Abstract
Objective
Meniscus injury and the hypoxia-inducible factor (HIF) pathway are independently linked to osteoarthritis pathogenesis, but the role of the meniscus HIF pathway remains unclear. We sought to identify and evaluate HIF pathway response in normal and osteoarthritic meniscus and to examine the effects of Epas1 (HIF-2α) insufficiency in mice on early osteoarthritis development.
Methods
Normal and osteoarthritic human meniscus specimens were obtained and used for immunohistochemical evaluation and cell culture studies for the HIF pathway. Meniscus cells were treated with pro-inflammatory stimuli, including interleukins (IL)-1β, IL-6, transforming growth factor (TGF)-α, and fibronectin fragments (FnF). Target genes were also evaluated with HIF-1α and HIF-2α (Epas1) overexpression and knockdown. Wild-type (n = 36) and Epas1+/− (n = 30) heterozygous mice underwent destabilization of the medial meniscus (DMM) surgery and were evaluated at 2 and 4 weeks postoperatively for osteoarthritis development using histology.
Results
HIF-1α and HIF-2α immunostaining and gene expression did not differ between normal and osteoarthritic meniscus. While pro-inflammatory stimulation significantly increased both catabolic and anabolic gene expression in the meniscus, HIF-1α and Epas1 expression levels were not significantly altered. Epas1 overexpression significantly increased Col2a1 expression. Both wild-type and Epas1+/− mice developed osteoarthritis following DMM surgery. There were no significant differences between genotypes at either time point.
Conclusion
The HIF pathway is likely not responsible for osteoarthritic changes in the human meniscus. Additionally, Epas1 insufficiency does not protect against osteoarthritis development in the mouse at early time points after DMM surgery. The HIF pathway may be more important for protection against catabolic stress.
Introduction
The hypoxia-induced factor (HIF) pathway has been implicated in cartilage homeostasis and osteoarthritis pathogenesis. HIF-1α is a transcription factor that serves many anabolic functions by maintaining metabolic homeostasis through cell growth and protection, while HIF-2α is implicated in an array of anabolic and catabolic functions.1-8 HIF-2α (encoded by the gene Epas1) shares many metabolic targets with HIF-1α, but its regulation is cell-type specific.3,9-14 Both HIF-1α and HIF-2α are present in articular cartilage.4-7,12-25 In articular chondrocytes, HIF-1α is reported to be active in both normoxia and hypoxia, where this transcription factor controls many chondrocytic genes and has a major role in maintaining cartilage homeostasis.4,6,16,17,20,23
HIF-2α is linked to anti-catabolic processes in human chondrocytes, including increased expression of several matrix-related genes.13,14,23,24 HIF-2α was also linked to the development of osteoarthritis in human, rabbit, and mouse chondrocytes with increased expression and catabolic activity.7,12,21,22,26,27 Heterozygous deletion of Epas1 in Epas1+/− mice provided resistance to surgically induced osteoarthritic changes using the destabilized medial meniscus (DMM) model, but the reason is unknown.7,12,28 Transgenic mice overexpressing Epas1 demonstrated increased osteoarthritic changes in the knee cartilage. Levels of HIF-2α were increased in osteoarthritic cartilage of the DMM mice. 7 Similar trends were seen in human osteoarthritic cartilage: levels of HIF-2α and its targets were present in damaged human osteoarthritic cartilage, while levels were lower in normal and undamaged osteoarthritic cartilage compared to damaged regions of the osteoarthritic cartilage. 7 The role of HIF-1α was not explored in the mouse and is variably reported in human osteoarthritis studies.7,21,22
Osteoarthritis is an enigma in joint disease, but it is increasingly apparent that the disease is a function of whole joint pathology and not solely the articular cartilage. 29 A growing body of evidence supports the link between meniscus injury and osteoarthritis. 30 Although the HIF pathway appears to be active in the meniscus, 15 little is known about the HIFs in meniscal osteoarthritis pathogenesis, and to date, no study has investigated the role of HIF-2α expression and the relationship to osteoarthritis in the meniscus.
The purpose of this study was to identify both HIF-1α and HIF-2α in normal and osteoarthritic meniscus cells and then to identify a potential role for the HIF pathway in the osteoarthritis pathogenesis. Pro-inflammatory stimulation implicated in osteoarthritis pathogenesis was used to determine meniscus responses in context of the HIF pathway. A second goal of this research was to study the meniscus in the context of whole joint pathology in surgically induced osteoarthritis in wild-type and Epas1 knockdown mice. By identifying a potential role for the HIF pathway in osteoarthritis, it may be possible to modify the HIF pathway in multiple joint tissues to modify osteoarthritis pathogenesis.
Methods
Knee Tissue Acquisition
The Wake Forest School of Medicine institutional review board and animal care and use committee approved these protocols. Normal human meniscus specimens (n = 16, age range 25-58 years) were procured through the National Disease Research Interchange (NDRI, Philadelphia, PA) or the Gift of Hope Organ and Tissue Donor Network (Itasca, IL) while osteoarthritic menisci were obtained from patients undergoing total knee arthroplasty for osteoarthritis (n = 31, range 44-83 years). Meniscus tissue was macroscopically graded according to a modified International Cartilage Research Society Cartilage Morphology Score (Table S1).
Human Meniscus Immunohistochemistry
Human meniscus samples were fixed in 4% paraformaldehyde and processed for paraffin embedding. Radial sections (4 µm) were used for histologic evaluation. Samples were deparaffinized and stained for immunohistochemistry or immunofluorescent evaluation. Antigen retrieval was performed with proteinase K (5 mg/mL) diluted 1:250 in proteinaske K buffer (50 mM Tris, 5 mM CaCl2, pH 7) and incubated at 37°C for 15 minutes. The samples were washed with TBS-Tween and then blocked with Avidin/Biotin Blocking Kit (SP-2001, Vector Laboratories). Meniscus sections were incubated overnight at 4°C with either 1:300 anti-HIF-1α (R&D Systems) or anti-HIF-2α (R&D systems). For diaminiobenzidine (DAB) development, samples were washed with TBS-Tween, and the appropriate biotinylated secondary antibody (anti-mouse for HIF-1α antibody and anti-goat for HIF-2α antibody at 1:300 dilution; Cell Signaling Technology) was used and developed according to the manufacturer’s protocol (DAB Peroxidase Substrate Kit, #SK-4100, Vector Laboratories). Sections were then counterstained with hematoxylin for nuclear identification. Negative control shown in Supplementary Fig. 2. Slides were imaged using either light microscopy. Cells that stained positively in the nucleus with peroxidase staining were considered positive for nuclear HIF-1α or HIF-2α for quantification.
Cell Culture and Stimulation
Normal and osteoarthritic human meniscus cells were isolated and cultured as previously described. 31 Briefly, meniscus sections were digested in a sequential manner with Pronase (Calbiochem) and then overnight with collagenase. 32 Monolayer cultures were established by plating cells in 6-well plates at 2 × 106 cells/mL in Dulbecco’s modified Eagle’s medium (DMEM)/Ham’s F-12 medium supplemented with 10% fetal bovine serum. Plates were maintained for approximately 5 days, with feedings every 48 hours until cells reached 100% confluence. Primary cultures were used to preserve the heterogeneous composition of the meniscal cell population. Primary cultures were then incubated overnight in serum-free media (DMEM/F12) and then treated for 6 or 24 hours with one of the following: 10 ng/mL interleukin (IL)-1β; 10 ng/mL IL-6 with 25 ng/mL soluble IL-6 receptor; transforming growth factor (TGF)-α 20 ng/mL (all from R&D Systems); hypoestoxide (25 µM); or 1 µM fibronectin fragments (FnF), a recombinant fragment of fibronectin protein containing domains 7 to 10 of full length fibronectin prepared as previously described. 33 For viral and small-interfering RNA experiments, cells were plated and allowed to acclimate for 48 hours prior to infection.
The Epas1 virus was generated by Vector BioLabs using a cDNA plasmid for Homo sapiens endothelial PAS domain protein 1 (GenBank ID: BC051338.1). The adenoviral shuttle vector contained a GFP coexpression reporter for confirmation of infection and expression in cell culture. The virus was serial amplified in HEK293 cells and then was harvested, titrated, and used in cell culture studies. The HIF-1α virus (ADV-100, Cell Biolabs, Inc.) also contained a GFP reporter and was expanded in culture using HEK293 cells and was titered using QuickTiter Adenovirus Quantitation Kit (VPK-106, Cell Biolabs, Inc.).
