Abstract
Objective
Our aim is to explore the candidate pathogenesis genes and pathways of developmental dysplasia of hip (DDH).
Design
Proliferating primary chondrocytes from hip cartilage were used for total RNA extraction including 5 DDH patients and 5 neck of femur fracture (NOF) subjects. Genome-wide mRNA and microRNA (miRNA) were then sequenced on the Illumina platform (HiSeq2500). Limma package was used for difference analysis of mRNA expression profiles. edgeR was used for difference analysis of miRNA expression profiles. miRanda was used to predict miRNA-target genes. The overlapped DDH associated genes identified by mRNA and miRNA integrative analysis were further compared with the differently expressed genes in hip osteoarthritis (OA) cartilage.
Results
Differential expression analysis identified 1,833 differently expressed mRNA and 186 differently expressed miRNA for DDH. Integrative analysis of mRNA and miRNA expression profiles identified 175 overlapped candidate genes (differentially expressed genes, DEGs) for DDH, such as VWA1, TMEM119, and SCUBE3. Further gene ontology enrichment analysis detected 111 candidate terms for DDH, such as skeletal system morphogenesis (P = 4.92 × 10−5) and skeletal system development (P = 8.85 × 10−5). Pathway enrichment analysis identified 14 candidate pathways for DDH, such as Hedgehog signaling pathway (P = 4.29 × 10−5) and Wnt signaling pathway (P = 4.42 × 10−2). Among the identified DDH associated candidate genes, we also found some genes were detected in hip OA including EFNA1 and VWA1.
Conclusions
We identified multiple novel candidate genes and pathways for DDH, providing novel clues for understanding the molecular mechanism of DDH.
Keywords
Introduction
Developmental dysplasia of hip (DDH) is the most common congenital abnormality of lower limbs, including hypoplasia of shallow acetabulum to subluxation and frank dislocation. 1 Previous studies have demonstrated that the incidence of DDH ranges from 1% to 13%, depending on demographic characteristics, risk factors, and diagnostic methods. 2 Hip osteoarthritis (OA) is caused by degeneration of articular cartilage and the underlying bone and can be divided into 2 types: primary (associated with advancing age) and secondary (mainly subsequent to avascular necrosis, developmental dysplasia). 3 DDH greatly increases the risk of severe hip OA, which leads to significant functional limitations in middle-aged to older individuals. 4 However, the association between the 2 diseases is not clear. Both conditions are known to have a strong genetic component. 5
To date, a large number of studies have been conducted for DDH, but most are focused on the clinical arthroplasty.6,7 Given the unpredictable changes in molecular mechanism, this field should be drawn more attention. Currently, some molecular studies identified multiple genetic loci associated with DDH. For instance, Baghdadi et al. 8 suggested that the abnormal methylation of GDF5 contributed to the development of DDH. A significant association between the D repeat polymorphism of ASPN and DDH was also reported. 9 However, the molecular mechanism of DDH remains largely unknown. MicroRNAs (miRNA) are involved in the pathogenesis of many human diseases, but their role in bone formation, regeneration and homeostasis has been apparent until recently. 10 miRNA controls the activity of many genes essential for bone metabolism. 11 Mahboudi et al. 12 suggested that miRNA-140 was a potent inducer of chondrogenic differentiation in bone marrow mesenchymal stem cells, and that overexpression of miRNA-140 could enhance the chondrogenic differentiation. Currently, the prediction method of miRNA target genes is widely used due to its low cost and little prior knowledge requirement. 13 Although the results can be obtained quickly through bioinformatics prediction, the method has high false positive and negative rate, which is easy to cause interference and misleading to downstream functional experiments. 14 In addition, there are few prediction algorithms that can identify miRNA functions or integrate data from different experimental high-throughput technologies. Undoubtedly, a computational approach that effectively combines mRNA expression profiles with miRNA expression profiles is essential for reliable prediction of miRNA targets. 15
In this study, 5 DDH cartilages and 5 neck of femur fracture (NOF) cartilages were obtained to perform genome-wide mRNA and miRNA sequencing. The mRNA and miRNA expression profiles were then utilized to conduct differential expressions and gene set enrichment analyses, respectively. Integrative analysis of mRNA and miRNA was finally conducted to identify the candidate genes, gene ontology (GO) terms and pathways for DDH. The DDH-associated differentially expressed genes (DEGs) were further compared with hip OA–related DEGs. Our results provide new clues for understanding the molecular mechanisms of DDH.
Materials and Methods
Samples Collection
Human hip articular cartilage was obtained from the Honghui Hospital of Xi’an city (Shaanxi Province, China) from patients following informed consent and approval by the institutional review board at Xi’an Jiaotong University. Articular cartilage was obtained from 10 Han Chinese subjects, including 5 adult DDH patients and 5 unrelated NOF subjects. All the study subjects underwent careful clinical examination and radiography collection. The DDH hip joints were taken from patients with Crowe type II who did not have previous surgery, OA, or other musculoskeletal disease. All the DDH patients had severe hip joint pain, limb shortening and limited hip function before total hip replacement. 16 The NOF subjects were diagnosed as Garden III-IV, and they had neither family history nor clinical evidence of any inflammatory disorders and arthritis. The basic characteristics of the study subjects are summarized in Table 1 .
The Basic Characteristics of the Study Subjects.

DDH = developmental dysplasia of hip; NOF = neck of femur fracture.
