Introduction
The type of infection that results from Salmonella enterica is determined by the virulence factors of the bacterium and the host’s factors.1–3 Salmonella infection could manifest clinically as gastroenteritis (diarrhea, abdominal cramps, and fever) or a fatal febrile systemic infection (typhoid) that needs to be treated with antibiotics.4–7 Focal conditions and asymptomatic carriers are possible and are significant sources of continued infection transmission.
8
Salmonella is a gram-negative, flagellated, with O, H, and Vi antigens. More than 1800 Salmonella serovars are known.2,9 Salmonellae infection is possible when the bacteria gets through the gastric acid barrier, into the mucosa of the small and large intestines, and makes toxins.
10
Invasion of the epithelial cells causes the release of cytokines that cause inflammation.
11
It results in diarrhea, leading to ulceration and the destruction of mucosal cells. Also, if this spread in the intestines persists, it could result in systemic infection.
11
Horizontal gene transfers shape bacterial genomic diversity.
12
Some pathogens have genomic islands or islets (GI) with functionally linked genes.13,14 Salmonella pathogenic islands (SPIs) contain virulence genes, distinguishing them from nonpathogenic types.15–18 Virulence factors determine a host’s pathogenicity.
19
The adhesion and invasion of host cells by pathogenic S. enterica are aided by pef, spv, invA, and fim genes, respectively. Magnesium transport C (mgtC), Salmonella toxin (stn), and pip A, B, and D help the bacterium survive in the host system.20,21
The burden of invasive bloodstream infection due to S. enterica is increasing, especially in developing countries.
5
Globally, 1.2 million deaths attributable to S. enterica are recorded annually, with the vast majority occurring in resource-limited countries.
22
In resource-limited countries, non-typhoidal Salmonella infections cause bacteremia in immune-compromised and malnourished adults and children.
16
Most morality from Salmonella infection is connected to poor diagnostic infrastructures leading to misdiagnosis and drug misuse.
23
It is established that most pathogenic bacteria are acquired from the environment, food, and water sources.24,25 Although typhoidal Salmonella is a human host-adapted strain, recent literature has found typhoidal Salmonella in the food chain and water sources.26–30
The worldwide rise of multi-drug resistance is a major health concern.
31
It is becoming increasingly important to routinely apply antimicrobial susceptibility testing to select the antibiotic of choice and to screen for emerging multi-drug resistant (MDR) strains,
32
as several recent studies have reported the emergence of multi-drug resistant Salmonella pathogens from various origins, including humans,
33
birds,
34
cattle,
31
and fish.
35
Pathogens’ high antimicrobial resistance has been attributed to antimicrobial misuse in human and animal husbandry.36–39 Comparative genomic data from invasive Salmonella data and those from the environment and food chain have shown relatedness between clinical isolates and other sources.10,11,40 It has caused great concern as clinical S. enterica are resistant to the commonly used antibiotic, and some have been found to harbor extended-spectrum beta-lactamases (ESBL) genes which could make treatment difficult.41–46 Recent data from surveillance has found non-typhoidal Salmonellae to be highly resistant to antimicrobials.4,47–49 Thus, clinical care for individuals with invasive Salmonella infection is expensive, increases their hospital stay, and burdens them financially.50,51 Studies have been carried out on Salmonella virulence factors, but information on invasive isolates is scarce, especially from the pediatric population. This study aims to investigate the virulence and antimicrobial resistance genes pattern of S. enterica from invasive bloodstream infection in children from north-central Nigeria.
Materials and methods
Study design
This is a secondary cross-sectional analysis of isolated S. enterica in children with bloodstream infection in the Federal Capital Territory (FCT) and Nasarawa State, Nigeria.
Collection of Salmonella isolates analyzed in the study
Eighty-three gram-negative bacilli isolates were collected from blood cultures recovered from the study conducted at seven hospitals in FCT and Nasarawa State, Nigeria. Presumptively identified Salmonella isolates from four thousand and sixteen blood cultures were processed from June 2015 to June 2018. The study was part of Community-Acquired Bacteremia Syndrome in Young Nigeria Children conducted in north-central Nigeria from 2008 to 2018 by the International Foundation Against Infectious Diseases in Nigeria. The outcomes from 2008 to 2015 had been previously reported by Obaro et al.,
5
and those previously reported isolates were excluded from this study.
Isolation of S. enterica of positive blood culture
Bacterial analysis, including gram staining and biochemical analysis using the analytical profile index (API20E) (Biomerieux, SA Lyon, France), was used to identify the Salmonella pathogens. Obaro et al.
