, the difference observed in antibiotic susceptibility profile might relate to irrational drug use, empiric treatment, laboratory method used, and source of bacteria.
Limitations
The current study was conducted for only a few months and cannot represent the bacterial profile of the sewage systems of all seasons of the year. The study was conducted in only seven health institutions, and the finding of this study cannot be generalized to all health institutions found in Hawassa city. We did not assess factors that could be associated with bacterial contamination of the sewage system. Sample size was not calculated in this study.
Conclusion
All sewage systems of health institutions included contained different types of pathogenic bacteria. Most bacteria isolated belong to Gram-negative bacteria. The most prevalent bacteria were E. coli followed by Shigella species, Salmonella species, Pseudomonas species, S. aureus, and Klebsiella species. All bacteria were susceptible to azithromycin. More than 80% of the identified bacteria were susceptible to norfloxacin, ciprofloxacin, and gentamicin. Most bacteria were resistant to ampicillin.
Footnotes
Acknowledgements
We would like to thank all health institutions that participated in the study, and Mr. Tamirat Legesse and Mr. Mohammed Aynie for their contribution during data collection.
Author contributions
B.M.M. conceived and designed the study, and performed the laboratory work and analysis of the data. S.H. involved in protocol development, data analysis, and supervision. M.M.A. conceived and designed the study, and involved in supervision, data analysis, and manuscript preparation. All authors read and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Availability of data
The data used during the current study are available from the corresponding author upon request.
Ethical approval
The study was obtained ethical clearance from the Institutional Review Board of the College of Medicine and Health Sciences, Hawassa University, with reference number IRB/022/13.
