Abstract
Objectives:
Molecular approaches to identifying resistance-conferring mutations suggest a revolution in the field of tuberculosis. The aim of the study was to determine the association between resistance-conferring mutations with fitness loss in Mycobacterium tuberculosis clinical isolates and HIV co-infection in the Amhara region of Ethiopia.
Methods:
A laboratory-based cross-sectional study was conducted between September 2022 and June 2023. A line probe assay was performed on 146 culture-positive clinical isolates. Logistic regression analysis was used to measure the strength of the association between the drug-resistance-conferring mutations with fitness loss in M. tuberculosis isolates and tuberculosis/HIV co-infection. A p-value ⩽ 0.05 was considered statistically significant.
Results:
A total of 11 distinct mutations at four genetic loci among 19 resistant isolates were detected. The frequency of rifampicin, isoniazid, and fluoroquinolones resistance-conferring mutations was identified in 12 (8.2%), 17 (11.6%), and 2 (1.4%) of the isolates, respectively. The most prominent specific mutations were S450L (5/9, 55.6%), S315T (11/11, 100%), C-15T (4/4, 100%), and D94G (1/1, 100%). Double mutations were observed in 10 (52.6%) multidrug-resistant tuberculosis isolates; the most common were detected in both the rpoB and katG genes (8/10, 80.0%). The HIV-co-infected tuberculosis patients carried a higher proportion of low fitness of non-rpoB S450L variants than those tuberculosis patients without HIV (80.0% vs 14.3%) and showed a significant association (cOR = 0.042, 95% CI: 0.002–0.877, p = 0.041), but not with the low fitness of non-katG S315T variants (cOR = 3.00, 95% CI: 0.348–25.870, p = 0.318).
Conclusion:
This study provides valuable information on the genetic variants with fitness loss associated with HIV co-infection, but requires further whole-genome-based mutation analysis.
Introduction
Tuberculosis (TB) is a contagious disease that continues to remain a major health problem and a major cause of mortality and morbidity. 1 The timely diagnosis of TB and adequate treatment regimens contribute to controlling the transmissibility of drug-resistant TB (DR-TB) and patients’ mortality. 2 In Ethiopia, the diagnostic methods include acid-fast smear staining, GeneXpert MTB/RIF assay, BACTEC MGIT 960 broth culture system, and line probe assays (LPAs), which are currently implemented in some parts of the country but are not routine tests in each hospital except smear staining. This could permanently affect the accurate diagnosis of DR-TB patients.3–7 The extent of occurrence of DR-TB and its burden vary significantly from country to country and from region to region, but it is highly prevalent in resource-limited settings, mainly in Asia and Africa. 1 The rising number of 10.6 million global TB cases in 2021 could be due to immune suppressive disorders like the HIV co-epidemic (6.7%), 1 which synergistically magnifies the burden of the disease. 8 There are other reasons such as bad quality of drugs including an increase in drug resistance, poor quality of National Tuberculosis Programs, and lack of patient compliance with the therapeutic regimen. 9
Resistance to anti-TB drugs in M. tuberculosis isolates is obtained mainly through mutations in drug resistance-related genes, such as rpoB gene encoding the β subunit of RNA polymerase in the 81-bp hotspot region of the rifampicin (RIF) resistance determining region (RRDR) 10 ; responsible for resistance; katG, inhA, and fabG promoters that confer isoniazid (INH) resistance; gyrA and gyrB genes linked to fluoroquinolones (FQs) resistance; rrs, eis, and tlyA genes are responsible for the injectable agent (IA) resistance.11,12 The accumulation of point mutations in coding regions for drug targets and/or drug-converting enzymes is a major mechanism for acquiring resistance by reducing the bacterium’s susceptibility to drugs 13 because the tubercle bacilli have no known efficient mechanism for horizontal gene transfer. 14
When bacteria develop drug-resistant mutations, there is often an associated biological cost of M. tuberculosis isolates. Biological costs were determined by comparing the growth curves of drug-resistant isolates with drug-susceptible strains. Although drug-resistant strains often suffer an initial reduction in fitness, they continue to evolve by acquiring one or more secondary-site mutations that can improve or even restore fitness over time. 15 This bacterial biological fitness is variable as it may depend on the type of resistance-conferring variants in the essential rpoB and katG genes in clinical isolates in the absence of drugs. 16 Furthermore, although the acquisition of drug resistance determinants is often associated with a loss in fitness, some strains with the most frequent resistance-conferring rpoB S450L and katG S315T mutations correlated to a high resistance toward RIF and INH, respectively, confer relatively low fitness costs, can grow faster as well as or better than drug-susceptible clinical isolates, and likely facilitate increased transmission ability.15–19 Nevertheless, the level of fitness effect of inhA mutations associated with low-level INH resistance was unclear, 17 but the inhA mutation does not affect katG activity. 16 The fitness cost of known mutations in DR-TB isolates of M. tuberculosis is mostly influenced by strain genetic background 18 and HIV co-infection in the absence of drugs.16,19 The low-fitness cost of resistance-conferring mutations to RIF and INH in M. tuberculosis is more frequent in HIV-co-infected patients due to their increased susceptibility. 16 To the best of our knowledge, identifying the drug resistance and conferring mutations with fitness cost in M. tuberculosis is helpful in controlling the spread of DR-TB. Limited studies have taken into account that the prevalence of genetic variants likely varies by setting as defined somewhere 20 ; however, the association of HIV co-infection with the fitness cost of genetic variants is unknown. 21 The aim of the study was to assess the association between the drug resistance-conferring genetic mutations with fitness cost and TB/HIV co-infection as well as to determine the performance characteristics of a diagnostic MTBDRplus v2.0 test.