For the viral infection, 4 × 109 viral particles in 1 mL serum-free DMEM/F-12 media (Gibco) plus 25 µL of 1 M CaCl2 (Sigma) were used per 106 cells with control samples receiving an empty vector. Cells were incubated for 2 hours at 37°C and then washed twice in 10% media. Fresh 10% media was added to the cells, which were allowed to incubate for 2 days. For the siRNA mediated knockdown, cells were rinsed twice with PBS and changed to 1 mL 10% media without antibiotics for 24 hours at 70% confluence. Cells were transfected with Lipofectamine 2000 (#11668, Invitrogen) according to the manufacture’s protocol. Briefly, 100 pmol of siRNA for nonspecific control, HIF-1α or Epas1 (Cell BioLabs), and 5 µL lipofectamine 2000 were diluted separately in 250 µL Opti-MEM + glutamax 1× reduced serum media 1× (Invitrogen). A total of 250 µL of each diluted siRNA and lipofectamine was incubated together for 20 minutes, and each well of cells received 500 µL transfection reagent and siRNA added to existing media and incubated for 24 hours. Protein lysates of transfected cells were obtained using the Pierce NE-PRE Nuclear and Cytoplasmic Extraction Reagents (78833; Thermo Scientific). RNA was extracted from parallel experiments using the TRIzol method (Invitrogen) according to the manufacture’s protocol.
RNA was quantified (Nanodrop, ThermoScientific) and verified (BioAnalyzer Chip, Agilent) to ensure high-quality RNA (RNA integrity number >6). The reverse-transcription polymerase chain reaction (PCR) generated cDNA (RetroScript Kit, Ambion). Real-Time PCR was performed using the Applied Biosystems 7900HT thermocycler with TaqMan Universal PCR MasterMix and Taqman Gene Assay (Life Technologies; see Supplementary Table S2, primers used are listed in the Appendix). Data were analyzed using the ΔΔCT method normalized to endogenous housekeeping genes in array-specific analysis templates (SABiosciences; http://www.sabiosciences.com/pcrarraydataanalysis.php) in Microsoft Excel (Microsoft).
Immunoblot Analysis
For protein analysis, nuclear and cytoplasmic lysate samples containing equal amounts of total protein (1:1 sample in Lamelli Sample Buffer, 5% 2-mercaptoethanol; BioRad) were loaded onto 10% polyacrylamide gels, separated by SDS-PAGE, transferred to nitrocellulose (Odyssey, Invitrogen) and probed with the primary antibody (anti-Hif-1α [MAB1536, R&D Systems]; anti-Hif-2α [ab20654, Abcam]) and appropriate secondary antibody (anti-mouse for HIF-1α antibody and anti-goat for HIF-2α antibody; Cell Signaling Technology). Blots were visualized with chemiluminescence (Amersham ECL, GE Life Sciences) as previously described 34 or the SuperSignal West Femto Substrate system (#34095; Thermo Scientific). To verify loading of equal amounts of protein, blots were stripped and re-probed with antibodies to Lamin B (#101-B7; Millipore) for nuclear fractions or LDH (#20-LG22; Fitzgerald) for cytoplasmic fractions with the appropriate secondary antibodies (anti-mouse for HIF-1α antibody and anti-goat for HIF-2α antibody; Cell Signaling Technology).
Mouse Destabilized Medial Meniscus Model
C57/BL6129S7 mice (male, 10-12 weeks old, Jackson Laboratories) were genotyped for wild type (WT; Epas1+/+) or Epas1 heterozygote (Epas1+/−) using the primers for allele Epas1tm1Rus (Fig. S1). The heterozygote was chosen because the homozygous Epas1 deletion is embryonic lethal. 3 All controls were wild-type mice littermates. Male mice (WT = 36; Epas1+/− = 30) underwent destabilization of the medial meniscus (DMM) surgery. Mice were anesthetized using 2% isoflurane gas and prepared for aseptic surgery. A parapatellar arthrotomy was performed under microscope visualization, and the medial meniscus and meniscotibial ligament was identified and transected, followed by a layered closure. Animals were monitored and allowed unrestrained movement after surgery. No additional pain medication was needed postoperatively. Mice were euthanized at postoperative week 2 (WT = 17, Epas+/− = 13) and 4 (WT = 19, Epas1+/− = 17) and knees prepared for histology according to the methods previously described.35,36 The evaluating histology expert was blinded to the treatment arm.
Histopathology of DMM Mouse Knees
Knee joints were fixed in 10% formalin, decalcified in 10% ethylenediaminetetraacetic acid (EDTA), processed, embedded intact into paraffin, and sectioned in a coronal plane. Sectioning, staining, grading, and measuring of these sections was performed as previously described in detail.35,36 Representative sections (sections of high quality in a midcoronal location within 100 µm of each other) were evaluated. Two to 4 sections were stained with hematoxylin and eosin (H&E) for evaluation. Evaluations were confined to the tibial plateaus. Measurements included articular cartilage area and thickness (AC thick); subchondral bone area (SCB area) and thickness (SCB thick); areas of chondrocyte cell death (CCD, defined as areas of articular cartilage occupied by 2 or more dead chondrocytes as determined by H&E staining) and percentages of CCD (CCD%) with respect to total articular cartilage area; number of viable chondrocytes (#chond); and total area (Men) and percentage of CCD within the meniscus (MenCCD%). Areas with full-thickness cartilage loss were given “0” for area of CCD, “100%” for CCD%, and “0” for #chond. Last, articular cartilage structure (ACS) was evaluated using semiquantitative scores that ranged from 0 to 12.
Statistical Analysis
All gene expression experiments and mouse data were analyzed using multivariate general linear models with appropriate post hoc testing when indicated (Tukey and Holm-Sidak). Correlations were determined using Spearman’s rho. Statistical calculations were performed in SPSS (IBM) and graphs generated in Prism v6.0 (GraphPad Software, Inc.). Statistical significance was set at P ≤ 0.05.
