Abstract
Pancreatic ductal adenocarcinoma (PDAC) is characterized by early recurrence following pancreatectomy, rapid progression, and chemoresistance. Novel prognostic and predictive biomarkers are urgently needed to both stratify patients for clinical trials and select patients for adjuvant therapy regimens. This study sought to determine the biological significance of RABL6A (RAB, member RAS oncogene family-like protein 6 isoform A), a novel pancreatic protein, in PDAC. Analyses of RABL6A protein expression in PDAC specimens from 73 patients who underwent pancreatic resection showed that RABL6A levels are altered in 74% of tumors relative to adjacent benign ductal epithelium. Undetectable RABL6A expression, found in 7% (5/73) of patients, correlated with improved overall survival (range 41 to 118 months with 3/5 patients still living), while patients with RABL6A expression had a worse outcome (range 3.3 to 100 months, median survival 20.3 months) (P = 0.0134). In agreement with those findings, RABL6A expression was increased in pancreatic cancer cell lines compared to normal pancreatic epithelial cells, and its knockdown inhibited pancreatic cancer cell proliferation and induced apoptosis. Moreover, RABL6A depletion selectively sensitized cells to oxaliplatin-induced arrest and death. This work reveals that RABL6A promotes the proliferation, survival, and oxaliplatin resistance of PDAC cells, whereas its loss is associated with extended survival in patients with resected PDAC. Such data suggest RABL6A is a novel biomarker of PDAC and potential target for anticancer therapy.
Introduction
Pancreatic ductal adenocarcinoma (PDAC) is the fourth leading cause of cancer deaths in the United States and has an overall 5-year survival of less than 5%.1,2 Pancreatic resection combined with adjuvant therapy remains the only curative treatment for PDAC, 1 although outcome remains poor with a 5-year survival rate of 25% to 35%. 3 Notably, most patients (80%) present with inoperable metastatic disease at diagnosis and have few effective therapeutic options. The most common chemotherapeutic treatments are gemcitabine and 5-FU-based therapies, which provide only minimal benefit to patient survival.4,5 A new regimen of chemotherapy including 5-FU, leucovorin, irinotecan, and oxaliplatin (FOLFIRINOX) has been demonstrated to improve median survival compared to gemcitabine therapy (11.1 vs. 6.8 months, respectively) 6 for patients with metastatic disease; however, this regimen has significant toxicity. Currently, there are no reliable biomarkers that can predict which patients will respond to chemotherapy. There is clearly an unmet need for reliable biomarkers of PDAC that will predict patient survival and/or response to therapy and ideally improve the clinical management of this disease.
RABL6A (RAB, member RAS oncogene family-like protein 6 isoform A) was originally discovered in a yeast 2-hybrid screen for ARF (Alternative Reading Frame) interacting proteins. 7 ARF, a product of the INK4a/ARF tumor suppressor locus, 8 is inactivated in most human cancers and functions through multiple anticancer pathways (p53-dependent and p53-independent) to prevent tumorigenesis. 9 The RABL6 gene (originally named Parf [Partner of ARF], also called RBEL1 or c9orf86) was later identified from in silico analyses of Ras/Rab-containing sequences and found to encode 4 isoforms (A-D) due to alternative splicing.10,11 All forms of RABL6 have GTPase activity, while only the largest form, RABL6A, contains the ARF binding domain.7,10,11 The biological function of RABL6 proteins remains unclear, although knockdown of RABL6A or all 4 isoforms caused significant death in breast and colon cancer cell lines. 11 Those results suggested RABL6 may promote tumor cell survival.
Several observations suggested a role for RABL6A in PDAC. First, its partner ARF is a powerful suppressor of PDAC that is inactivated in 40% of tumors. 12 Second, RABL6A mRNA is highly expressed in normal pancreas compared to other tissues, implying RABL6A has relevance in pancreas physiology. 7 Third, published microarray studies reveal frequent alteration (both up- and down-regulation) of RABL6A mRNA levels in PDAC tumors.13-16 In this study, we evaluated RABL6A expression levels in human PDAC cell lines and in resected PDAC tumors from patients and correlated the latter with clinicopathological variables and survival. We also assessed how its depletion affected pancreatic cancer cell growth, survival, and responsiveness to PDAC chemotherapeutic drugs. Our findings show that RABL6A loss correlates with improved patient outcome and represents a potential novel biomarker for survival in patients with PDAC.
Results
RABL6A is highly expressed in pancreatic cancer cell lines
To begin examining the biological importance of RABL6A, we first assessed its normal tissue distribution. RABL6A protein was most highly expressed in the normal mouse pancreas and brain (Fig. 1A), consistent with semiquantitative RT-PCR analyses, which showed elevated RABL6A mRNA levels in the same tissues in humans (data not shown). Because of its high expression in normal pancreas, as well as evidence from cancer microarrays that RABL6 mRNA is altered in human PDAC tumors,13-16 we measured RABL6A levels in 4 PDAC-derived cell lines (Panc-1, MiaPaCa-2, BxPC-3, and CAPAN-1). Quantification of western blots showed that RABL6A protein levels are increased in the tumor cells (2- to 7.4-fold) relative to its expression in immortalized human pancreatic epithelial, Nestin-expressing (HPNE) cells (Fig. 1B).
