Abstract
Background:
In recent years, driven by the rapid advancement of proteomics research, numerous scholars have investigated the intricate associations between plasma proteins and various diseases. Thus, this study aimed to identify novel therapeutic targets for preventing and treating metabolic-related diseases through Mendelian randomization (MR).
Methods:
This study primarily utilized the MR method, leveraging genetic data from multiple large-scale publicly available genome-wide association studies. We employed two-sample MR within this framework to assess the associations between 1001 plasma proteins and 5 metabolism-related diseases. Finally, we strengthen the robustness and reliability of the MR results by conducting a series of sensitivity analyses, including bidirectional MR, colocalization analysis, Cochran’s Q test, and the MR-Egger intercept test.
Results:
The results from the inverse variance weighted method revealed that, following false discovery rate correction, many plasma proteins are significantly associated with metabolic-related diseases. Genetically predicted risks vary across diseases: for coronary artery disease, from 0.82 FGR proto-oncogene, Src family tyrosine kinase (FGR) to 1.13 (interleukin-6); for obesity, from 0.992 (POLR2F) to 1.005 (PRKAB1); for osteoporosis, from 0.998 (AIF1) to 1.001 (CLC); for stroke, from 0.71 (TNFRSF1A) to 1.47 (TGM2); and for type 2 diabetes, from 0.79 (KRT18) to 1.47 (RAB37).
Conclusion:
Our findings reveal numerous plasma proteins linked to metabolic-related diseases. These findings offer fresh insights into the etiology, diagnostics, and treatment of these conditions.
Introduction
The endocrine metabolic system is recognized as one of the most vital systems in the human body and plays a pivotal role in maintaining homeostasis. 1 Metabolic disorders frequently contribute to the onset of diseases such as diabetes, obesity, hyperuricemia, osteoporosis, stroke, and coronary heart disease. As research into metabolic-related diseases progresses, numerous studies have indicated that multiple metabolic-related diseases frequently co-occur, implying potential internal correlations. 2 Therefore, in 1999, the World Health Organization introduced the concept of metabolic syndrome, characterized by central obesity and concurrent conditions such as dyslipidemia and hyperglycemia. 3 Furthermore, stroke, coronary heart disease, and osteoporosis are also recognized as components of metabolic-related diseases. Studies indicate that their occurrence and development are interconnected, sharing common underlying causes, mechanisms, and characteristics. 4 As the primary driver of non-communicable diseases, metabolic-related conditions have emerged as a leading cause of death worldwide, affecting a growing number of patients. 5
Plasma proteins have been studied for disease diagnosis and treatment for more than a century, yet traditional proteomics techniques have limited their progress. 6 Advances in high-throughput detection and quantitative techniques for serum proteins have spurred further research into the relationship between proteins and disease risk.7,8 Plasma proteins can also act as binding sites for clinical drugs, modifying downstream protein activities to impede disease progression. 9 Previous studies have categorized plasma proteins into five distinct groups according to their potential pharmacological roles: substitution of deficient or aberrant proteins, enhancement of existing channels, provision of novel functions or activities, interference with molecular or organismal processes, and delivery of other compounds or proteins. 10 To date, numerous plasma proteins have been identified as potential therapeutic targets for various diseases.11,12 A large-scale population study identified 47 plasma proteins linked to type 2 diabetes. 13 Another study identified three plasma proteins linked to stroke. 14 Research suggests that choosing drug targets supported by direct genetic evidence in drug development trials can double the success rate of clinical drug development. 15 However, existing studies often face challenges such as small sample sizes, numerous confounding factors, and uncertainties about factors influencing proteomics, potentially leading to biased results.
