Abstract
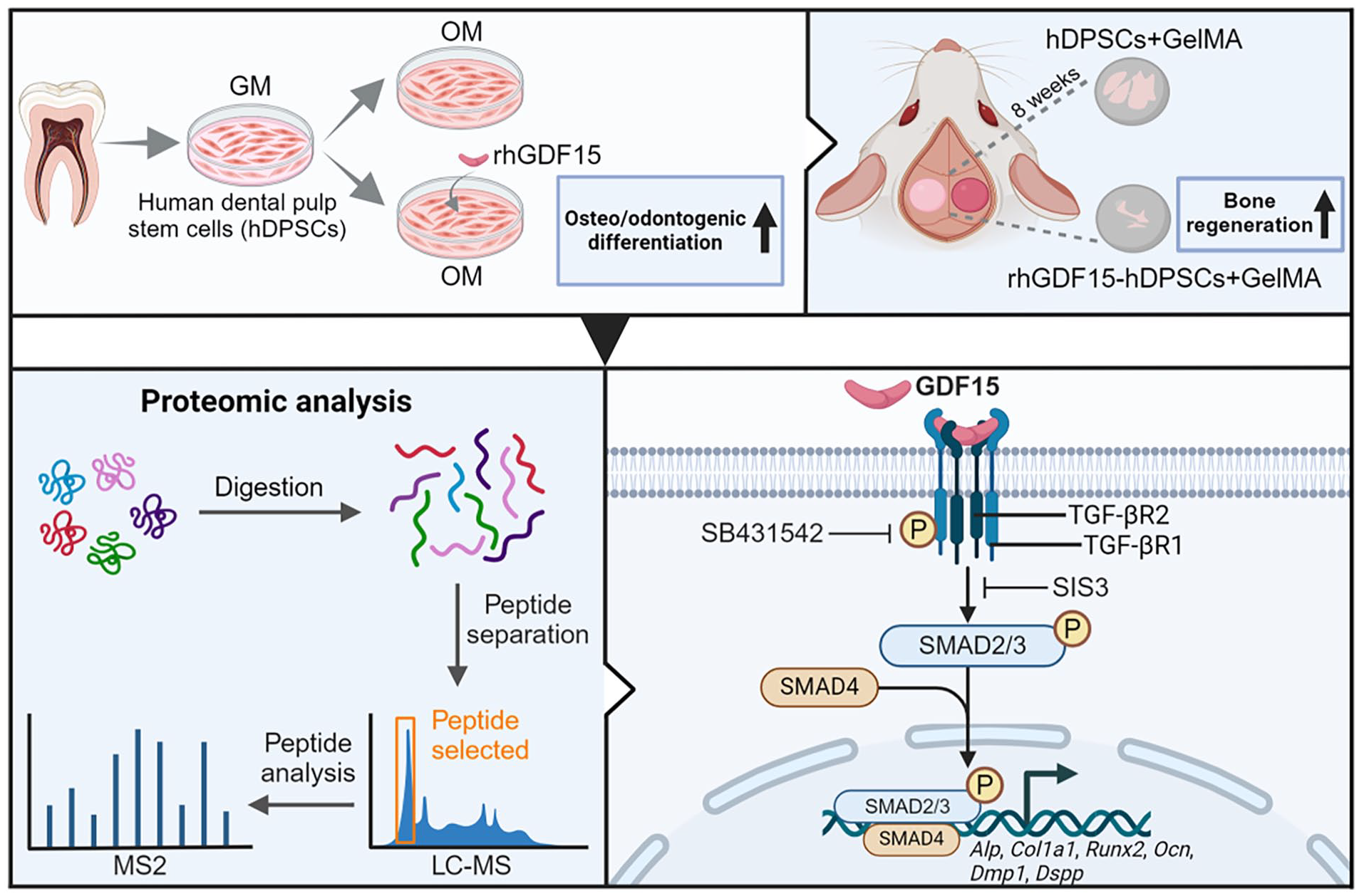
Mesenchymal stem cell-mediated bone tissue engineering strategies, including human dental pulp stem cells (hDPSCs), represent an effective therapeutic approach for bone defect repair, particularly in maxillofacial bone defects. Growth differentiation factor 15 (GDF15), a multifunctional cytokine, plays a critical role in bone tissue formation and remodeling. This study aims to investigate the effects of GDF15 on the osteogenic differentiation of hDPSCs and elucidate the underlying molecular mechanisms. Our findings demonstrate that GDF15 expression and secretion are upregulated during the osteogenic differentiation of hDPSCs. Both
Keywords
Background
Human dental pulp stem cells (hDPSCs), characterized by their multidirectional differentiation potential, low immunogenicity, easy accessibility, absence of ethical controversies, and high proliferative capacity, represent an ideal stem cell source for cell therapy, regenerative medicine, and tissue engineering applications.1,2 As evidenced by previous studies, hDPSCs exhibit robust osteogenic differentiation potential under osteogenic microenvironment stimulation in both
The osteogenic differentiation of hDPSCs is orchestrated through sequential regulation and synergistic interplay of a cascade of growth factors and cytokines.7,8 Therefore, the identification and characterization of critical growth factors regulating the osteogenic differentiation of hDPSCs will facilitate advancements in bone tissue engineering and its clinical translation. Researchers have identified that multiple members of the transforming growth factor-β (TGF-β) superfamily play a facilitating role during the osteogenic differentiation of hDPSCs.9–11 Growth differentiation factor 15 (GDF15), a member of the TGF-β superfamily, is critically involved in prenatal development, postnatal growth, and the maintenance and remodeling of diverse tissues and organs.