Results
HIF Expression in Normal and Osteoarthritic Human Meniscus Cells
The presence of HIF-1α and HIF-2α protein in normal and osteoarthritic menisci was evaluated by immunohistochemistry. Both HIF proteins were identified in normal and osteoarthritic tissue (
Fig. 1
). The percentage of positively stained cells was not different between normal and OA: HIF-1α normal (55.3 ± 2.6%, n = 2), osteoarthritic (75.3 ± 11.7%, n = 5); HIF-2α: normal (75.4 ± 10.3%, n = 2), osteoarthritic (83.9 ± 13.7%, n = 5). Results reported mean ± standard deviation. There was also no difference in basal HIF-1α or HIF-2α RNA levels when comparing unstimulated normal and OA meniscus cells in primary cell culture (Group: P = 0.409; Time P = 0.285;
Fig. 2A
and
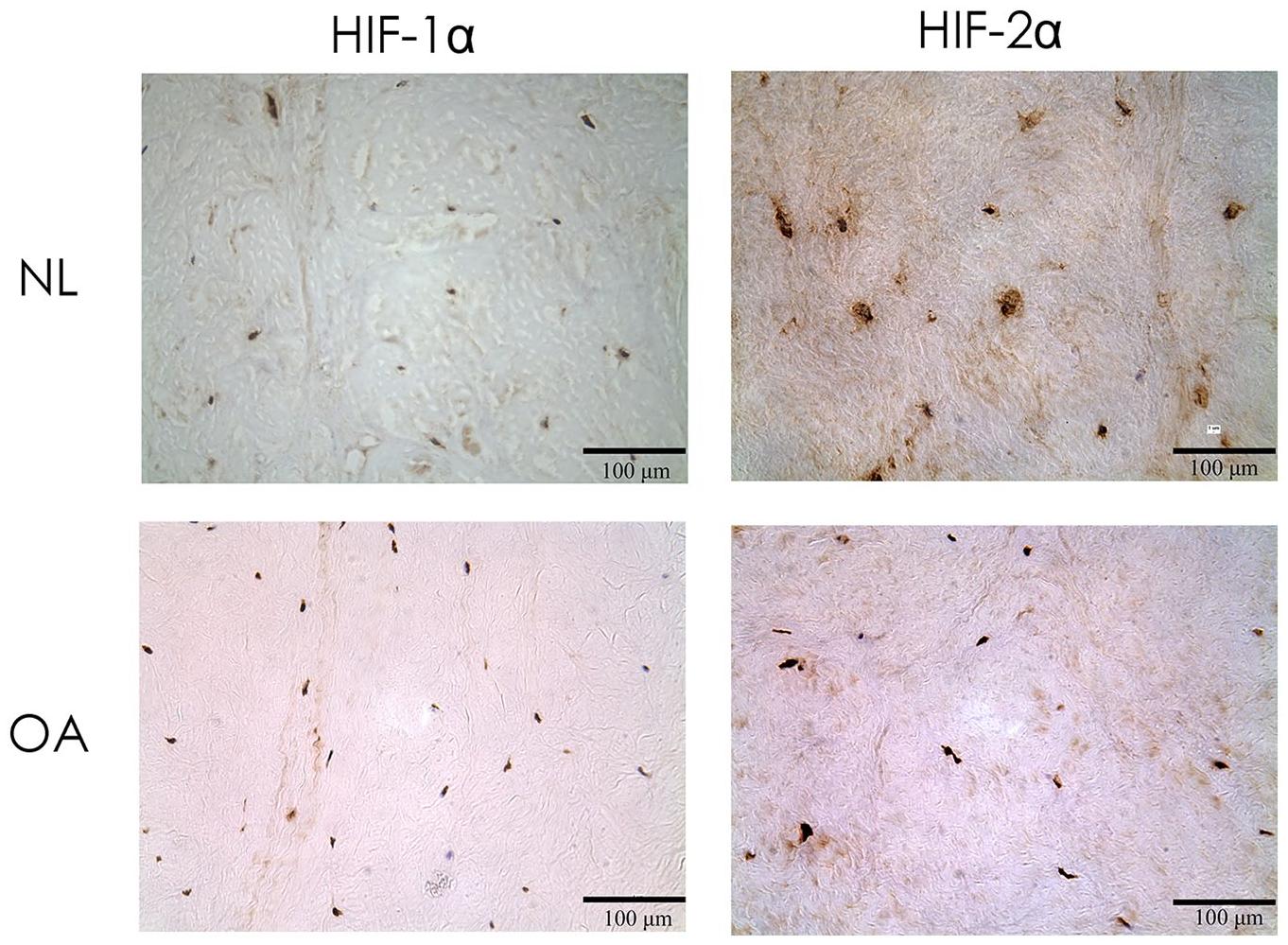
HIF protein staining in normal and osteoarthritic human meniscus. Light microscopy views of HIF-1α and HIF-2α stained sections of normal (NL) and osteoarthritic (OA) meniscus. Protein staining was developed with DAB and counterstained with eosin.
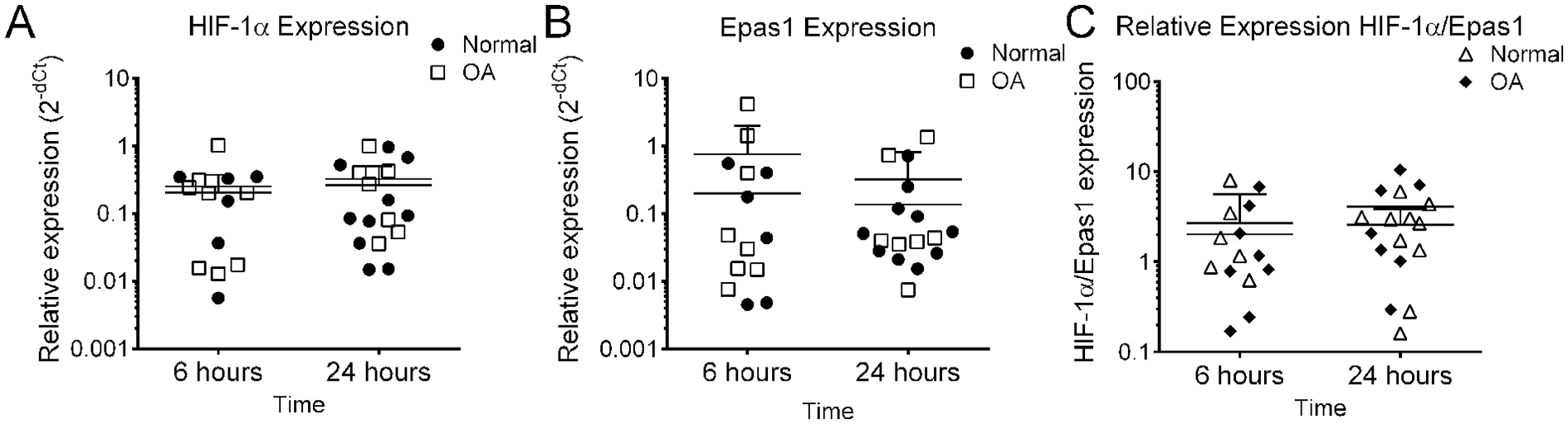
Relative expression of HIF genes in normal and osteoarthritic human meniscus cells. Unstimulated samples were evaluated in concordance with meniscus cell stimulation experiments. (
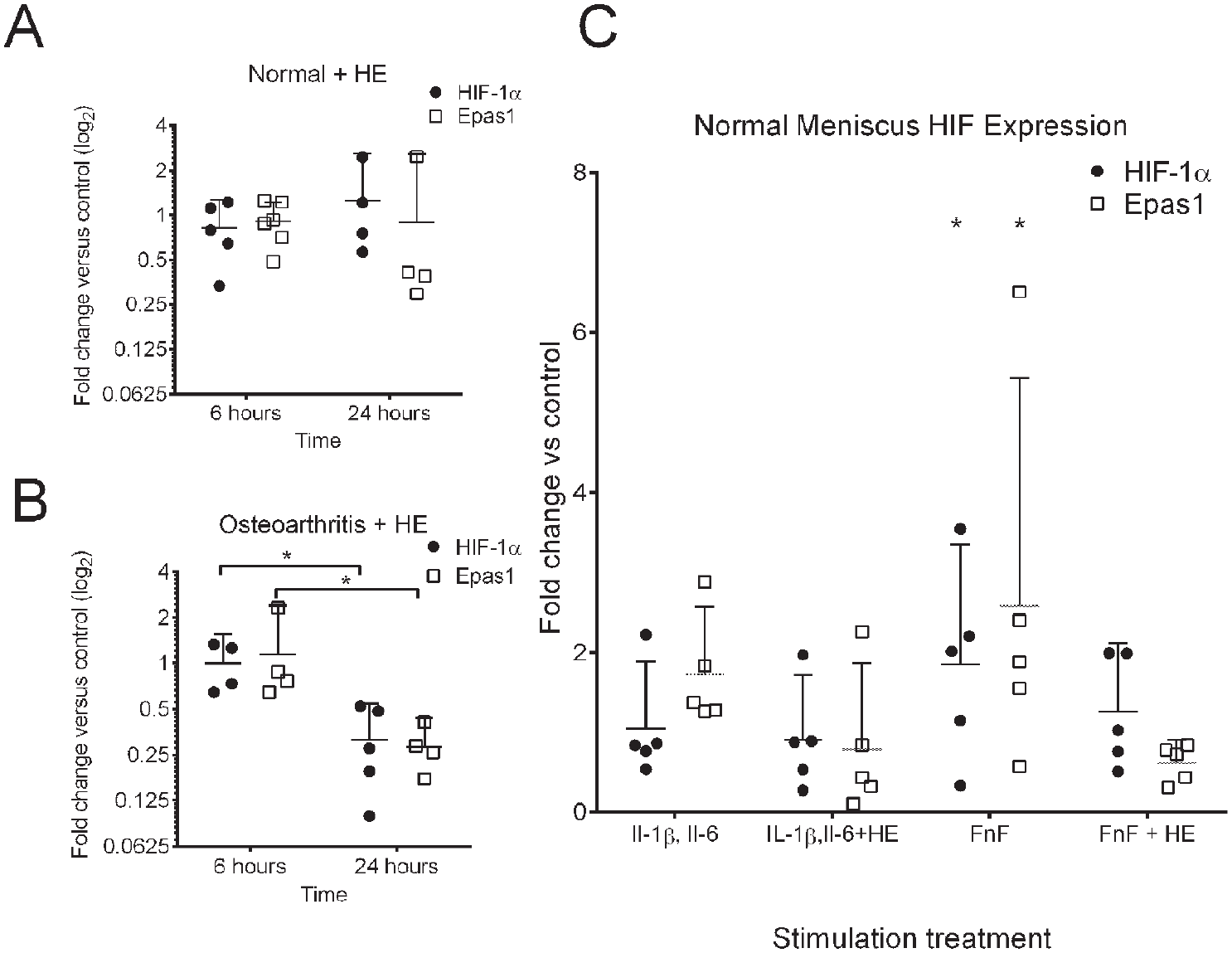
Pro-inflammatory mediators implicated in cartilage osteoarthritis pathogenesis were used to stimulate osteoarthritic meniscus cells to evaluate effects on HIF-1α and Epas1.29,37 Primary cell cultures were stimulated with the pro-inflammatory mediators IL-1β, IL-6, and TGF-α and assessed at 24 hours ( Fig. 3 ). There were no significant effects of any of these stimulants on HIF-1α or Epas1 expression in osteoarthritic meniscus cells.