Mendelian randomization (MR) was initially regarded as a promising alternative to randomized controlled trials, employing genetic tools as credible proxies for exposure and disease. According to Mendelian genetics, genetic information is randomly distributed at conception, occurring before disease onset, thereby minimizing confounding biases from various factors. 16 Furthermore, MR in conjunction with proteomics is increasingly recognized as a powerful approach for identifying potential therapeutic targets in the context of various diseases.17–19 For example, the study conducted by Bourgault et al. 20 identified 29 plasma proteins that may serve as potential therapeutic targets for acute pancreatitis, whereas Rasooly et al. 21 highlighted seven plasma proteins that could offer new therapeutic opportunities for heart failure.
This study explored the relationship between plasma protein levels and five metabolic-related diseases—coronary artery disease (CAD), obesity, osteoporosis, stroke, and type 2 diabetes—through MR. In previous studies, we utilized the most recent plasma proteomics data to identify additional drug targets for treating metabolic-related diseases.
Methods
Study design
This study utilized the MR method to investigate the causal relationships between the levels of 1001 plasma proteins and 5 metabolic-related diseases—CAD, obesity, osteoporosis, stroke, and type 2 diabetes—across multiple independent large-scale human genome-wide association studies (GWAS). The reliability of the findings was subsequently assessed via various sensitivity analysis methods and bidirectional MR. In all MR analyses, three fundamental assumptions must be rigorously followed: first, the selected genetic instruments must strongly correlate with the exposure factor; second, they must remain unaffected by confounding variables; and third, their effect on the outcome must be entirely mediated by the specified exposure factor. 22 Figure 1 depicts the foundational design framework of this study.

Study flow chart.
Data sources
All the data utilized in this study were obtained from global GWASs conducted within European populations, which were exclusively sourced from publicly accessible aggregated datasets (see Table 1).
Data sources for studied phenotypes.

GWAS, genome-wide association study.
Genetic data for plasma proteins were obtained from a recent GWAS, in which where researchers utilized Olink proteomics data from 54,000 samples in the UK Biobank to identify genetic loci associated with 1001 different plasma protein levels. 23 Genetic data for CAD were sourced from a GWAS involving 407,746 samples. 24 Data on obesity and osteoporosis were obtained from publicly available studies conducted by the MRC-IEU on the UK Biobank. 25 Genetic data for stroke and type 2 diabetes were obtained from a comprehensive study involving the UK Biobank and FinnGen research. 26 Importantly, minimal overlap in samples was observed when the datasets of plasma proteins and the five metabolic-related diseases were compared.
Instrument selection
To ensure a sufficient number of single-nucleotide polymorphisms (SNPs) for subsequent statistical analysis, we established a significance threshold for plasma protein levels of 1 × 10−5. We then utilized the European 1000 Genomes panel to select SNPs exhibiting relative independence, specifically within a genomic span of 10,000 base pairs, with a linkage disequilibrium threshold (r2) less than 0.01. Furthermore, we evaluated the statistical robustness of all the SNPs via F-statistics, excluding those with an F-statistic less than 10 to mitigate bias arising from weak instrumental variables. 27
Statistical analysis
This study primarily employs the inverse variance weighting (IVW) method to evaluate the association between plasma protein levels and type 2 diabetes. It is complemented by four sensitivity analysis techniques—weighted median, simple mode, weighted mode, and MR-Egger tests—to ensure the robustness of the IVW analysis findings. The study proceeded as follows: first, the relationships between the levels of 1001 plasma proteins and 5 metabolic-related diseases were evaluated, and the overall effect size was ultimately calculated. Cochran’s Q test was used to assess the heterogeneity of genetic instruments (p < 0.05), whereas the MR-Egger intercept test was used to assess directional pleiotropy (p < 0.05). 28 The false discovery rate (FDR) was applied for multiple testing corrections, with IVW-FDR < 0.05 set as the significance threshold. The inverse associations of identified plasma proteins linked to metabolic-related diseases were subsequently investigated. Finally, Bayesian colocalization analysis was employed to assess the probability of two traits sharing causal variants. Genes with a posterior probability (PP4) of ⩾0.8 were classified as having strong evidence of colocalization, whereas those with a PP4 of ⩾0.5 were considered to have moderate evidence supporting colocalization. 29
All the statistical analyses were conducted via the TwoSampleMR package (version 0.5.6) and the coloc package within the R software environment (version 4.3.3). Significance was evaluated via rigorous statistical criteria for two-tailed tests.