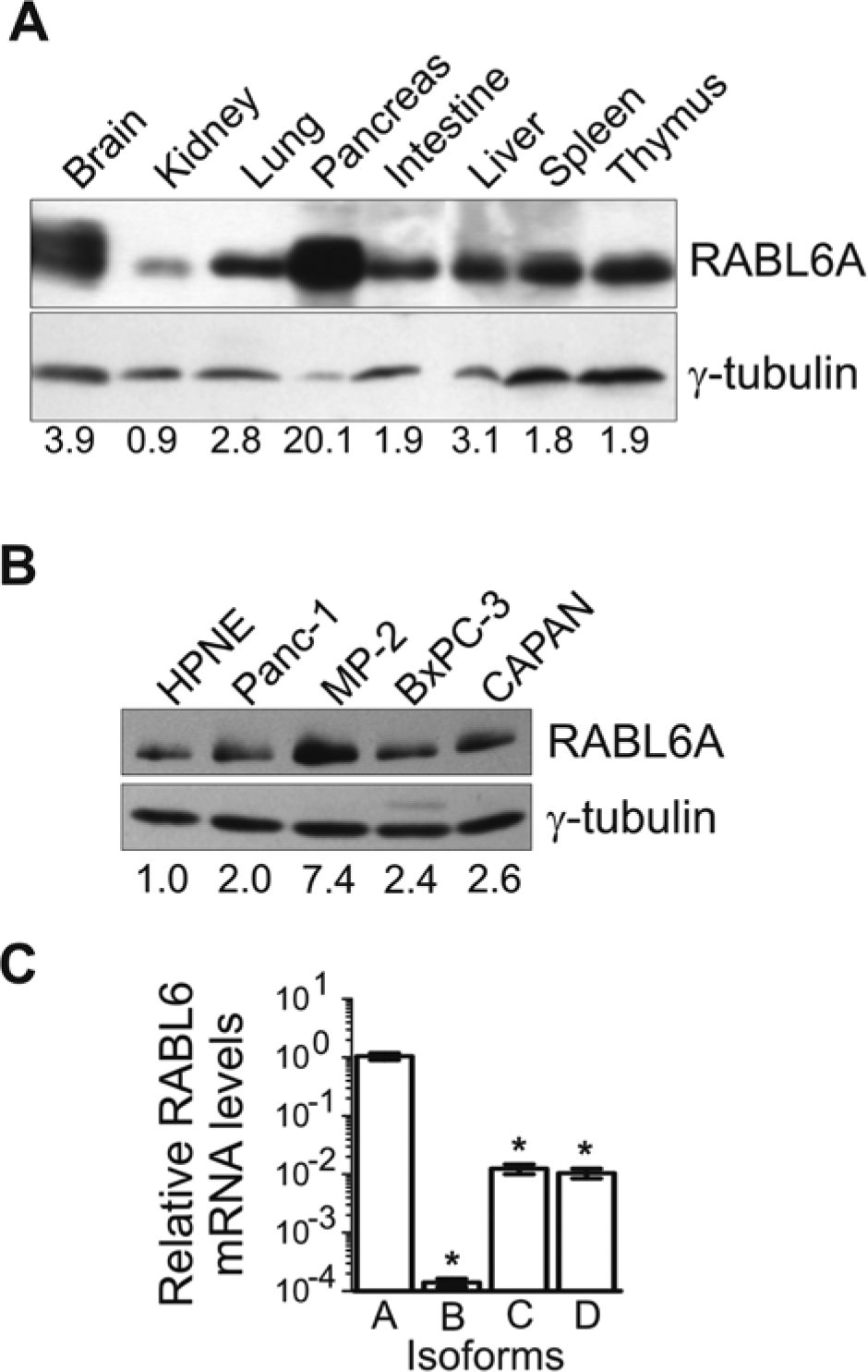
RABL6A is a pancreatic protein with elevated levels in PDAC cell lines. (
Current antibodies only recognize RABL6A; therefore, qRT-PCR was employed to examine expression of the other RABL6 isoforms in PDAC cells. Panc-1 cells were chosen as a model system for our study because they displayed a moderate up-regulation of RABL6A protein relative to HPNE cells, similar to the majority of PDAC lines. As shown in Figure 1C, RABL6A is the most highly expressed isoform, whereas RABL6C and -D transcripts are present at 100-fold lower levels. High Ct values (>35) for RABL6B mRNA suggested it is expressed at negligible levels, if at all. A similar pattern of mRNA expression for the RABL6 isoforms was observed in MiaPaCa-2 cells (data not shown). These results indicate RABL6A is the most abundant RABL6 isoform expressed in PDAC cells.
RABL6A expression is altered in human PDAC tumors
We examined RABL6A protein levels in human PDAC tumors by immunohistochemistry (IHC). Initially, western blot analyses of Panc-1 cells expressing a control (CON) or short hairpin RNAs (shRNA) targeting RABL6 for knockdown (KD) was performed to test the specificity of 2 antibodies to RABL6A, a rabbit polyclonal antibody 7 and a newly generated mouse monoclonal antibody (MAb). Both antibodies effectively detected the 130 to 135 kDa RABL6A protein by immunoblotting, but the MAb showed the highest specificity (Suppl. Fig. S1A). Using formalin-fixed paraffin-embedded (FFPE) Panc-1 CON and KD cells, conditions were optimized for the immunohistochemical detection of endogenous RABL6A (Suppl. Fig. S1B). RABL6A staining was cytoplasmic, as expected,7,10 and specific, as demonstrated by the significant signal reduction in KD cells. Moreover, the staining was sensitive to even low levels of residual RABL6A expression present within KD cells.
Endogenous RABL6A levels in pancreatic resections from 73 PDAC patients were assessed by IHC using the RABL6A MAb (Fig. 2). Tumor-associated RABL6A expression was evaluated relative to adjacent benign pancreatic ducts within the same tissue sections, and 5 different scores for RABL6A staining in tumor cells were used: −2 (undetectable), −1 (modestly decreased), 0 (equivalent), +1 (modestly increased), and +2 (markedly increased). Figure 2A shows representative results from IHC analyses of PDAC tumors assigned a −2 or +2 staining score. Consistent with published microarray data, RABL6A protein expression was altered in the majority (74%) of PDAC patient tumors with half of the tumors showing increased levels (Fig. 2B).