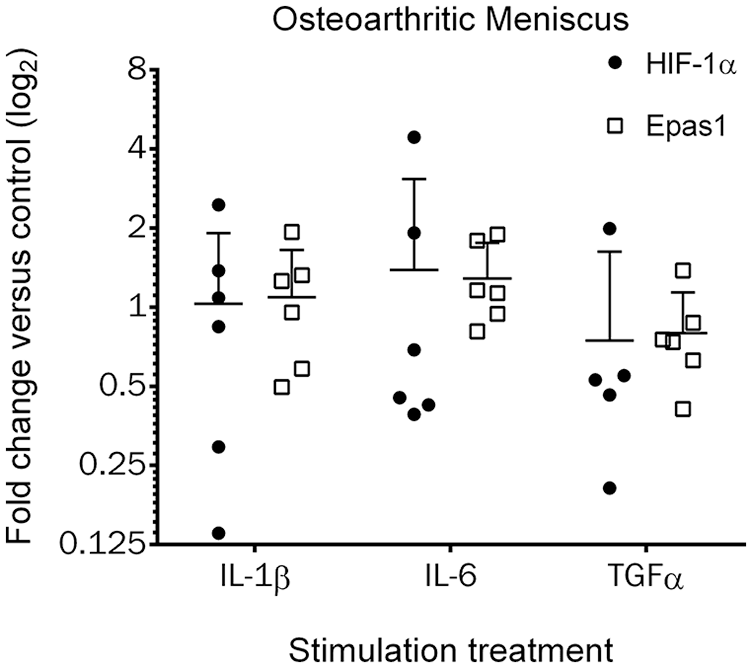
HIF expression in human osteoarthritic meniscus cells following pro-inflammatory stimulation. Primary cell cultures were stimulated with IL-1β, IL-6, or TGF-α and cells were harvested 24 hours after stimulation (n = 6 unique donors; n = 5 unique donors for HIF-1α stimulated with TGF-α). Differences between stimulation were not significant. The line indicates the mean and the error bars the 95% confidence interval.
Gene expression in normal meniscus cells was also evaluated following pro-inflammatory stimulation. In this set of experiments, fibronectin fragments (FnF) were also included as a stimulus since these were previously demonstrated to be pro-inflammatory in normal meniscus and articular chondrocytes. 38 Pro-inflammatory stimulation in normal meniscus cells did not significantly increase HIF-1α or Epas-1 expression ( Fig. 4A ). The effects of pro-inflammatory stimulation on normal meniscal cells were also evaluated on selected genes reported as targets of HIF-1α and/or Epas1. Pro-inflammatory stimulation significantly increased MMP1 expression with fold increases of: IL-1β, 21.6 ± 9.8 (P = 0.002); IL-6, 17.5 ± 7.1 (P = 0.005); and FnF, 24.8 ± 8.4 (P = 0.001; Fig. 4B . MMP3 expression was also increased: IL-1β, 14.4 ± 6.1-fold (P = 0.003); IL-6, 7.46 ± 1.5-fold (P = 0.139); and FnF, 20.0 ± 7.9 fold (P = 0.001; Fig. 4B . Fewer significant increases were identified in anabolic genes, but TGF-α increased collagen type II expression (5.65 ± 2.4-fold, P = 0.010) and FnF significantly increased Sox9 expression (10.3 ± 4.2-fold, P < 0.001; Fig. 4C ).
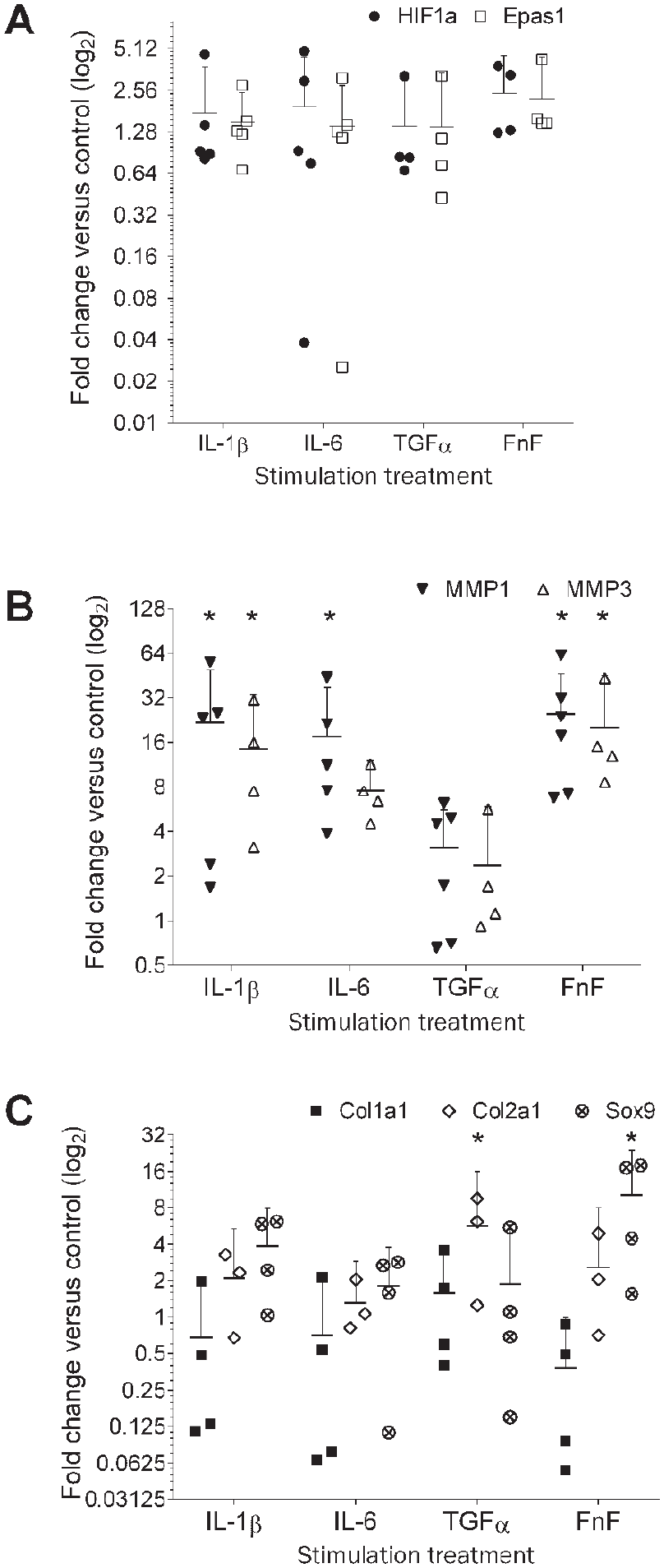
Effects of pro-inflammatory stimulation on HIF, MMP, and anabolic gene expression in normal human meniscus. Primary cell cultures were stimulated with IL-1β, IL-6, TGF-α, or fibronectin fragments (FnF). Cells were harvested 24 hours after stimulation (n ≥ 4 unique donors). (
Meniscus HIF expression in response to pro-inflammatory stimulation was correlated to gene response ( Table 1 ). HIF-1α expression positively correlated with Epas1 expression (R = 0.662, P < 0.001). HIF-1α and Epas1 expression positively correlated with both MMP1 (R = 0.403, P > 0.001; R = 0.408, P < 0.001) and MMP3 expression (R = 0.641, P < 0.001; R = 0.526, P < 0.001). Both HIF-1α and Epas1 positively correlated with Sox9 expression (R = 0.515, P < 0.001; R = 0.700, P < 0.001) and the Sox9 target, collagen type II (R = 0.472, P = 0.003; R = 0.501, P = 0.002). Sox9 was also correlated with collagen type II (R = 0.491, P = 0.002).
