Abstract
Objectives:
RNA isolation is necessary for the evaluation of gene expression. Due to the nature of its extracellular matrix, RNA isolation from articular hyaline cartilage is difficult and thus the tissue is commonly enzymatically digested in order to extract RNA from the obtained chondrocytes. We hypothesized that the digestion process affects the expression levels of common cartilage-associated genes.
Design:
Expression of cartilage-associated genes was compared between intact cartilage and digested chondrocytes from weight bearing and non-weight bearing regions of the equine fetlock joint.
Results:
The gene expression of SOX9, COL1A2, COL2A1, ACAN, and COLX were analyzed. Digested cartilage showed a significant decrease in the expression of COL1A2, COL2A1, and ACAN compared to intact cartilage in both joint regions, and an increase in COLX expression in non-weight bearing cartilage only.
Conclusions:
Enzymatic digestion of cartilage significantly impacts gene expression profile. We conclude that while RNA isolation from intact cartilage is more technically difficult, determination of gene expression should be conducted on intact cartilage if true representation of the in vivo processes is sought.
Introduction and Summary
Assessment of cartilage gene expression is fundamental to many aspects of cartilage biology: development, disease progression and repair processes, as well as tissue engineering. Unfortunately, isolation of good-quality RNA from articular cartilage remains a key challenge to investigators. Low chondrocyte content (1%-5% of tissue mass), highly cross-linked extracellular matrix (ECM) molecules, and a high proteoglycan content 1 make it particularly difficult to extract RNA from cartilage tissue using routine procedures. To work around this, ECM is often enzymatically digested to release chondrocytes, increasing RNA yield. 2 However, digestion of cartilage may alter gene expression. Bovine articular cartilage subjected to varying collagenase digestion times showed variations in key genes such as COL2A1, ACAN, and GAPDH. 3 Similar findings have also been reported in bovine meniscus tissue, where gene expression patterns more characteristic of a fibrocartilage phenotype were observed in enzymatically digested versus intact tissue. 4 Therefore, RNA isolation from enzymatically digested cartilage may not be representative of real-time chondrocyte gene expression in vivo, though a direct gene expression comparison between intact and digested articular cartilage is yet to be performed.
In this study, we evaluated the expression of cartilage-associated genes from intact and digested equine articular cartilage from the fetlock joint. We hypothesized that enzymatic digestion would result in differences in cartilage-associated gene expression compared with nondigested cartilage. Digested cartilage from 2 distinct regions showed a significant decrease in the expression of COL1A2, COL2A1, and ACAN compared with intact cartilage, whereas COLX expression was increased in digested cartilage in one region only (nonweightbearing region). Thus, enzymatic digestion of equine cartilage significantly impacts gene expression profile. We conclude that while RNA isolation from intact cartilage is more technically difficult, determination of gene expression should be conducted on intact cartilage if true representation of the in vivo processes is sought.
Methods, Results, and Discussion
Equine articular cartilage samples were collected from weightbearing (distal medial/lateral trochlear surfaces and sagittal ridge) and nonweight bearing regions (dorsal and palmar medial/lateral trochlear surfaces and sagittal ridge) of the fetlock joint of cadaver limbs of adult horses (n = 5, ages 3-20 years) euthanized for reasons not related to this study. Samples procured postmortem do not require an approved animal care utilization protocol and hence the cartilage collection adhered to institutional guidelines. Cartilage samples were either snap frozen in liquid nitrogen for RNA isolation from intact cartilage or minced and immediately digested at 37°C overnight in Dulbecco’s modified Eagle medium (DMEM)/F12 medium (Sigma-Aldrich, Oakville, Ontario, Canada) containing 1% type I collagenase (Sigma). Chondrocytes were retrieved by straining the digested sample through a 70-µm filter and immediately subjected to RNA isolation. Snap-frozen cartilage samples were placed in a precooled stainless-steel canister with a 25-mm stainless steel grinding ball, then immersed in liquid nitrogen for 2 minutes. The tissue was then disrupted using an oscillating milling machine (Retsch Mixer Mill MM 400; RETSCH; Haan, Germany) for 2 to 3 minutes at 30 Hz.
