Abstract
Objective
To find out the pathways and key genes and to reveal disc degeneration pathogenesis based on bioinformatic analyses.
Design
The GSE70362 dataset was downloaded from the GEO (Gene Expression Omnibus) database. Differentially expressed genes (DEGs) between the patients having disc degeneration and healthy controls were screened by Limma package in R language. Critical genes were identified by adopting gene ontologies (GOs), Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways, and protein-protein interaction (PPI) networks.
Results
We identified 112 DEGs, including 60 genes which were upregulated and 52 that were downregulated. Analyses, such as GO and KEGG demonstrated that the DEGs got enriched in 4 biological processes and 2 signaling pathways, mainly related to disc degeneration. The PPI network analyses identified 5 key proteins, CCND1 (cyclin D1), GATA3, TNFSF11, LEF1, and DKK1 (Dickkopf related protein 1).
Conclusion
In this study, the DEGs and pathways determined promoted us understand the disc degeneration mechanisms. Also, the study may contribute novel biomarkers for the diagnosis and prevention of disc degeneration, and seek new treatment methods to repair and even regenerate degenerative intervertebral disc.
Introduction
Low back pain is a common disabling condition that places a huge economic burden on society. 1 Intervertebral disc degeneration (IVDD) is a critical source of lower back pain; however, the mechanisms related to IVDD are not fully understood.2,3 It is currently believed that IVDD is caused by multiple factors, such as the environment, genetics, aging, sex, and susceptibility to injury.4,5 IVDD is characterized by apoptosis of nucleus pulposus cells (NPCs) and extracellular matrix degradation. Current therapeutic modalities for disc degeneration mainly include conservative and surgical treatment that provide symptomatic and supportive measures but do not cure the disease.6,7 Recently, there have been an increasing number of investigations into biological regeneration therapy for IVDD that mainly focus on intervertebral disc repair / regeneration.8 -10 However, due to the complex mechanisms of disc degeneration, such therapeutic techniques have not been widely used in the clinic. Therefore, the identification of key genes and pathways that cause IVDD will help unravel the underlying mechanisms and provide novel targets that will increase the feasibility of intervertebral disc repair/regeneration.
In this study, we identified the differentially expressed genes (DEGs) between normal and degenerated nucleus pulposus (NP) from transcriptome sequencing gene data in the GEO (Gene Expression Omnibus) database. Critical pathways and proteins were identified through gene ontologies (GOs), Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways, and protein-protein interaction (PPI) networks, which provided novel clues for unraveling the pathogenesis of IVDD and a theoretical basis for intervertebral disc repair and regeneration.
Research Materials and Methods
Retrieving Data
The GEO database (http://www.ncbi.nlm.nih.gov/geo) was used to download the GSE70362 microarray dataset, which includes 48 samples from 12 patients with disc degeneration. The GSE70362 dataset consisted of 24 samples from NP source and 24 samples from annulus fibrosus source. as published by Kazezian et al. 11 in 2015, and the patients were genotyped using the GPL570 Affymetrix Human Genome U133 Plus 2.0 Array.
Data Processing and Identification of DEGs
Gene expression data from NP samples were obtained from the organized and optimized GSE70362 microarray data. The affy packet (https://bioconductor.org/biocLite.R) in the R software bioconductor was used to read the data. The robust multiarray averaging (RMA) algorithm was used to correct the background of the data and normalize the data. Based on Thompson grade criteria, grades I and II samples were considered as nondegeneration and grades III to V as disc degeneration. The Limma package (Limma package R 3.4.3) was used to screen DEGs. The GSE70362 dataset consisted of 16 degenerated and 8 nondegenerated samples. DEGs from these samples were identified (P < 0.05 and |log2 FC| ≥ 1), and a volcano plot was constructed.
The Analyses of GO and KEGG
The GO functional enrichment and KEGG pathway analyses were performed using clusterProfiler (R 3.12.0). 12
PPI Analysis
The DEGs PPI networks were constructed via STRING (Search Tool for the Retrieval of Interacting Genes; http://string-db.org), with confidence scores ≥0.4. 13 PPI networks were drawn using Cytoscape 3.7.1 software.