5
have previously described the protocol used to culture and isolate the S. enterica used in this work. Briefly, all positive bottles were subcultured onto MacConkey agar (Oxoid, London, UK) and Salmonella shigella agar (Oxoid) plates and then incubated at 36°C for 24 h. The isolates were frozen at −80°C in 10% skim milk glycerol (Hardy Diagnostics, Santa Maria, California, USA) until used.
52
In conducting the study, the previously collected isolates were grown onto S. shigella agar (Oxoid) and incubated at 37°C for 24 h. In checking for Salmonella isolates, colonies were looked for on the plates. The morphological traits and characteristics of Salmonella species were selected, and gram staining of the selected colonies from each plate was examined.
52
Biochemical assays, including reactions on triple sugar iron agar, lysine iron agar, indole synthesis in tryptone broth, and urea splitting ability, were then conducted using the Phoenix MD (Beckon-Dickson systems, San Jose, California, USA). Molecular invA gene detection was used to validate the authenticity of the isolates. Polyvalent Salmonella antisera, A-G, A-S surface antigen, flagellar H (phase 1 and phase 2) (Beckon-Dickson systems) according to Kauffman-White Scheme
53
were used in the serotyping of the Salmonella isolates.
By Antimicrobial Susceptibility Testing and Multiple Antimicrobial Resistance (MAR) Index according to Clinical and Laboratory Standards Institute recommendations,
54
the antibiotic susceptibility of the isolates was determined. The antibiotic discs (ampicillin (10 μg), amoxicillin-clavulanate (30/10 μg), piperacillin (30 μg), piperacillin-tazobactam (30/10 μg), streptomycin (10 μg), trimethoprim-sulfamethoxazole (10/25 μg), chloramphenicol (30 μg), tetracycline (30 μg), aztreonam (30 μg), gentamicin (10 μg), amikacin (30 μg), cephalothin (30 μg), cefuroxime (30 μg), cefotaxime (30 μg), ceftazidime (30 μg), cefepime (30 μg), ceftriaxone (30 μg), levofloxacin (5 μg), meropenem (10 μg), imipenem (10 μg), tigecycline (25 μg), cefotaxime-clavulanate (30/10 μg), and ceftazidime-clavulanate (30/10 μg) that were utilized in a disk diffusion assay were from Oxoid. The BD PhoenixTM M50 system (Beckon-Dickson systems) was used for minimal inhibitory concentration (MIC) testing. For ciprofloxacin, MIC values >0.064 g/mL were viewed as reduced susceptibility, while MIC values 4 g/mL were interpreted as resistant; for azithromycin, MIC values >16 g/mL were interpreted as resistant. According to Davis and Brown,
55
the MAR indexes were derived as the ratio of antibiotics to which resistance was demonstrated to the number of antibiotics for which the isolate was screened for susceptibility. According to Algammal et al.,
32
the resistance profiles were classified as MDR, extensive drug resistant (XDR), or pan-drug resistant (PDR).
Molecular detection of resistance and virulence genes
Genomic DNA was extracted using the Maxwell 16-cell DNA purification kit (Promega, Madison, Wisconsin, USA) on an automated machine (Maxwell 16 extraction system, Madison, Wisconsin, USA). The real-time polymerase chain reaction (PCR) assays were performed on the AriaMx system (Agilent Inc., Santa Clara, California, USA). Primers and probes were purchased from LGC, Biosearch (Novato, California, USA) for the different genes based on primers and probes used by Ibrahim et al.
56
for invA; Bugarel Weil et al.
57
for sopB, ssaQ, mgtC, spi4D, spvC, and ljsGI-1; Roschanski et al.
58
for blaTEM, blaSHV; Vien et al.
59
for qnrA; Singh and Mustapha
60
for floR and tetG; and Guarddon et al.
61
for tetA and tetB. Supplemental Tables 1 and 2 show the genes sequences and amplification conditions used. A quality-controlled positive and negative internally characterized known resistant and susceptible Salmonella typhi strains from International Typhoid Consortium
62
were used as controls for amplification for detecting the resistance genes and virulence during PCR. Also, no template controls were incorporated into the PCR as an additional method of internal control in the PCR.