Results
Instrument selection
As indicated in Table S1, 4–573 SNPs were chosen on the basis of their respective selection criteria to investigate the associations between 1001 plasma proteins and metabolic-related diseases. All the selected genetic instruments presented F-statistics greater than 10, demonstrating the study’s ability to minimize bias from weak instrumental variables.
Proteome-wide MR analysis identified eight proteins that circulate in CAD
Table S1 displays the IVW results revealing significant associations of 138 plasma proteins with CAD (p < 0.05). However, following FDR correction, only 34 plasma proteins were consistently associated with CAD (IVW-FDR < 0.05). Among them, 18 plasma proteins were negatively associated with CAD risk, whereas 16 plasma proteins were positively associated with CAD risk. The genetic predictions of CAD risk varied from 0.82 for FGR to 1.13 for interleukin-6 (IL6; Figure 2). Despite Cochran’s Q test identifying heterogeneity in certain results, the consistent directionality observed with IVW by MR-Egger, weighted median, simple mode, weighted mode, and other sensitivity analysis methods indicates that heterogeneity can be ignored. Furthermore, the pleiotropy test revealed horizontal pleiotropy in the outcomes for CD5, LAG3, and PROC (Table S2). During the replication phase, following FDR correction and the exclusion of results exhibiting horizontal pleiotropy, we successfully validated eight plasma proteins associated with CAD (Tables S3 and S4 and Figure 3(a)). IL6 (odds ratio (OR): 1.13; 95% confidence interval (CI): 1.06, 1.21), LAYN (1.06; 1.03, 1.10), and CDH1 (1.06; 1.03, 1.09) were positively associated with CAD, whereas CEACAM8 (0.91; 0.87, 0.96), HYAL1 (0.87; 0.83, 0.91), LGALS7_LGALS7B (0.94; 0.91, 0.97), NCR1 (0.91; 0.88, 0.95), and NRP2 (0.92; 0.88, 0.97) were negatively associated with CAD.

Volcano plot of the MR analysis results.

Forest plot of MR analysis. (a) Forest plot of MR analysis in CAD. (b) Forest plot of MR analysis in obesity. (c) Forest plot of MR analysis in osteoporosis. (d) Forest plot of MR analysis in stroke. (e) Forest plot of MR analysis in T2DM.
Proteome-wide MR analysis identified one protein associated with obesity
Table S1 shows the IVW results, which revealed significant associations of 245 plasma proteins with obesity (p < 0.05). After FDR correction, 146 plasma proteins were consistently associated with obesity (IVW-FDR < 0.05). Among them, 64 plasma proteins were negatively associated with obesity risk, whereas 82 plasma proteins were positively associated with obesity risk. The genetic prediction of obesity risk ranged from 0.992 for POLR2F to 1.005 for PRKAB1 (Figure 2). Despite Cochran’s Q test identifying heterogeneity in certain results, the consistency in directionality observed with IVW by MR-Egger, weighted median, simple mode, weighted mode, and other sensitivity analysis methods indicates that heterogeneity can be ignored. Furthermore, the pleiotropy test revealed horizontal pleiotropy in the outcomes for ADA2, ADM, AIF1, and 46 other plasma proteins (Table S2). During the replication phase, following FDR correction and the exclusion of results exhibiting horizontal pleiotropy, we successfully validated only one plasma protein associated with obesity (Tables S3 and S4 and Figure 3(b)). Specifically, HYAL1 (0.998; 0.997, 0.999) was negatively associated with the risk of obesity.