Results
DEGs Screening
Based on P < 0.05 and |log2 FC| ≥ 1, we analyzed 16 degenerated and 8 nondegenerated NPs in the GSE70362 dataset and identified 112 DEGs, including 60 upregulated and 52 downregulated genes. A volcano plot was used to visualize the changes between normal and degenerated NPs, using the ggplot2 of R software (
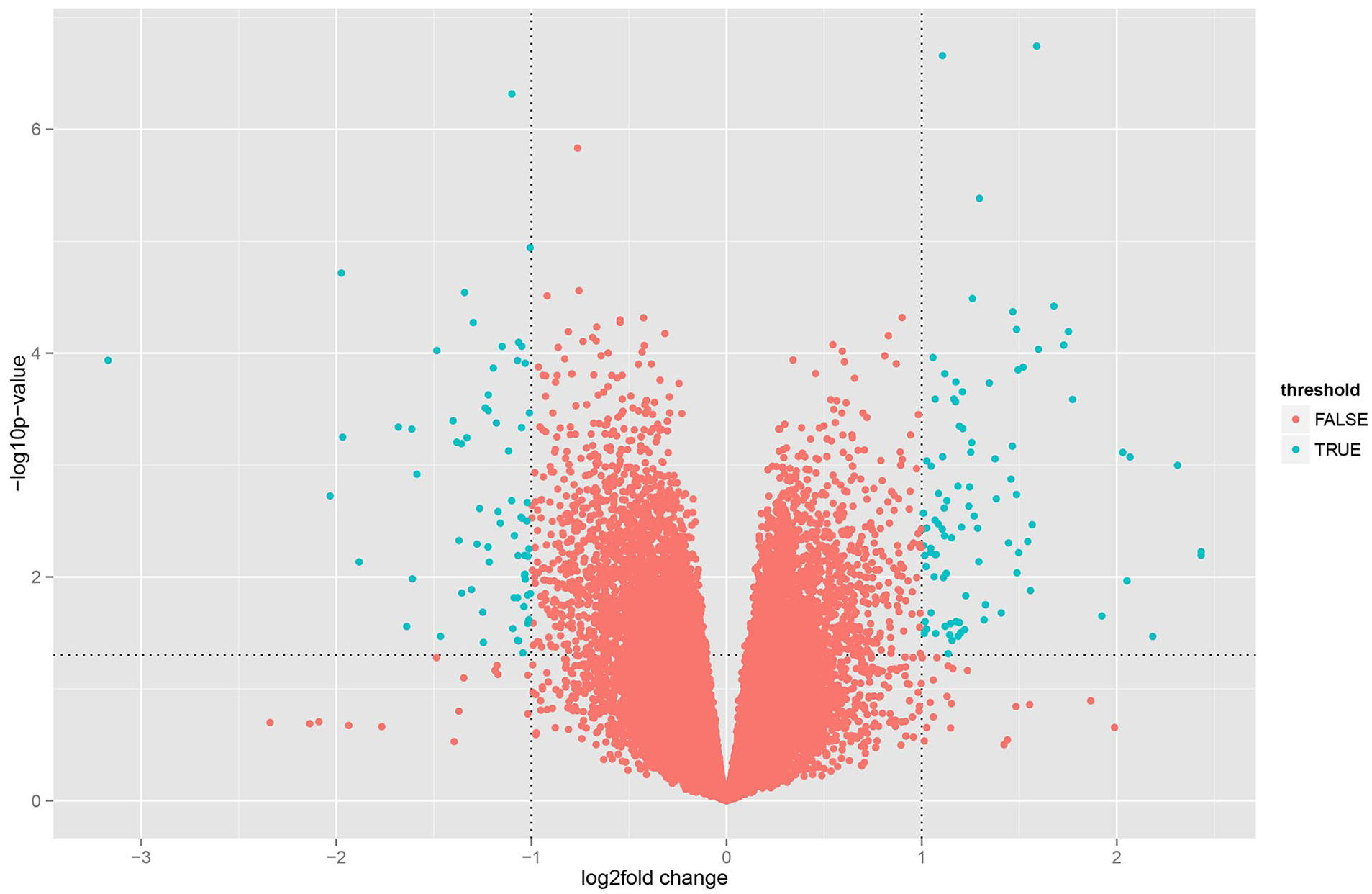
Volcano plot of differentially expressed genes (DEG). Green dot: fold change >2, P < 0.05; red dot: other genes.