12
Recently, studies have identified dysregulated expression of GDF15 in multiple bone-related pathologies, including multiple myeloma, primary myelofibrosis, and neoplastic bone metastasis.13–15 Current literature reveals conflicting findings regarding the role of GDF15 in bone-related processes. While some studies demonstrate its pro-osteogenic effects in bone development, bone metabolic homeostasis, and bone pathological remodeling,16–18 others report inhibitory or context-dependent outcomes.
19
This study was designed to evaluate the effects of GDF15 on the osteogenic differentiation of hDPSCs
Materials and methods
Isolation and cultivation of hDPSCs
Intact third molars were removed from donors aged 17–22 year. The experiments were approved by the Ethics Committee, performed in compliance with the Declaration of Helsinki, and written informed consent was obtained from all participants. The procedure for isolating hDPSCs aligns with our prior research. 27 Briefly, the pulp was removed from the teeth and diced into tiny fragments (1 mm3). Then, it was digested with 3 mg/mL collagenase type I solution (Sigma-Aldrich, St. Louis, MO, USA) at 37°C for 30 min. Next, the digested pulp sediments were spread evenly in T-25 culture flasks (Jet Bio, Guangzhou, China). After 8–10 h of incubation in the cell culture incubator, growth medium (GM) was carefully introduced into the culture flasks to promote cell growth. GM was composed of alpha-minimum essential medium (Hyclone, South Logan, UT, USA) containing 10% Fetal Bovine Serum (FBS; ExCell Bio, Shanghai, China) and 1% penicillin-streptomycin 100× solution (Hyclone, South Logan, UT, USA). Replace the medium every 3 days. The hDPSCs at passages 3–5 were used in the subsequent experiments.
Colony-forming assay
In culture dishes (Jet Bio, Guangzhou, China) with a 6 cm diameter, 500 cells/well of hDPSCs were plated. The hDPSCs were cultivated in GM for 10 days before being fixed for 15 min with 4% paraformaldehyde (PFA; Beyotime, Shanghai, China), and then stained for 10 min with crystal violet staining solution (Beyotime, Shanghai, China). Image acquisition was conducted using an Epson Perfection scanner (version V330; Epson Co., Ltd., Shanghai, China) and an inverted phase-contrast microscope (Leica, Wetzlar, Germany). Moreover, the rate of colony formation was measured by ImageJ software (Media Cybernetics, Silver Springs, MD, USA).
Cell counting kit-8 (CCK-8) assay
hDPSCs were implanted in 96-well plates (Jet Bio, Guangzhou, China) at 2 × 103 cells/well and incubated for 12 h to allow adhesion. This time point was day 0. The proliferation rates were measured on days 0, 1, 3, 5, and 7. At the specified time intervals, hDPSCs were treated with 10% CCK-8 reagent (Dojindo, Kumamoto, Kyushu, Japan) at 37℃ for 2 h. A multipurpose microplate luminescence detector (PerkinElmer, Inc., Waltham, MA, USA) was accustomed to measure the reaction solution’s optical density (OD) at 450 nm. In addition, hDPSCs were treated for 0, 1, 3, 5, and 7 days with varying doses of rhGDF15 (PeproTech; Rocky Hill, NJ, USA; 0–100 ng/mL) to assess the impact of GDF15 on the proliferative ability of hDPSCs.
Crystal violet assay
The hDPSCs were adhere-wall cultivated for 12 h after being introduced into 12-well plates (Jet Bio, Guangzhou, China) at a count of 1 × 104 cells/well, then cultured in GM for 0, 1, 3, 5, and 7 days, respectively. At the designated intervals, a 4% PFA was applied to fix the cells, after that, they were dyed with a crystal violet staining solution for 10 min. The images were recorded in the same manner as for the colony-forming assay. The stained cells were lysed with 33% acetic acid (Chongqing Chuandong Chemical (Group) Co., Ltd, Chongqing, China) and the OD of the lysate was detected at 570 nm using the multifunctional microplate luminescence detector.
Flow cytometry
For surface marker identification of hDPSCs, trypsin digestion was used to harvest the cells, and then incubated for 30 min, shielded from light, with CD105-FITC, CD90-FITC, CD73-PE, CD45-FITC, CD19-FITC, and CD14-FITC (BD Pharmingen, San Diego, CA, USA). PE or FITC served as the isotypic controls. Flow cytometry analysis was conducted utilizing BD flow cytometry (BD Biosciences, San Jose, CA, USA).
Osteogenic differentiation of hDPSCs
hDPSCs were seeded into culture plates at a density of 1 × 10⁵ cells/mL. Upon reaching 80% confluence, GM was replaced with osteogenic induction medium (OM), where OM was made by adding 5 mM β-glycerophosphate (Sigma-Aldrich, St. Louis, MO, USA), 50 μg/mL ascorbic acid (Solarbio, Beijing, China) and 100 nM dexamethasone (Solarbio, Beijing, China) to GM. Following
Alkaline phosphatase (ALP) staining
hDPSCs were fixed with 4% PFA for 15 min. According to the manufacturer’s instructions, after rinsing with distilled water to remove residual fixative, the hDPSCs were stained using the BCIP/NBT Alkaline Phosphatase Color Development Kit (Beyotime, Shanghai, China) for 10 min. The images were recorded in the same manner as for the colony-forming assay.