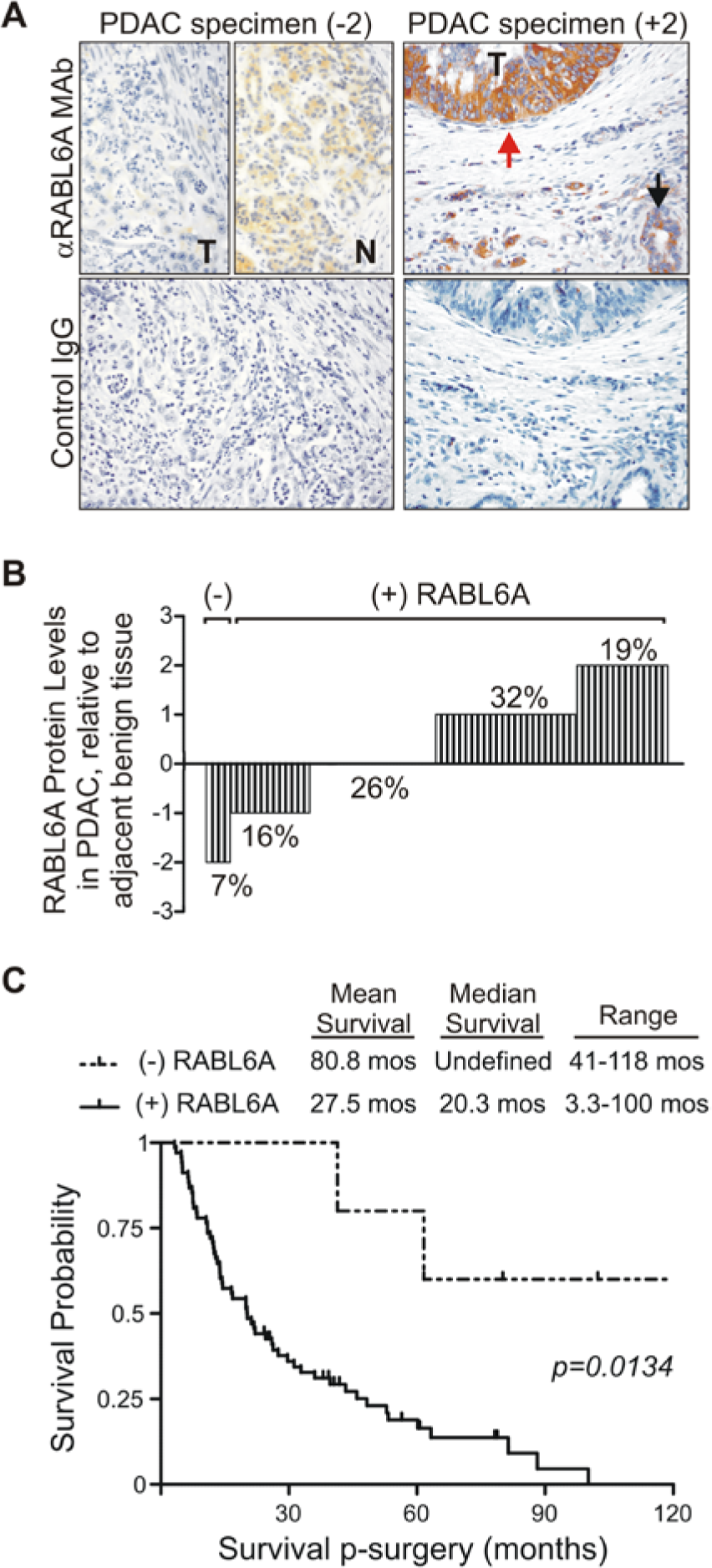
RABL6A expression is altered in PDAC tumors and its loss is associated with increased survival in resected PDAC patients. (
RABL6A loss is associated with improved survival following pancreatic resection
Patient data were evaluated for possible correlations between RABL6A expression levels and clinicopathologic variables as well as survival. The absence of RABL6A expression in patient tumors (IHC score of −2, henceforth called (−) RABL6A) was associated with improved survival (Fig. 2C). Mean survival of this group was 80.8 months (range = 41 to 118 months), with 3 of 5 patients still living. Median survival of patients lacking RABL6A expression was not estimable as survival curves did not fall below 50% during the study period. By comparison, patients whose tumors expressed RABL6A (IHC scores of −1, 0, +1, and +2, collectively grouped as (+) RABL6A) had a worse outcome, with a median survival of 20.3 months (P = 0.0134 by Cox regression model; hazard ratio = 6.16; 95% confidence interval [CI] = 1.46-26.01). Survival curves for patients expressing any level of detectable RABL6A (−1, 0, +1, +2) in tumors were statistically indistinguishable from each other. No significant correlations were observed between RABL6A expression levels and the AJCC T and N stages (Table 1). A total of 14 out of 73 patients analyzed were alive at the end of our follow-up (19%). Notably, 80% (4 of 5) of patients with RABL6A loss (−2 group) lived longer than 5 years, whereas only 37.5% of the other patients surpassed the 5-year survival mark (P = 0.0415; hazard ratio = 7.87; 95% CI = 1.08-57.25). These data demonstrate that RABL6A loss is a marker of extended survival in resected PDAC patients.
Association Between RABL6A Levels and Clinicopathological Variables of PDAC.
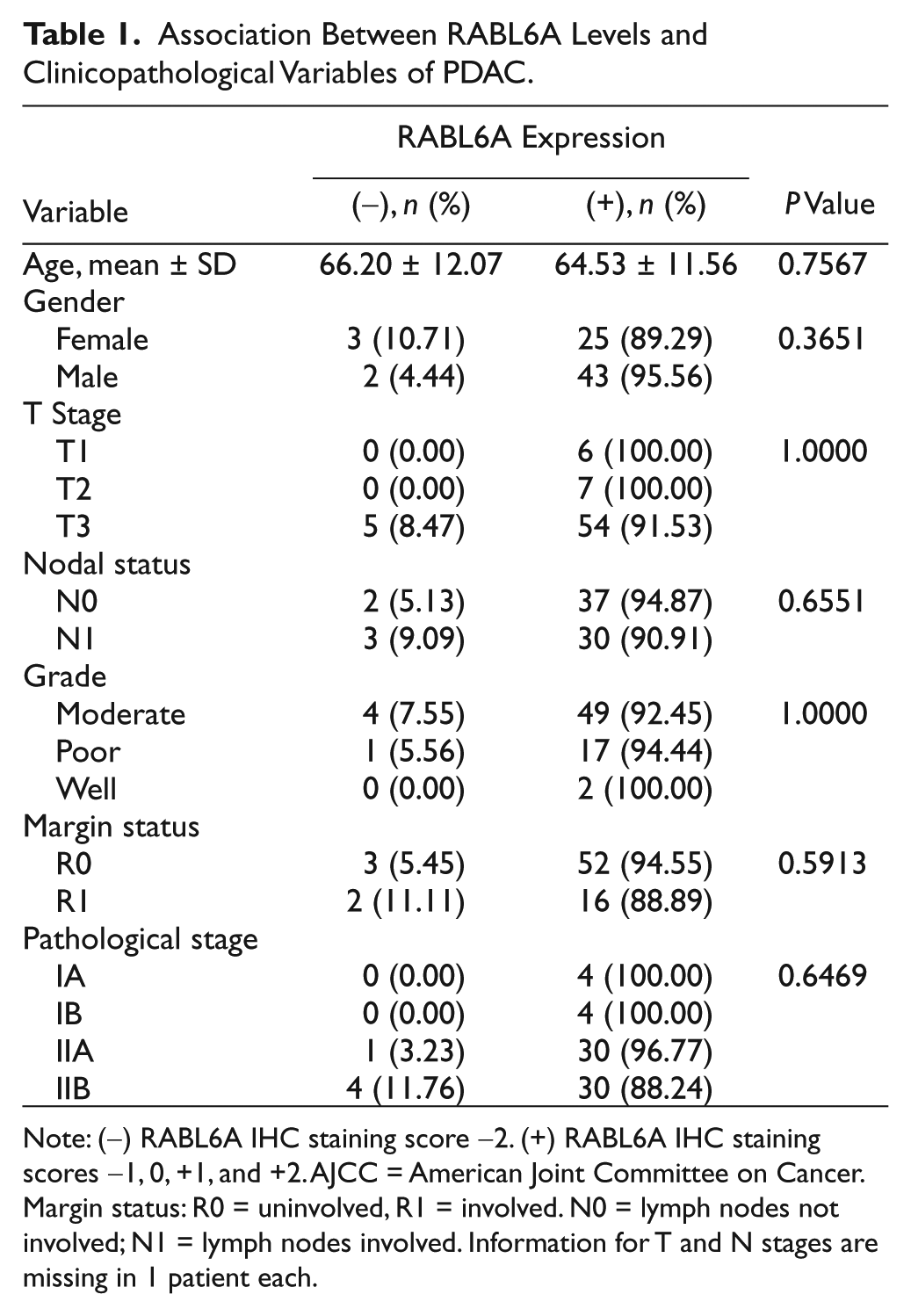
Note: (−) RABL6A IHC staining score −2. (+) RABL6A IHC staining scores −1, 0, +1, and +2. AJCC = American Joint Committee on Cancer. Margin status: R0 = uninvolved, R1 = involved. N0 = lymph nodes not involved; N1 = lymph nodes involved. Information for T and N stages are missing in 1 patient each.