GO Functional Enrichment of DEGs
GO functional enrichment of DEGs was analyzed using clusterProfiler (R 3.12.0) with P < 0.05, and “regulation of leukocyte differentiation indicating positive,” “interferon-alpha cellular response,” “BMP response,” and “cellular response to BMP stimulus,” and so on were identified (
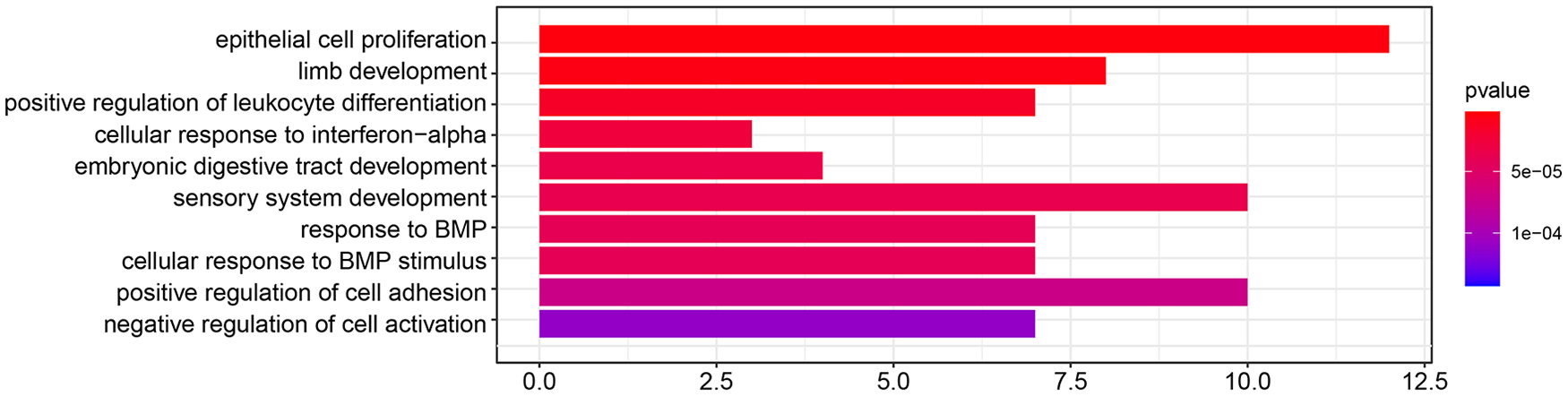
Gene ontology (GO) functional enrichment of differentially expressed gene (DEG). Select important GO entries with P < 0.05.
KEGG Pathway Analysis
ClusterProfiler (R 3.12.0) was adopted to analyze KEGG pathways with P < 0.05, and “pathway of the Hippo signaling” and “pathway of the Wnt signaling” were identified, which are associated with disc degeneration (
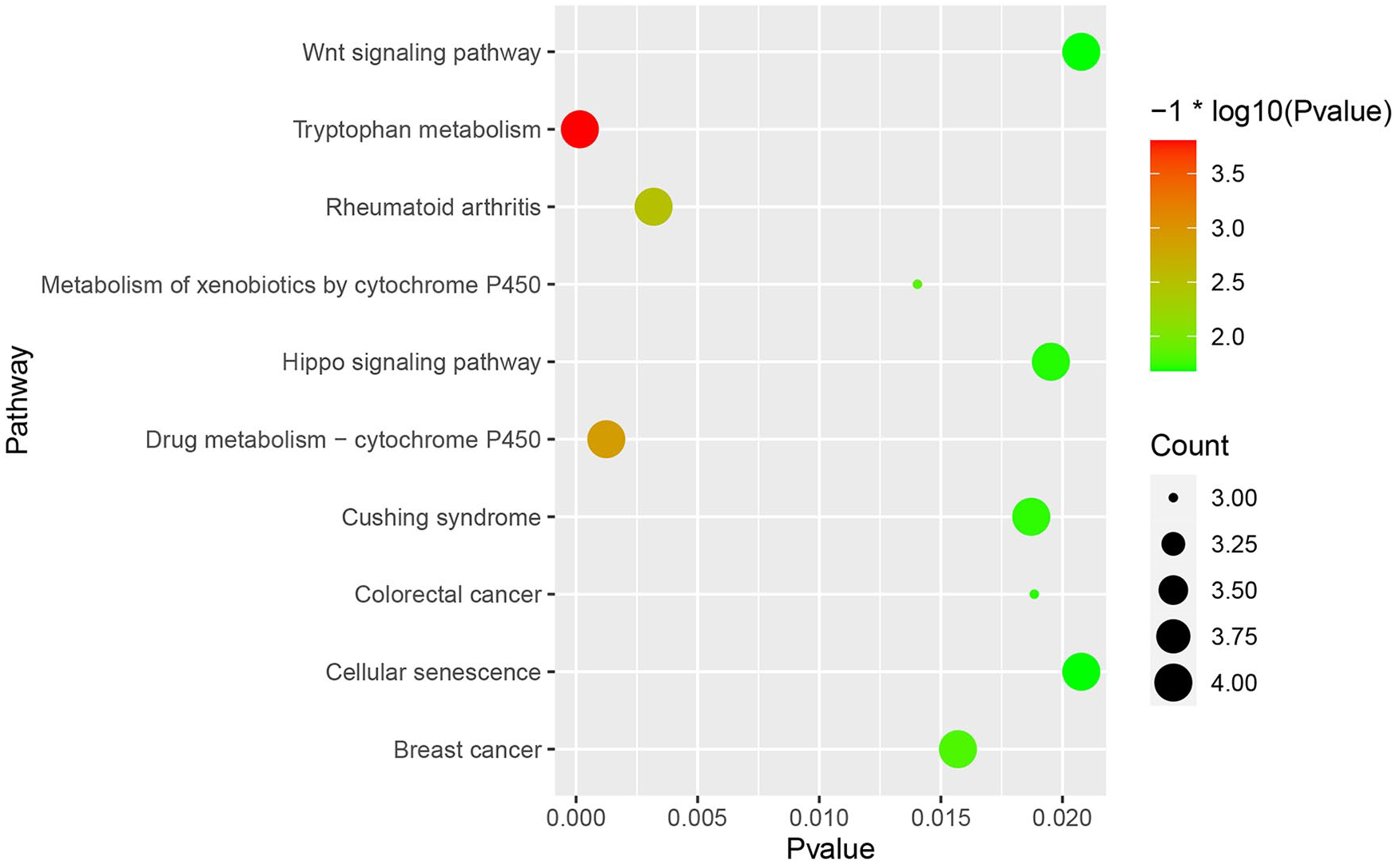
Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways of differentially expressed gene (DEG.) Select significant enrichment pathways with P < 0.05.
PPI Network and Module Analysis
STRING was used to construct DEGs PPI networks with confidence scores ≥0.4 and visualized using Cytoscape 3.7.1 software. We identified 77 nodes and 111 connections, with red and blue colors representing up- and downregulated genes, respectively. The more connections in the network, the greater the degree of importance of the protein. We identified 5 proteins (degree ≥8), including upregulated CCND1, GATA3, TNFSF11, LEF1, and downregulated DKK1 (
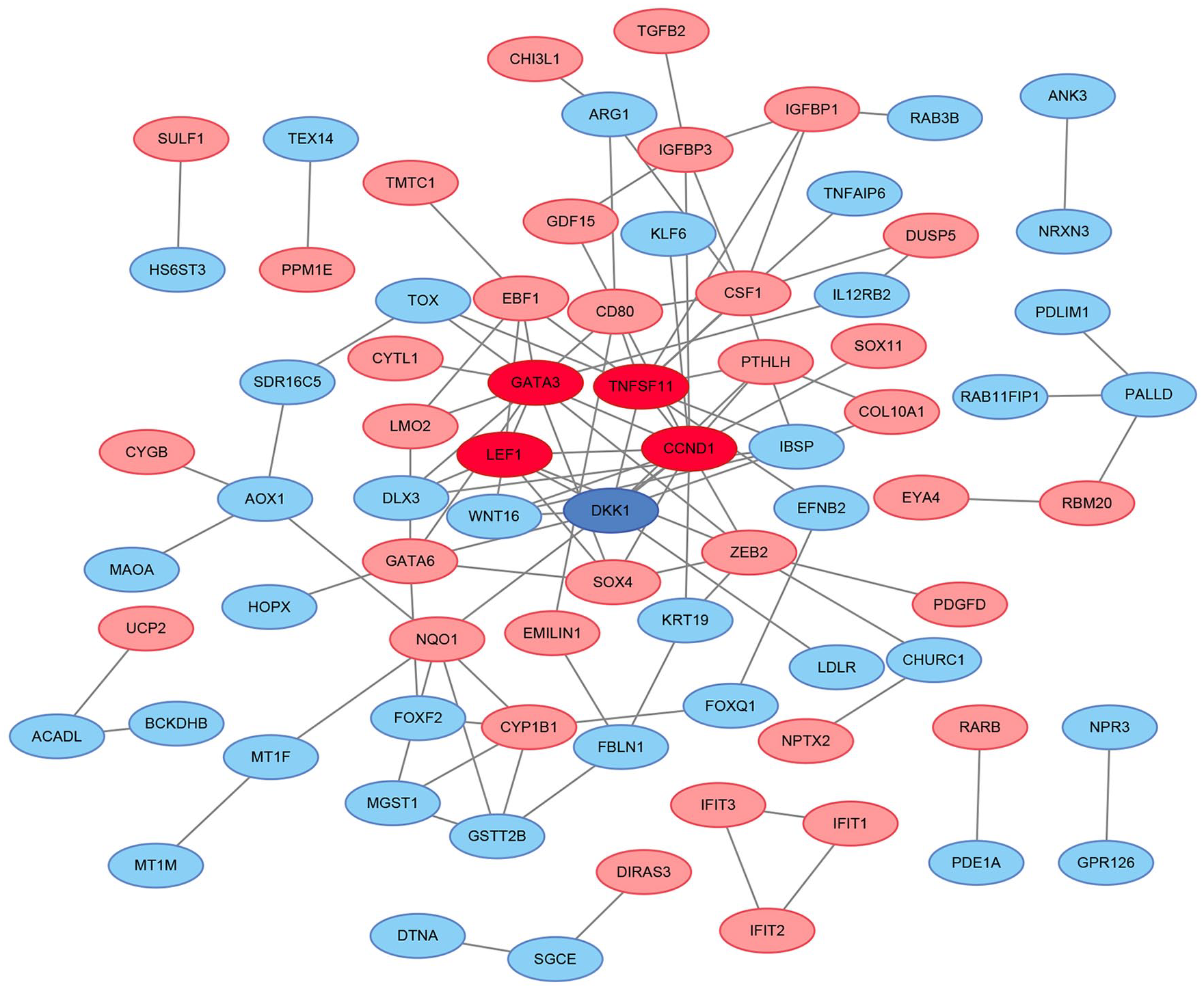
Protein-protein interaction (PPI) network of defferentially expressed gene (DEG). Red color stands for genes that are upregulated and blue color indicates genes that are downregulated, the deepened color refers to the core genes.
Discussion
IVDD is a common condition with complicated etiology. Current treatment measures do not cure disc degeneration or reverse the progression of this condition. In our study, transcriptome sequencing and comparative analyses of DEGs between normal individuals and patients with disc degeneration were performed. GO functional enrichment highlighted “regulation of leukocyte differentiation indicating positive,” “interferon-alpha cellular response,” “BMP response,” and “BMP stimulus cellular response.” Notably, these biological processes are involved in the inflammatory response, immune reaction, and tissue regeneration in the disc. In the intervertebral disc, inflammatory reaction causes disc degeneration. In particular, the production of inflammation mediators, namely interleukin-6, tumor necrosis