Correlation of HIF Expression with Selected Catabolic and Anabolic Genes in Normal Human Meniscus Following Pro-Inflammatory Stimulation.
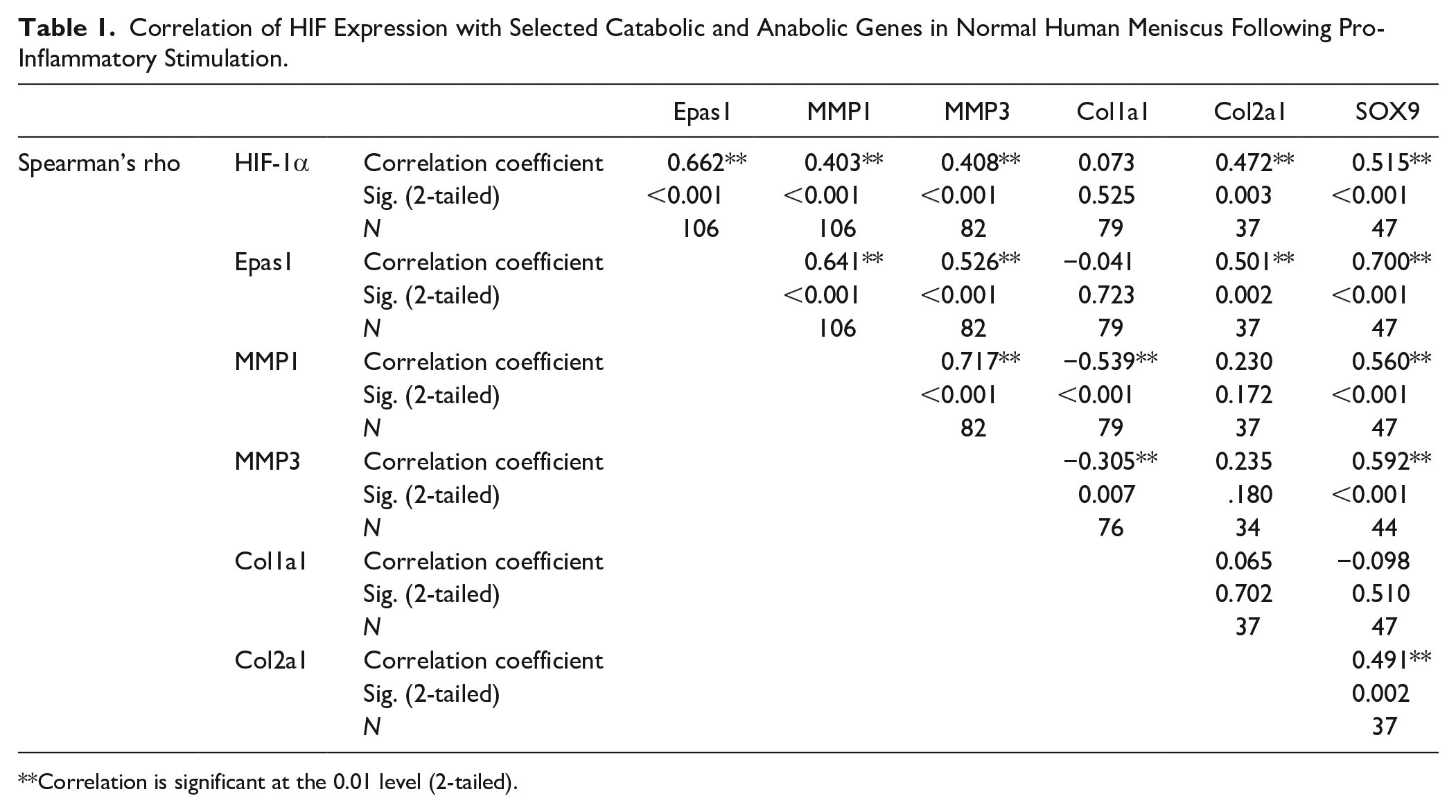
Correlation is significant at the 0.01 level (2-tailed).
HIF Expression and the NF-κB Pathway
We examined HIF expression in normal and osteoarthritic meniscus cells treated with the NF-κB pathway inhibitor hypoestoxide. Treatment with hypoestoxide did not significantly change HIF expression in normal cell cultures ( Fig. 5A ), but by 24 hours HIF expression was significantly reduced in osteoarthritic meniscus cells ( Fig. 5B ; P = 0.023). Normal meniscus cells were then treated with pro-inflammatory stimuli known to increase NF-κB activity with and without the addition of hypoestoxide. Stimulation group effects were significant (P = 0.036) and FnF significantly increased HIF-1α and Epas1 (P = 0.009) expression. The addition of hypoestoxide reduced expression to a level not significantly different from control. The differences between HIF-1α and Epas1 responses were not significant (P = 0.250).
Effects of hypoestoxide on HIF expression. HIF expression in normal (
HIF Overexpression and Knockdown in Normal Human Meniscus Cells
The relationship between HIF-1α and Epas1 and their downstream transcription targets was evaluated with HIF-1α or Epas1 gene overexpression and knockdown. Following gene overexpression or knockdown of HIF-1α or Epas1, nuclear and cytoplasmic fractions of cell lysates were examined for the presence of the target proteins (Supplementary Fig. S3). The adenoviral transduction of HIF-1α or Epas1 successfully increased target protein production with the presence of the protein product located in the nucleus where it is active. Since HIF proteins are difficult to visualize due to their transient nature, siRNA knockdown was confirmed by RT-PCR ( Fig. 6 ).
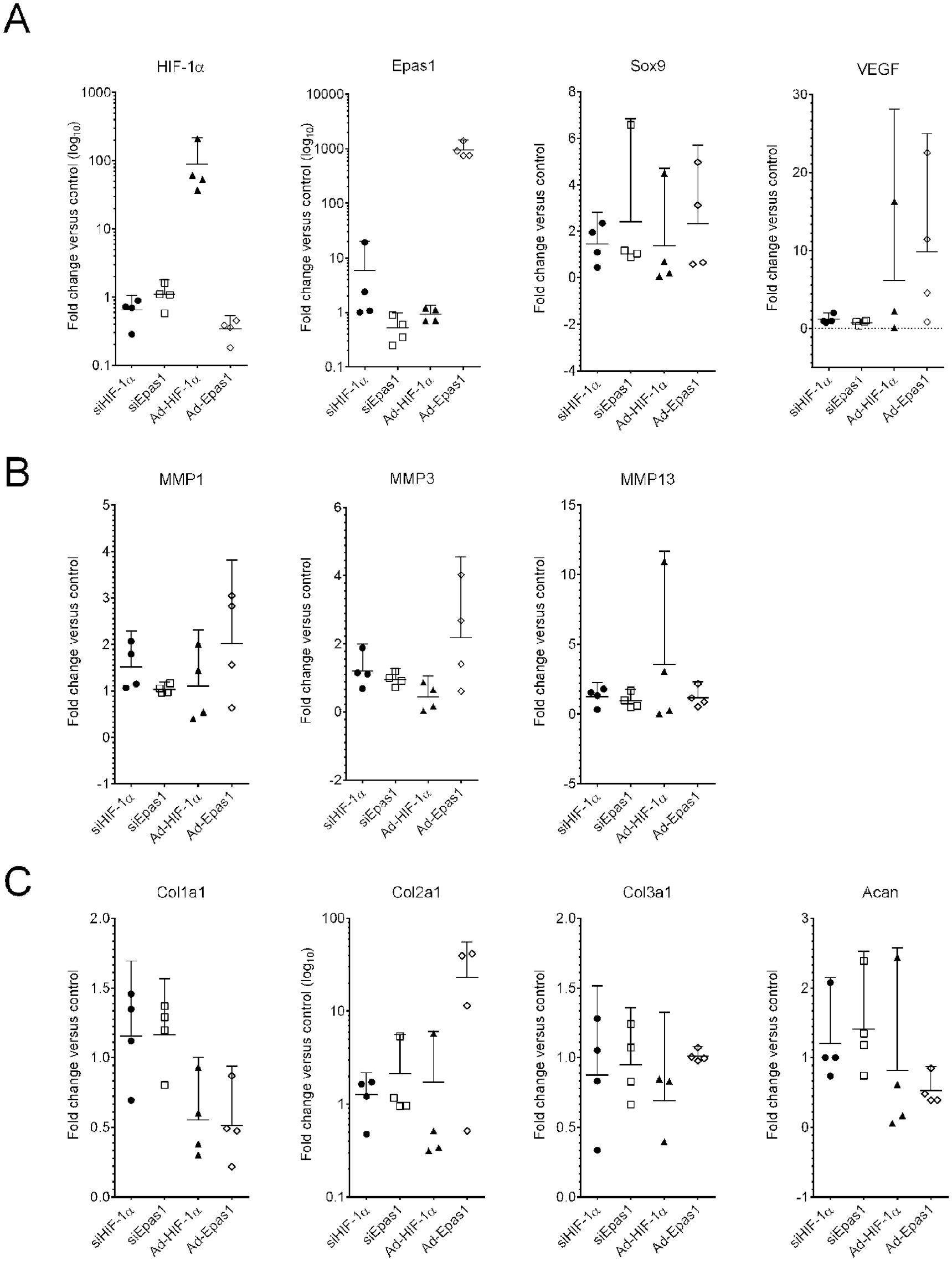
Human meniscus cell response of HIF family and target genes to HIF adenoviral overexpression or siRNA knockdown. (
For overexpression or knockdown experiments, the target genes were expanded to include additional transcriptional targets of HIF-1α or Epas1. Adenoviral overexpression and siRNA knockdown significantly altered the respective expression of HIF-1α (P = 0.009) and Epas1 (P < 0.001) as expected ( Fig. 6 ). While the expression of MMP1, MMP3, or MMP13 were not significantly influenced by HIF knockdown or overexpression, the expression of collagens type I and II were significantly altered (P = 0.014, P = 0.042; Fig. 6 ). Collagen type I expression was decreased by both HIF-1α and Epas1 overexpression, while collagen type II expression was variably increased by Epas1 overexpression. Collagen type III, aggrecan, and Sox9 were not significantly altered.