Proteome-wide MR analysis identified three proteins that circulate in osteoporosis
Table S1 shows the IVW results revealed significant associations of 106 plasma proteins with osteoporosis (p < 0.05). Following FDR correction, only nine plasma proteins were consistently associated with osteoporosis (IVW-FDR < 0.05). Among them, seven plasma proteins were negatively associated with osteoporosis risk, whereas two plasma proteins were positively associated with osteoporosis risk. The genetic prediction of osteoporosis risk ranged from 0.998 for AIF1 to 1.001 for CLC (Figures 2 and 3(c)). Despite Cochran’s Q test identifying heterogeneity in certain results, the consistency in directionality observed with IVW by MR-Egger, weighted median, simple mode, weighted mode, and other sensitivity analysis methods indicates that heterogeneity can be ignored. Furthermore, the pleiotropy test revealed horizontal pleiotropy in the outcomes for PRSS27 only (Table S2). During the replication phase, following FDR correction and the exclusion of results exhibiting horizontal pleiotropy, we successfully validated three plasma proteins associated with osteoporosis (Tables S3 and S4 and Figure 3(c)). Among these, only CLC (1.001; 1.000, 1.002) was positively associated with osteoporosis, whereas C2 (0.999; 0.998, 1.000) and AIF1 (0.998; 0.997, 0.999) demonstrated negative associations.
Proteome-wide MR analysis identified 42 proteins associated with stroke
Table S1 shows the IVW results revealed significant associations of 238 plasma proteins with stroke (p < 0.05). After FDR correction, 138 plasma proteins were consistently associated with stroke (IVW-FDR < 0.05). Among them, 85 plasma proteins were negatively associated with stroke risk, whereas 53 plasma proteins were positively associated with stroke risk. The genetic prediction of stroke risk ranged from 0.71 for TNFRSF1A to 1.47 for TGM2 (Figures 2 and 3(d)). Despite Cochran’s Q test identifying heterogeneity in certain results, the consistency in directionality observed with IVW by MR-Egger, weighted median, simple mode, weighted mode, and other sensitivity analysis methods indicates that heterogeneity can be ignored. Furthermore, the pleiotropy test revealed horizontal pleiotropy in the outcomes for 47 plasma proteins (Table S2). During the replication phase, following FDR correction and the exclusion of results exhibiting horizontal pleiotropy, we successfully validated 42 plasma proteins associated with stroke (Tables S3 and S4 and Figure 3(d)). The genetically predicted stroke risk ranged from 0.78 for FSTL3 to 1.22 for SEZ6L2.
Proteome-wide MR analysis identified 11 proteins associated with type 2 diabetes
Table S1 displays the IVW results, which revealed significant associations of 160 plasma proteins with type 2 diabetes (p < 0.05). After FDR correction, 55 plasma proteins were consistently associated with type 2 diabetes (IVW-FDR < 0.05). Among them, 26 plasma proteins were negatively associated with type 2 diabetes risk, whereas 29 plasma proteins were positively associated with type 2 diabetes risk. The genetic prediction of type 2 diabetes risk ranged from 0.79 for KRT18 to 1.47 for RAB37 (Figures 2 and 3(e)). Despite Cochran’s Q test identifying heterogeneity in certain results, the consistency in directionality observed with IVW by MR-Egger, weighted median, simple mode, weighted mode, and other sensitivity analysis methods indicates that heterogeneity can be ignored. Furthermore, the pleiotropy test revealed horizontal pleiotropy in the outcomes for 18 plasma proteins, including ABHD14B and ADA2 (Table S2). During the replication phase, following FDR correction and the exclusion of results exhibiting horizontal pleiotropy, we successfully validated 11 plasma proteins associated with type 2 diabetes (Tables S3 and S4 and Figure 3(e)). Specifically, CCL27 (1.18; 1.11, 1.26), SEZ6L2 (1.15; 1.08, 1.22), OMD (1.13; 1.07, 1.20), and SPON1 (1.09; 1.03, 1.15) were positively associated with type 2 diabetes. In contrast, CSF1 (0.87; 0.80, 0.94), FLT3LG (OR: 0.94; 95% CI: 0.90, 0.98), IL15 (0.91; 0.87, 0.95), IL18BP (0.88; 0.83, 0.94), KIR2DL3 (0.96; 0.95, 0.97), MSR1 (0.90; 0.86, 0.94), and TNFRSF21 (0.88; 0.82, 0.94) were negatively associated with type 2 diabetes.