Methods and materials
Study settings, design, and period
A laboratory-based cross-sectional study was conducted between September 2022 and June 2023 in the Northwest Amhara region of Ethiopia. The Amhara area, one of Ethiopia’s 13 regions, is located in Northwestern Ethiopia between 36° 20′ and 40°20′ E longitude and 9°20′ and 14°20′ N latitude. According to the central statistics agency of Ethiopia’s 2015 census, the region has an estimated area of 154,708.96 km2 and a total population of 20,018,988 of which the study area has a total area of 146,705.14 km2 and a population of 7,086,962 people. 22
Sample size determination
A total of 259 study participant sample sizes was calculated using a single population proportion formula, n = z2p (1 − p)/d2 in the previous study. 21 However, out of which, only 146 stored culture-positive clinical isolates of M. tuberculosis at the University of Gondar Comprehensive Specialized Hospital (UoGCSH) TB culture laboratory were used in the present study due to resource constraints. The clinical isolates were collected from GeneXpert-positive pulmonary TB patients with HIV co-infected and/or HIV-negative cases, drug-susceptible and/or drug-resistant TB, new and/or previously treated TB cases, etc. The UoGCSH is fully furnished and certified to manipulate TB cultures and perform phenotypic and genotypic susceptibility testing. Besides, it serves MDR-TB patients, mostly from neighboring zones and woredas.
Eligibility criteria
All culture-positive M. tuberculosis complex clinical isolates existing with the correct patient identification number and having sociodemographic data were included in this study. However, culture-negative M. tuberculosis complex clinical isolates, non-TB mycobacterium, and extra-pulmonary isolates were excluded from this study.
Study variables
The INH and RIF resistance-conferring mutations with fitness cost in drug-resistant M. tuberculosis clinical isolates were considered dependent variables, whereas the sociodemographic and clinical characteristics of the patients were considered independent variables.
Laboratory procedures
The N-acetyl-L-cysteine-sodium hydroxide (NALC-NaOH) method was used to process the sputum from patients with pulmonary TB who tested positive for GeneXpert. These samples were then cultured on Lowenstein–Jensen egg medium slants at 37°C for up to 8 weeks while their growth was monitored once a week. 23 The growth of M. tuberculosis complex isolates was confirmed by the antigenic method (SD Bioline TB Ag MPT64, Diagnostics, Inc., South Korea) 24 and acid-fast bacilli smear staining according to the Ethiopian Minister of Health and Global Laboratory Initiative Standard Protocols.23,25
The phenotypic drug susceptibility testing (DST) against the first-line streptomycin (STR), INH, RIF, and ethambutol (EMB) drugs was performed on stored clinical isolates of M. tuberculosis using an automated BD BACTEC MGIT 960 growth detection system (BD, Sparks, MD, USA). Each drug’s final concentration in MGIT media was as follows: STR 1.0, INH 0.1, RIF 1.0, and EMB 5.0 in µg/mL as described in the previous study. 21 The genotypic DST and molecular characterization of genetic mutations in the rpoB responsible for RIF resistance, katG and inhA genes confer high-level and low-level INH resistance, respectively, was performed on the 146 culture-positive M. tuberculosis clinical isolates using GenoType® MTBDRplus v.2.0 LPAs (Hain Life Science, Nehren, Germany).26,27 MDR-TB isolates (also known as resistant to both INH and RIF) were once again examined by GenoType® MTBDRsl v.2.0. The MTBDRsl LPA was performed to characterize known mutations in the gyrA and gyrB genes linked to FQs resistance, and the rrs and eis genes conferring resistance to second-line injectable drugs (SLIDs).26,28 The LPA experiments were carried out at the UoGCSH TB culture laboratory by following the steps of DNA-kit strip technology, including genomic DNA extraction, multiplex polymerase chain reaction amplification, reverse hybridization, and interpretation of results.
Quality control
All laboratory analyses were carried out following standard operating procedures. The data were manually double-checked for completeness and consistency and entered twice independently. For each run of the test, the M. tuberculosis H37Rv reference strain (ATCC 27294) susceptible to all tested drugs and molecular-grade water were used as positive and negative controls, respectively. 26
Statistical analysis
A pre-tested, structured questionnaire was used to collect sociodemographic and clinical data of the patients. The questionnaire used in this study was validated (Ref: VP/RTT/05/829). The data were first entered using Epidata V. 3.1 and analyzed using SPSS V. 23.0. Descriptive statistics (sociodemographic characteristics as well as resistance-conferring mutations) are described in frequencies and percentages. Tables were used to present the results. The sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV) of MTBDRplus v.2.0 were evaluated for the detection of RIF, INH, and MDR-TB compared to a gold standard phenotypic MGIT 960. Categorical data differences were evaluated using Pearson’s chi-square (χ2) or Fisher’s exact test when the expected cell frequencies were less than five. A logistic regression analysis was carried out to explore the association of host-related risk factors with the fitness cost of rpoB and katG genetic variants in M. tuberculosis clinical isolates. A p-value ⩽ 0.05 was considered statistically significant.
Results
Patients’ sociodemographic and clinical characteristics
A total of 146 stored culture-positive M. tuberculosis clinical isolates from pulmonary TB patients were included in the study. Of the 146 isolates, 125 (85.6%) were from new cases of TB, while 21 (14.3%) were from patients who had previously undergone TB treatment. Among the patients, 86 (58.9%) were male and 60 (41.1%) were female. The majority of the isolates 104 (71.2%) were obtained from patients in the 18–44 age groups and the mean age of the patients was 32.3 ± 2 years. The proportion of TB patients with HIV co-infection was 20.5% (30/146), while the percentage of TB cases without HIV was 79.5% (116/146). Significant differences were found between the group of pulmonary TB patients with HIV co-infection and the group of TB patients without HIV in terms of contact with previous TB cases (p = 0.048). However, no statistically significant differences were found between the two groups regarding sociodemographic and clinical characteristics, such as sex, age group, previous history of TB treatment, or cigarette and alcohol consumption habits (p > 0.05) (Table 1).