A 12.5 μL of Perfecta master mix low ROX kit (Quanta Bioscience Inc., Madison, Wisconsin, USA), 1.0 μL of each 10 mM primers and probes, 7.5 μL of Nuclease free water (Sigma-Aldrich, St Louis, Missouri, USA), and 2.0 μL of DNA template make up a 25 μL PCR reaction mixture. Thermal conditions were those described by the referenced authors (Tables 1 and 2). After the amplification experiments were completed, the cycle thresholds were determined by identifying the fluorescence signal by analyzing the amplification plots in AriaMx system software version 3.1.
Statistical analysis
Data were imputed and validated in Excel 2016. Descriptive statistics were computed for the multiple antibiotic resistance index. Agreement between the values of antimicrobial resistance phenotypes and their corresponding genotypes was established by κ value (coefficient of agreement) according to Jeamsripong et al.
63
Chi-square and Fisher’s exact test were used to test association as appropriate in every case. p < 0.05 was taken as statistically significant. Statistical Package for Social Science Version 20 (IBM, Santa Barbara, California, USA) was used.
Discussion
Increased resistant S. enterica has continued to pose a significant threat to human health and animal protection, and their spread is being increasingly found in clinical, food, and animal samples. Our study found typhoidal Salmonella serovars and non-typhoidal Salmonella serovars similar to the report by Awol et al.
64
in a multicenter study. Ke et al.
65
and Stanaway et al.
66
show that children in poor and middle-income countries with sub-optimal water, sanitation, and hygiene have a considerable and progressive increase in invasive non-typhoidal Salmonella (iNTS) infection. Invasive typhoid and non-typhoidal Salmonella have been previously reported in Nigerian children.5,8,27,67,68
Regarding the serovars found in this study, S. typhi was the highest, followed by S. choleraesuis, S. enteritidis, and S. typhimurium. Salmonella typhi is the most common serovar in invasive Salmonella infection in children, according to previous studies conducted in Nigeria and elsewhere.5,8,67,69–71 Serovars S. enteritidis and S. typhimurium are not frequently observed in invasive non-typhoidal infections in industrialized countries. Still, in sub-Saharan Africa, they are becoming a reoccurring decimal. Two African authors have previously reported them in invasive Salmonella infection.72,73
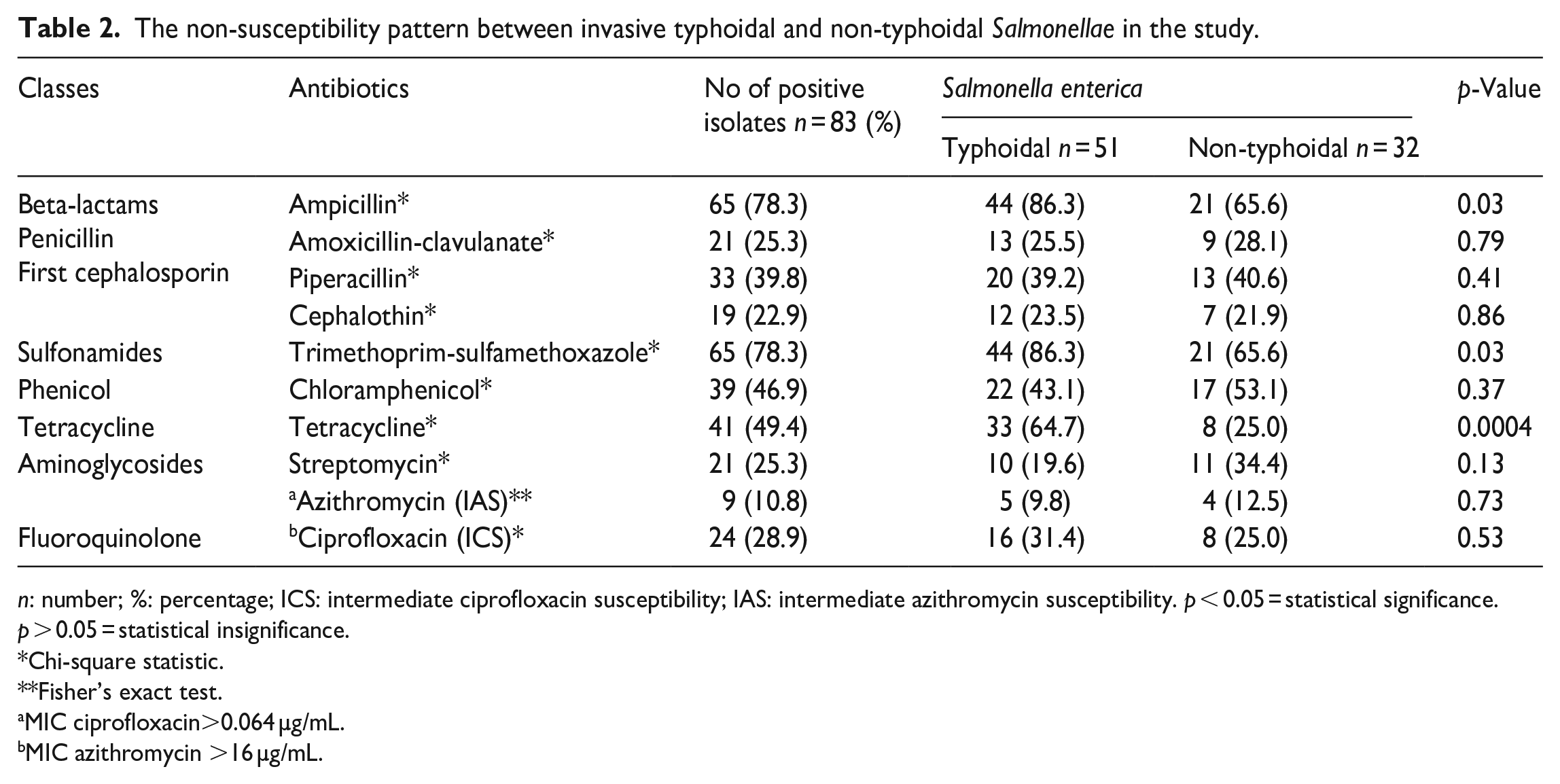
High levels of resistance to ampicillin, trimethoprim-sulfamethoxazole, tetracycline, and other routinely used antibiotics were found in our investigation. Salmonella strains isolated from invasive environments resist many of the most widely used antibiotics.5,67,70,71 In particular, multidrug-resistant iNTS caused life-threatening invasive disease outbreaks in children in Nigeria, Rwanda, and Malawi.67,74,75