Total RNA was isolated from both intact and digested cartilage tissue using the mirVana total RNA isolation kit (Life Technologies) and purified using the RNA Clean and Concentrator Kit (Zymo Research; Irvine, CA, USA). RNA was DNase-treated using the PerfeCTa DNase I Kit (Quanta Biosciences; Beverly, MA, USA) then reverse transcribed using the qScript cDNA Synthesis Kit (Quanta BioSciences; Beverly, MA, USA) following the protocols as outlined by the manufacturer.
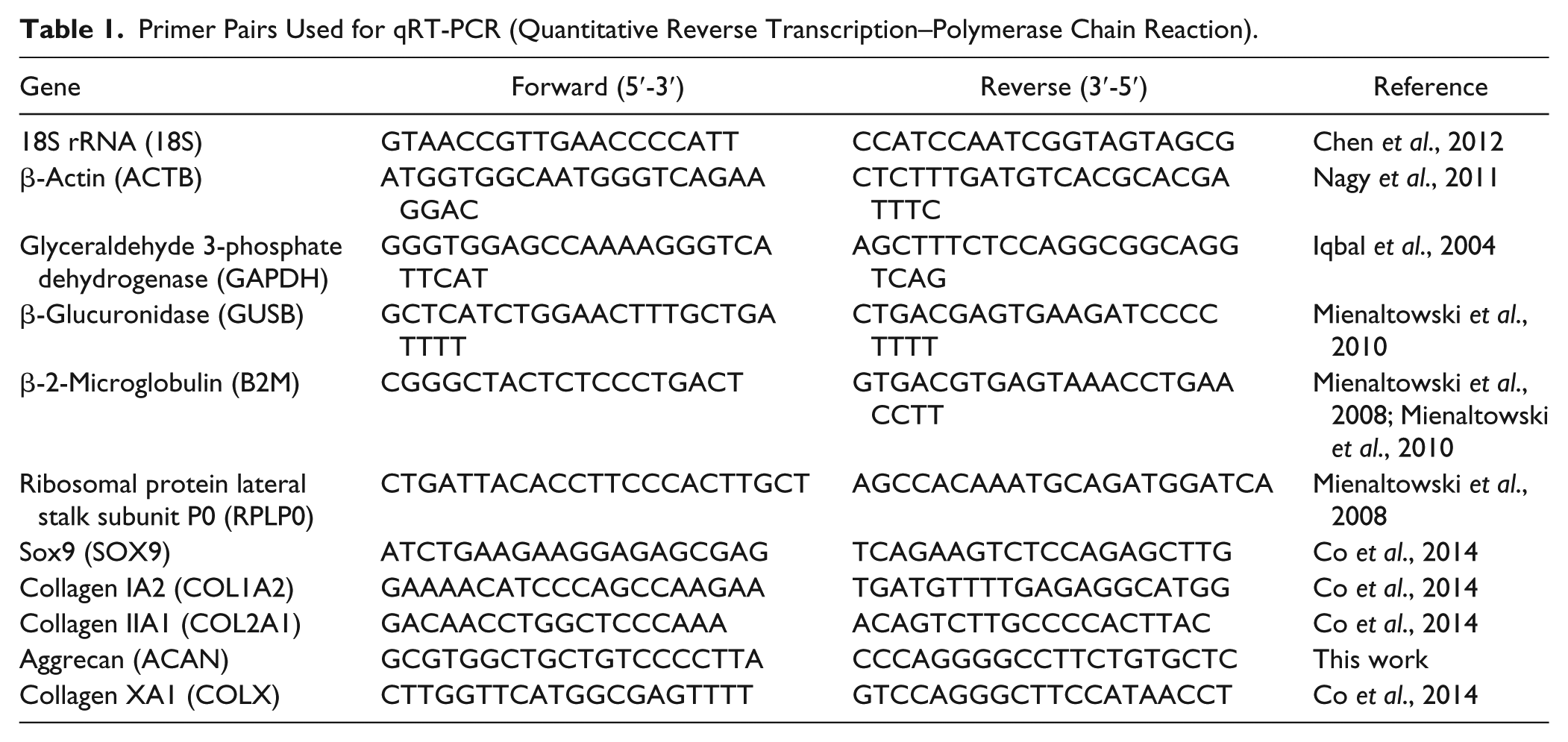
Gene expression of SOX9, COL1A2, COL2A1, COLX, and ACAN was assessed in digested and intact cartilage via qRT-PCR (quantitative reverse transcription–polymerase chain reaction) on the CFX96 Touch Real-Time PCR detection system (Bio-Rad, Hercules, CA, USA; primers listed in Table 1 ). Expression levels of genes of interest were calculated relative to the reference genes previously established as being stable for the selected samples using NormFinder (RPLP0 and B2M; Supplementary Fig. 2). Expression levels were normalized to intact cartilage samples, and gene expression analysis was performed using the 2−ΔΔCT method. Data were analyzed using a 2-way analysis of variance and presented as means ± the 95% confidence interval. Significance was assigned at P < 0.05. Comparisons were made between intact and digested cartilage samples in weightbearing and nonweightbearing regions. The Tukey-Kramer method was used for post hoc analysis.