Alizarin red S (ARS) staining
After 15 min of 4% PFA fixation, hDPSCs were stained for 10 min with 1% ARS (Sigma-Aldrich, St. Louis, MO, USA). The cells were rinsed repeatedly with distilled water and air-dried before acquiring images. Additionally, the stained cells were lysed with cetylpyridinium chloride monohydrate (Solarbio, Beijing, China), and the OD of the lysate was determined at 562 nm.
Quantitative real-time PCR (qPCR)
Total RNA was isolated utilizing RNAiso Plus (TaKaRa, Tokyo, Japan), and complementary DNA was synthesized employing the PrimeScript™ RT kit (TaKaRa, Tokyo, Japan). qPCR was executed in compliance with the experimental protocols described in the handbook using the QuantiNova SYBR Green PCR Kit (QIAGEN, Duesseldorf, Germany) and the fluorescence quantitative PCR instrument system (Bio-Rad, Hercules, CA, USA).
Sense and antisense primers for qPCR.

Western blotting (WB)
hDPSCs were lysed using a cold RIPA lysis solution (Beyotime, Shanghai, China) with a mixture of protease and phosphatase inhibitors (Beyotime, Shanghai, China). Cells were further lysed using an ultrasonic cell crusher (SCIENTZ, Zhejiang, China) at low temperature. Protein samples were then standardized using a BCA Protein Assay Kit (Beyotime, Shanghai, China). For the extraction of cytoplasmic and nuclear proteins, nuclear and cytoplasmic extraction reagents (Thermo Fisher Scientific, Waltham, MA, USA) were employed to separate them following the manufacturer’s instructions. For plasma membrane protein extraction, the Plasma Membrane Protein Extraction Kit (Proteintech, Wuhan, China) was employed according to the manufacturer’s instructions. After being separated utilizing TGX Stain-Free™ FastCast™ Acrylamide Solutions (Bio-Rad, Hercules, CA, USA), the protein samples were subsequently transferred on polyvinylidene fluoride membranes (Merck Millipore, Billerica, MA, USA). The membranes were blocked for 1 h at room temperature by placing them in 5% skim milk (BioSharp, Hefei, China) or 5% bovine serum albumin (BSA; BioFroxx, Einhausen, Germany) solution. Next, the membranes were incubated in primary antibody solution overnight at 4°C and in HRP-conjugated secondary antibody (Bio-Rad, Hercules, CA, USA) for 2 h. Finally, membranes were visualized with a WB analytical imaging system (Bio-Rad, Hercules, CA, USA). Table 2 lists the primary antibody concentrations that were employed.
Antibody source and application concentration for WB analysis.

Co-immunoprecipitation (Co-IP)
Following stimulation of hDPSCs with rhGDF15 (20 ng/mL) for 30 min, membrane proteins were extracted. The GDF15 antibody (Santa Cruz Biotechnology, Dallas, USA; sc-377195) was conjugated to protein A/G magnetic beads (MedChemExpress, New Jersey, USA) using a rotary mixer (Bio-Rad, Hercules, CA, USA) under sequential conditions: 30 min at room temperature and 2 h at 4°C. Subsequently, the extracted membrane proteins were incubated with the antibody-bead complexes in the rotary mixer under identical conditions (30 min at room temperature and 2 h at 4°C). After thorough washing, the protein samples were eluted by heat-denaturation (95°C for 5 min) and subjected to Sodium Dodecyl Sulfate Polyacrylamide Gel Electrophoresis analysis.
Enzyme-linked immunosorbent assay (ELISA)
Upon reaching 80% confluence, hDPSCs were designated as day 0 cultures, at which point the GM was replaced with OM for subsequent cultivation. The supernatants from the cell cultures were obtained on days 0, 1, 4, 9, 14, and 21 of OM culture (the old medium was substituted with a new one the day before sample collection, and the medium was collected when the cells were incubated for 24 h) and stored at −80℃. The samples were assayed following the instruction manual of the human GDF-15 ELISA Kit (Abclonal, Wuhan, China), and the GDF15 concentration of the corresponding samples was calculated.
Gdf15 overexpression by lentiviral transfection
Lentiviruses expressing
Gdf15 knockdown by siRNAs
The small interfering RNAs (siRNAs) targeting
Sense and antisense sequences for

Animals
Nude (BALB/cA-nu) mice and Sprague-Dawley rats were all obtained by Chongqing Tengxin Biotechnology Co., Ltd. (Chongqing, China). All animal experiments were approved by the Ethics Committee and conducted according to the ARRIVE guidelines.
Subcutaneous transplantation in nude mice
After treating hDPSCs with rhGDF15 (20 ng/mL) for 5 days, 4 × 106 hDPSCs were mixed with 30 mg of hydroxyapatite/β-tricalcium phosphate (HA/β-TCP) granular scaffolds (Sichuan Baimeng Bioactive Materials Co., Ltd.; Sichuan, China), and immediately implanted into the dorsal subcutis of 10-week-old male nude mice. After 8 weeks, the implants were obtained, fixed overnight in 4% PFA, decalcified in 10% Ethylene Diamine Tetraacetic Acid, then paraffin-embedded and sectioned.