The HIF target genes were compared to HIF-1α and Epas1 for potential correlations following overexpression and knockdown experiments ( Table 2 ). Significant correlations included a negative correlation between HIF-1α and Epas1 (R = −0.609, P = 0.012). Epas1 positively correlated with the known HIF target, VEGF (R = 0.546, P = 0.035). Both aggrecan (Acan) and collagen type II correlated with Sox9 (R = 0.756, P = 0.001; R = 0.582, P = 0.018). The remainder of the correlation coefficients and significance values are delineated in Table 2 .
Correlation of Selected Genes Following HIF-1α and Epas1 Overexpression and Knockdown in Normal Human Meniscus Cells.
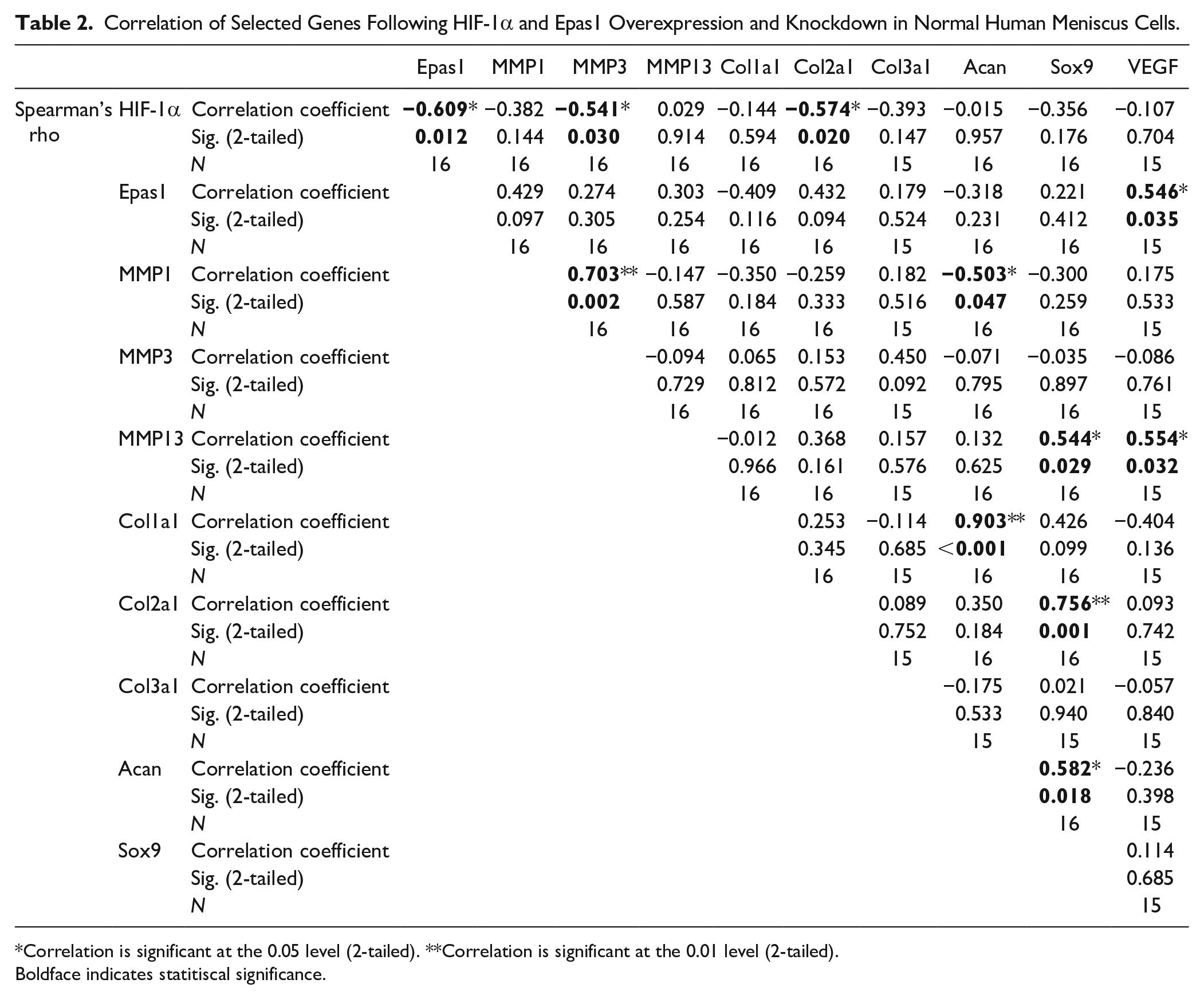
Correlation is significant at the 0.05 level (2-tailed). **Correlation is significant at the 0.01 level (2-tailed).
Boldface indicates statitiscal significance.
Effect of Epas1 Heterozygous Deletion on the Mouse Destabilized Medial Meniscus Model of Osteoarthritis
The role of HIF-2α in the development of osteoarthritis was further explored using the established mouse DMM model in Epas1 heterozygous deficient mouse. Knee histology sections were evaluated 2 and 4 weeks postoperatively, and representative histologic images from the 4-week time point are shown in Fig. 7A . As evidenced by both the histology sections and the ACS scores, the DMM model was successful in producing osteoarthritic changes ( Fig. 7A , Table S3). Cartilage thinning, fissuring, and chondrocyte cell death with nuclear dropout were all evident in the medial compartment. The main effects were not significant for mouse genotype or time point. Osteoarthritis severity was significantly different for the medial versus lateral tibial plateaus (P < 0.001). Compartment differences were significant for the following variables: ACS score (P < 0.001), articular cartilage area (P < 0.001), articular cartilage thickness (P = 0.001), subchondral bone area (P < 0.001), subchondral bone thickness (P < 0.001), chondrocyte cell density (P < 0.001), chondrocyte number (P < 0.001), and meniscus area (P < 0.001). There were no differences between compartments in meniscus cell density or meniscus cell viability. These differences were not significant between gene type. A complete list of results for all variables is presented in Table S3.