Primer Pairs Used for qRT-PCR (Quantitative Reverse Transcription–Polymerase Chain Reaction).
We observed marked differences in gene expression between intact and digested cartilage in 4 of the 5 cartilage-associated genes evaluated. SOX9 expression was unaffected by enzymatic digestion; expression levels were similar between intact and digested cartilage samples in both weightbearing and nonweightbearing regions of the joint (P = 0.690 and P = 0.806, respectively; Fig. 1 ). In contrast, digested cartilage showed a decrease in COL1A2, COL2A1, and ACAN expression compared with intact cartilage samples in both weightbearing and nonweightbearing regions (P < 0.05 for all 3 genes in both regions), indicating that 24 hours of digestion changed the expression or detection of mRNAs encoded by these genes. No significant interaction was found between RNA isolation method and joint region on the expression of these genes. SOX9 is a key transcription factor in chondrogenic development and has important physiological roles in mature cartilage tissues.5,6 In contrast to its downstream targets COL1A2, COL2A1, and ACAN, SOX9 expression appears to be less affected by ECM disruption, whereas enzymatic digestion appeared to cause a decrease in the expression of these genes. Several plausible explanations exist with regards to the causes of the observed differences in expression levels. These include both ex vivo events that may occur during the digestion period in living cells and/or differences in global transcript levels purified by the RNA extraction protocol in intact versus digested tissue. For example, there are multiple regulators and effectors of COL1A2, COL2A1, and ACAN independent of SOX9 (such as SOX5/SOX6) that may be affected by digestion, possibly causing their observed decrease in expression.7-9 These findings are consistent with another study that observed no difference in SOX9 expression in cartilage and chondrocytes in monolayer culture. 10 This and another study also observed lower levels of COL2A1 and ACAN in chondrocytes cultured in monolayer,10,11 though this was accompanied by an increase rather than a decrease in the levels of COL1A2. However, these observations were made after 7 days or multiple passages in culture,10,12 whereas our healthy chondrocytes were immediately subjected to RNA isolation and not cultured. It is possible that enzymatic digestion induces an early dedifferentiated chondrocyte phenotype, as the cells show decreased COL2A1 and ACAN, but not yet an increase in COL1A2.