Healing of calvarial defects in rats
In 8-week-old rats, circular bone fragments with a diameter of 5 mm were removed from both sides of the calvarial apex of the rat skull using a surgical ring drill at a speed of 1000 rpm. Saline cooling was maintained throughout the procedure to prevent thermal damage. When hDPSCs were treated with GM for 5 days with or without adding 20 ng/mL rhGDF15, the cells were used in
Micro-CT analysis
Samples were imaged and analyzed using a viva CT 40 micro-CT scanner (SCANCO Medical AG, Wangen-Bruttisellen, Switzerland). The scanning resolution for the rat calvaria was 17.5 μm. The scanned data were reconstructed in three dimensions using the viva CT 40 correspondence analysis software and bone volume (BV), BV/ total volume (TV) (BV/TV), trabecular number (Tb.N), and trabecular separation (Tb.Sp) were calculated for each sample.
Hematoxylin & eosin (H&E) staining
Staining was implemented utilizing the H&E staining kit (Solarbio, Beijing, China): hematoxylin for 3 min, differentiation in hydrochloric acid alcohol for 30 s, and eosin for 10 min. The sections were scanned using a Research Grade Whole Slide Scanning System SLIDEVIEW VS200 (Olympus, Tokyo, Japan). To quantify the new bone generated within the grafts, the percentage of the bone formation area was computed using ImageJ software for five images from different regions of the implant.
Masson staining
In accordance with the manufacturer’s instructions, sections were stained employing a Masson staining kit (Solarbio, Beijing, China). The sections were analyzed in a manner consistent with H&E staining.
Immunohistochemical staining
Paraffin sections were dewaxed and then hydrated, heat repair was used to expose the antigenic epitopes, and the primary antibody was incubated with the samples at 4°C for a whole night after being blocked for 10 min with an endogenous peroxidase blocker (ZSGB-BIO, Beijing, China) and 1 h with 10% goat serum (ZSGB-BIO, Beijing, China) at room temperature. Next, the samples were subjected to incubation by an enzyme-labeled sheep anti-mouse IgG polymer (ZSGB-BIO, Beijing, China) for 1 h. Subsequently, freshly prepared DAB color solution (ZSGB-BIO, Beijing, China) was dropped into the samples and allowed to incubate for 1–5 min. The nuclei were then stained for 30 s using a hematoxylin staining solution. Images were acquired using a Research Grade Whole-Slide Scanning System SLIDEVIEW VS200. The concentrations of primary antibodies used are listed in Table 4.
Antibody source and application concentration for immunohistochemical staining.

Immunofluorescence staining
The hDPSCs were fixed with 4% PFA for 15 min, incubated with 0.3% Triton X-100 (Beyotime, Shanghai, China) for 10 min, and blocked with 3% BSA for 1 h. Primary antibodies are incubated with cells for a whole night at 4℃. Subsequently, the cells were incubated with a fluorescent-labeled secondary antibody (Thermo Fisher Scientific, Waltham, MA, USA) at room temperature for 1 h. Finally, nuclear staining was performed with DAPI (Beyotime, Shanghai, China) for 10 min.
For the new bone tissue in calvarial defects in rats, after the tissue was made into paraffin sections, the samples on the sections were deparaffinized, rehydrated, and repaired at high temperatures. The samples were permeabilized with 0.3% Triton X-100 for 10 min and blocked with 10% goat serum (ZSGB-BIO, Beijing, China) for 1 h. The primary antibody was incubated at 4℃ overnight, after which the secondary antibody was incubated for 1 h at room temperature. Subsequently, nuclear staining was performed with DAPI. Images were observed and recorded under the fluorescent microscope Zeiss Axio Observer 7 (Zeiss, Aalen, Germany). Fluorescence intensity was evaluated quantitatively utilizing ImageJ tools.
For immune co-localization analysis of the subcutaneous implants in nude mice, multiplex fluorescence immunohistochemical staining was performed using the Multiplex Immunohistochemistry Kit (Panovue, Beijing, China) following the manufacturer’s protocol. Briefly, tissue sections were deparaffinized and rehydrated, followed by heat-mediated antigen retrieval. Sequential incubations with primary and secondary antibodies were conducted using reagents provided in the kit. Antibody stripping was subsequently performed using the included elution buffer to remove bound antibodies, allowing sequential re-incubation with additional primary and secondary antibodies for multiplex staining. Table 5 lists the main antibody concentrations that were employed.
Antibody source and application concentration for immunofluorescence staining.

Proteomic analysis
Liquid chromatography-tandem mass spectrometry (LC-MS/MS; Shanghai Zhongke New Life Biotechnology Co., Ltd, Shanghai, China) was utilized for proteomic analysis of hDPSCs treated with OM or OM containing 20 ng/mL rhGDF15 for 6 h. Sample lysis of hDPSCs was performed using SDT buffer (4% SDS, 100 mM Tris-HCl, pH 7.6). The protein samples were prepared using filter-assisted sample preparation. Briefly, the samples were mixed in 10 mM dithiothreitol (DTT) for 1.5 h; after being exposed to 20 mM iodoacetamide, the samples were dark-stored for 30 min. Ultimately, the peptides that were produced were gathered as filtrates after trypsin was introduced to the samples and digested overnight (15–18 h) at 37°C. The filtrate was desalted on C18 Cartridges (Empore™ SPE Cartridg C18). LC-MS/MS analysis was conducted using a Q mass spectrometer (Thermo Fisher Scientific). Utilizing MaxQuant software (version 1.6.14), the MS raw data for every sample was combined and retrieved for identification and quantitative analysis. Hierarchical cluster analysis was conducted using Cluster (version 3.0) and Java Treeview software. Motifs were analyzed using MeMe, and protein subcellular localization was predicted using CELLO, a multi-class SVM classification system. The sequences of the differentially expressed proteins were analyzed utilizing InterProScan and the NCBI BLAST+ client software to discover homologous sequences. The Blast2GO program was utilized to map Gene Ontology (GO) terms and annotate sequences. The proteins under investigation were also compared against the online Kyoto Encyclopedia of Gene Genomes (KEGG) database for pathway analysis.