factor-α, and interleukin-1β, promote matrix degradation, accelerate nucleus pulposus cell (NPC) senescence and death, recruit immune cells, and eventually lead to biomechanical dysfunction of the intervertebral disc.14,15 Compared to patients with disc herniation, Weber et al. 16 observed elevated levels of proinflammatory cytokines, including interferon α2 (IFN-α2), in patients with narrowed and degenerated discs. Ye et al. 17 demonstrated that bone morphogenetic protein–2 (BMP-2) increases aggrecan and type II collagen in NP and annulus fibrosus cells, which suggests its anticatabolic function in these cells. 17
The KEGG pathways analysis identified 2 pathways related to disc degeneration, the Hippo and Wnt signaling pathways. The Hippo signaling pathway is evolutionarily conserved and participates in many biological processes, such as cell proliferation, organ size control, and tissue regeneration. 18 It has been demonstrated that the Hippo signaling pathway plays an important role in the repair and regeneration of the liver, 19 lungs, 20 and heart. 21 When tissues and organs reach a terminal volume, the phosphorylation of LATS1/2 (large tumor suppressor kinases 1/2) in the Hippo signaling pathway leads to the phosphorylation of YAP (yes-associated protein) and cytoplasmic YAP degredataion, and this causes the tissues and organs to stop growing. When tissues and organs are in a state of injury and repair, YAP protein phosphorylation decreases and YAP enters the nucleus, from the cytoplasm, to interact with the TEAD (TEA domain) transcription factor family. This process drives the expression growth-promoting transcriptional genes, such as CTGF (connective tissue growth factor) and Cyr61 (cysteine-rich angiogenesis inducer 61), to mobilize cell proliferation and repair damage. 22 The function of the Hippo signaling pathway in disc degeneration has not been extensively studied. Zhang et al. 23 suggested a correlation between the Hippo pathway and spontaneous disc degeneration. YAP activation and dephosphorylation have also been studied in the young rat intervertebral discs, which gradually weaken with age. YAP overexpression inhibits the Hippo pathway in acute disc injury, and YAP suppression promotes Hippo activation and enhances senescence in NPCs. These findings show that the Hippo pathway plays an important role in intervertebral disc dynamic balance maintenance and NPC proliferation control. 