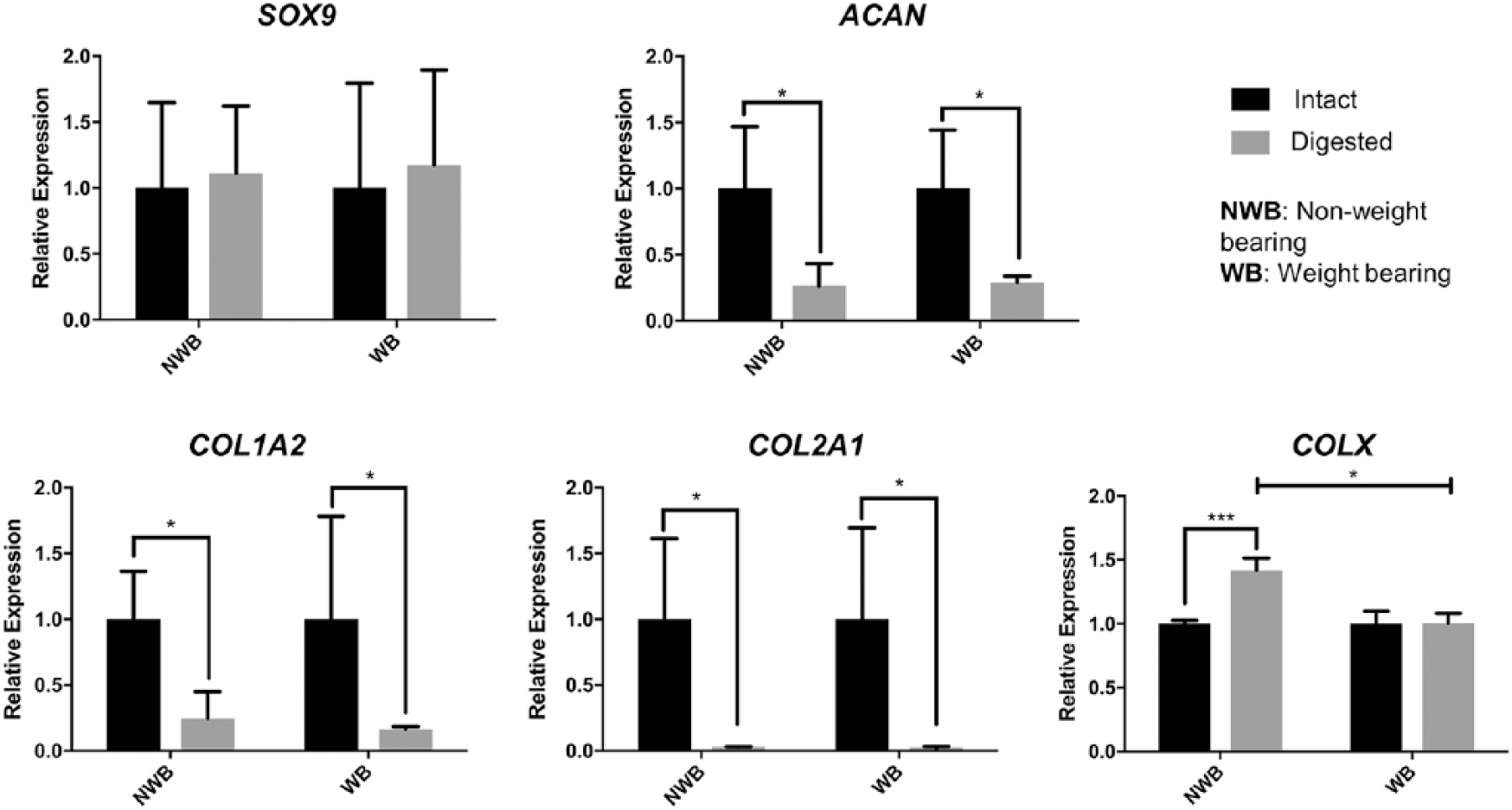
Gene expression analysis of SOX9, ACAN, COL2A1, COL1A2, and COLX in intact and digested cartilage from weightbearing and nonweightbearing regions of articular cartilage from equine fetlock joints. *P < 0.05, ***P < 0.001.
Interestingly, COLX showed increased expression in nonweightbearing digested cartilage compared to intact cartilage (P < 0.0001), but there was no difference in expression between intact and digested weightbearing cartilage (P = 0.949; Fig. 1 ). A significant interaction was noted between joint region and RNA isolation method for COLX expression (P = 0.001). COLX is a marker of hypertrophic cartilage and its upregulation is correlated with osteoarthritis. 13 Hypertrophic cartilage is associated with increases in matrix metalloproteinase 13, which is a collagenase precursor that aids in the physiological digestion of the ECM in hypertrophic cartilage to allow for calcification. 14 Similarly, digestion of the ECM using collagenase in this study may have caused the chondrocytes to begin expressing COLX as a compensatory mechanism. This may have been an immediate response due to degradation of the ECM, as this is a part of the mechanism in which OA begins. 15 This response to degradation may also account for the difference seen in COLX expression in nonweightbearing versus weightbearing digested cartilage. Eggli et al. 16 reported a decrease in numerical volume density of cells in weightbearing regions of articular cartilage in rabbits compared to less weightbearing regions. Nonweightbearing regions thus likely have less ECM than weightbearing regions, rendering them easier to enzymatically digest, resulting in increased COLX expression over weightbearing regions.