The antimicrobial resistance of iNTS is a big problem because it can cause bacteremia in immunocompromised people.
76
The high prevalence of antimicrobial-resistant Salmonella is a serious concern for public health.
58
Salmonella typhi was the most common cause of invasive typhoidal Salmonella infection, and its high MDR and low ciprofloxacin susceptibility rates were the most striking findings of our investigation. This result is consistent with patterns seen in Cambodia by Vlieghe et al.
77
when describing MDR in S. typhi. The MDR has been observed in S. enterica in several countries in literature.78–80 Salmonella typhi with multi-drug resistance was found in an assessment of typhoid fever cases in Pakistan, Vietnam, India, China, Indonesia, and Nigeria.73,81–85Our finding regarding MDR in Salmonella isolates in Nigeria agreed with previous reports and the assertion that Nigeria is in the vicinity region referred to as a hotspot for antimicrobial overuse.19,86
The MAR index, a cost-effective and valid method of bacteria origin tracking, was also calculated. It has become possible to tell bacteria apart by their resistance to the most popular antibiotics in human medicine by doing a MAR analysis.87–89 Compared to other methods of bacteria source tracking, such as genotypic characterization, the MAR indexing method is cost-effective, rapid, and easy to perform.
90
MAR index values greater than 0.2 indicate a high-risk source of contamination and index for high antibiotics usage.
88
This study reported a high MAR index >0.2 for S. enterica from invasive sources, which concerns the efficacy of treatment options available. High MAR in S. enterica has been attributed to plasmids containing one or more resistance genes,91–93 each encoding a single antibiotic resistance phenotype.
94
This study did not find XDR and PDR support in literature.91–93
Our results revealed significant positivity of blaTEM in the S. enterica studied. The blaTEM gene found in this study was slightly higher in invasive typhoidal strains than iNTS. The presence of the blaTEM gene in most of the S. enterica in our study supported earlier assertion, which suggested that blaTEM genes code for beta-lactam drug resistance like ampicillin.
95
Beta-lactamase produced by gram-negative bacteria remains the primary mechanism by which they develop resistance to beta-lactam antibiotics. Additionally, ESBLs are increasingly common among S. enterica serovars, and their frequency and prevalence have been reported to rise.96–98 Although blaCTX-M and blaSHV were not found in this study, recent studies from Asia and some parts of Africa have reported them,96,99 specifically reports of the blaSHV gene from clinical S. enterica from India.100–103 Therefore, monitoring the incidence of blaTEM in S. enterica isolates is a crucial public health tool in combatting this threat.