Chondrocyte Isolation
Proliferating primary chondrocytes at passage 0 from the hip cartilage were used in this study. Chondrocytes were isolated as described previously. 17 In brief, the collected cartilage tissue was sliced and digested in phosphate buffered saline (PBS) containing 0.2% trypsin in a 37°C incubator for 30 minutes, and then incubated with 0.025% collagenase II in Dulbecco’s modified Eagle medium (DMEM)/F-12 media for 8 hours. The chondrocytes were counted to 1 × 105 and resuspended in T25 flasks. The chondrocytes were incubated (5% CO2, 37°C) in DMEM/F-12 media with 10% fetal bovine serum (FBS) and 1% antibiotics (streptomycin and penicillin) for 5 days.
RNA Isolation and Sequencing
Total RNA specimens were extracted from chondrocyte using the TRIzol Reagent (TIANGEN, China). 18 The integrity of RNA samples was assessed using the Nanodrop spectrophotometer (NanoDrop Technologies) and Agilent Bioanalyzer 2100 (Agilent, USA). RNA samples with A260/A280 ratios ranging from 1.9 to 2.1, A260/A230 ratios ≥ 2, and RNA integrity values ≥ 8 were used in this study. The NEBNext Multiplex Small RNA Library Prep Set (NEB #E7300S/L, Illumina) was used for library preparation and sequencing. Briefly, 5 µg RNA was used for library preparation. HiSeq2500 Illumina platform was then used to generate sequenced reads. Paired-end read (150 bp) was performed on mRNA sequencing while single read (50 bp) was performed on small RNA sequencing. The contaminated samples, the reads pair with high N content (the whole pair of reads was removed if the content of N in a read exceeds 5%), reads less than 50 bp in length and unpaired reads were removed to obtain the clean reads. Human genome GRCh38 was used for alignment.
Differential Expression and Functional Gene Set Enrichment Analysis
Limma packages (version 3.32.10) were used for differential expression analysis of mRNA expression profiles.19,20 The screening criteria for DEGs were as follows: |log2FC| ≥ 1 and P ≤ 0.05. GO and pathway enrichment analyses for DEGs were performed by the KOBAS server (version 3.0). 21 Benjamini and Hochberg (BH) procedures were used for multiple testing corrections to correct the P value, and the corrected P < 0.05 was considered significant. For miRNA, Bowtie 1.1.1 and miRDeep2 2.0.0.6 were used for short sequence alignment 22 and miRNA prediction, respectively. 23 The edgeR package 3.8.6 was used for differential expression analysis of miRNA expression profiles. 24 The upregulated miRNA were identified as P ≤ 0.05 and log2 (FC) ≥ 1, while the downregulated miRNA were identified as P ≤ 0.05 and log2 (FC) ≤ −1. The miRanda software 3.3a was used to predict the target genes of identified miRNA, with score ≥140 and energy ≤ −20 kcal/mol. 13 The functional gene set enrichment analysis of the target genes for identified miRNAs were performed as same as mRNAs.
Integrative Analysis of mRNA and miRNA Expression Profiles
The expression correlation coefficients between differential expressed mRNA and miRNA were calculated to screen for significant negative correlation pairs (P < 0.05 and expression correlation coefficient < −0.80). The target genes of identified miRNA were intersected with significant negative correlation pairs, and then constructed the miRNA-gene expression regulatory network. Based on the DEGs in the regulatory network, GO and pathway enrichment analyses were performed using KOBAS 3.0. P < 0.05 was considered significant. 21
The Gene Expression Profiling Data of Hip Osteoarthritis
In order to explore the difference of gene expression profiles between DDH and hip OA cartilage, a recent gene expression profiling data of hip OA was used here. 25 Briefly, hip cartilages from NOF (n = 10) and hip OA (n = 9) patients were used for RNA extraction. RNA samples with RNA integrity values ≥7 were selected for whole-genome expression array experiments. Illumina whole-genome expression array Human HT-12 V3 was used to profile gene expression of RNA samples according to the manufacturer’s protocol. Detailed information of mRNA expression profile can be found in the published study. 25
Results
Difference Expression and Gene Set Enrichment Analysis
Differential expression analysis of mRNA expression profiles identified 1,833 significant DEGs for DDH (Supplemental Table S1), such as AKR1C2 (log FC = 1.681, P = 3.54 × 10−5), IL16 (log FC = 1.691, P = 4.16 × 10−5), and PDCD1LG2 (log FC = −1.123, P = 4.39 × 10−5). GO enrichment analysis of mRNA expression profiles identified 982 significant GO terms with P < 0.01 for DDH (Supplemental Table S2), such as organelle (P = 2.47 × 10−28), protein binding (P = 3.70 × 10−25), and microtubule cytoskeleton organization (P = 2.78 × 10−10). Pathway enrichment analysis of mRNA expression profiles identified 175 significant pathways with P < 0.01 for DDH, such as metabolic pathways (P = 1.18 × 10−6), generic transcription pathway (P = 2.03 × 10−6), and metabolism of proteins (P = 2.50 × 10−6) (Supplemental Table S3).
Differential expression analysis of miRNA expression profiles identified 186 significant miRNA for DDH (Supplemental Table S4), such as hsa-miR-195-5p (log FC = −2.543, P = 2.04 × 10−6), hsa-miR-1247-5p (log FC = 3.512, P = 9.70 × 10−5), and hsa-miR-374c-5p (log FC = 2.526, P = 1.23 × 10−5). A total of 65 upregulated and 86 downregulated miRNAs were predicted from the 186 miRNA. GO enrichment analysis of miRNA expression profiles identified 11 significant GO terms for DDH, such as regulation of cell projection organization (P = 0.016) and negative regulation of translation (P = 0.016) (Supplemental Table S5). Pathway enrichment analysis of miRNA expression profiles identified one significant pathway for DDH, axon guidance (P = 0.032) (Supplemental Table S6).