23 Furthermore, regulating YAP protein activity through the Hippo signaling pathway may be an effective IVDD treatment strategy. The Wnt/β-catenin pathways plays a dominat role in cell proliferation, differentiation, degeneration, and regeneration.24,25 Holguin et al. 26 found that disc degeneration is associated with loss of Wnt signaling and is accompanied by an increase in a β-catenin regenerative response. In lumbar disc herniation (LDH), Chen et al. 27 observed elevated β-catenin and WNT1 in NP tissues, compared to normal tissues. As disc degeneration progresses, β-catenin-positive cells increase. Further investigations have revealed that TUG1 siRNA might prevent NPC apoptosis and senescence that are induced by tumor necrosis factor-α, improve cell proliferation, and alleviate extracellular matrix metabolism imbalances by inhibiting the Wnt/β-catenin pathway. These observations provide an experimental basis for clinical IVDD treatments. 27
The PPI network analysis identified 5 core proteins: CCND1, GATA3, TNFSF11, LEF1, and DKK1. CCND1 is a positive cell cycle progression regulator that is responsible for the G-to-S phase cell cycle transition. As an important regulator, the abnormal expression of CCND1 is associated with tumor initiation and progression. 28 Deregulation of CCND1 severely affects cell cycle progression, leading to abnormal cell proliferation, differentiation, and apoptosis.29,30 In the cartilage tissue of patients with osteoarthritis (OA), Cheng et al. 29 observed that SNHG16 and CCND1 were significantly upregulated, and that SNHG16 promoted the development of OA via the miR-93-5p/CCND1 axis. Studies have shown that RMRP expression in degenerated NP tissue is higher than in normal NP tissue. RMPP increases the CCND1 expression in NPCs, and high expression of RMRP positively correlated with the degree of disc degeneration. 31 Furthermore, Tan et al. 32 found that SNHG1 in IVDD tissues was upregulated, compared with control tissues, and that SNHG1 increased in tissues with a higher degree of degeneration, compared to tissues with a lower degree of degeneration. In addition, they also found that ectopic SNHG1 expression enhanced CCND1 in NPCs. Additionally, Zheng et al. 33 found that psoralen promotes chondrocyte prolideration through the CCND1-Wnt/β-catenin signaling pathway. Last, Cao et al. 34 found that high FOXD2-AS1 expression promoted chondrocyte growth by regulating the miR-206/CCND1 axis. Whether these treatments can also be used to repair IVDD needs to be determined by further research.