RABL6A promotes pancreatic cancer cell proliferation and survival
RNA interference was employed to silence RABL6A and determine its role in PDAC cell proliferation and survival. Two different shRNA constructs were used that are predicted to target all 4 RABL6 isoforms (KD1) or selectively knockdown RABL6A (KD2) (Fig. 3A, top). Lentiviruses expressing the KD1 or KD2 shRNAs versus an empty vector control (CON) were expressed in Panc-1 cells and qRT-PCR performed to assess efficiency of knockdown. Compared to control cells, KD1 effectively silenced all RABL6 isoforms whereas KD2 specifically depleted RABL6A mRNA expression (Fig. 3A, bottom). Western blotting confirmed robust down-regulation of RABL6A expression by both shRNAs with greater knockdown consistently seen in KD1 cells (Fig. 3B).
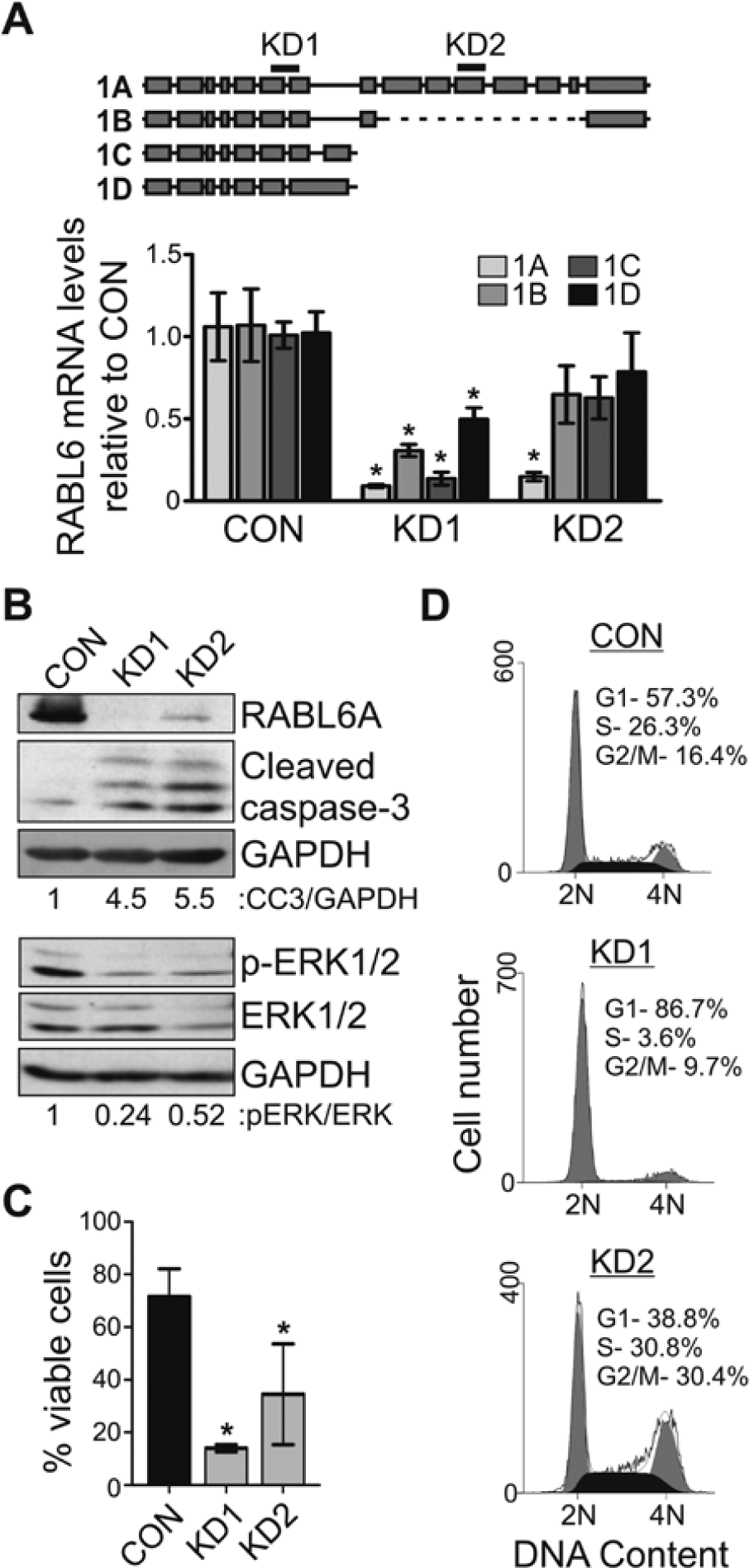
RABL6A loss promotes cell death through apoptosis. (
Cell viability was significantly reduced by RABL6 knockdown (Fig. 3C), which was associated with increased levels of cleaved caspase-3 and decreased phosphorylation of extracellular-signal-related kinase 1 (ERK1) and ERK2 (Fig. 3B). The same effects were seen in KD1 and KD2 cells. To determine the effect of RABL6 knockdown on the cell cycle, DNA content was measured by flow cytometry (Fig. 3D). Interestingly, unique cell cycle arrest phenotypes were induced in Panc-1 cells by depletion of all RABL6 isoforms (KD1) versus RABL6A alone (KD2). KD1 cells arrested in G1 phase whereas KD2 cells accumulated in G2/M. Notably, an identical loss in viability and G2/M arrest was observed in Panc-1 cells expressing another shRNA specific to RABL6A (denoted KD3, Suppl. Fig. S2). While these data may suggest distinct roles for RABL6 isoforms in cell cycle progression, the simplest conclusion from these findings is that RABL6A mediates PDAC cell survival and proliferation.
Given that RABL6A is required for PDAC viability and cell cycle progression, we wondered how PDAC tumors lacking the protein can arise and exist among our patient group. To address this question, we propagated RABL6 knockdown cultures up to 8 weeks to examine the long-term consequences of its loss. Cells required constant antibiotic selection during this time to maintain shRNA expression. Acute RABL6 knockdown continued to cause significant Panc-1 cell death during the first several weeks after knockdown. A similar cell death phenotype was observed in MiaPaCa-2 and BxPC-3 PDAC cells following acute RABL6 silencing (Suppl. Fig. S3). During extended culture, however, viable proliferating populations for all 3 PDAC lines arose that maintained RABL6A loss (Fig. 4). The lengthy time required to generate those knockdown lines suggests the cells sustained genetic alterations that allowed them to bypass the initial cell cycle arrest and death evoked by RABL6A loss. Notably, all 3 PDAC lines expressing the KD1 shRNA proliferated at significantly slower rates than CON cells. In contrast, cells with selective RABL6A knockdown (KD2 cells) generally proliferated at similar rates as the controls. The exception was MiaPaCa-2 KD2 cells, which displayed an intermediate rate of growth between CON and KD1 cells (Fig. 4, middle panel). MiaPaCa-2 cells express much higher basal levels of RABL6A compared to Panc-1 and BxPC-3 (see Fig. 1) and as such may be more dependent on it for growth.