Reverse MR analysis of metabolic-related diseases with the levels of the 56 proteins
As depicted in Table S5 and Figure 4, CAD has inverse causal associations with 9 plasma proteins such as CCL27 and DLL1; obesity has inverse causal associations with 10 plasma proteins including FSTL3 and AIF1; stroke has inverse causal associations with 11 plasma proteins such as CD46; and type 2 diabetes has inverse causal associations with 6 plasma proteins, including GZMA and C2. Owing to a shortage of suitable SNPs for MR regarding osteoporosis, we refrained from exploring their reverse causal relationships with these 56 plasma proteins.
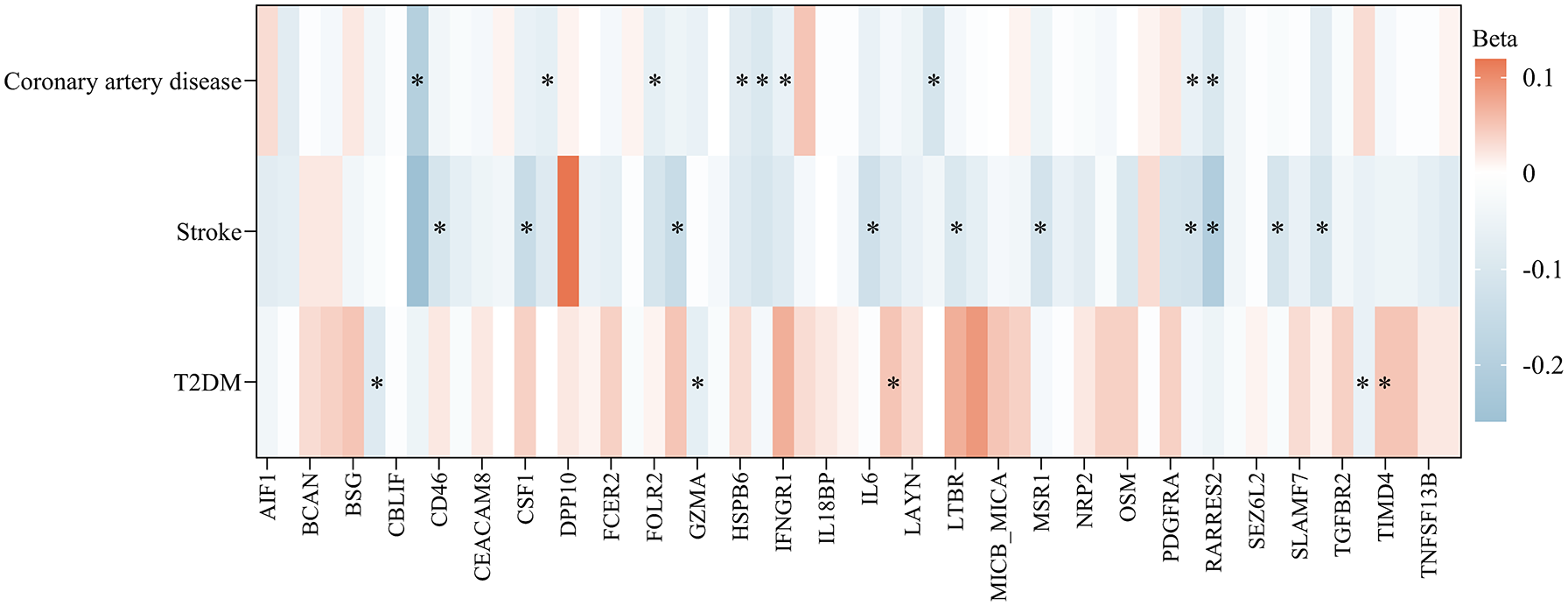
MR analysis of the associations of metabolic-related diseases with metabolic-related disease-related plasma proteins.