Molecular docking analysis
The proteins used for docking were GDF15 (UniProt ID: Q99988) and TGF-βR2 (UniProt ID: P37173), with structural models retrieved from the AlphaFold Protein Structure Database. Protein-protein molecular docking was performed using the HDOCK server. 28 The protein-protein binding sites are shown in Supplemental Table S1. Binding free energy calculations and residue-level energy decomposition were conducted via the MM/GBSA module of the HawkDOCK server, enabling ranking of amino acid residues based on their energetic contributions (Supplemental Table S2). 29 The docking score was used as the evaluation criterion for molecular docking. Among the 10 output conformations, the optimal docking model was selected, and its binding energy was calculated. Finally, the top-ranked binding poses exhibiting favorable interaction energies were visualized using PyMOL 2.4 software.
Statistical analysis
All experiments were performed with a minimum of three independent replicates. The data are presented as mean ± standard deviation. To do statistical studies, GraphPad Prism 9 (GraphPad, San Diego, CA) was utilized. One-way ANOVA or the Student’s
Results
The secretion of GDF15 was upregulated during the osteogenic differentiation of hDPSCs
In the extracted hDPSCs, it was observed that the cells had a spindle-shaped morphology (Figure 1(a)), strong clone formation (Figure 1(b)), proliferation (Figure 1(c) and (d)) and osteogenic differentiation (Figure 1(e), (f), (h), and (i)), and were positive for the cell surface markers CD73, CD90, and CD105 (Figure 1(g)). During induction of differentiation, mRNA expression of

GDF15 secretion was significantly upregulated during the osteogenic differentiation of hDPSCs. (a) Morphology of primary hDPSCs. Scale bars (white): 100 μm. Morphology of passage 3 hDPSCs at 24 h of culture. Scale bars (black): 250 μm. P0 = Passage 0; P3 = Passage three. (b) Colony-forming assay to assess the self-renewal ability of hDPSCs. Scale bars: 250 μm. (c) The CCK-8 assay was employed to assess the proliferation of hDPSCs. (d) Proliferation of hDPSCs detected by crystal violet staining. Scale bars: 250 μm. (e) ALP staining of hDPSCs grown in OM for a week. Scale bars: 250 μm. (f) ARS staining of 21-day-cultured hDPSCs in OM and quantitative analysis of mineralized nodule deposition. Scale bars: 250 μm. (g) Flow cytometry demonstrated that the hDPSCs highly expressed CD105, CD90, and CD73; and lowly expressed CD45, CD19, and CD14. (h) During induction, intracellular mRNA levels of
Gdf15 overexpression promoted osteogenic differentiation of hDPSCs in vitro

G
Gdf15 knockdown inhibited osteogenic differentiation of hDPSCs in vitro
Three si
RhGDF15 promoted the osteogenic differentiation of hDPSCs
For proliferation, Ki67 immunofluorescence staining exhibited that GDF15 significantly promoted hDPSC proliferation. From day 3 to 7, the cell proliferation rate of rhGDF15 treatment groups were faster than that of control group, and the 20 ng/mL treatment group was the fastest (Supplemental Figure S1A, B). Moreover, the results of the CCK-8 test are consistent with this (Supplemental Figure S1C). These results suggested that rhGDF15 can promote hDPSC proliferation