Since RNA isolation from healthy, adult equine cartilage tissue is extremely difficult, many researchers use enzymatic digestion to release the tissue’s chondrocytes for a greater RNA yield. While several RNA isolation parameters were assessed in this study (ball size in tissue homogenization, different RNA isolation kits, and incubation in lysis buffer) to determine the conditions for the greatest yield from intact tissue, it was significantly less than the amount extracted using enzymatic digestion (Supplementary Fig. 1). However, as gene expression changes may result from the digestion itself, this method may not accurately reflect the actual levels of gene expression in intact tissue. This study demonstrates that enzymatic digestion of equine articular cartilage affects the expression of key cartilage-associated genes. We have determined that chondrocyte retrieval for RNA isolation, while improving RNA yield, has significant effects on transcript levels compared with intact cartilage, even without chondrocyte culture. This finding has important implications in studying cartilage development/disease and/or tissue engineering, as endogenous expression levels of matrix-associated genes must be appropriately evaluated to provide accurate benchmarks for research. The results presented here strongly suggest that optimizing RNA isolation from intact tissue for gene expression studies is likely preferable to isolating RNA from chondrocytes following enzymatic digestion of the cartilage tissue.
Supplemental Material
Supplementary_Methods_and_Results – Supplemental material for Gene Expression Profile Is Different between Intact and Enzymatically Digested Equine Articular Cartilage
Supplemental material, Supplementary_Methods_and_Results for Gene Expression Profile Is Different between Intact and Enzymatically Digested Equine Articular Cartilage by Sarah I. M. Lepage, Rishi Sharma, David Dukoff, Leanne Stalker, Jon LaMarre and Thomas G. Koch in CARTILAGE
Supplemental Material
Supp_Fig_1 – Supplemental material for Gene Expression Profile Is Different between Intact and Enzymatically Digested Equine Articular Cartilage
Supplemental material, Supp_Fig_1 for Gene Expression Profile Is Different between Intact and Enzymatically Digested Equine Articular Cartilage by Sarah I. M. Lepage, Rishi Sharma, David Dukoff, Leanne Stalker, Jon LaMarre and Thomas G. Koch in CARTILAGE
Supplemental Material
Supp_Fig_2 – Supplemental material for Gene Expression Profile Is Different between Intact and Enzymatically Digested Equine Articular Cartilage
Supplemental material, Supp_Fig_2 for Gene Expression Profile Is Different between Intact and Enzymatically Digested Equine Articular Cartilage by Sarah I. M. Lepage, Rishi Sharma, David Dukoff, Leanne Stalker, Jon LaMarre and Thomas G. Koch in CARTILAGE
Footnotes
Author Contributions
Experimental design and approach were developed by Thomas Koch, Jon LaMarre, Sarah Lepage, Rishi Sharma, and David Dukoff. Tissue preparation and experiment execution was performed by Rishi Sharma and David Dukoff. Data collection, assembly and analysis were performed by Rishi Sharma, Sarah Lepage, and Leanne Stalker. Statistics were analyzed and assembled by Rishi Sharma in consultation with Ms. Gabrielle Monteith. Funding was obtained by Thomas Koch. All authors were involved in drafting, revising and/or approving the final manuscript.
Acknowledgments and Funding
We wish to acknowledge Gabrielle Monteith for her help with statistics, as well as Allison Tscherner for her technical assistance, intellectual contributions, and guidance in RNA isolation from difficult tissues and reference gene analysis. The Equine Guelph Research Foundation and the National Science Engineering Research Council Discovery Grant Program of Canada generously provided operating funds. The Ontario Veterinary College graduate scholarship provided partial support for Rishi Sharma, and the NSERC undergraduate scholarship research assistant program provided support for David Dukoff.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