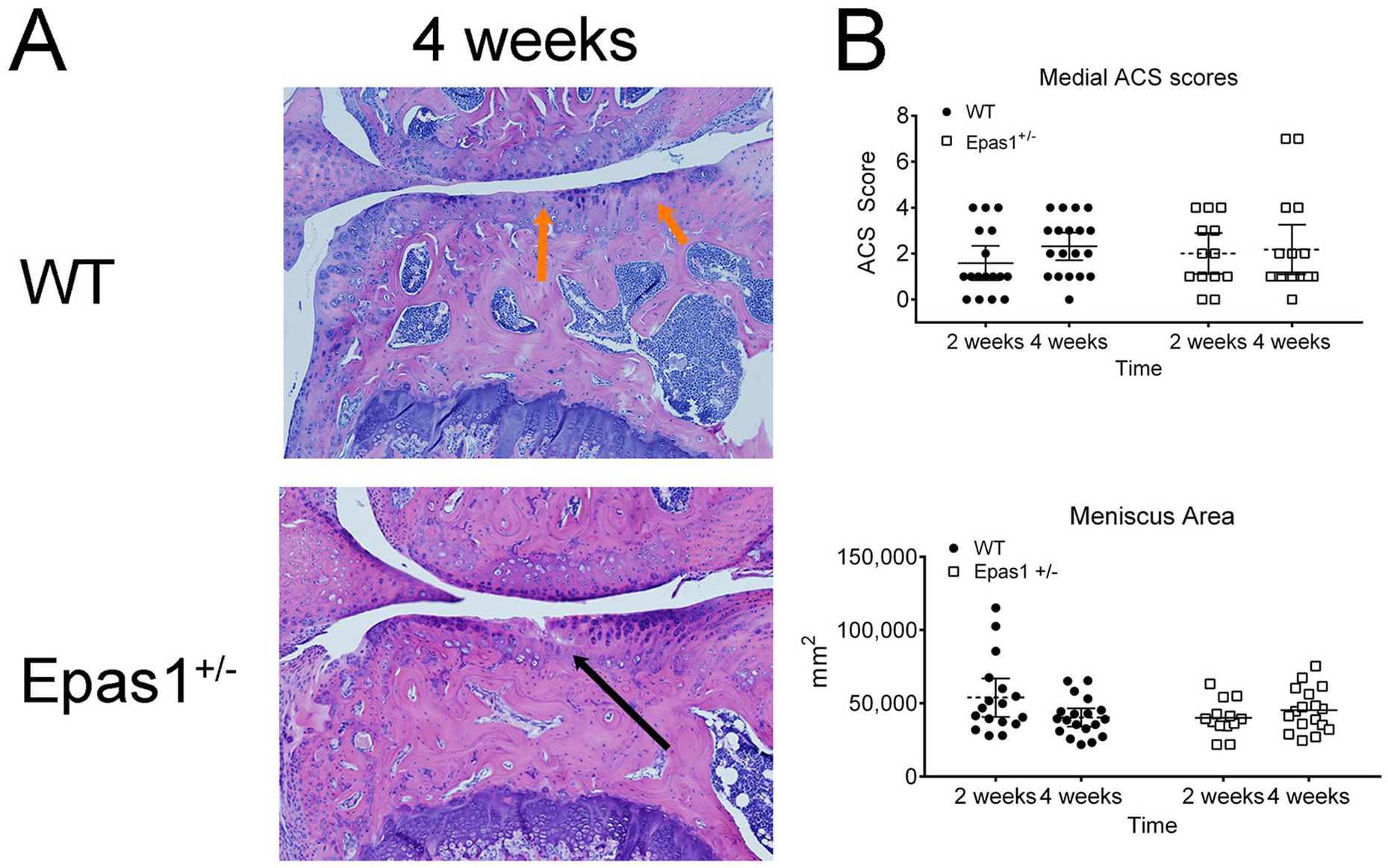
Representative medial compartment histology and analysis of following destabilized medial meniscus surgery in mice. (
Discussion
The HIF pathway and its regulation in joint tissues is complex and poorly defined, particularly in the meniscus. The principle finding of the present study was that HIF-1α and HIF-2α could not be directly linked to the development osteoarthritis in human meniscus cells and HIF-2α (Epas 1) knockdown was not protective in the DMM model of OA at the time points tested. We found similar expression of HIF-1α and HIF-2α in both normal and osteoarthritic meniscus tissue, which was contradictory to the reports in osteoarthritic human and mouse cartilage.7,21,22 Furthermore, we identified that HIF-1α and HIF-2α regulation were not strictly inversely related, but instead demonstrated variable responses to catabolic stimuli and complementary responses in downstream targets of catabolism and anabolism.
Pro-inflammatory stimulation was expected to increase Epas1 to a greater degree than HIF-1α expression; however, pro-inflammatory cytokines IL-1β and IL-6 did not increase HIF-1α or Epas1 expression in meniscus cells. We found that treatment with FnF increased both Epas1 and HIF-1α expression. Cytokine stimulation did not alter the expression level of Epas1 in meniscus cells as was previously described in chondrocytes.7,12,26 Yang et al. described Epas1 induction in rabbit articular chondrocytes by multiple pro-inflammatory cytokines, including TNF-α, IL-1β, IL-17, and IL-21, but did not place these results in the context of HIF-1α. 7 HIF-1α is also demonstrated to be increased by pro-inflammatory stimulation.17,23,25,39 While some studies linked the increased expression of Epas1 to increasing catabolic activity,7,12,21,22 other authors also identified catabolic stress induction of HIF-1α as a protective and possible repair response in chondrocytes.4,6,17,20,23-25 Both HIF-1α and HIF-2α were demonstrated to control energy production and share many of the same targets which was theorized to improve tissue repair.3,40,41 Furthermore, HIF-1α was predominantly thought to control anabolic expression in chondrocytes and meniscus cells. 15
HIF-1α expression positively correlated with Epas1 following stimulation, which suggests a common pathway of induction or synergistic effects. Both HIF-1α and Epas1 positively and significantly correlated with MMP1 and MMP3 expression in the present study, which were previously reported to be targets only of Epas1 in osteoarthritic chondrocytes7,12,26; however, the increase in stimulated meniscus cells may be attributable to a HIF-independent pathway. We previously established that pro-inflammatory stimulation in normal and osteoarthritic meniscus cells increased MMP1 and MMP3 transcription and translation, and MMP increases were linked to multiple inflammatory pathways, including NF-κB.29,34,37,42 The level of increased MMP expression following pro-inflammatory stimulation far exceeded the level of increase seen with Epas1 overexpression in normal meniscus. The lack of correlation in stimulation studies and the more profound increases in MMP expression with pro-inflammatory stimulation suggests that HIF-independent pathways are likely responsible for enhanced MMP production.
The HIF relationship to anabolic factors exhibited a more complex regulation than expected. HIF-1α and Epas1 expression in normal meniscus cells was positively correlated with both collagen type II expression and Sox9 expression following pro-inflammatory stimulation. Epas1 overexpression increased collagen type II expression. Collagen type II is a major constituent of cartilage extracellular matrix, but is also reported in the inner zones of the meniscus. 27 Sox9 is an important developmental gene found to be responsible for the phenotypic development of both meniscus and cartilage cells and upregulates chondrocytic matrix genes, including Col2a1, Col9a1, Col11a2, and aggrecan.13,24 The regulation of Sox9 in both meniscus cells and articular chondrocytes is unclear. HIF-1α has been demonstrated to have a significant role in Sox9 regulation but much of this work was either completed in osteoarthritic cartilage and meniscus or in the mouse.5,15,20 More complete studies in human articular cartilage suggested that human tissue responds differently from that of mice.23,24,28 Sox9 induction was dependent on HIF-2α and not on HIF-1α in human articular chondrocytes. 13 Increased HIF-2α in human chondrocytes is also linked to increased Sox9 expression and increased expression of chondrocytic extracellular matrix genes, including collagen type II and aggrecan.13,24 These increases in cartilage specific genes were seen even in normoxia. 24 HIF-2α is also linked to the control of additional anti-catabolic responses in healthy human articular chondrocytes through upregulation of Sox9 independent genes, chondrocyte regulators Mig6 and inhibin A (InhbA), cartilage matrix protein chondromodulin-1 (Chm-1), and cartilage growth factor Gdf10.14,43 These factors are linked in varying degrees to anabolic functions in chondrocytes and meniscus cells. Our findings placed in context of the reported HIF family targets support that HIF activity could be induced by catabolic stress but respond with an anabolic and anti-catabolic response.