The effect of rhGDF15 on osteogenic differentiation of hDPSCs was investigated
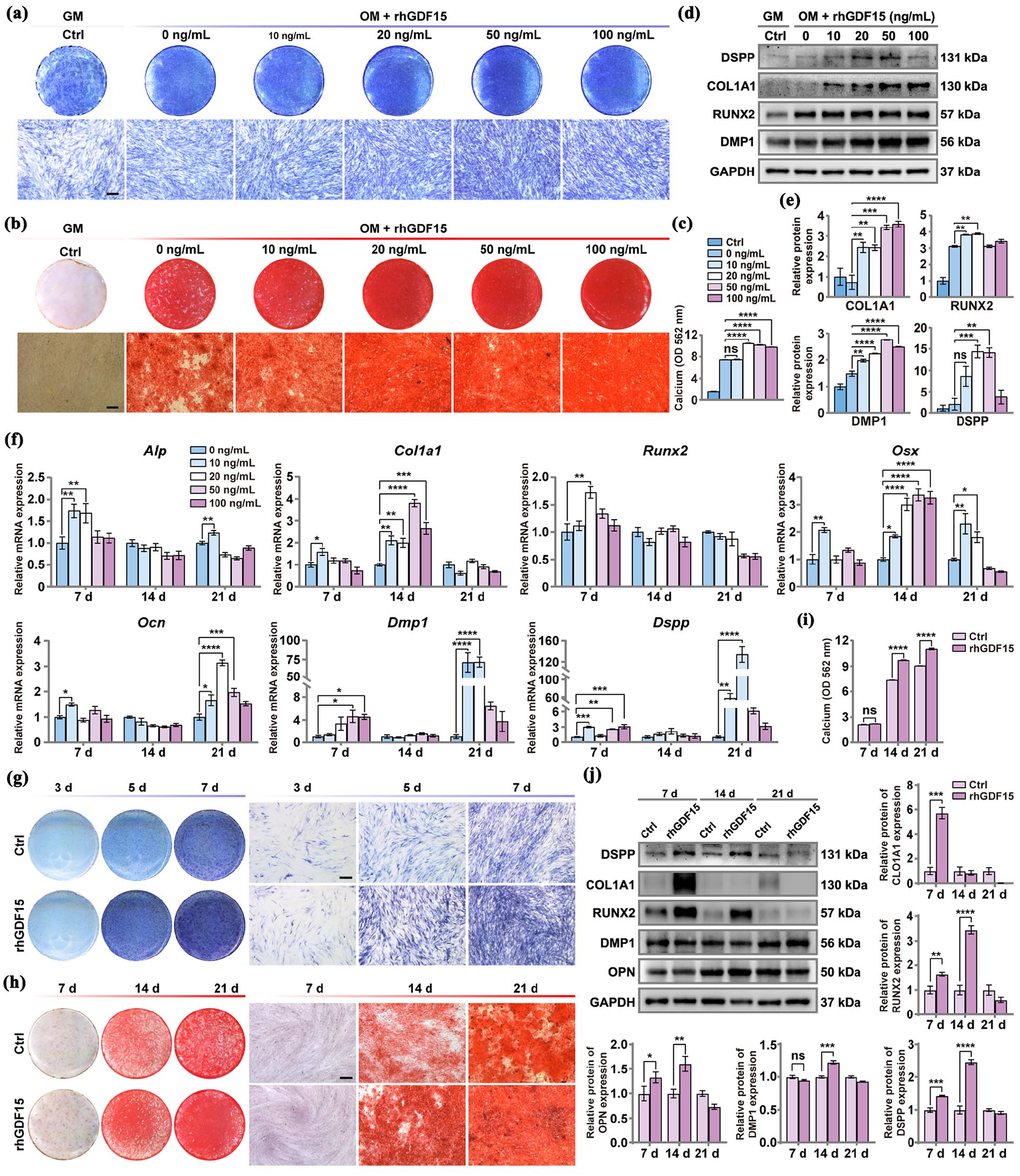
rhGDF15 stimulated the osteogenic differentiation of hDPSCs
GDF15 promoted osteogenesis in vivo
To evaluate the effects of GDF15 on bone formation
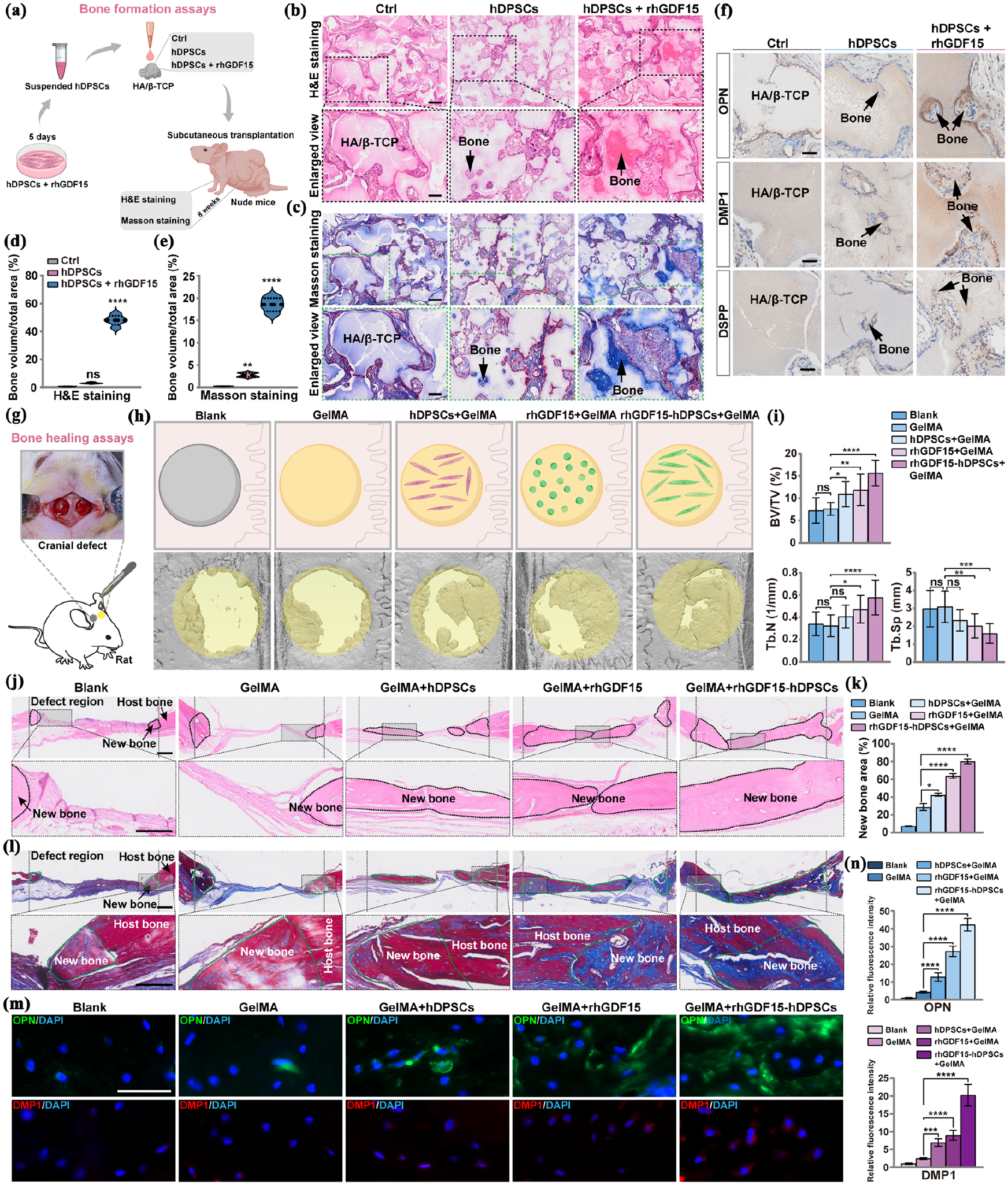
rhGDF15 promoted bone formation and bone repair
RhGDF15 activated TGF-β/SMAD cellular signaling of hDPSCs
Studies have demonstrated that glial cell-derived neurotrophic factor family receptor alpha-like protein (GFRAL) identified in the brain is a receptor for GDF15.30,31 However, in the present study,

Proteomic analysis identified significant enrichment of the TGF-β/SMAD signaling pathway. (a) Proteins that are differently expressed in the OM and OM + rhGDF15 groups are displayed in a heatmap. (b) Volcano map. Blue means comparatively little expression, whereas red exhibits comparatively strong expression. (c) Venn Diagram Analysis. (d) Subcellular localization analysis. (e) Enriched GO terms ranked top in molecular functions, cellular components, and biological processes. (f) Enrichment KEGG analysis of the top 20 in the bubble chart. The
Ligands attach to target cell membrane receptors in the TGF-β/SMAD signaling pathway to initiate downstream signaling.
32
Through qPCR analysis, we detected that

rhGDF15 activated TGF-β/SMAD signaling pathway in hDPSCs. (a and b)
Next, we detected the expression of critical proteins of the pathway in rhGDF15-stimulated hDPSCs by WB. The results exhibited that GDF15 stimulation increased the expression of TGF-βR2, p-SMAD2/SMAD2, and p-SMAD3/SMAD3 in hDPSCs, particularly when hDPSCs were exposed to 20 ng/mL of rhGDF15 for a duration of 30 min (Figure 6(f) and (g)). Additionally, a significant nuclear translocation of p-SMAD2 and p-SMAD3 was observed in rhGDF15-treated hDPSCs (Figure 6(h)). In the nude mouse subcutaneous transplantation model, immunofluorescence co-localization analysis detected significantly elevated expression of TGF-βR2, p-SMAD2, and p-SMAD3 within the bone formation regions of rhGDF15-treated implants compared to those with untreated hDPSCs (Figure 6(i)). Previous research has revealed that GDF15 also belongs to the BMP subfamily. 18 Accordingly, we investigated the BMP/SMAD signaling pathway and found that the phosphorylation ratio of p-SMAD1/5/9/SMAD1/5/9 exhibited no significant change in expression after rhGDF15 treatment of hDPSCs (Supplemental Figure S3A, B). These findings further confirmed that GDF15 promoted osteogenic lineage differentiation of hDPSCs via the TGF-β/SMAD signaling pathway, rather than through the BMP/SMAD cascade.
Increased osteogenic differentiation of hDPSCs due to GDF15 stimulation was partially reversed by inhibitors of the TGF-β/SMAD signaling
