Abstract
Since the discovery of Mdm2, the contribution of this RING E3 ubiquitin ligase to the pathobiology of cancer has focused almost exclusively on its role as a negative regulator of the p53 tumor suppressor. Under normal conditions, Mdm2 promotes the ubiquitin- and proteasome-dependent degradation of p53. Levels of p53 are thus kept sufficiently low to allow for cell survival and cell cycle progression. In the context of such insults as DNA damage or ribosomal stress, however, the Mdm2-p53 interaction is disrupted and p53 is stabilized. The myriad intracellular outcomes of p53 activation together comprise a robust program of tumor suppression that is short-circuited in cancer. Over half of all human malignancies are known to have lost p53 expression or sustained
Introduction
The
Mdm2 binds the N-terminal of p53 and masks the transactivation domain required for p53 transcriptional activity.4,5 Mdm2 also functions as a RING finger E3 ubiquitin ligase that targets p53 for degradation via the ubiquitin-proteasome system.6-8 The RING finger further comprises a binding site for MdmX, a closely related heterodimerization partner of Mdm2 that has been reviewed elsewhere. 9 Multiple cellular stresses are capable of disrupting the Mdm2-p53 interaction. Proteins involved in the DNA damage response can promote p53 stability and activation following genotoxic stress, for example, through a series of posttranslational modifications on both Mdm2 and p53.10,11 In response to oncogenic signaling stress, the p14ARF protein is transcriptionally upregulated and binds the central domain of Mdm2, impairing the ability of Mdm2 to regulate p53. 12 Similarly, nucleolar stress can result in the release of ribosomal proteins from the nucleolus, several of which have been shown to bind Mdm2 and stabilize p53 (Fig. 1). 13
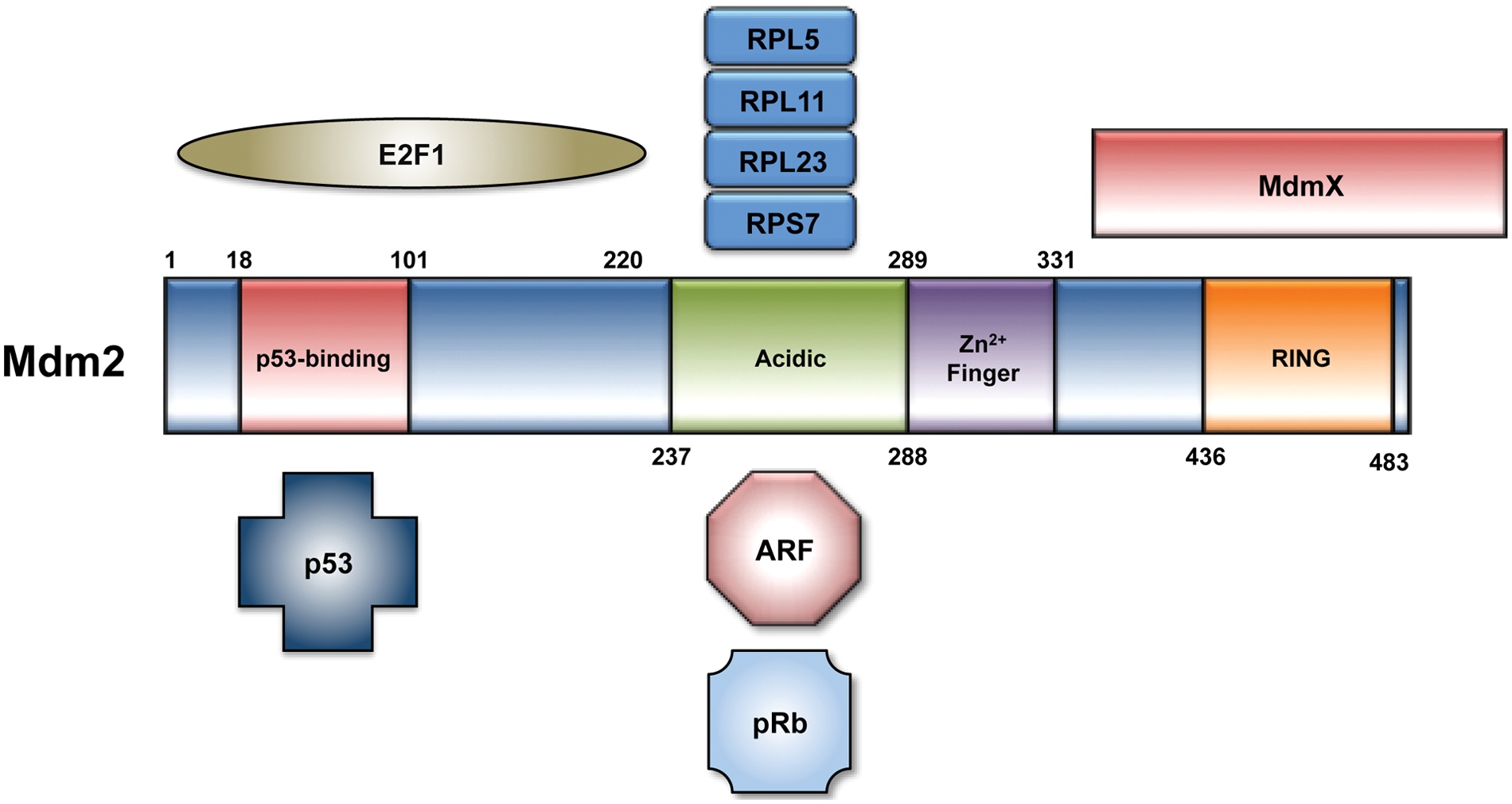
The 491 amino acids that comprise the Mdm2 protein can be subdivided into an N-terminal p53 binding domain, a central acidic domain and adjacent zinc finger domain, and a C-terminal RING finger E3 ubiquitin ligase domain. There is considerable overlap across the N-terminal domain of Mdm2 that binds p53 and E2F1. Key regulators of Mdm2 activity, including ARF and ribosomal proteins, interact with Mdm2 at its acidic domain. Mdm2 has also been shown to require residues 254 to 264 in this region for its interaction with pRb.
When activated, p53 positively regulates Mdm2 expression.
14