The role for the NF-κB pathway in regulating the HIF pathway remains poorly defined. We observed a decrease in both HIF-1α and Epas1 expression following NF-κB inhibition in osteoarthritic meniscus. The decrease is consistent with previous descriptions of NF-κB regulation, since both genes have been shown to be stabilized or upregulated at least in part through the NF-κB pathway.7,12,17,18 While HIF-1α and Epas1 do have interactions with the NF-κB pathway, the decrease in HIF expression following NF-κB inhibition may be the result of inhibiting a larger cellular repair response in diseased or stressed tissue and less of consequence of NF-κB activation.19,23
The variability in HIF gene expression and the lack of clear correlations with reported targets is likely multifactorial. While HIF expression is reported to be influenced by cytokines,7,22,44 much of the regulation may occur at the protein level and not the expression level. 39 Proteosome degradation of the HIFs is well described,19,24 but its methods for activation appear complex. 40 HIF levels can be heavily influenced by prolyl hydroxylation as demonstrated in some studies,23,24,28 and may not be stabilized by inflammatory stimulation to the degree originally proposed in recent studies.5,7,12,16,25 The sensitivity in HIF regulation likely accounts for the reciprocal changes in HIF-1α and Epas1 expression following viral infection.
The heterogeneous composition of the meniscus may also contribute to our observed variability. The meniscus is composed of both chondrocytic and fibroblastic cell types. If the primary role of the HIF family is to promote chondrocyte differentiation, then responses of the HIF family within the individual cell types may be confounded by the behavior of inner versus outer meniscus cells and fibroblasts.15,45,46 The response of collagens to HIF-2α and induction of Sox9 expression may promote a more chondrocytic phenotype in meniscus cells, while HIF-1α downregulation of collagen type I, the primary meniscus extracellular matrix component may further drive a chondrocytic phenotype.5,14,16,17 Although the HIF family proteins may not be as important in osteoarthritis pathogenesis in meniscus cells, it appears that they have a very significant role in chondrocyte phenotype and differentiation.13,14,19,23,24,28,41 Future studies in human meniscus cells may place a greater emphasis on the role of the HIF pathway in development, maintenance, and repair of inner versus outer meniscus cells.
The DMM model of osteoarthritis was used to examine the effect of Epas1 deficiency on preventing osteoarthritic changes. We did not see a significant difference between genotypes in joint protection at either 2 or 4 weeks after DMM surgery. Our results were inconsistent with the chondroprotective effects of Epas1 insufficiency reported by previous authors at 8 weeks after DMM surgery.7,12,22 Osteoarthritis development by 4 weeks in our study confirmed technical adequacy of the DMM surgery, and early osteoarthritic changes were both temporally and histologically congruent with previous reports.35,36 Our histologic grading system was more rigorous than that previously used and consequently identifies tissue specific and more subtle early disease alterations.35,36 The quantitative measurements acquired from H&E staining are more sensitive than those from safranin-O staining and are extensively characterized35,36,47-49; since the meniscus is fibrocartilage, the meniscus does not stain well with safranin-O staining.35,36 The chondroprotective effects of Epas1 insufficiency could theoretically be more evident at 7 to 8 weeks7,12; however, spontaneous reversal of cartilage degradation is unlikely. Joint destruction in mice following local Epas1 transfection may be related to a complete distortion and overload of the HIF pathway, which is tightly regulated and highly sensitive.3,40 Reports in mice also linked HIF-2α’s role in regulating the pro-inflammatory cytokine IL-6 production and articular chondrocyte apoptosis.21,22 Epas1 may also not be as important in human osteoarthritic changes as evidenced by a large cohort study that refuted linking a functional single nucleotide polymorphism in the human Epas1 gene to osteoarthritis in a Japanese population.50,51 Although the objective of our mouse model was not to examine all effects of Epas1 insufficiency, our results suggest that Epas1 insufficiency does not significantly offer protection of the chondral surfaces nor influence meniscus appearance in early disease.
The current study carries common limitations of laboratory models. Cell culture responses may not directly mimic those of the intact meniscus in its native environment, but the use of primary cell culture provided an effective modality for precise stimuli control and measurement of gene expression and protein production. Cell culture studies were conducted in normoxia, which does impact hypoxia sensitive genes. Previous reports suggested that HIF proteins are stabilized by pro-inflammatory stimulation and was still active under normoxic conditions.7,16,17 Cell culture experiments performed by Yang et al. in rabbit articular chondrocytes responded similarly to pro-inflammatory stimuli in both normoxia and hypoxia. 7 It is possible to miss rapid transcription alterations, but our evaluation time points were selected based on HIF increases in chondrocytes by 6 to 24 hours.4,5,17,25 Not all gene targets were confirmed with protein translation, but the HIF family proteins are transcription factors and should be able to demonstrate their activity by increase in target gene expression. In our analysis of downstream factors, we structured the statistical analysis as normalized to controls with the goal of finding a difference. Despite using a less conservative approach, we did not find an association with downstream targets. Our limited sample size for this analysis precluded alternative statistical measures, and we recognize this strategy as a limitation, albeit an intention one. As aforementioned, meniscus cells are a heterogeneous population and our results cannot be directly extrapolated to articular chondrocytes.
The regulation and influence of the HIF family genes is exceedingly complex. Multiple factors have been demonstrated to influence both their expression and activation, including catabolic stress, hypoxia and normoxia, and tissue types.1,2,4-11,13-25,28,35,40,41 The balance between HIF-1α and HIF-2α is much more likely to be responsible for effecting significant transcriptional changes, and this balance is controlled by a number of factors, including oxygen tension, tissue type, catabolic stress, and perhaps most importantly by species.1,2,4,6,8,10,13-16,18-20,23-26,28,40 Furthermore, the role of HIFs in the meniscus may be more related to phenotype differentiation, based on the number of chondrocytic targets it has than meniscal degeneration. Much of the evidence that examined HIF-1α and HIF-2α together, identified that the HIFs were responsible for both anabolic and anti-catabolic effects.3,13,14,19,23,24,41 Our results are more congruent with the theory that the 2 HIFs are dynamic and are not primarily responsible for osteoarthritic changes in the meniscus. Future emphasis should be placed on exploring the potential repair functions in HIF-mediated anabolism and anti-catabolism in both meniscus and cartilage tissue. HIF-1α and HIF-2α could not be directly linked to the development osteoarthritis in human meniscus cells and HIF-2α (Epas 1) knockdown was not protective in the DMM model of OA at the time points tested.
Supplemental Material
HIF_SupplementaryInformation_Revised – Supplemental material for Role of the Hypoxia-Inducible Factor Pathway in Normal and Osteoarthritic Meniscus and in Mice after Destabilization of the Medial Meniscus
Supplemental material, HIF_SupplementaryInformation_Revised for Role of the Hypoxia-Inducible Factor Pathway in Normal and Osteoarthritic Meniscus and in Mice after Destabilization of the Medial Meniscus by Austin V. Stone, Richard F. Loeser, Michael F. Callahan, Margaret A. McNulty, David L. Long, Raghunatha R. Yammani, Sara Bean, Kadie Vanderman, Susan Chubinskaya and Cristin M. Ferguson in CARTILAGE
Footnotes
Acknowledgments and Funding
The authors would like to thank Dr. Cathy Carlson at the University of Minnesota for her assistance in mouse specimen processing. We would also like to thank the Wake Forest School of Medicine Orthopaedic Surgery Joint Service for their assistance in meniscus specimen acquisition. We would like to thank the National Disease and Research Interchange (NDRI) and Dr. Arkady Margulis at Rush Medical Center for procuring meniscus from the Gift of Hope Organ and Tissue Donor Network. We also would like to acknowledge donor’s families. The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by the Young Investigator Grant from the American Orthopaedic Society for Sports Medicine (Stone) and the clinician scientist grant from the Orthopaedic Research and Education Foundation (Stone). Additional support was received from the NIH/NIAMS K08AR059172 (Ferguson) and NIH/NIAMS R37 AR049003 (Loeser and Rush Klaus Kuettner Endowed Chair for Osteoarthritis Research; Chubinskaya).
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
The Wake Forest School of Medicine institutional review board approved the protocols.
Animal Welfare
The Wake Forest School of Medicine animal care and use committee approved the protocols.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