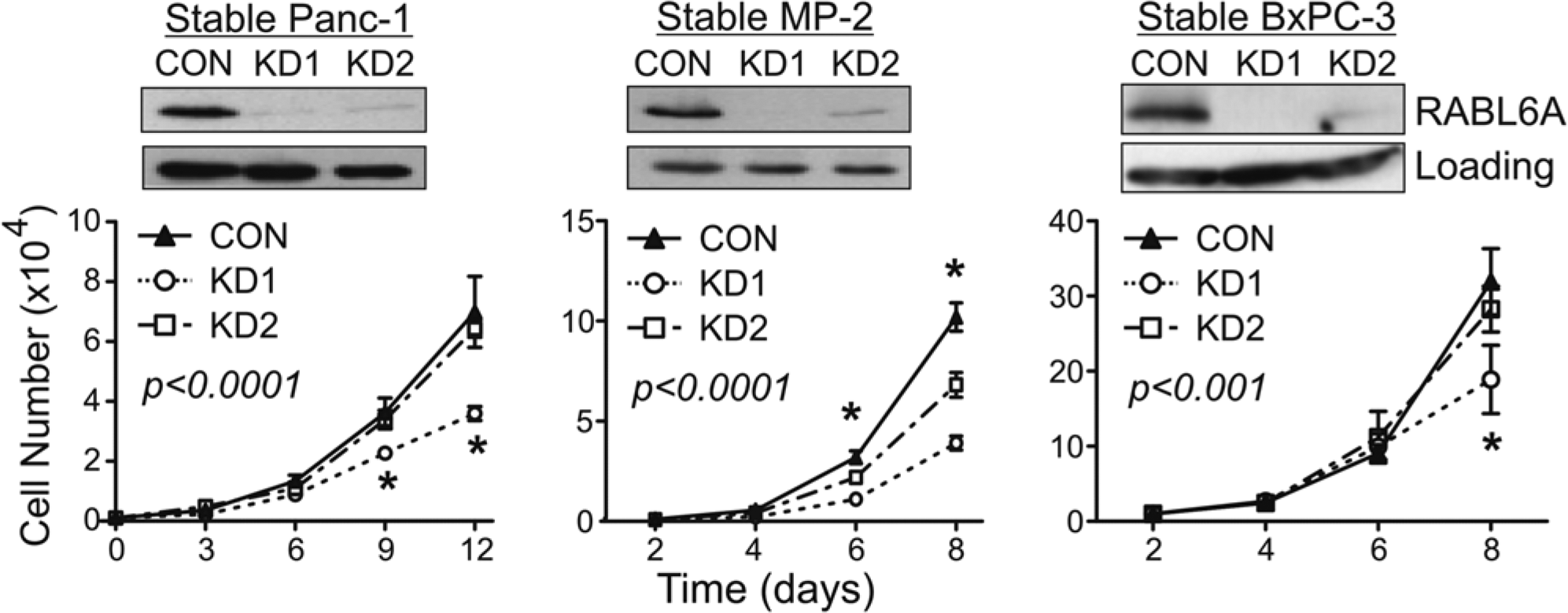
Effects of sustained RABL6A depletion on PDAC cell proliferation. (
RABL6A levels dictate chemosensitivity to oxaliplatin in PDAC cells
The contribution of RABL6A to PDAC cell survival and proliferation suggested its loss might sensitize cells to chemotherapy-induced death and arrest. Stable proliferating Panc-1 populations expressing control, KD1, or KD2 shRNAs (see Fig. 4, left panel) were exposed to varying doses of PDAC relevant drugs, gemcitabine or agents used in the FOLFIRINOX combination (5-FU, irinotecan, and oxaliplatin). Treatment with gemcitabine, irinotecan, or 5-FU had no effect on the cell number or viability of RABL6A-depleted cells compared to CON cells (Suppl. Fig. S4). In contrast, treatment with oxaliplatin induced greater arrest and death in RABL6A knockdown cells compared to CON cells. At low concentrations (0.125-0.5 µg/mL), oxaliplatin caused a significant reduction in cell number for both knockdown populations (Fig. 5A) while having a minimal effect on their viability (Fig. 5B). These data demonstrated that RABL6A loss enhanced the antiproliferative effects of oxaliplatin at low doses. Conversely, higher concentrations of oxaliplatin (2-4 µg/mL) combined with RABL6A depletion resulted in significantly greater cell death than that observed in CON cells (Fig. 5B). From these data, we conclude that endogenous RABL6A confers resistance to oxaliplatin therapy in PDAC cells.
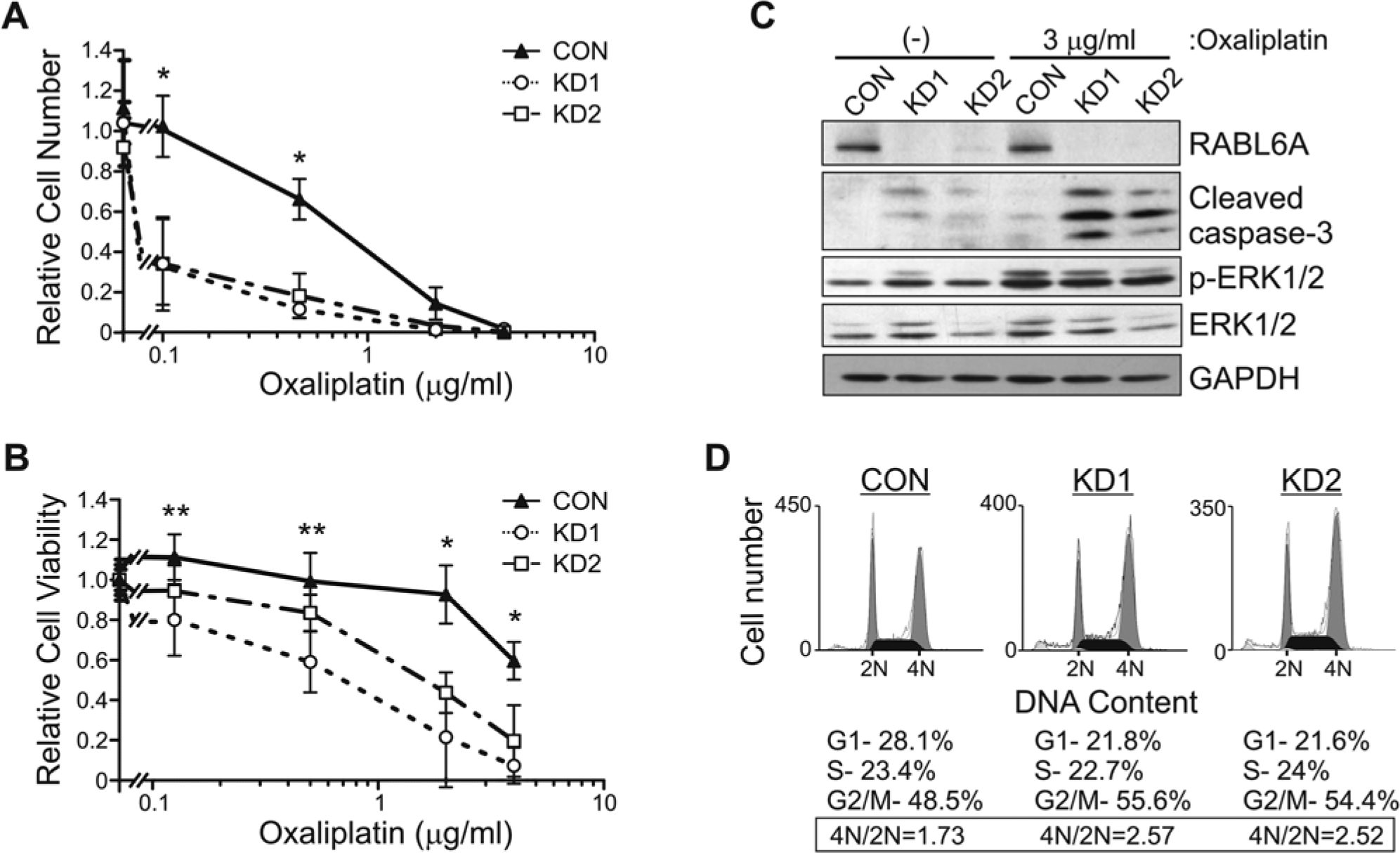
Sustained RABL6A depletion sensitizes Panc-1 cells to oxaliplatin-induced cell cycle arrest and apoptosis. Established Panc-1 CON, KD1, and KD2 cells were exposed to the indicated concentrations of oxaliplatin for 4 days and examined for cell number (
Oxaliplatin stimulates apoptotic cell death and G2/M phase cell cycle arrest in cancer cells lacking functional p53,17,18 and the PDAC cells studied here express mutant p53. It is also known that RABL6A silencing in PDAC (Fig. 3) and other cancer cells 11 coincides with increased caspase-3 cleavage and reduced ERK phosphorylation; therefore, we tested if those changes were exacerbated in RABL6A-depleted cells exposed to oxaliplatin (Fig. 5C). Caspase-3 cleavage was significantly increased in oxaliplatin-treated KD1 and KD2 cells (lanes 5 and 6) compared to levels induced by RABL6A depletion alone (lanes 2 and 3) or by oxaliplatin treatment of CON cells (lane 4). RABL6 knockdown combined with exposure to oxaliplatin also induced a striking accumulation of mitotically arrested cells (Fig. 5D). In contrast, the down-regulation of ERK phosphorylation caused by transient RABL6 depletion (Fig. 3B) was lost in established RABL6 knockdown cultures (Fig. 5C, lanes 1-3) and no changes in ERK signaling were evident following oxaliplatin treatment (lanes 4-6). These findings suggest that impaired mitotic progression and increased caspase-dependent cell death contribute to the sensitization of RABL6A-depleted PDAC cells to oxaliplatin whereas ERK signaling is unlikely to play a role.
Discussion
Cancer is a complex, heterogeneous disease for which personalized therapies tailored to the unique genetic makeup of each patient and their tumors hold the greatest promise for improved survival and quality of life. The key to personalized anticancer treatments is the identification, characterization, and validation of molecular biomarkers that will predict tumor development (diagnostic markers), survival (prognostic markers), and/or response to particular therapies (predictive biomarkers).19,20 Here, we discovered that RABL6A loss is a new marker of extended survival in patients with resected PDAC. Moreover, RABL6A may be a predictive marker for PDAC response to oxaliplatin since it selectively conferred resistance to that drug in cultured PDAC cells.