Sociodemographic and clinical characteristics of patients (n = 146).

χ2: Pearson’s chi-square test; HIV: human immunodeficiency virus; TB: tuberculosis.
Statistically significant.
Patterns of resistance to first-line and second-line drugs
The GenoType® MTBDRplus LPA was utilized to identify susceptibility profiles and detect mutations in the rpoB, katG, and inhA genes that confer resistance to first-line RIF and INH drugs. Among the 146 clinical isolates tested for susceptibility, the study revealed that 134 (91.8%) were susceptible to RIF and 129 (88.4%) were susceptible to INH. Also, 12 (8.2%) showed resistance related to RIF, 17 (11.6%) showed resistance related to INH, and 19 (13.0%) demonstrated resistance to at least one or two drugs. The overall proportion of MDR-TB isolates was 52.6% (10/19). Moreover, the study found that seven isolates (41.2%) had INH-mono-resistant status and 10 (58.8%) were MDR-TB isolates among the INH-related resistant isolates. Concerning the RIF-related resistant isolates, the proportion of MDR-TB isolates was higher (83.3%, 10/12) in comparison to the RIF-mono-resistant isolates (16.7%, 2/12). There was a statistically significant difference in the proportion of TB with HIV-co-infected patients and HIV-negative cases, as well as among previously treated and new TB cases in terms of drug resistance profile (p = 0.05) (Table 2).
Distribution of drug resistance profiles based on TB/HIV co-infection and history of TB treatment status (N = 146).
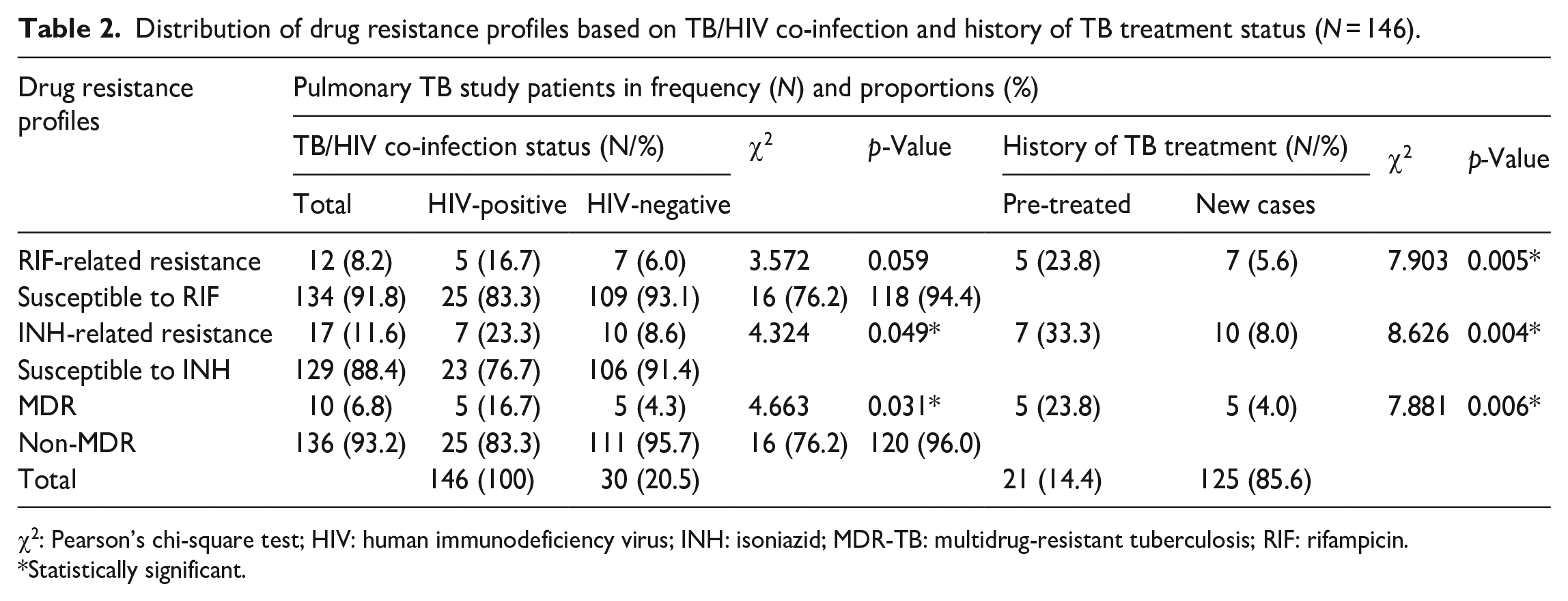
χ2: Pearson’s chi-square test; HIV: human immunodeficiency virus; INH: isoniazid; MDR-TB: multidrug-resistant tuberculosis; RIF: rifampicin.
Statistically significant.
In the study, all of the MDR-TB isolates were tested again to detect mutations in the gyrA, gyrB, rrs, and eis genes that are linked to resistance to second-line drugs. The MTBDRsl LPA assay was used for this purpose. Results showed that 8 (80.0%) isolates were susceptible to SLDs, while 2 (20.0%) were resistant to at least one of the second-line FQs drugs. Among the resistant isolates, a single FQ-resistant isolate (also known as pre-extensively drug resistance (XDR) TB) was identified. A single isolate (50.0%) was found in both HIV-co-infected patients who were new treatment cases and in HIV-uninfected individuals who had previously taken anti-TB drugs. Nevertheless, no pre-XDR-TB isolates with SLID resistance or XDR isolates were identified (Table 2).