The transcription factor GATA3 is responsible for the differentiation of Th2 (T helper type 2) cells. 35 In the resting state, Th1 (T helper type 1) and Th2 cells stay in a state of relative balance. Disruption of the Th1/Th2 cell balance may interfere with the cytokine network and lead to the occurrence and development of many diseases. 36 For example, asthma that is triggered by allergies usually presents a Th2 phenotype that is abnormally polarized for a particular allergen and leads to an inflammatory response. 37 Furthermore, Th1-/Th2-related transcription factors are elevated in and associated with various disease, such as systemic lupus erythematosus (SLE). 38 Additionally, insulin-dependent diabetes mellitus (IDDM) is a disease that is mediated by Th1 and Th2 cells. 39 Little is known, however, about the relationship between Th cell polarization and IVDD.
The NP is the body’s largest avascular tissue structure 40 and is surrounded by fibrous rings and cartilage endplates. This structure is not connected to the immune system of the body and also hides its own antigens. In patients with LDH, NP exposure to the body’s immune system causes local inflammation around nerve roots. In these patients, Yao et al. 41 showed that Th2 cells and related cytokines significantly increase, while Th1 cells and related cytokines remained unchanged in the NP. Furthermore, immunofluorescence staining showed fewer IFN-γ+ CD4 cells than IL-4+ CD4 cells, which suggests that Th cells may differentiate into Th2 cells, and the Th1/Th2 imbalance may be involved in the pathogenesis of pain in LDH. 41 Yang et al. 42 demonstrated that GATA3-positive macrophages are absent in the normal heart, but accumulate in the myocardium after acute myocardial infarction. Furthermore, cardiac function significantly improved after myocardial infarction in mice lacking GATA3 myeloid cells, which suggests that GATA3 may be a valuable marker in macrophages for the development of immunotherapies for heart disease. 42 Zhang et al. 43 found that elevated GATA3 is a poor prognosticator for patients with PTCL (viz. peripheral T cell lymphoma). Furthermore, lymophoma T cells enhance M2 macrophage differentiation via a mechanism that is GATA3-dependent. 43 Chen et al. 44 found that miR-466a-3p directly inhibited GATA-3 and alleviated allergic rhinitis in mice by reducing the secretion of the Th2 cytokine IL-4 and OVA (ovalbumin)-specific IgE.