There are currently no clinically meaningful biomarkers that guide PDAC treatment 21 ; however, there are some promising developments.22 -26 Most studies have focused on identifying factors with predictive value for response to gemcitabine chemotherapy since it remains the most commonly used drug for PDAC treatment.23,27,28 However, the recent observation that FOLFIRINOX therapy nearly doubles median survival of PDAC patients compared to gemcitabine alone 6 strongly warrants the identification of biomarkers that predict sensitivity to oxaliplatin, 5-FU, and/or irinotecan. We found no changes in the response of pancreatic cancer cells with altered RABL6A expression to treatment with gemcitabine, 5-FU, or irinotecan, whereas RABL6A loss selectively sensitized cells to oxaliplatin-induced cell arrest and apoptosis. Only a few genes have been identified that influence the cellular response to oxaliplatin, including ERCC1, 29 MSH3, 30 survivin,31,32 and NF-kB. 31 Notably, those studies were performed in other cancer cell types and the importance of those genes to PDAC is not established. RABL6A appears to be the first gene identified that specifically influences response to oxaliplatin in PDAC cells. Interestingly, cells depleted of all 4 RABL6 isoforms (KD1) were consistently more sensitive to oxaliplatin treatment than cells lacking RABL6A alone (KD2). That may reflect more effective silencing of RABL6A in KD1 cells or imply a role for the other RABL6 isoforms in oxaliplatin responsiveness.
Oxaliplatin has significant hematological and neurological side effects, either alone or in the more recently established FOLFIRINOX treatment.33 -36 Indeed, FOLFIRINOX treatment causes greater toxicity in patients compared to gemcitabine, and high doses of oxaliplatin are usually required to produce the desired clinical effect.6,33,37 We found that RABL6 loss reduced the concentration of oxaliplatin required to arrest or kill PDAC cells in culture. Thus, loss of RABL6 in PDAC tumors may enable those patients to be treated with lower doses of oxaliplatin in FOLFIRINOX therapy and may promote better quality of life during treatment without compromising outcome. In our study, PDAC patients with undetectable RABL6A were few in number (n = 5), and all were given different types of therapy with only 1 patient receiving oxaliplatin treatment. Hence, no correlation between therapy regimen and survival was evident. Oxaliplatin is now being given to PDAC patients on a more frequent basis; as a result, it will be feasible in future studies involving larger patient cohorts to examine potential correlations between RABL6A status, oxaliplatin therapy, and survival.
An important question raised by our study is how RABL6A loss improves PDAC patient survival. The simplest explanation is that acute loss of RABL6A or all RABL6 isoforms causes a rapid and significant death and arrest of PDAC cells, which would effectively impair tumor development in vivo. Over time, however, our data suggest that cells lacking RABL6A sustain genetic alterations that enable them to circumvent the early death and arrest provoked by RABL6A loss. This would explain how PDAC tumors lacking RABL6A can arise and be identified among our patient group, and why their development would be infrequent in vivo (just 7% of tumors in this study). Moreover, the fact that all stable KD1 (and some KD2) knockdown populations continued to proliferate at reduced rates compared to control cells predicts slower progression of RABL6 null tumors. This could also contribute to the extended survival of patients lacking RABL6A in their tumors.
Decreased ERK phosphorylation has been shown to predict longer survival in PDAC patients. 38 In PDAC cells, the robust death induced by acute depletion of RABL6A or all RABL6 isoforms coincided with decreased ERK phosphorylation. Those data mimic effects of RABL6 knockdown in breast and colon cancer cell lines, 11 which led Montalbano et al. to propose that RABL6 promotes tumor cell viability and proliferation via MEK/ERK signaling. Our findings agree with the importance of RABL6A to tumor cell survival but challenge the involvement of ERK because established PDAC cultures with stable RABL6A knockdown displayed no impairment in ERK phosphorylation. Moreover, ERK phosphorylation was unchanged by oxaliplatin treatment of RABL6A-depleted cells despite the increased death under those conditions. These results strongly argue that RABL6A controls PDAC tumor cell survival through mechanisms other than MEK/ERK signaling.