Performance of MTBDRplus in detecting resistance to INH, RIF, and MDR-TB
As presented in Table 3, the performance characteristics of the GenoType MTBDRplus LPA were evaluated for identifying clinical isolates resistant to RIF, INH, and MDR-TB compared to the phenotypic MGIT 960 system. Out of the 146 M. tuberculosis complex isolates tested, there were concordant outcomes from both DST methods, indicating that 133 isolates (91.1%) were RIF susceptible, while 11 (7.5%) were RIF-resistant isolates and 126 isolates (86.3%) were INH susceptible, with 17 (11.6%) showing INH-related resistance. However, five (3.4%) isolates had discordant results. One (0.7%) and three (2.1%) isolates were susceptible to RIF and INH, respectively, as per the genotypic LPA assay but showed resistance in the phenotypic DST. In addition, one isolate (0.7%) was resistant to RIF according to the genotypic LPA but susceptible following the phenotypic MGIT 960.
Diagnostic performance of MTBDRplus for detecting RIF, INH, and MDR-TB resistance compared to MGIT 960 DST (N = 146).

DST: drug-susceptibility testing; INH: isoniazid; k: kappa value; LPA: line probe assay; MDR-TB: multidrug resistance tuberculosis; MGIT: mycobacterium growth indicator tube; NPV: negative predictive value; PPV: positive predictive value; RIF: rifampicin.
For the detection of MDR-TB, there were concordant results for 133 (91.1%) isolates which revealed susceptible to either INH or RIF, and nine isolates (6.2%) were resistant to both INH and RIF by both methods. Nevertheless, three isolates (2.1%) showed a discordant result with resistance to both INH and RIF by phenotypic DST but were susceptible to genotypic LPA, and one (0.7%) isolate was resistant to genotypic LPA but susceptible to phenotypic MIGT 960. The sensitivity, specificity, PPV, and NPV of the MTBDRplus LPA assay were 91.7%, 99.3%, 91.7%, and 99.3% for identifying RIF-related resistance; 85.0%, 100%, 100%, and 97.7% for detecting INH-related resistance; and 75.0%, 99.3%, 90.0%, and 97.8% for MDR-TB, respectively. There was an excellent agreement between the LPA and culture-based phenotypic DST in detecting resistance related to RIF and INH (0.91) as well as MDR-TB (0.80) (Table 3).
Frequencies of mutations associated with resistance to anti-TB drugs
The frequencies of multiple gene mutations associated with resistance to first-line drugs (such as the rpoB gene for RIF, and the katG and inhA genes for INH) and second-line drugs (such as the gyrA and gyrB genes for FQs, and the rrs and eis genes for SLIDs) were determined as depicted in Figure 1 and Table 4.

Representative patients based on the first-line LPA result.
Frequency of mutations in the rpoB, katG, inhA, and gyrA genes conferring resistance to RIF, INH, and FQs drugs (n = 19).
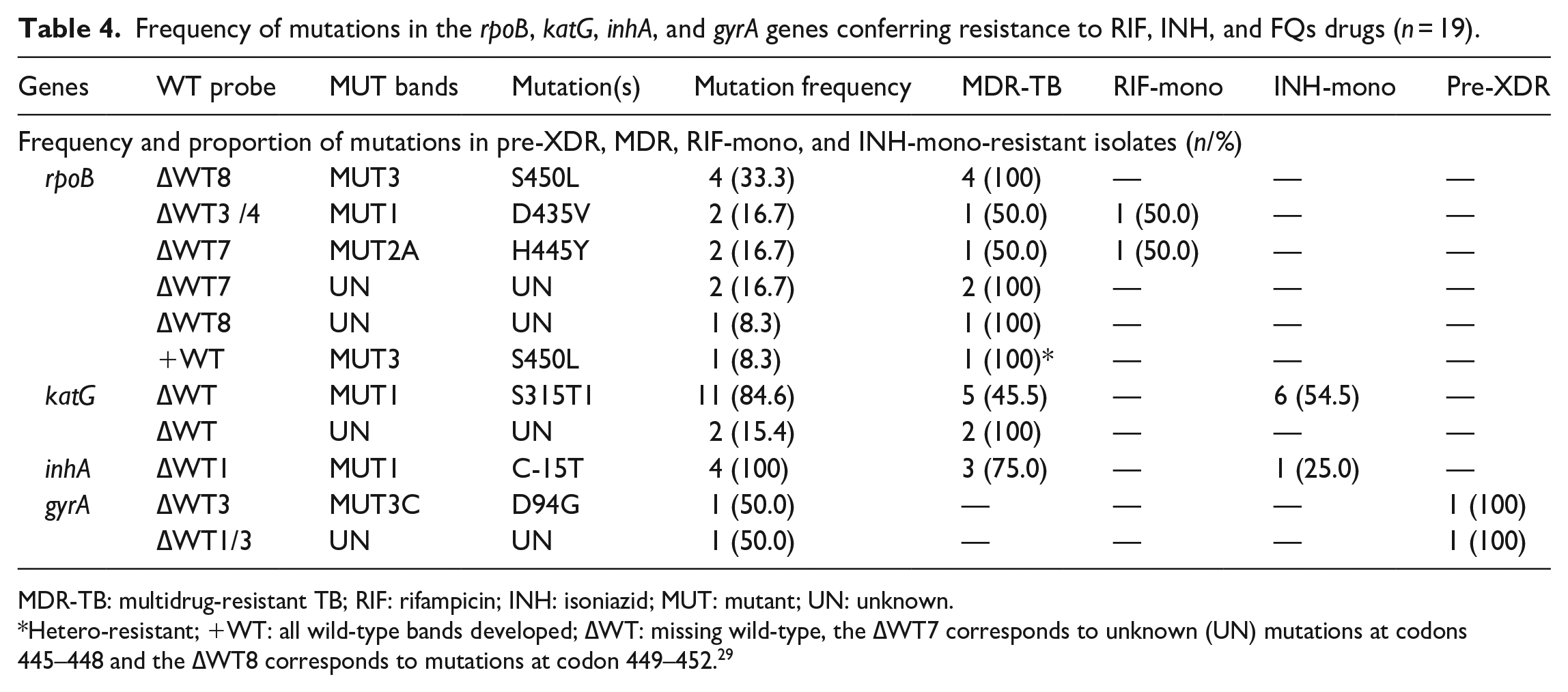
MDR-TB: multidrug-resistant TB; RIF: rifampicin; INH: isoniazid; MUT: mutant; UN: unknown.