Integrative Analysis of mRNA and miRNA Expression Profiles
We detected 175 differential expressed genes from the miRNA-mRNA expressed regulatory networks, such as CDKN1C (PmRNA = 0.011, PmiRNA = 3.88 × 10−10), VWA1 (PmRNA = 0.039, PmiRNA = 4.98 × 10−2), TMEM119 (PmRNA = 3.91 × 10−3, PmiRNA = 2.48 × 10−3), and SCUBE3 (PmRNA = 3.28 × 10−3, PmiRNA = 3.92 × 10−5) (Supplemental Table S7). Fig. 1 presents the miRNA-gene expression regulation network.
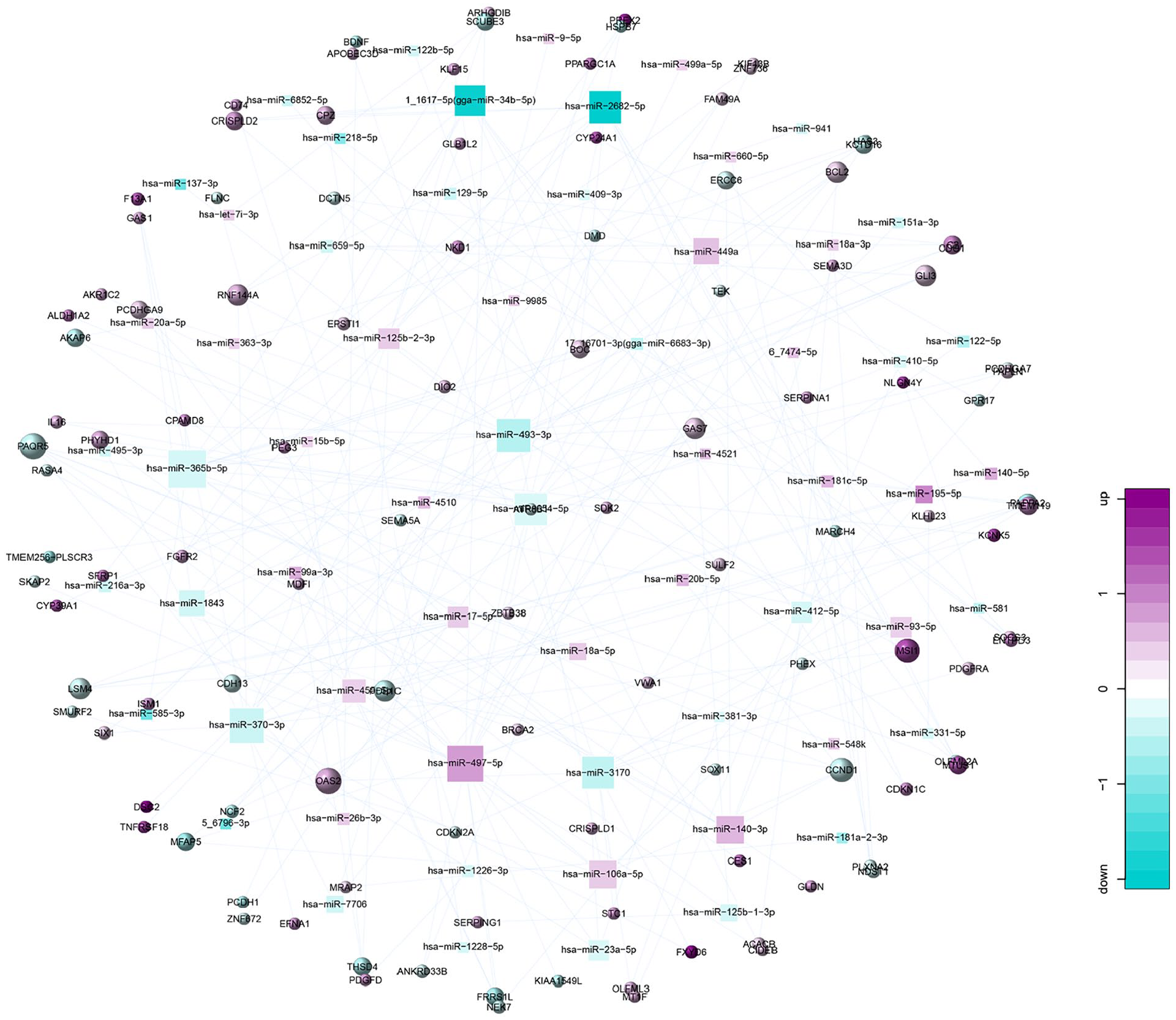
miRNA-gene expression regulation network.
GO enrichment analysis identified 111 significant GO terms for DDH, such as skeletal system morphogenesis (P = 4.92 × 10−5), skeletal system development (P = 8.85 × 10−5), embryonic skeletal system development (P = 1.55 × 10−4), and regulation of osteoblast proliferation (P = 8.60 × 10−4) ( Fig. 2 , Supplemental Table S8). Pathway enrichment analysis identified 14 significant pathways for DDH, such as Hedgehog signaling pathway (P = 4.29 × 10−5), Wnt signaling pathway (P = 0.044), PI3K-Akt signaling pathway (P = 0.024), and Ras signaling pathway (P = 0.022) ( Fig. 3 , Supplemental Table S9).