Mitotic catastrophe is considered an important onco-suppressive mechanism that reduces genomic instability by eliminating mitosis-incompetent cells through cell senescence or death. 39 In that regard, altered expression of apoptotic factors, such as the B-cell lymphoma 2 (Bcl-2) family member, Bax, has been shown to influence the incidence of death on mitotic catastrophe. 40 In RABL6A-depleted Panc-1 cells, we have found reduced expression of the anti-apoptotic protein, Bcl-2 (Suppl. Fig. S5). Cells lacking RABL6A also exhibited a significant mitotic block associated with enhanced caspase-3 cleavage following oxaliplatin treatment. These data provide new, compelling evidence that RABL6A promotes PDAC cell survival and chemoresistance, at least, in part, by suppressing mitotic catastrophe and caspase-mediated apoptosis.
We consistently observed unique cell cycle arrest phenotypes in Panc-1 cells following depletion of all RABL6 isoforms (KD1) versus RABL6A alone (KD2). While KD1 cells arrested in G1 phase, KD2 cells accumulated in G2/M. At face value, these results may suggest that RABL6A is necessary for mitotic progression while other isoforms (likely RABL6C or D since B is not detectable) are required for G1 progression. However, we currently cannot exclude the possibility that the different arrest profiles merely reflect more effective silencing of RABL6A by KD1 than KD2 shRNAs (see Fig. 3B). We are comparing the effects of RABL6A silencing by KD1 and KD2 shRNAs in additional human tumor cell lines and in most cases find that both shRNAs cause a G1 arrest with concomitant loss of cells in S phase. This has been seen in BxPC-3 cells (Suppl. Fig. S6), as well as U251 glioblastoma and BON-1 pancreatic neuroendocrine tumor cells (J. Hagen and D.E. Quelle, unpublished results). When considered with the G2/M arrest seen in Panc-1 knockdown cells, these findings suggest that RABL6A is required for both G1 and G2/M phase activities. Consistent with that conclusion, microarray analyses of RABL6A knockdown in both mouse embryonic fibroblasts and BON-1 cells showed significantly altered expression of genes required for both G1 and G2/M progression (J. Hagen and D.E. Quelle, unpublished results). Which checkpoint is engaged following RABL6A depletion may depend on the genetic makeup of the tumor cell (i.e., integrity of the specific checkpoint pathways) and/or the magnitude of RABL6A loss.
The ARF tumor suppressor plays a key role in PDAC, 41 and RABL6A is a novel ARF binding partner. 7 A functionally relevant interplay between RABL6A and ARF during PDAC tumorigenesis in patients is possible because RABL6A interacts with the essential growth inhibitory domains of ARF. 7 As such, it will be worthwhile to determine how RABL6A affects ARF activity in PDAC cells and if correlations exist between RABL6A and ARF status in human PDAC tumors. In that regard, we have preliminary evidence that RABL6A can inhibit ARF function by relocalizing it to the cytoplasm (V.P. Muniz, S.M. Reed, and D.E. Quelle, unpublished data), consistent with an oncogenic role of RABL6A. It is important to emphasize, however, that all PDAC cell lines used in this study lack ARF due to gene deletion or promoter silencing. The cells also express mutated, functionally compromised p53. Therefore, the oncogenic activities observed herein for RABL6A in PDAC cells are independent of ARF and p53, revealing that RABL6A plays an important role in the biology of PDAC regardless of ARF and p53 status.
In summary, we show that RABL6A is a new marker of survival in resected PDAC patients. Expanded analyses of a larger cohort of patients will be needed to firmly establish RABL6A biomarker status; nonetheless, the similarity of our patient population to others previously studied strengthens our conclusions. Specifically, only 37.5% of our patients (excluding those lacking RABL6A) surpassed the 5-year survival mark, closely reflecting the reported rate of 25% to 35% 4 and underscoring the remarkable extension in lifespan found for patients lacking RABL6A tumor expression. Significantly, our studies also suggest RABL6A may be a predictive marker of patient responsiveness to oxaliplatin therapy. Given the dismal outcome and absence of clinically meaningful biomarkers for PDAC, it will be important to evaluate RABL6A status in PDAC biopsies and resected tissues from larger patient cohorts undergoing therapy with oxaliplatin versus other neoplastic agents. Such information may improve PDAC patient treatment and outcome by stratifying patients for neo-adjuvant treatments, enrollment in new clinical trials, and adjustments in the dosing regimens for FOLFIRINOX therapy.
Material and Methods
Cell culture
Panc-1 cells were maintained in standard Dulbecco’s modified Eagle’s medium (DMEM) containing 10% fetal bovine serum (FBS), 4 mM glutamine, and 100 µg/mL penicillin/streptomycin. MiaPaCa-2 cells were grown in the same media containing 2.5% equine serum. BxPC-3 cells were maintained in RPMI-1640 medium, 10% FBS, 2 mM glutamine, 100 µg/mL penicillin/streptomycin, 2.5 g/L
RNA interference, lentivirus production, and infection
RABL6-specific shRNA constructs in the pLKO.1 lentiviral vector were purchased from Open Biosystems (Huntsville, AL). Four shRNAs were initially screened for efficiency of RABL6A knockdown. Two yielded greater than 90% knockdown and were used for the subsequent experiments. The RABL6 targeted sequences are shown below (with an order of sense, loop [underlined], and antisense): KD1, gcagtggaccttcaattacatctcgagatgtaattgaaggtccactgc; KD2, ccagtcaagacatcactctttctcgagaaagagtgatgtcttgactgg.
To produce lentiviruses encoding human RABL6 shRNAs, 293T cells (1 × 106) in 35 mm dishes were transfected with 2 µg total of the lentiviral construct, packaging plasmid pCMV-dR8.74psPAX2, and envelope plasmid pVSVG (ratio of 1:0.9:0.1, respectively) using Polyfect reagent (Qiagen, Valencia, CA). Media containing virus was harvested 2 days posttransfection, polybrene added to 8 µg/mL, and sterile filtered (0.45 µM). Panc-1, BxPC-3, and MiaPaCa-2 cells were transduced for 2 days sequentially using 1.5 mL fresh virus for 8 to 10 hours each day. Cells were harvested 3 to 5 days after infection for assays or selected with 1 µg/mL puromycin for the indicated time periods prior to analyses, as indicated in figure legends.