Hetero-resistant; +WT: all wild-type bands developed; ∆WT: missing wild-type, the ∆WT7 corresponds to unknown (UN) mutations at codons 445–448 and the ∆WT8 corresponds to mutations at codon 449–452. 29
Mutations in the rpoB, KatG, and inhA are associated with resistance to first-line drugs
Among the 146 clinical isolates, a total of 29 resistance-conferring mutations correlating with resistance to first-line drugs in three genes (rpoB, katG, and inhA) were identified. A total of 12 (8.2%) RIF-related resistant isolates showed any of the resistance-conferring mutations in the rpoB gene at different amino acid positions. Ten out of 12 RIF-related resistant isolates (83.3%) were MDR-TB strains, while two isolates (16.7%) were RIF mono-resistant isolates. Nine (75.0%) of the tested clinical isolates showed known mutations in the rpoB mutant gene, which were determined by missing at least one WT band and developing known MUT bands. These isolates were referred to as RIF-resistance-detected isolates. The most prominent specific mutation pattern was found at codon 450 substitution S > L, which was detected in five out of nine isolates (55.6%). Along with this, a hybridization of ∆WT8/MUT3 (4/5, 80%) and +WT/MUT3 (1/5, 20%) was observed. All S450L mutations were found in MDR-TB strains. The next most frequent mutations in the rpoB gene were H445Y and D435V substitutions, which were detected by failing ∆WT7/MUT2A and ∆WT3&4/MUT1 hybridization with an equal frequency of two out of nine (22.2%) RIF-resistant isolates carried specific rpoB gene mutations.
Three unknown mutations, which were caused by the absence of ∆WT7 and ∆WT8 probes resulting in the lack of corresponding MUT bands, were detected in two isolates (66.7%) and one isolate (33.3%), respectively. These isolates were designated as RIF-inferred resistant isolates, and all three mutations were observed in MDR-TB isolates. Hetero-resistance or a mixed infection pattern to RIF was identified in a single isolate (8.3%) through the presence of all +WT probes with the detection of the MUT3 band in the rpoB gene. No mutation of MUT2B at codon H445D was observed in any of the tested isolates in our findings.
A mutation in the katG gene and inhA promoter region is responsible for resistance to INH. Among the 17 isolates (11.6%) that showed resistance to INH, 13 were found with katG variants. The frequency of these katG variants was 6/17 (35.3%) in INH-mono-resistant isolates and 7/17 (41.2%) in MDR-TB isolates. The known mutations in the katG gene at codon 315 that confer high-level INH resistance were detected in 11/13 (84.6%) of the isolates by missing the ∆WT probe coupled with the corresponding hybridization of MUT probe bands. The most common variant was katG MUT1, which occurred at codon S315T1 (100%). It was present in 54.5% of INH-mono-resistant isolates and 45.5% of MDR-TB isolates. A proportion of 15.4% unknown mutations in the katG mutant gene that confers INH resistance were observed in two isolates because they were not included in the LPA strips. All the unknown mutations in the katG gene were detected in MDR-TB isolates. No INH hetero-resistant isolates were found.
Mutations in the regulatory region of inhA, which correlate with low-level INH resistance, were detected in 4 (23.5%) isolates. All of these mutations were identified by missing the ∆WT1 probe along with the well-known MUT1 mutation at codon C-15T. Among low-level INH-resistant isolates that exhibited alterations in the inhA promoter region, 3 (75.0%) were MDR-TB and 1 (25.0%) were INH-mono-resistant. There were no isolates that exhibited hetero-resistance to INH in the inhA mutant region.
According to gene mutation analysis, dual mutations in the rpoB and katG genes and the rpoB and inhA genes were detected in 70.0% and 30.0% of MDR-TB isolates, respectively. Nevertheless, no double mutations have been found in the katG and inhA genes in any of the INH-resistant isolates and the triple rpoB, katG, and inhA genes in any of the MDR-TB isolates (Table 4).
Mutations in the gyrA, gyrB, rrs, and eis genes are linked with second-line drugs
In all, 10 clinical isolates of MDR-TB were analyzed to identify mutations in the gyrA, gyrB, rrs, and eis genes to cause resistance to second-line FQs and injectable drugs. Two out of the 10 MDR isolates (20%) were pre-XDR isolates with a combination of mutations at rpoB, katG, and gyrA genes. A single pre-XDR isolate was detected by the S450L rpoB, only katG ∆WT and gyrA ∆WT3 along with the corresponding D94G-specific mutations, while the remaining isolate was identified by the D435V rpoB, C-15T inhA, and only ∆WT1&3 gyrA without the known mutations. There were no isolates that had mutations in the gyrB gene, which would cause resistance to FQs, as well in the rrs and eis target genes that would cause resistance to SLIDs.