RANKL (receptor activator of nuclear factor-kappa B ligand), which is enciphered by TNFSF11, was initially thought to be part of the tumor necrosis factor ligand superfamily. RANKL is critical to bone metabolism45,46 and was recently found to function in IVDD pathogenesis.47 -50 Using immunohistochemistry, Takegami et al. 50 found that RANKL was elevated in intervertebral disc (IVD) tissues of humans that showed greater degeneration levels, compared with those in an early stage. Furthermore, Sano et al. 51 showed that catabolic factor expression was upregulated by RANKL in the presence of pro-inflammatory stimuli, such as pro-inflammatory cytokines and matrix-degrading enzymes. Additionally, anti-RANKL antibodies inhibited the inflammation-induced upregulation of catabolic factors in human NPCs, which suggests that targeting RANKL under pathological conditions may alleviate IVDD by downregulating catabolic factors, namely proinflammatory cytokines and matrix degrading enzymes. 51
Consistently, TNF-α and IL-6 levels in injured discs have been significantly improved after the application of an anti-RANKL antibody. 52 Moreover, inhibition of RANKL has shown greater specificity than targeting IL-1β or TNF-α in the inflammatory response. Nonspecific immunosuppressants, such as a TNF-α inhibitor, have shown various adverse reactions during clinical applications.53,54 The anti-RANKL antibody denosumab has been applied in the clinic to treat patients with osteoporosis and has exhibited a good safety profile55 -57 and cost-effectiveness.58,59 Therefore, an anti-RANKL antibody may be an effective treatment for patients with IVDD.
LEF1 is a sequence-specific DNA-binding protein that is present in adult mouse pre-B and T lymphocytes of the midbrain, neural crest, hair follicles, and tooth germs during embryonic development.60,61 LEF1-deficient mice exhibit developmental defects in their teeth, hair follicles, breast, and brain. 61 Ji et al. 62 found that LEF1-knockdown inhibits chondrocyte proliferation and that PBXIP1/HPIP (pre-B-cell leukemia homeobox interacting protein 1, or PBX1 interacting protein of humans) is of great importance for OA development. HPIP promotes Wnt target gene transcription by enhancing LEF1 transcription activity, and this leads to OA. 62 Wang et al. 63 further confirmed that LEF1 is elevated in IVDD and that miR-22 inhibits LEF1 expression. Notably, LEF1 is a transcriptional activator for the Wnt/β-catenin pathway, and H19 functions as a competing endogenous RNA that prevents miR-22 from inhibiting LEF1, thereby attenuating the miR-22/Wnt/β-catenin axis and enhancing NPC regeneration, which is induced by the H2O2. These findings indicate that the H19/miR-22/LEF1 system may provide a novel target to counter NPC degeneration due to H2O2 and human IVDD. 63
Dkk1 is a blocker of the Wnt/β-catenin pathway, and Huang et al. 64 demonstrated that Dkk1 is negatively correlated with OA severity and various pro-inflammatory cytokines, which suggests that Dkk1 may be a specific biomarker for joint diseases. Patience et al. 65 detected decreased DKK1 and increased LEF1 in a mouse model of OA. Additionally, Oh et al. 66 found that the overexpression of Dkk1 had a protective effect on cartilage degeneration in OA mice. In addition, Ye et al. 67 demonstrated that activation of the Wnt/β-catenin pathways may upregulate MMP13 (matrix metallopeptidase-13) expression and lead to NPC degeneration, while DKK1 overexpression specifically blocked the Wnt pathway and inhibited MMP13, to cause protective or even reverse effect on disc degeneration.