Protein expression analyses
Frozen cell pellets were lysed directly in SDS-PAGE loading buffer and identical cell equivalents loaded onto polyacrylamide gels prior to western blotting and protein detection via enhanced chemiluminescence (ECL, Amersham, Buckinghamshire, UK), as described.7,42 Antibodies that detect RABL6A were generated, including a rabbit polyclonal made against an internal peptide [NH2-MVAGFQDDVDIEDQC-COOH] (used at 1.5 µg/mL) and a mouse monoclonal made against recombinant GST-tagged human RABL6A (Iowa State University Hybridoma Facility, Ames, IA). Other antibodies used include those against cleaved caspase-3 (1:500; Cell Signaling, Downers, MA), γ-tubulin (1:10,000; clone GTU-88 mouse monoclonal, Sigma-Aldrich, St. Louis, MO), p44/p42 ERK 1 and 2 (1:1,000; 137F5, Cell Signaling), phosphorylated ERK 1 and 2 (1:1,000; E-4, Santa Cruz Biotechnology, Santa Cruz, CA), Bcl-2 (1:1,000; Santa Cruz Biotechnology), and GAPDH (1:20,000; Ab8245 mouse monoclonal, Abcam, Cambridge, MA).
qRT-PCR analyses
Quantitative RT-PCR analysis of all RABL6 isoforms expression in control and knockdown Panc-1 cells was performed with 1 µg of total RNA using a High Capacity cDNA Reverse Transcription Kit (Applied Biosystems Inc., Foster City, CA) and the Power SYBR Green PCR Master Mix (Applied Biosystems Inc.). PCR reactions using RABL6 primers specific for each isoform (A forward 5′-aatgctgtccttcgtcatgga-3′, A reverse 5′-gcacgggaaagtcatccg-3′; B forward 5′-ggcgcagcatcatctctagg-3′, B reverse 5′-ctagagctcctcgtagtcgcca-3′; C forward 5′-ttgcagcttcagaaggatgtca-3′, C reverse 5′-gacagcacagcccctttcc-3′; D forward 5′-acctggacagacctccaggtt-3′, D reverse 5′-cacgtggtgagtgcttacctgaa-3′) were performed as follows: denaturation at 95°C for 10 minutes, followed by 40 cycles of 95°C for 15 seconds, 60°C for 1 minute on an ABI Prism 7000 real-time detection system (Applied Biosystems Inc.). Fold differences in RABL6 mRNA levels were calibrated to GAPDH mRNA expression and computed using the 2−ΔΔCt method. 43
Cell cycle analyses
PDAC cells were harvested and suspended in propidium iodide solution (0.05 mg/mL in 0.1% sodium citrate) containing 0.1 µg/mL RNase A. Cells were incubated for 30 minutes in the dark and analyzed by flow cytometry using a FACScan cytometer (Benton Dickinson, Franklin Lakes, NJ). DNA content and cell cycle profiles were analyzed using the MOD Fit LT software (Verity Software House, Topsham, ME).
Chemosensitivity assay
Chemotherapeutic drugs gemcitabine, oxaliplatin, irinotecan, and 5-FU were purchased from Sigma (St. Louis, MO) and dissolved either in sterile ddH2O (gemcitabine and oxaliplatin) or DMSO (irinotecan and 5-FU). Panc-1 cells with or without RABL6 knockdown were seeded at 3 to 5 × 103 cells per well in triplicate in 48-well culture plates. After overnight incubation, an equal volume of 2× concentrated drug was added to the cells and incubated for 4 to 7 days. Drug doses used were based on previously published dose–response curves.22,34,35,44,45 Cells (both attached and floating) were harvested and cell number and viability were assessed by Trypan blue staining (1:1 v/v) and counting on a hemocytometer. Experiments were repeated at least 3 times independently and statistical analyses (ANOVA or Student’s t test) performed to assess significance of the data.
Patient selection and biomarker statistical analysis
This retrospective study was approved by the Institutional Review Board at the University of Iowa. Archival FFPE PDAC tissue samples from patients who underwent pancreatic resection at the University of Iowa Hospitals and Clinics between 2000 and 2010 were used for RABL6A IHC analyses. Clinicopathologic variables were collected from Pathology and SEER (Surveillance, Epidemiology and End Results) Databases, which included survival as well as American Joint Committee on Cancer (AJCC) T stage and N status, margin status, and tumor grade. Patients with M1 disease or death within 90 days from resection were excluded. A total of 73 patients met criteria for analysis.
Kaplan–Meier methods were used to estimate and plot overall survival as a function of time relative to RABL6A expression levels in the tumors. Subjects who were alive at the end of follow-up were treated as censored observations in the analysis. Estimated hazard ratios and 95% confidence intervals were generated using a Cox proportional hazard model to compare survival between the RABL6A tumor score groups. Chi-square tests were used for tests of associations between tumor scores and categorical clinicopathologic variables, and ANOVA for continuous variables. All tests were 2-sided and carried out at the 5% level of significance. Analyses were performed with the SAS 9.3 software package (Cary, NC).
Immunohistochemistry
The diagnostic PDAC archival patient tumor specimens were originally fixed in 10% neutral FFPE. Tissue sections cut at 4 µm were placed on Superfrost Plus (Fisher Scientific, Waltham, MA) slides and IHC was performed using the Envision+ System-HRP (DAKO, Carpinteria, CA) method. For staining with the RABL6A polyclonal antibody (pAb), antigen unmasking was accomplished by microwaving the sections in citrate buffer pH 6.0 at 1,000 W for 2 cycles, 4 minutes followed by 3 minutes. After blocking the endogenous peroxidase activity with 3% hydrogen peroxide for 8 minutes, the sections were stained with the RABL6A pAb. Slides were then incubated with Envision+ System HRP labeled polymer anti-rabbit detection system (DAKO). For staining with a mouse monoclonal antibody (MAb) to human RABL6A, antigen unmasking was accomplished using citrate buffer pH 6.0 in a Decloaking Chamber (Biocare Medical, Concord, CA). After blocking the endogenous peroxidase activity with 3% hydrogen peroxide for 8 minutes, the sections were blocked using the Avidin/Biotin kit (Vector Labs, Burlingame, CA) and stained with the RABL6A MAb. The slides were then incubated in Envision+ System-HRP labeled polymer anti-mouse detection system (DAKO).
The primary antibody signal for RABL6A staining with both pAb and MAb was demonstrated with DAB+ Substrate Chromogen System (DAKO) and enhanced with DAB Enhancer (DAKO). The sections were counterstained with Harris hematoxylin. Matching IgG isotype was substituted for the primary antibody to serve as the negative control. All washes were performed in DAKO 1× buffer. IHC stains of RABL6A in FFPE pellets of Panc-1 cells expressing CON or RABL6A knockdown shRNA were included as positive and negative controls, respectively, for stains of patient samples.
Two anatomic pathologists independently and blindly evaluated the RABL6A IHC staining (RWA, JAW). IHC staining intensity in the ductal adenocarcinoma was quantified relative to adjacent benign ducts within the same slide for each tumor. Scoring for RABL6A staining was as follows: −2 (undetectable compared to adjacent benign ducts), −1 (detectable but modestly decreased compared to adjacent benign ducts), 0 (equivalent to adjacent benign ducts), +1 (modestly increased compared to adjacent benign ducts), and +2 (markedly increased compared to adjacent benign ducts).
Footnotes
Acknowledgements
We are grateful for assistance from the University of Iowa Flow Cytometry Facility, Holden Comprehensive Cancer Center Population Research Core, Trent Place, and Dr. Frederick Domann for contributions to PDAC cell growth curve and qRT-PCR analyses; Dr. Chuck Lynch for assistance accessing SEER data; and Drs. John Koland and John Colgan for providing ERK and Bcl-2 antibodies.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by NIH grants R01-CA090367 (DEQ) and R21-CA127031 (DEQ) as well as The American Surgical Association Foundation Fellowship (JJM) and an American Cancer Society Seed Grant, Holden Comprehensive Cancer Center (JJM).