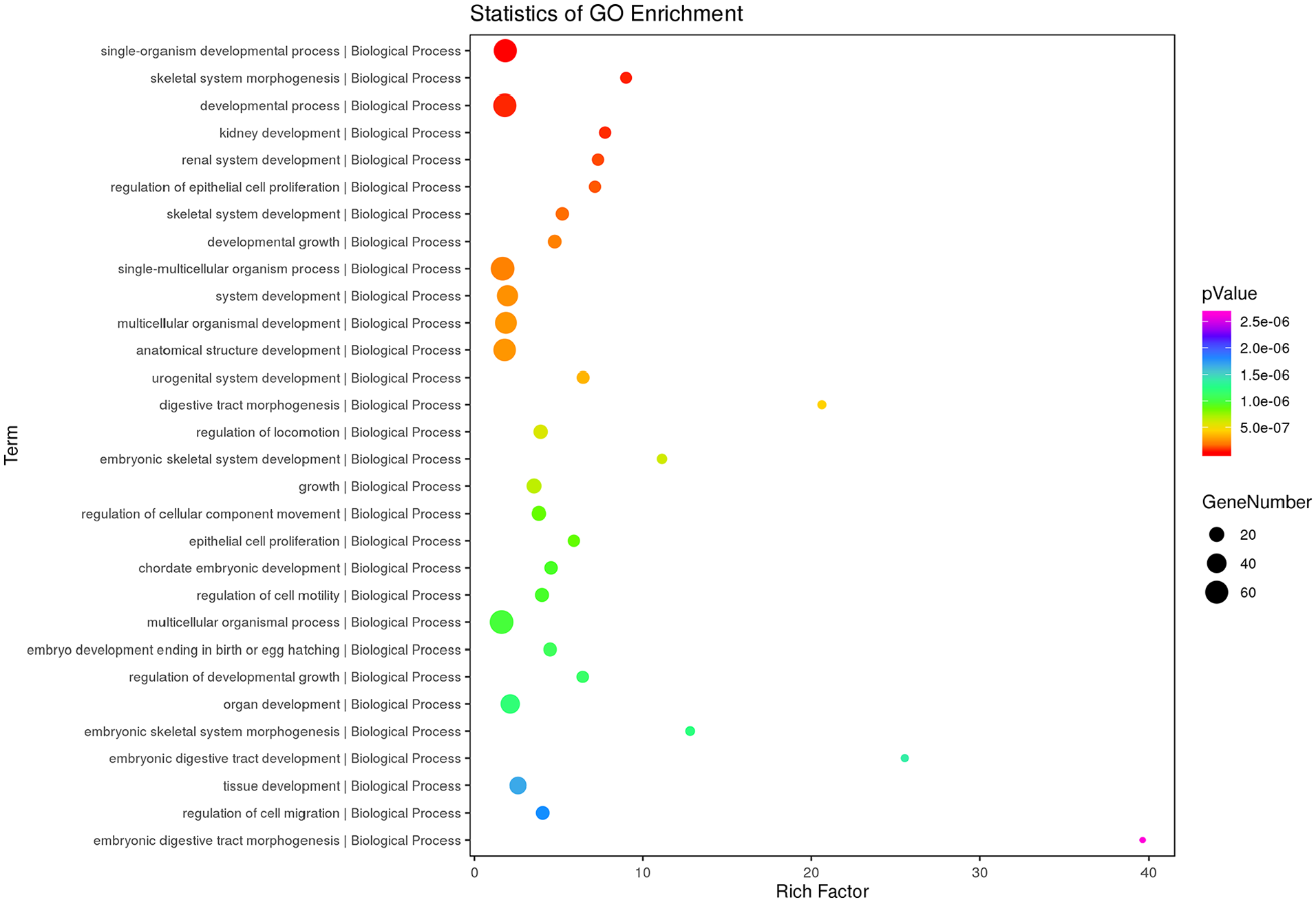
Top 30 terms of gene ontology (GO) enrichment analysis results of the differential expression of mRNAs and genes (DEMs-DEGs).