Salmonella enterica isolates with phenotypically intermediate ciprofloxacin susceptibility (ICS) were found to harbor plasmid-mediated quinolone resistance (PMQR) genes (qnrA). This observation agrees with a report from India
104
with observations in human samples (51.4%), food-producing animals (28.6%), environmental samples (11.4%), and animal samples (8.6%), respectively.89,105–107 Ciprofloxacin has become prominent in treating severe infections caused by S. enterica, especially those resistant to nalidixic acid, which has increased significantly in recent years. Still, high resistance levels to ciprofloxacin are rare, but its resistance is foundational for other resistance mechanisms.75,108,109 Mutants resistant to fluoroquinolones are being rapidly selected due to the spread of PMQR genes.
110
Moreover, interactions between mutations in the QRDR and PMQR genes might result in high fluoroquinolones MIC. However, a study
111
speculated that the qnr genes could increase fluoroquinolone resistance.
In literature, tetA, tetB, and tetG were consistently found in S. enterica of human origin. The tetB gene is predominant among the phenotypic tetracycline-resistant strains in the literature. Our findings agreed with two studies that have reported tetA and tetB in S. enterica from a human with gastroenteritis in India and Nigeria.112,113 tetG has also been reported in humans with bacteremia in Nigeria.
113
However, studies examining tetracycline resistance in multiple isolates reported tetA, tetB, and other types in S. enterica isolates from humans and those from animals, environments, and poultries.113,114 Regarding the family Enterobacteriaceae, the tetB and tetA tetracycline resistance determinants have historically been the most prevalent.
112
However, tet (C, D, E, M, O) associated with tetracycline resistance in Salmonella species and other bacteria are less frequently found.
Salmonella isolates that have developed phenotypic resistance to chloramphenicol are strongly linked to the development and expression of efflux that pumps the drug out of the bacteria’s cells,115,116 encoded by floR or cml genes. In our study, all invasive S. enterica phenotypically resistant to chloramphenicol had floR gene. These findings agree with reports of floR gene detection from S. enterica in the literature.117,118 Also, it has been asserted that the floR gene of Salmonella pathogenicity island-1 contributes to S. enterica infectivity.
92
Chloramphenicol was one of Nigeria’s most common drugs of choice in treating Salmonella-related infections. A survey revealed 72.4%–89.2% increased resistance from 1997 to 2007, thus limiting its therapeutic value.117,118 In Iraq, chloramphenicol-associated genes were highly occurring in S. enterica strains isolated from clinical samples.119,120 In literature, chloramphenicol was the first-line drug used to treat typhoid fever, but its recurrent use limits its therapeutic value due to resistance development.119,120
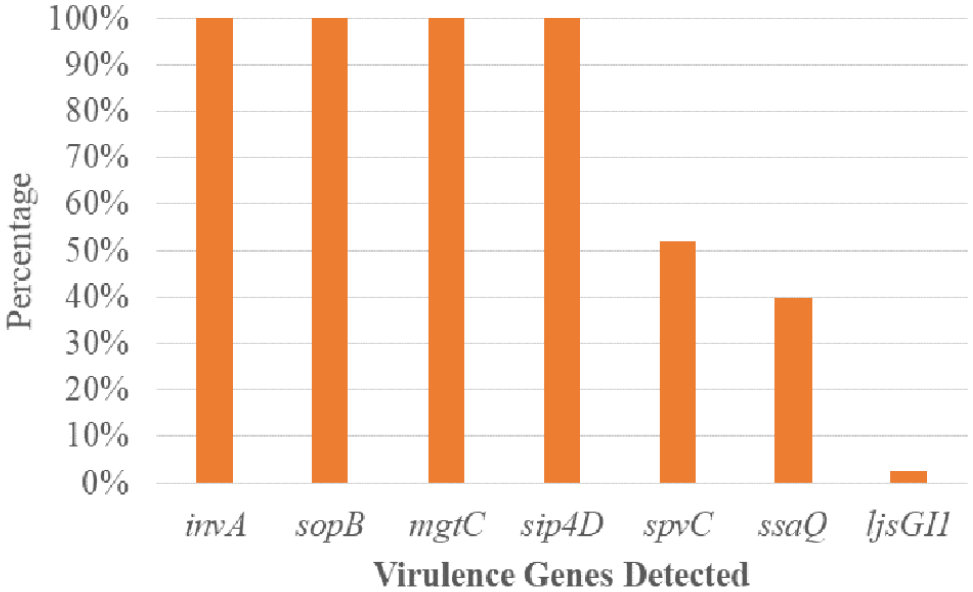
The presence of invA, sopB, spi4D, and mgtC genes in all the tested isolates agreed with the literature’s earlier evidence.13,121,122 Salmonella invasion gene (invA) is involved in the invasion of the intestinal epithelium cells and is found in pathogenic S. enterica.