To investigate whether the TGF-β/SMAD signaling pathway plays a regulatory role in GDF15-mediated osteogenic differentiation of hDPSCs, we utilized specific TGF-β/SMAD pathway inhibitors, SB431542 and SIS3. The inhibitor concentrations (SB431542 (10 μM) and SIS3 (3 μM)) were selected based on WB (Figure 7(a)). After pretreatment of hDPSCs with SB431542 or SIS3 for 1 h, rhGDF15 treatment significantly reduced the expression of p-SMAD2/SMAD2 and p-SMAD3/SMAD3 (Figure 7(b) and (c)). Moreover, the results of immunofluorescence staining analysis were consistent with WB analysis (Figure 7(d)). hDPSCs were treated with rhGDF15 to induce osteogenic differentiation, in combination with the TGF-β/SMAD pathway inhibitors SB431542 or SIS3. After 7 days, the upregulation of mRNA levels of osteogenic-specific genes (

Activation of the TGF-β/SMAD signaling pathway by GDF15 is partially reversed by the inhibitors. (a) The concentration of the inhibitor was selected based on the analysis of the protein level ratios of phosphorylated to total SMAD2 or SMAD3. (b and c) The protein expression levels and quantitative analysis of p-SMAD2/SMAD2 and p-SMAD3/SMAD3 were assessed in hDPSCs pretreated with inhibitor for 1 h followed by rhGDF15 stimulation for 30 min. (d) Representative images and quantitative analysis of p-SMAD2 or p-SMAD3 immunofluorescence staining. Scale bars: 100 μm. (e and f) The osteogenic effects of inhibitor-treated hDPSCs were analyzed by qPCR. (g) ALP staining following 7 days of treatment. Scale bars: 250 μm. (h) ARS staining following 21 days of treatment and relative quantitative analysis. Scale bars: 250 μm. (i and j) The osteogenic effects of hDPSCs treated with inhibitors for 7 days were evaluated by WB, and the protein bands were analyzed in gray. Mean ± SD was employed to express all data (*

rhGDF15 promoted bone formation through TGF-β/SMAD signaling
Discussion
Studies have demonstrated the significant potential of hDPSCs in bone regeneration.2,33 Under appropriate growth factors and microenvironmental conditions, hDPSCs undergo osteogenic differentiation, forming mineralized nodules
Our study revealed that rhGDF15 enhances the proliferation of hDPSCs. Previous studies have identified the pro-proliferative effects of GDF15 in diverse cell types, including cardiac fibroblasts, 36 senescent chondrocytes, 37 bone marrow stromal cells (BMSCs), 14 and myeloma cells. 13 Our findings confirm that hDPSCs are responsive to GDF15-mediated proliferative regulation, expanding its therapeutic potential in regenerative medicine applications. As reported in the literatures, GDF15 was secreted by cells such as macrophages, 38 BMSCs 39 and periodontal ligament cells.20,40 In this study, we observed that GDF15 is synthesized intracellularly and subsequently secreted extracellularly during the osteogenic differentiation of hDPSCs. Based on these results, we hypothesize that GDF15 may not only enhance hDPSCs proliferation but also play an active role in their osteogenic differentiation through autocrine/paracrine signaling mechanisms.