Colocalization analysis supported the causality of plasma proteins with metabolic-related diseases
As indicated in Table S6, among the plasma proteins identified through proteome-wide MR related to metabolic-related diseases, only the association between CCL27 and type 2 diabetes received robust genetic colocalization support (PP4 ⩾ 0.8). In contrast, HYAL1 and CAD presented moderate support (0.8 ⩾ PP4 ⩾ 0.5). Regrettably, other associations between plasma proteins and their respective diseases did not demonstrate significant colocalization support in the MR analysis.
Selected plasma proteins currently under investigation as drug targets
Table 2 presents a selection of plasma proteins that are either in development or already available on the market, on the basis of data from DrugBank. IL6-related drugs are the most commonly used primarily for treating conditions such as rheumatic polymyalgia and rheumatoid arthritis. Notably, the majority of potential plasma proteins are currently being developed as therapeutic targets for cancer treatment.
Selected plasma proteins currently under investigation as drug targets.

Discussion
This MR study assessed the associations between 1001 plasma proteins and the risk of 5 metabolic-related diseases via multiple sensitivity analysis methods. Thirty-one, 100, 8, 91, and 37 plasma proteins were significantly associated with CAD, obesity, osteoporosis, stroke, and type 2 diabetes, respectively. It also explored the inverse associations of these plasma proteins with the respective diseases. These findings provide new insights into the treatment of these diseases.
Our study initially validated associations between several plasma proteins and diseases previously identified in the literature. For example, a meta-analysis underscored the association between IL6 and CAD, suggesting that inhibiting IL6 could effectively target cardiovascular disease prevention. 30 IL6 is recognized as a proinflammatory cytokine that induces the liver to produce acute-phase proteins in response to tissue damage and infection. 31 Multiple prospective studies have demonstrated that elevated plasma IL6 levels are associated with increased CAD risk, 32 which aligns with our study findings. In our study, we observed a negative association between CCL4 and CAD risk. However, mouse studies have suggested that inhibiting CCL4 can lower the circulating levels of inflammatory factors such as IL6 and TNF-α, thus reducing atherosclerotic plaque size, slowing progression, and improving plaque stability.33,34 PPP3R1, a critical calcineurin subunit, was shown to increase energy expenditure and mitigate diet-induced obesity in genetically modified mice. 35 While our study identified pleiotropy in its association, given the involvement of various calcineurin structures in energy metabolism, 36 there remains reason to suspect a potential link between PPP3R1 and obesity.
Our study identified eight plasma proteins linked to osteoporosis. An animal study suggested that miR-181a-5p inhibits the differentiation of preosteoblastic MC3T3-E1 cells in vitro, partially through disrupting Runx1-dependent suppression of AIF1 transcription. 37 AIF1, a calcium-binding protein originating from monocytes and macrophages, may be implicated in diverse metabolic disorders, including atherosclerosis and diabetes. 38 MMPs constitute a diverse group of endopeptidases that facilitate extracellular matrix degradation or remodeling, which are pivotal in processes such as wound healing and angiogenesis. 39 Multiple MR studies have established a link between elevated plasma MMP levels and reduced stroke risk. 40 The associations of TYRO3, ARG1, and MICB with type 2 diabetes have been previously documented in certain studies, 41 underscoring the relative reliability of the data sources employed in this study.