It has recently been proposed that phosphorylation of Mdm2 at serine 395, conferred by the ATM kinase in the setting of DNA damage, promotes allosteric remodeling of the Mdm2 RING domain, binding of p53 mRNA, and enhanced p53 protein translation.15,16 The 2 proteins thus participate in an autoregulatory feedback loop that both restrains p53 function under normal conditions and drives p53 activation under stress. The importance of this interplay, at least from a development standpoint, is evidenced by the finding that
These and additional data from human tumors collectively support an oncogenically relevant role for Mdm2 that rests on its ability to negatively regulate the stability and transcriptional activity of p53, thereby abrogating the tumor-suppressive effects of this pathway. Nevertheless, investigations in this field warrant reexamination. Methods employed to drive Mdm2 overexpression in cell culture, for example, have not yielded consistent results. Divergent experimental strategies have yielded data supporting discrepant conclusions regarding the role of Mdm2 in tumorigenesis that remain unresolved. These and other aspects of this work will be discussed in greater detail in the sections that follow.
Modeling Mdm2 Overexpression in Cells: Different Contexts, Different Methods, and Contradictory Conclusions
Initial studies of Mdm2 in cell culture suggested that Mdm2 was an oncogene.2,3 These investigations employed a cosmid vector harboring the bacterial
Although these early findings en- dorsed an oncogenic role for Mdm2, several aspects of these studies merit further discussion. First, the number of foci observed in REFs that received both
Results of more recent studies investigating the effects of Mdm2 overexpression in cells suggest that Mdm2 may be antiproliferative under certain circumstances (Table 1). Nontransformed human and rodent cells can select against Mdm2 expression in the formation of stable clones.
19
The same effect has not been consistently observed in tumor lines, although the reasons for discrepancies between studies are not clear.
7
Nevertheless, nontransformed cells transiently transfected with full-length Mdm2 cDNA arrest at G0/G1 when analyzed by flow cytometry, whereas tumor-derived cells do not.
19
Through the use of deletion mutants, growth inhibitory domains of Mdm2 have been mapped to amino acid residues 155 to 220 (ID1) and 270 to 324 (ID2). The absence of one or both of these domains has been shown to enhance the tumorigenicity of transfected 3T3 cells in nude mice, and neither domain appears to be directly involved in the physical interaction of Mdm2 with p53.