123
Therefore, for Salmonella infection to occur, invasion of the cells must occur, aided by invA gene.124–126 The invA gene influences the type of Salmonella infection that could result in either systemic or localized.
127
This gene is a transcriptional regulator required to express several genes encoding type III secretion system SPI-1 effector proteins.57,121,128 The invA gene was previously hypothesized to be widely distributed among the S. enterica isolates irrespective of their serovars or source of isolation. Thus, the invA gene is a suitable target for detecting S. enterica from different biological specimens, as documented in the literature.56,60,121,127,129–131
The inositide phosphate phosphatase (sopB) gene is an effector protein that induces macropinocytosis. Gene sopB is an actin-binding protein that interacts with the host cell actin cytoskeleton. It is required for efficient bacterial internalization by the host cell.
132
The sopB gene in all isolates is instructive because sopB has been reportedly involved in micropinocytosis.
133
Macrophages have been considered the main target of Salmonella during infection, and these cells are responsible for bacterial dissemination and control.134–136 In addition to macrophages, other immune system cells are targets of S. enterica pathogens, dendritic cells, and neutrophils.
Furthermore, B cells have also been targeted by S. enterica through the expression of sopB.
137
The sopB genes are necessary for intracellular survival in the host, so the presence of sopB gene is suggested to contribute to the invasiveness of S. enterica pathogens,138,139 found in all the isolates in this study. The sopB gene is also involved in host cell survival by activating the Akt signaling pathway, including activation of the host innate immune system and cell death.
133
The presence of bacterial effector sopB in our study supported the earlier assumption that activation of Akt pathway is mediated through the expression of sopB.
In literature,
126
Salmonella’s SPI-3 island is associated with intra-macrophage invasion, which supports survival when Mg2+, required in the bacteria transported system, is of limited amount. The presence of mgtC gene in all the Salmonella in this study supported the proposition that S. enterica uses the expression of mgtC gene to circumvent the lack of Mg2+ in the bacteria. They, therefore, initiate Mg2+ production without depending on the host for Mg2+. Our findings are supported in earlier literature.13,137,139,140 Salmonella enterica contains several transport systems, both inducible and constitutive.
140
These transport systems have functional complementarities to adjust the Mg2+ concentration in different environmental conditions. In addition, these systems are controlled by transcriptional and post-transcriptional regulatory networks to maintain strict control of the Mg2+ balance.
126
Regarding maintaining Salmonella viability and development in environments with low Mg2+ levels, mgtC appears to be the most crucial SPI-3 component, as reported in some S. enterica isolates
140
and S. typhi.
13
Since mgtC is encoded in a region of SPI-3 that is highly conserved, it plays a crucial role in virulence that is not met by any other factor encoded in SPI-3 or anywhere else on the S. enterica chromosome.121,141
The ssaQ gene was detected in 33 (39.8 %) Salmonella isolates examined. The importance of this gene is relevant in the surveillance of S. enterica, which has been involved in systemic infection in the past. It has been found to produce proteins for the bacteria that bind to and stabilize the larger protein, which is important for the overall efficiency of the secretory system.
142
Essential for virulence in host cells, survival in macrophages, and biofilm development is the ssaQ gene, which codes for proteins in the SPI-2 type III secretion system.
143
The Salmonella plasmid virulence (spvC) gene was significantly higher in iNTS than in typhoid Salmonella isolates. By eliminating their beta-subunits, the spvC gene renders inactive the host’s dual-phosphorylated mitogen-activated protein kinases. It is also hypothesized to play a role in systemic S. enterica infection due to its anti-inflammatory effector effects and attenuation of the intestinal inflammatory response.
144
The spvB gene may collaborate with spvC and other Salmonella effectors to play a role in pathogenesis by triggering apoptosis in human macrophages. The spv genes increase the virulence of non-typhoid Salmonella serovars to induce extra-intestinal illness, as shown by experimental models and human epidemiological data.
145
Intestinal infections caused by non-typhoid Salmonella, typically present as self-limiting gastroenteritis, can be terminated by spv genes.
129
In mice, a study
145
discovered that the spv locus in Salmonella serovars is a crucial distinction in the pathogenesis of typhoid fever compared to that of non-typhoid Salmonella bacteremia.