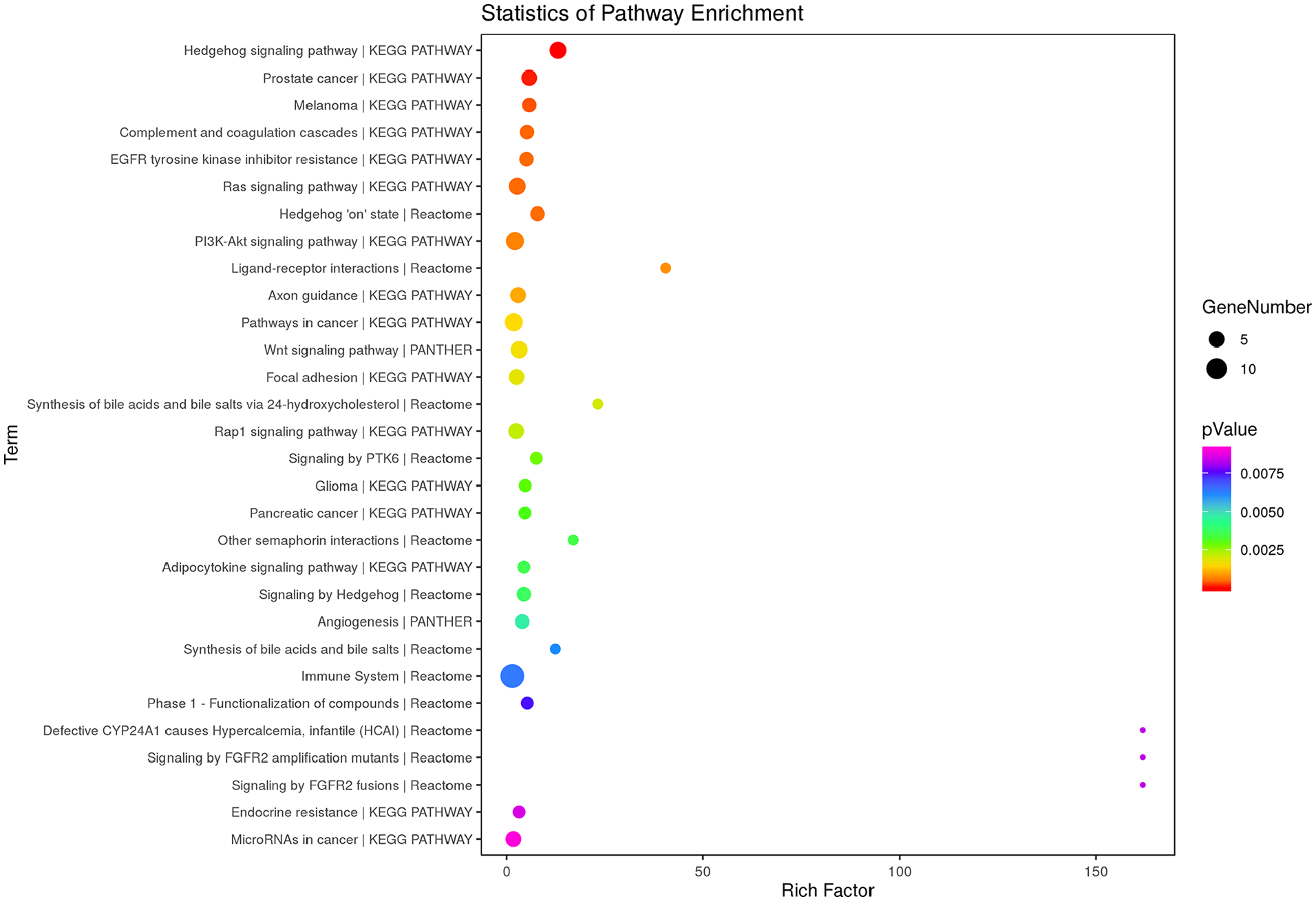
Top 30 terms of pathway enrichment analysis results of the differential expression of mRNAs and genes (DEMs-DEGs).
Comparison of Differential Expressed Genes between DDH and Hip OA
Among the overlapped genes between DDH and hip OA, only 1% showed regulation in the same direction ( Fig. 4 ). For example, CDKN1C (PDDH = 0.011, Phip OA = 5.58 × 10−3), VWA1 (PDDH = 0.039, Phip OA = 5.58 × 10−3), and ACACB (PDDH = 5.32 × 10−4, Phip OA = 8.75 × 10−3) were upregulated in both DDH and Hip OA. EFNA1 (PDDH = 0.010, Phip OA = 5.58 × 10−3) was upregulated in DDH but downregulated in hip OA ( Table 2 ).
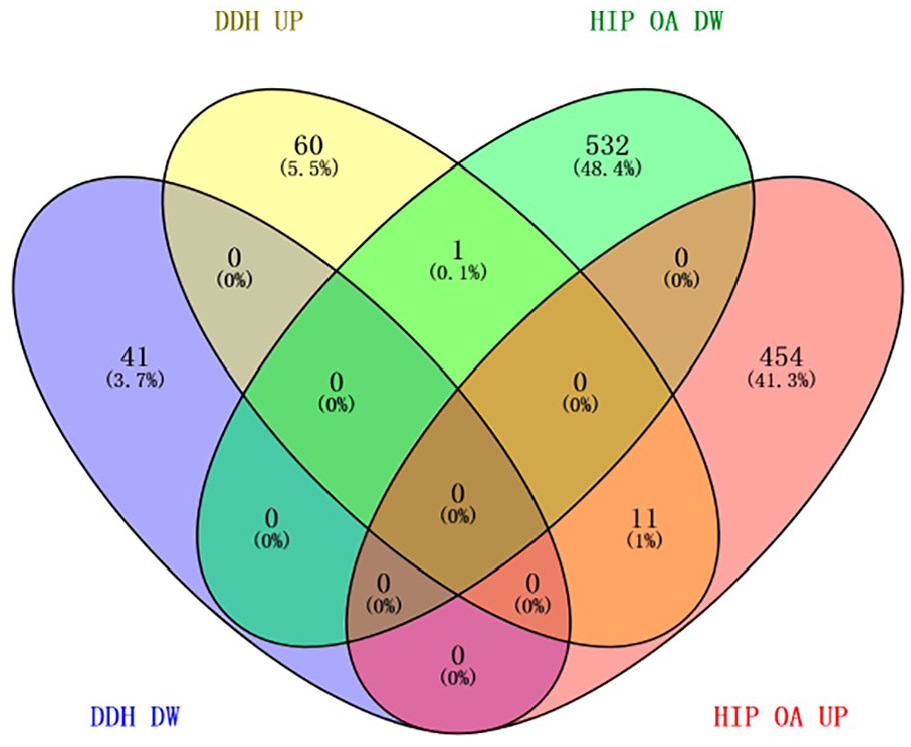
Four-way Venn diagram of DDH and hip OA DEGs. DDH, developmental dysplasia of hip; OA, osteoarthritis; DEGs, differentially expressed genes; DW, downregulated; UP, upregulated.