Risk factors associated with the fitness loss of genetic variants in isolates
To investigate the correlation between host-related risk factors and the drug resistance-conferring mutations with a fitness cost in clinical isolates of M. tuberculosis, a total of 12 RIF-resistant and 17 INH-resistant strains having specific mutations in the rpoB, katG, and inhA genes were used. Out of which, an equal proportion of 41.7% (5/12) RIF-resistant isolates and 41.2% (7/17) INH-resistant isolates were from HIV-co-infected cases and previous TB treatment cases. Among the 12 RIF-related resistant and 17 INH-related resistant isolates, six distinct mutations in the rpoB and three distinct mutations in the katG and inhA genes were detected, respectively. In this study, the essential genes for RIF and INH resistance mutations were divided into two groups based on the extent of fitness loss in M. tuberculosis. The most common mutations at the S315T codon in the katG gene, which causes high-level INH resistance, and the S450L codon in the rpoB gene, which confers high-level RIF resistance, were considered to be high fitness or variants without fitness loss (most commonly cited as no or low fitness cost of variants), while any mutations conferring resistance at other codons (non-rpoB450 and non-katG315 mutations observed rarely among clinical isolates) were considered to be low fitness (exhibiting a high fitness loss or higher fitness cost), as previously documented.15,16,18,19 Specific mutations at codon rpoB450 that confer resistance to RIF were found in 5 out of 12 isolates (41.7%), while mutations at non-rpoB450 codons were observed in 7 out of 12 isolates (58.3%). In the case of INH resistance, 11 isolates (64.7%) had the katG315 variant, whereas 6 isolates (35.3%) carried variants at non-katG315 codons. The study noted that there was a higher proportion of low-fitness rpoB variants in TB/HIV co-infected patients (4/5, 80.0%) compared to HIV-negative patients (1/7, 14.3%). A bivariate analysis demonstrated a significant correlation between the low fitness of non-rpoB S450L variants and TB/HIV co-infection (cOR = 0.042, 95% CI: 0.002–0.877, p = 0.041). This study found that the low fitness of rpoB variants was more prevalent among previously treated TB cases (3/5, 60.0%) than new TB cases (2/7, 28.6%) and showed no significant differences (p = 0.286). The low fitness of katG variants was not significantly related to any of the host-related factors such as HIV co-infection status and TB treatment status (p > 0.05) (Table 5).
Bivariate analysis of factors associated with and without fitness cost of mutations in TB patients.

cOR: crudes odds ratio; CI: confidence interval; HIV: human immunodeficiency virus; INH: isoniazid; RIF: rifampicin.
Statistically significant.
Discussion
In the present study, a higher rate of MDR-TB was detected among previously treated patients than among newly diagnosed TB patients (23.8% vs 4.0%), as well as among patients co-infected with HIV compared to those without HIV (16.7% vs 4.3%). Our findings are supported by previous studies that recognized a higher prevalence of MDR-TB cases among previously treated TB cases than among newly diagnosed TB cases in Addis Ababa, Ethiopia (20.3% vs 5.2%), 30 China (52.53% vs 36.65%) 11 and (34.5% vs 6.8%), 12 and India (12.21% vs 7.81%). 9 In addition to acquired resistance, person-to-person transmission of DR-TB strains is the main mechanism of drug resistance in M. tuberculosis during the course of treatment. Therefore, managing DR-TB requires increased attention to high-risk patients, such as those co-infected with HIV or those who have had prior TB therapy, as well as in high-risk settings like hospitals.
The study identified resistance-conferring mutations in seven target genes responsible for resistance to FLDs and SLDs. Among any drug-resistant isolates, the overall 63.2% (12/19) RIF-related resistant isolates had resistance-conferring specific mutations resulting in amino acid changes and/or unknown mutations in the rpoB gene. Moreover, 10 out of 12 (83.3%) were MDR-TB isolates. This suggests that RIF resistance is considered a surrogate marker for the detection of MDR-TB. The study found that among the resistant isolates related to RIF, there was an overall proportion of 75.0% of known mutations in the rpoB gene that confer RIF resistance through amino acid changes within the 81-bp hotspot region. This finding is consistent with reports from Northwest Ethiopia (68.5%) 7 and India (79.0%). 9 However, the values in our finding are lower than the findings from studies in the Tigray region of Ethiopia (92.5%), 31 China (91.7%), 12 Pakistan (84.4%), 32 and Burkina Faso (84.5%). 33 This lower result in our study was due to the methodology used (LPA method) which only detects the specific mutations in the 81 bp hotspot region, whereas there are other mutations located outside this region that were conferring resistance to drugs, and also a small sample size.
The most common genetic variant in the rpoB gene associated with high levels of resistance to RIF was found to be at codon S450L (55.6%), followed by H445Y and D435V (22.2%). Our findings in S450L are consistent with the high rates reported in other studies across various regions such as Eastern Ethiopia (59.1%), 34 Northwest Ethiopia (64.0%), 7 Iran (66%), 35 the Central Africa Republic (53.4%), 36 Pakistan (55.6%) 32 and (64.1%), 2 India (58.8%), 9 and China (62.5%) 12 and (58.2%). 37 By contrast, the frequency of mutation at codon S450L in our study was higher than the previous report from Iran (40%) 38 and China (31.25%) 39 but lower than studies conducted in the Tigray region of Ethiopia (70%), 31 Addis Ababa Ethiopia (81.3%), 30 and Pakistan (85%). 40 The high-frequency occurrence of the S450L mutation in the rpoB gene associated with RIF resistance may indicate that drug resistance has been acquired, resulting in a reduced fitness cost. The possible reasons for the differences in the occurrence of specific mutations in the rpoB gene could be due to the patterns of drug resistance-conferring mutations, geographic locations, and sociodemographic characteristics of the study participants, including HIV co-infection and prior TB therapy. This assumption is supported by the research of Welekidan et al. 31