Next, we investigated the osteogenic differentiation potential of GDF15 in hDPSCs
To comprehensively evaluate the osteogenic potential of GDF15, both ectopic and in situ bone regeneration models were employed. For the subcutaneous transplantation model in nude mice, the HA/β-TCP scaffold was chosen based on its well-documented osteoconductive properties, structural stability, and ability to mimic the mineralized extracellular matrix of bone.44,45 The immunodeficient nature of nude mice further eliminates confounding immune responses, ensuring reliable assessment of hDPSC-mediated osteogenesis in isolation.
46
In our nude mouse subcutaneous transplantation model, the constructs loaded with rhGDF15-activated hDPSCs exhibited robust osteogenic capacity, corroborating previous reports on the rhGDF15-driven osteoblastic lineage commitment of BMSCs
GDF15 binds to target cell surface receptors, and then initiates downstream signaling cascade responses.
49
Researchers have established that GFRAL as the sole known endogenous receptor that exhibits a strong affinity for GDF15.50,51 GDF15 regulated energy metabolism and body weight primarily through appetite suppression and its physiological pathways dependent on interactions with GFRAL.
30
However, the researchers only found GFRAL expression in the hindbrain and prostate.15,52 GDF15 may bind to receptors other than GFRAL.
25
Our research found that
While this study provides mechanistic insights into GDF15-driven osteogenic differentiation of hDPSCs, certain limitations warrant consideration. As a stress-responsive cytokine, GDF15 exhibits low baseline expression in human tissues under physiological conditions but is markedly upregulated in diverse pathological contexts, including hypoxia, inflammation, tissue injury, and cancer.17,58 Our experimental models focused exclusively on GDF15′s osteoinductive effects in a controlled
Conclusion
In summary, our findings demonstrate that GDF15 enhances bone regeneration by promoting the osteogenic differentiation of hDPSCs, a process mediated at least partially through activation of the TGF-β/SMAD signaling pathway. These mechanistic insights position GDF15 as a promising candidate growth factor in regenerative medicine, offering a novel therapeutic strategy for bone tissue engineering that leverages the intrinsic differentiation potential of hDPSCs.
Supplemental Material
sj-docx-1-tej-10.1177_20417314251357752 – Supplemental material for GDF15 promotes osteogenic differentiation of human dental pulp stem cells by activating the TGF-β/SMAD signaling pathway
Supplemental material, sj-docx-1-tej-10.1177_20417314251357752 for GDF15 promotes osteogenic differentiation of human dental pulp stem cells by activating the TGF-β/SMAD signaling pathway by Pingmeng Deng, Bin Yang, Chuling Huang, Yuejia Li, Ziyi Mei, Yong Li and Jie Li in Journal of Tissue Engineering
Footnotes
Ethical considerations
All human teeth sample collection experiments and animal experiments were approved by the Research Ethics Committee of the Affiliated Hospital of Stomatology, Chongqing Medical University (CQHS-REC-2021 (LSNo. 040) and CQHS-REC-2023 (LSNo.087)). The study was performed in accordance with the Declaration of Helsinki and the ARRIVE guidelines.
Consent to participate
Written consent was secured from all participants.
Author contributions
PD designed the study, performed most of the experiments, analyzed the data and wrote the manuscript. BY assisted in the rat calvarial defect experiments and data analysis. CH performed proteomics sample preparation and data analysis. YL and ZM participated in the cell experiments and related data analysis. YL supervised the experimental methodology and provided intellectual contributions to the experimental design and analysis. JL conceived the project, supervised the experiments, performed the manuscript review and obtained the funding. All authors read and approved the final manuscript.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by National Natural Science Foundation of China (Grant No. 82071072), Natural Science Foundation of Chongqing (Grant No. CSTB2022NSCQ-LZX0004), and CQMU Program for Youth Innovation in Future Medicine (Grant No. W0179).
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability statement
All data are available from the corresponding authors upon reasonable request.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