Comparison of Differentially Expressed Genes between Developmental Dysplasia of Hip (DDH) and Hip Osteoarthritis (OA). a
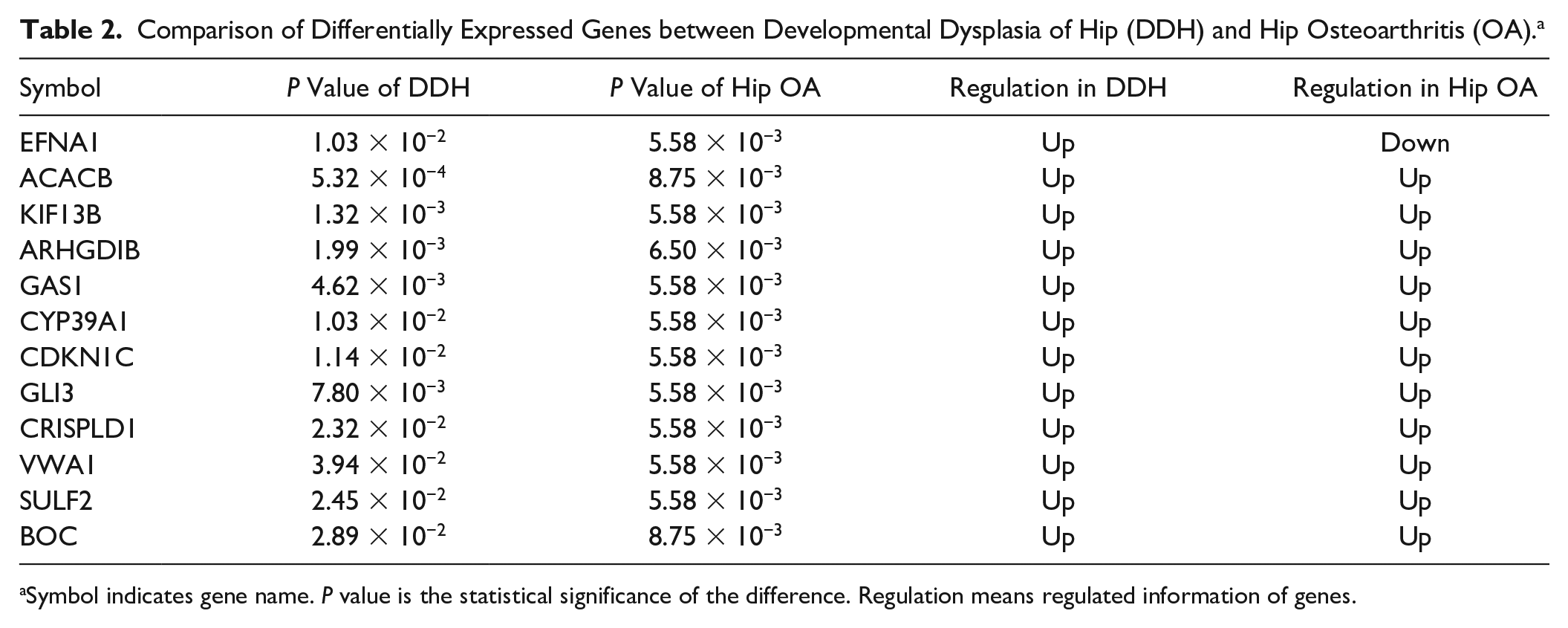
Symbol indicates gene name. P value is the statistical significance of the difference. Regulation means regulated information of genes.
Discussion
To gain novel insight into the molecular mechanism of DDH, five paired cartilages were utilized to conduct the genome-wide mRNA and miRNA sequencing, respectively. Integrative analysis of mRNA and miRNA expression profiles identified multiple DDH associated genes, GO terms and pathways.
We identified several novel candidate genes for DDH, such as CDKN1C and VWA1. CDKN1C is a key negative regulator of cell growth encoded by a paternally imprinted/maternally expressed gene in humans, and associated with metaphyseal dysplasia and intrauterine growth retardation. 26 Lord et al. 27 found that CDKN1C mRNA was expressed by only the immature chondrocytes within the precartilage region in red deer. VWA1 belongs to the von Willebrand factor, A (VWFA) domain superfamily of extracellular matrix proteins, and plays a role in cartilage structure and function. 28 Klatt et al. 29 demonstrated that the VWA1 domain of matrilin-3 was primarily responsible for the induction of proinflammatory cytokines interleukin (IL)-6 releases from primary human chondrocytes. In our study, CDKN1C and VWA1 were upregulated in both DDH and hip OA cartilage, indicating the implication of CDKN1C and VWA1 in the cartilage destruction of hip disorder.
DDH is a common musculoskeletal condition in newborns. Infants with DDH have a higher incidence of early-onset hip OA in adulthood. 30 The dysplastic hips is the main precursor for hip OA and is seen in 20% to 40% of patients with hip OA. 31 The clinical manifestations of DDH were similar to those of hip OA. Genetic association has also been reported between DDH and OA. 32 Besides, similar pathways and genes were found in this study including skeletal system development pathway. Nevertheless, DDH and hip OA have differential genetic regulation and pathogenic factors. 33 In addition to hip OA development, patients with DDH can develop other musculoskeletal and biomechanical changes. 34 In our study, we also found some differences in genetic component between DDH and hip OA including EFNA1, which was upregulated in DDH cartilage but downregulated in hip OA cartilage. EFNA1 is one of the cell differentiation and proliferation regulation factors. 35 EFNA1 was expressed in normal bone, and it ligand binding induced MAPK activation which were closely related with OA cartilage injury. 36 Our results suggest that EFNA1 may play different roles in the progress of DDH and hip OA.