Furthermore, we identified numerous other plasma proteins potentially linked to metabolic disorders. For example, in epithelial ovarian cancer, DDK4 might enhance tumor invasion by activating the JNK pathway to promote actin filament formation. 42 In addition, FAP is linked to poor prognosis in multiple cancers, including gastric cancer. 43 The overexpression of FAP can remodel the extracellular matrix by cleaving substrate peptides or proteins, thereby enhancing tumor cell migration. 44 Furthermore, FAP is involved in various signaling pathways including the PI3K-AKT and TGF-β pathways, 45 and dysregulation of the PI3K-AKT pathway is closely associated with obesity and type 2 diabetes. 46
This study has several significant strengths. First, genetic data on plasma proteins and five metabolic-related diseases originate from extensive population-based GWASs, which are meticulously structured to minimize sample overlap and thereby mitigate the potential confounding effects arising from shared samples across datasets. Second, all the chosen genetic instruments present F-statistics greater than 10, indicating robust instrument strength that effectively mitigates bias from weaker instruments and bolsters the credibility of the instrumental variables utilized in the analysis. Third, employing diverse sensitivity analysis methods bolstered the reliability of the study outcomes. Finally, several disease risk factors identified through routine observational studies have been shown to lack a causal association with disease in randomized controlled trials, leading to costly trials that fail to produce the anticipated positive outcomes. In contrast to conventional observational studies and GWASs, MR associations are immune to causal inversion, are less prone to confounding effects, and incur significantly lower costs, thus preserving valuable resources for more impactful research. 47
While this study is relatively comprehensive, it is important to acknowledge several limitations that may impact our interpretation of the research findings. First, MR inherently faces limitations, such as trait heterogeneity and developmental compensation issues, which could impact the accuracy and applicability of our research findings. 48 Second, our reliance on summary-level data restricted our capacity to perform stratified analysis or a comprehensive investigation into individual-level data. Third, given that our study participants predominantly had European ancestry, caution is warranted in extrapolating the research findings to more ethnically diverse populations, such as Asians. Subsequent studies are crucial for validating the relevance of our research findings across diverse ethnic groups. Fourth, while uncovering causal relationships between numerous plasma proteins and metabolic-related diseases, our understanding of their underlying mechanisms remains incomplete, underscoring the need for additional basic research to elucidate these intricate pathways. Finally, we did not include all metabolism-related diseases, such as hypertension and metabolic-associated fatty liver disease, in our analyses because similar studies using the same methods and datasets have already been reported, or relevant data were unavailable. Nevertheless, by investigating five metabolism-related diseases as outcome events, our study helps to address existing research gaps to the greatest extent possible.
Conclusion
In conclusion, this comprehensive proteome-wide MR study identified numerous plasma proteins strongly linked to metabolic-related diseases, thus identifying promising targets for understanding their pathogenesis, diagnostics, and therapeutic interventions. Further fundamental experiments are essential to evaluate the efficacy of these candidate targets.
Supplemental Material
sj-docx-1-tae-10.1177_20420188251343140 – Supplemental material for Novel therapeutic targets for metabolism-related diseases: proteomic Mendelian randomization and colocalization analyses
Supplemental material, sj-docx-1-tae-10.1177_20420188251343140 for Novel therapeutic targets for metabolism-related diseases: proteomic Mendelian randomization and colocalization analyses by Yue-Yang Zhang, Bin-Lu Wang, Bing-Xue Chen and Qin Wan in Therapeutic Advances in Endocrinology and Metabolism
Supplemental Material
sj-xlsx-2-tae-10.1177_20420188251343140 – Supplemental material for Novel therapeutic targets for metabolism-related diseases: proteomic Mendelian randomization and colocalization analyses
Supplemental material, sj-xlsx-2-tae-10.1177_20420188251343140 for Novel therapeutic targets for metabolism-related diseases: proteomic Mendelian randomization and colocalization analyses by Yue-Yang Zhang, Bin-Lu Wang, Bing-Xue Chen and Qin Wan in Therapeutic Advances in Endocrinology and Metabolism
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