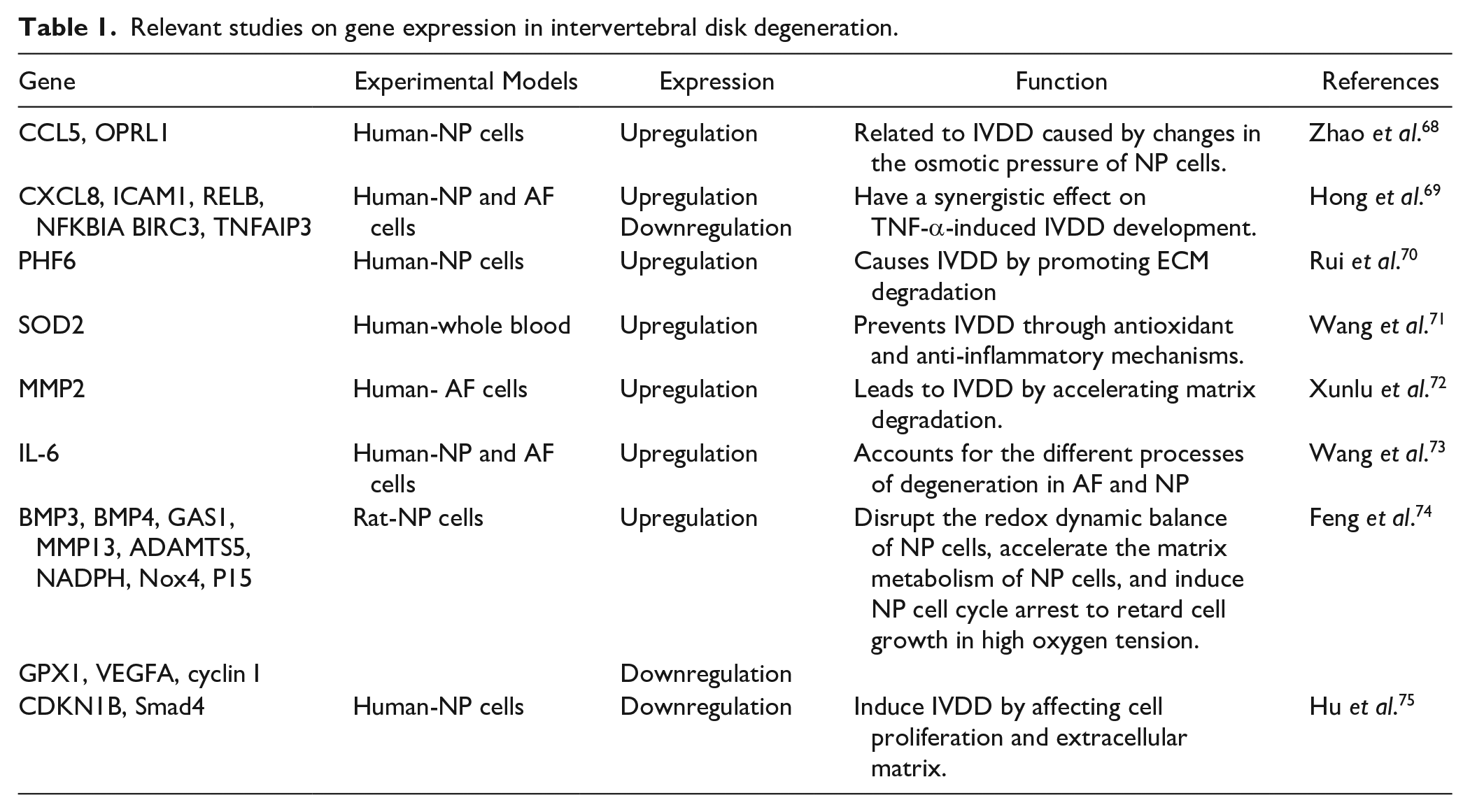
In addition, we have summarized the literature related to IVDD bioinformatic analyses that have been conducted over the past 5 years ( Table 1 ), to further deepen our understanding of IVDD diagnosis and treatment.
Relevant studies on gene expression in intervertebral disk degeneration.
In summary, we have identified several genes related to degenerative disc diseases: CCND1, GATA3, TNFSF11, LEF1, and DKK1. More studies are needed to confirm the therapeutic targets of these genes. Furthermore, bioinformatic analyses will provide new approaches for studying the pathogenesis and treatment of disc degeneration.
Footnotes
Acknowledgments and Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by “the Nature Science Foundation of China” (Grant No. 81672220) and “the Jiangsu Province Science and Technology Support Plan” (Grant No. BE2019669).
Author Contributions
This paper was accomplished based on collaborative work of the authors. Qi Yan: Writing—original draft and writing—review and editing. Quan Xiao: Data curation. Jun Ge, Cenhao Wu, Yingjie Wang, and Hao Yu: Formal analysis, methodology. Huilin Yang: Validation. Jun Zou: Project administration. All authors have read and agreed to the published version of the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
The design of the present study adhered to the tenets of the Declaration of Helsinki and the protocol was reviewed and approved by the Ethics Committee of The First Affiliated Hospital of Soochow University (ECSUFH). (Approval No. 168/2020).