GO enrichment analysis found that the identified candidate genes were functionally involved in the skeletal system morphogenesis, skeletal system development and embryonic skeletal system development. It was well known that DDH patients show abnormality of hip development, and the clinical manifestations depend on the age of the patient. 37 For example, DDH just may present with mild limitation of abduction in newborn. In the toddler, DDH may present with asymmetric gait, while in adults with hip pain and degenerative arthritis. 37 The abnormal of skeletal system development may be related to the fact that the patients with DDH also suffer from severe OA. 38
We also identified several candidate pathways for DDH, including Hedgehog signaling pathway and Wnt signaling pathway. The critical function of Hedgehog signaling and Wnt signaling pathway in chondrogenesis has been already well identified in the past 2 decades.39,40 Indian Hedgehog has an essential function in endochondral ossification and induces osteoblast differentiation in the perichondrium.41,42 Various Wnts are involved in the subsequent stages of chondrocyte differentiation during development, and deregulation of Wnt signaling was observed in cartilage degeneration. 43 Interactions between Hedgehog signaling and Wnt signaling can regulate cartilage development, endochondral bone formation and synovial joint formation. 44 Our results demonstrate that the synergistic effect of Hedgehog signaling pathway and Wnt signaling pathway is crucial for DDH progress, indicate that abnormal chondrogenesis may play a crucial role in the etiology of DDH.
DDH spans a broad spectrum of abnormal hip development during infancy and early development. The definition of DDH comprises a wide range of abnormalities, from acetabular dysplasia without hip dislocation to frank hip dislocation. 45 In this study, our sample comes from a small subset of DDH, Crowe II (subluxation of the femoral head of 50% to 75%). The results may not be applicable to acetabular dysplasia and dislocation. The etiology of hip OA is multifactorial, including biochemical, genetic, gene-environment interactions and acquired abnormalities.46,47 Besides, several features of hip joint architecture, such as acetabular dysplasia, pistol grip deformity, wide femoral neck, hip subluxation and true dislocation of the hip, appear to play a vital role in the pathogenesis of hip OA.38,48 The etiology of these different types of hip OA secondary to DDH may not totally the same.
To the best of our knowledge, this is the first study to explore the molecular pathogenesis of DDH using genome-wide trans-omics of miRNA and mRNA expression profiles. Notably, there are several limitations of this study. First, our study used 5 pairs of DDH and NOF samples. Similar sample sizes have been used in previous RNA sequencing studies.25,49 We integrated microRNA and mRNA sequencing to ensure the accuracy of gene identification and the value of the analysis. Second, it is almost impossible to collect young NOF samples that are completely matched with the age of DDH samples, which might be a confounding factor. Richard et al. 50 indicated that age affects the basal pattern of gene expression in joint tissues. However, Silva et al. 51 found that the cellular and epigenetic features of the intermediate and deep zones of aged NOF cartilage are similar to those of young healthy cartilage. Third, DDH is abnormal in bone, cartilage, and ligament, while this study only focuses on the mechanism of DDH cartilage damage. Further studies are needed to confirm our findings and reveal the potential roles of identified genes, GO terms, and pathways in the development of DDH.
In conclusion, we identified several candidate genes and pathways as the potential biomarkers for the development of DDH. The comparison of DEGs between DDH and hip OA identified several differential genetic components. Our study indicates potential regulatory mechanisms of the mRNA and miRNA in DDH, and aims to assist further molecular biological investigations between DDH and hip OA.
Supplemental Material
sj-xlsx-1-car-10.1177_1947603521990859 – Supplemental material for Integrative Analysis of MicroRNA and mRNA Sequencing Data Identifies Novel Candidate Genes and Pathways for Developmental Dysplasia of Hip
Supplemental material, sj-xlsx-1-car-10.1177_1947603521990859 for Integrative Analysis of MicroRNA and mRNA Sequencing Data Identifies Novel Candidate Genes and Pathways for Developmental Dysplasia of Hip by Bolun Cheng, Yumeng Jia, Yan Wen, Weikun Hou, Ke Xu, Chujun Liang, Shiqiang Cheng, Li Liu, Xiaomeng Chu, Jing Ye, Yao Yao, Feng Zhang and Peng Xu in CARTILAGE
Supplemental Material
sj-xlsx-2-car-10.1177_1947603521990859 – Supplemental material for Integrative Analysis of MicroRNA and mRNA Sequencing Data Identifies Novel Candidate Genes and Pathways for Developmental Dysplasia of Hip
Supplemental material, sj-xlsx-2-car-10.1177_1947603521990859 for Integrative Analysis of MicroRNA and mRNA Sequencing Data Identifies Novel Candidate Genes and Pathways for Developmental Dysplasia of Hip by Bolun Cheng, Yumeng Jia, Yan Wen, Weikun Hou, Ke Xu, Chujun Liang, Shiqiang Cheng, Li Liu, Xiaomeng Chu, Jing Ye, Yao Yao, Feng Zhang and Peng Xu in CARTILAGE