Out of the 17 INH-resistant isolates, a total of 64.7% had specific mutations at codon 315 of the katG gene, which is similar to the previously reported findings in Northwest Ethiopia (70.6%), 6 Northeastern Ethiopia (66.7%), 4 India (71.9%), 13 China (57.6%), 11 and South Africa (63.9%), 41 but lower than from Northwest Ethiopia (87.8%), 7 the Tigray region of Ethiopia (78%), 31 Eastern Ethiopia (96.5%), 34 India (75.3%), 9 China (85.4%), 12 and Pakistan (81.6%). 32 The KatG S315T1 substitution (100%) was the most frequent specific variant that caused high-level INH resistance in our study. This finding aligns with the previous studies that identified the S315T1 variant as the most common genetic variation in the katG gene in Northwest Ethiopia (70.6%) 6 and (87.8%), 7 Addis Ababa Ethiopia (76.9%), 5 Iran (70%), 35 India (86.3%), 9 China (73.8%), 37 Burkina Faso (77.3%), 33 and Pakistan (78.2%) 32 ; but higher S315T1 mutation reported in Eastern Ethiopia (91.8%), 34 Addis Ababa Ethiopia (100%), 30 India (96.2%), 13 central Africa Republic (94.2%), 36 and Mongolia (99.1%) 42 ; and lower proportions of S315T1-specific mutations were observed in Northeastern Ethiopia (25%), 4 China (62.9%), 12 Iran (32%), 43 and Pakistan (69.8%). 2 A specific C-15T mutation in the inhA regulatory region was detected in a higher proportion of 100% in our study, which is consistent with previously reported results in Eastern Ethiopia (100), 34 Mongolia (98.0%), 42 and India (88.9%) 9 ; however, lower rates were detected in Northwest Ethiopia (41%) 6 and (3.3%), 7 Addis Ababa Ethiopia (30.8%), 5 Tigray region of Ethiopia (5%), 31 China (5.7%) 12 and (24.6%), 37 South Africa (13.4%), 41 and Pakistan (14.9%) 32 and (5.6%). 2 The reasons for differences in the frequency of genetic mutations might be due to variations in the sample size, the methodology used to characterize mutations, and sociodemographic factors such as prior TB treatment and HIV co-infection, which is supported by Ranjan et al. 13
A total of 16.7% of INH-resistant isolates showed unknown mutations in the katG gene. Our finding is consistent with previously defined reports of uncertain katG mutations in the Tigray region of Ethiopia (17.1%), 31 China (20%), 12 Burkina Faso (3.6% katG), 33 and India (13%). 9 It is known that approximately 10%–25% of INH-resistant strains with uncertain mutations have mutations outside of the KatG and inhA genetic loci. 33
In the study, dual mutations were found at both the ropB and katG genes (70%) and the rpoB and inhA genes only in cases of MDR-TB (30.0%). However, no isolate showed mutations at both the katG and inhA genes, or no triple mutations at the rpoB, katG, and inhA genes. Our results are in line with previous studies conducted in Ethiopia, which also found no isolates with combined mutations in both katG and inhA genes7,31,34 or triple mutations at the rpoB, katG, and inhA genes from India. 9 By contrast, other studies have shown a higher rate of double mutations at both the katG and inhA genes of INH-mono-resistant isolates in the Central Africa Republic (4.2%), 36 China (13.4%) 11 and (3.4%), 12 Pakistan (14.7%) 40 and (1.1%), 32 South Africa (22.7%), 41 Burkina Faso (14.5%), 33 and India (3.4%). 9 The extent of INH resistance conferred by various mutations may vary depending on the genetic background of the strain. Earlier research revealed that strain lineage two (Beijing genotype) was associated with any resistance to at least one of the tested drugs and strain lineage one was linked to mutations in the inhA promoter at codon C15T. 44
In our study, a single isolate was detected in the gyrA gene with a common mutation at codon D94G associated with FQ-resistant isolates, but the high prevalence of gyrA mutations has been well described in different regions.2,7,34,45 This could be due to the low prevalence of FQ-resistant isolates in our study because of the insufficient sample size used. Besides, no mutations in the gyrB gene conferring resistance to FQs, rrs, or eis genes causing SLIDs were observed in our study. Supporting our results, previous studies in Ethiopia revealed that none of the isolates had mutations in the gyrB gene conferring resistance to FQs.31,34 Contrarily, studies reported resistance-conferring mutations of the gyrB gene in Uganda (0.98%), 45 an unknown mutation of the WT1 rrs gene in Eastern Ethiopia (4.1%), 34 the rrs gene at codon A1401G in Pakistan (71.4%), 2 Uganda (77.0%), 45 and China (92.3%). 37
In the present study, the rpoB gene mutations with low fitness (high fitness cost of non-rpoB S450L) were significantly associated with TB/HIV co-infection (cOR = 0.042, 95% CI: 0.002–0.877, p = 0.041), but not the low fitness of non-katG S315T variants (cOR = 3.00, 95% CI: 0.348–25.870, p = 0.318). This result is in line with previous findings that have demonstrated a strong association between low fitness of rpoB variants and HIV co-infection (OR = 4.58, 95% CI: 1.69–12.44, p = 0.003). 16 This remarkable association may be explained by the highly characterized rpoB variants with low fitness that confer RIF-resistant isolates in TB patients co-infected with HIV, as well as by lower bacillary loads in HIV-co-infected TB patients, which may lead to fewer strain replication events and fewer mutations available for selection. Although there is a selection for high-fitness drug-resistant M. tuberculosis strains in TB patients in clinical settings, such strains with low-fitness resistance mutations might propagate efficiently in immunocompromised hosts, such as HIV co-infected patients, which could partially explain why DR-TB has been associated with HIV co-infection. 16 However, the low fitness of the non-rpoB S450L variant was not significantly associated with HIV co-infection in Uganda (OR = 2.29, 95% CI: 0.83–5.77, p = 0.111) or the non-katG S315T variant (OR = 1.54, 95% CI: 0.81–2.90, p = 0.181). 19