Supplemental Material
sj-xlsx-3-car-10.1177_1947603521990859 – Supplemental material for Integrative Analysis of MicroRNA and mRNA Sequencing Data Identifies Novel Candidate Genes and Pathways for Developmental Dysplasia of Hip
Supplemental material, sj-xlsx-3-car-10.1177_1947603521990859 for Integrative Analysis of MicroRNA and mRNA Sequencing Data Identifies Novel Candidate Genes and Pathways for Developmental Dysplasia of Hip by Bolun Cheng, Yumeng Jia, Yan Wen, Weikun Hou, Ke Xu, Chujun Liang, Shiqiang Cheng, Li Liu, Xiaomeng Chu, Jing Ye, Yao Yao, Feng Zhang and Peng Xu in CARTILAGE
Supplemental Material
sj-xlsx-4-car-10.1177_1947603521990859 – Supplemental material for Integrative Analysis of MicroRNA and mRNA Sequencing Data Identifies Novel Candidate Genes and Pathways for Developmental Dysplasia of Hip
Supplemental material, sj-xlsx-4-car-10.1177_1947603521990859 for Integrative Analysis of MicroRNA and mRNA Sequencing Data Identifies Novel Candidate Genes and Pathways for Developmental Dysplasia of Hip by Bolun Cheng, Yumeng Jia, Yan Wen, Weikun Hou, Ke Xu, Chujun Liang, Shiqiang Cheng, Li Liu, Xiaomeng Chu, Jing Ye, Yao Yao, Feng Zhang and Peng Xu in CARTILAGE
Supplemental Material
sj-xlsx-5-car-10.1177_1947603521990859 – Supplemental material for Integrative Analysis of MicroRNA and mRNA Sequencing Data Identifies Novel Candidate Genes and Pathways for Developmental Dysplasia of Hip
Supplemental material, sj-xlsx-5-car-10.1177_1947603521990859 for Integrative Analysis of MicroRNA and mRNA Sequencing Data Identifies Novel Candidate Genes and Pathways for Developmental Dysplasia of Hip by Bolun Cheng, Yumeng Jia, Yan Wen, Weikun Hou, Ke Xu, Chujun Liang, Shiqiang Cheng, Li Liu, Xiaomeng Chu, Jing Ye, Yao Yao, Feng Zhang and Peng Xu in CARTILAGE
Supplemental Material
sj-xlsx-6-car-10.1177_1947603521990859 – Supplemental material for Integrative Analysis of MicroRNA and mRNA Sequencing Data Identifies Novel Candidate Genes and Pathways for Developmental Dysplasia of Hip
Supplemental material, sj-xlsx-6-car-10.1177_1947603521990859 for Integrative Analysis of MicroRNA and mRNA Sequencing Data Identifies Novel Candidate Genes and Pathways for Developmental Dysplasia of Hip by Bolun Cheng, Yumeng Jia, Yan Wen, Weikun Hou, Ke Xu, Chujun Liang, Shiqiang Cheng, Li Liu, Xiaomeng Chu, Jing Ye, Yao Yao, Feng Zhang and Peng Xu in CARTILAGE
Supplemental Material
sj-xlsx-7-car-10.1177_1947603521990859 – Supplemental material for Integrative Analysis of MicroRNA and mRNA Sequencing Data Identifies Novel Candidate Genes and Pathways for Developmental Dysplasia of Hip
Supplemental material, sj-xlsx-7-car-10.1177_1947603521990859 for Integrative Analysis of MicroRNA and mRNA Sequencing Data Identifies Novel Candidate Genes and Pathways for Developmental Dysplasia of Hip by Bolun Cheng, Yumeng Jia, Yan Wen, Weikun Hou, Ke Xu, Chujun Liang, Shiqiang Cheng, Li Liu, Xiaomeng Chu, Jing Ye, Yao Yao, Feng Zhang and Peng Xu in CARTILAGE
Supplemental Material
sj-xlsx-8-car-10.1177_1947603521990859 – Supplemental material for Integrative Analysis of MicroRNA and mRNA Sequencing Data Identifies Novel Candidate Genes and Pathways for Developmental Dysplasia of Hip
Supplemental material, sj-xlsx-8-car-10.1177_1947603521990859 for Integrative Analysis of MicroRNA and mRNA Sequencing Data Identifies Novel Candidate Genes and Pathways for Developmental Dysplasia of Hip by Bolun Cheng, Yumeng Jia, Yan Wen, Weikun Hou, Ke Xu, Chujun Liang, Shiqiang Cheng, Li Liu, Xiaomeng Chu, Jing Ye, Yao Yao, Feng Zhang and Peng Xu in CARTILAGE
Supplemental Material
sj-xlsx-9-car-10.1177_1947603521990859 – Supplemental material for Integrative Analysis of MicroRNA and mRNA Sequencing Data Identifies Novel Candidate Genes and Pathways for Developmental Dysplasia of Hip
Supplemental material, sj-xlsx-9-car-10.1177_1947603521990859 for Integrative Analysis of MicroRNA and mRNA Sequencing Data Identifies Novel Candidate Genes and Pathways for Developmental Dysplasia of Hip by Bolun Cheng, Yumeng Jia, Yan Wen, Weikun Hou, Ke Xu, Chujun Liang, Shiqiang Cheng, Li Liu, Xiaomeng Chu, Jing Ye, Yao Yao, Feng Zhang and Peng Xu in CARTILAGE
Footnotes
Supplemental Material
Authors’ Note
All the supplementary materials including the RNA seq data of this study are available from the corresponding authors on request.
Acknowledgments and Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Scientific Foundation of China (81673112, 81922059); the key projects of international cooperation among governments in scientific and technological innovation (2016YFE0119100); the Natural Science Basic Research Plan in Shaanxi Province of China (2017JZ024); and the Fundamental Research Funds for the Central Universities (xzy022019006).
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
This study was approved by the institutional review board at Xi’an Jiaotong University (approval number: 2016-263).
Informed Consent
All patients provided informed consent.
Trial Registration
Not applicable.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