One more finding in our study, the MTBDRplus LPA assay exhibited a high sensitivity of 91.7% and a specificity of 99.3% for detecting RIF resistance, which is comparable with results reported in earlier studies in Northeastern Ethiopia (100% and 98.3%), 4 China (98.7% and 88.9%), 37 Pakistan (98.8% and 92.9%), 32 and Iran (91% and 100%) 35 ; while there was a higher sensitivity than from Northwest Ethiopia (75.0% and 100%) 6 and higher specificity than from South Africa (95% and 75%). 46 This difference in sensitivity might be due to variations in sample size and the presence of strains that carried mutations outside the 81-bp hotspot region or in the amino-terminal end of the rpoB gene that was not detected by LPA strips in detecting RIF resistance. Moreover, some mutations within the hotspot region of the rpoB gene in clinical isolates cause the WT probe to fail to hybridize with MUT strands on a strip, leading to them being misinterpreted as RIF-resistant isolates. This can result in inappropriate treatment and the acquisition of further drug resistance.33,40,42 The LPA assay may occasionally detect false-positive RIF resistance due to the possibility of silent mutations in the hotspot region that do not result in amino acid changes. Furthermore, our study revealed high sensitivity of 85.0% and 100% specificity for the MTBDRplus assay for identifying INH resistance, which is consistent with other studies from Northwest Ethiopia (94.4% and 100%), 6 South Africa (93% and 95%), 46 Pakistan (90.6% and N/A), 32 and China (82.1% and 94.4%) 37 ; but, our study exhibited higher sensitivity than those from Northeastern Ethiopia (75%) 4 and Pakistan (71.7%) 2 and Iran (71%). 35 The reduced sensitivity of the MTBDRplus assay may be due to the presence of isolates carrying mutations outside the katG and inhA genomic regions causing resistance to INH. There may also be a lack of probes for a specific mutation in the other genes, such as ahpC, fabG1, kasA, furA, and the ndh region, which are not routinely targeted by the LPA assay in detecting INH resistance. Different unknown mechanisms of drug resistance may also contribute to the observed lower sensitivity.33,42 Technical errors in phenotypic and molecular assays could also be a potential reason. 2 Our study did not assess the utility of the MTBDRsl assay in detecting FQs and SLIDs resistance, as the SLDs phenotypic DST was not carried out due to a shortage of consumables and kits.
Limitations of the study
In this study, the significant association between genetic mutations with fitness cost in rpoB and katG genes and HIV co-infection, and the performance characteristic of MTBDRplus in detecting RIF and INH-resistant isolates was assessed. However, the study has certain limitations. A comparatively small sample size was involved. Second-line drug phenotypic DST was not performed and the diagnostic performance of MTBDRsl was not evaluated. The compensatory mutations were not analyzed due to the absence of DNA sequencing data, which could have been explored in relation to the fitness cost of mutations. Therefore, further study is needed using WGS to identify the diversity of strain lineages, disputed, novel and compensatory mutations throughout the genome that are relevant to predicting patterns of drug resistance and isolate characteristics. Identifying drug-resistant strains and understanding their susceptibility patterns and associated factors are crucial for effective TB management. WGS data will overcome the limitations of CCs-based phenotypic DST and probe-based characterization.
Conclusion
The study highlighted the usefulness of mutations at rpoB, katG and inhA, gyrA and gyrB, and rrs and eis genes as molecular markers for the rapid detection of resistance for RIF, INH, FLQs, and SLIDs, respectively. Such detections are essential for proper treatment, prevention, and management of the spread of DR-TB. The most common mutation associated with RIF and INH resistance could likely be at position S450L in the rpoB gene and S315T1 in the katG region, respectively. Besides, all the FQs-resistant cases were associated with the gyrA gene. Significant isolates with unknown mutations that cause drug resistance were detected, which could impact decision-making in patient management and controlling DR-TB and highlight the need to establish a WGS facility at the regional level to monitor DR mutations. The current study provides insight into the significant association between genetic mutations with fitness cost in clinical isolates of M. tuberculosis and HIV co-infection. Understanding the prevalence of specific genetic variants is crucial for the development of effective treatment strategies against drug-resistant strains of M. tuberculosis. The performance characteristic of the GenoType MTBDRplus assay has been confirmed to be highly sensitive and specific for the early detection of RIF and INH resistance. Although some isolates had discordant results between the genotypic MTBDRplus test and the phenotypic DST, the kappa agreement between genotypic DST and phenotypic DST was acceptable.
Supplemental Material
sj-docx-1-smo-10.1177_20503121231208266 – Supplemental material for Molecular characterization of genetic mutations with fitness loss in pulmonary tuberculosis patients associated with HIV co-infection in Northwest Amhara, Ethiopia
Supplemental material, sj-docx-1-smo-10.1177_20503121231208266 for Molecular characterization of genetic mutations with fitness loss in pulmonary tuberculosis patients associated with HIV co-infection in Northwest Amhara, Ethiopia by Aynias Seid, Meseret Kassa, Yilak Girma, Eseye Dereb, Semira Nureddin, Ayenesh Abebe and Nega Berhane in SAGE Open Medicine
Footnotes
Acknowledgements
First and foremost, the authors would like to acknowledge all the study participants for their willingness to participate in the study and for providing the necessary information. The authors are also very thankful to the UoGCSH TB Culture Unit, University of Gondar, Department of Medical Biotechnology, and the data collectors for their remarkable support during the study.
Authors’ contributions
All authors made a significant contribution to the work reported. AS and NB developed the conception, study design, and experiments. Author AS, MK, YG, ED, AA, and SN gathered reagents and materials and conducted all the experimental work. Author AS and SN analyzed and interpreted the data and also took part in the drafting of the article. All authors read, revised, and approved the final version of the manuscript to be submitted.
Data availability
The data used in this study are available from the corresponding author on request.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethics approval
Ethical approval for this study was obtained from the University of Gondar (UoG), Institutional Review Board (IRB) of (Ref: VP/RTT/05/829). A written letter of support was obtained from the Amhara Public Health Institute (APHI), Bahir Dar branch (Ref: APHI/W/D/03/146).
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was financially supported by the University of Gondar (UoG), Gondar, Ethiopia.
Informed consent
Written informed consent was obtained from all study participants before the study started. The privacy rights of study subjects were protected. It was carried out in line with the ethical standards laid down in the Declaration of Helsinki.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