19
Mutations in ID2 have also been observed in
Summary of Selected Studies of Mdm2 Overexpression in Cell Culture
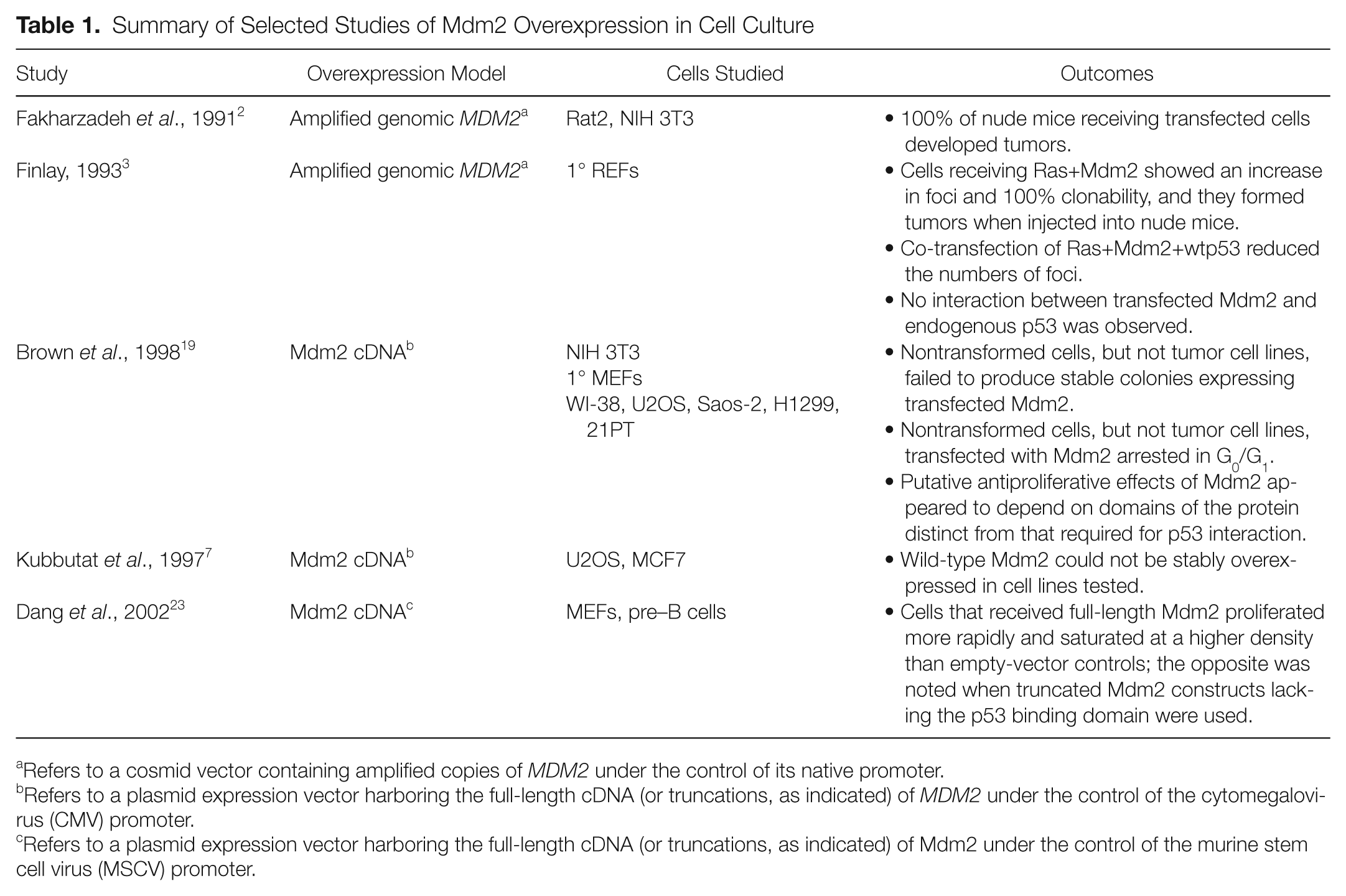
Refers to a cosmid vector containing amplified copies of
Refers to a plasmid expression vector harboring the full-length cDNA (or truncations, as indicated) of
Refers to a plasmid expression vector harboring the full-length cDNA (or truncations, as indicated) of Mdm2 under the control of the murine stem cell virus (MSCV) promoter.
The observation that exogenous Mdm2 can prevent progression through the cell cycle suggests that Mdm2 may be functioning more like a tumor suppressor than an oncogene. Although these findings are at odds with data from earlier investigations, the methods used to drive Mdm2 overexpression have not been uniform. Groups that embraced an oncogenic role for Mdm2 reported differences in the transformation ability of the
Many p53-independent functions of Mdm2 have been suggested in the wake of these findings. A role for Mdm2 in promoting S phase entry through its interactions with pRb and E2F1, for example, has been the subject of much study and is discussed below. Indeed, a growing body of work, including Mdm2 overexpression
Selected Mouse Models of Mdm2 Overexpression
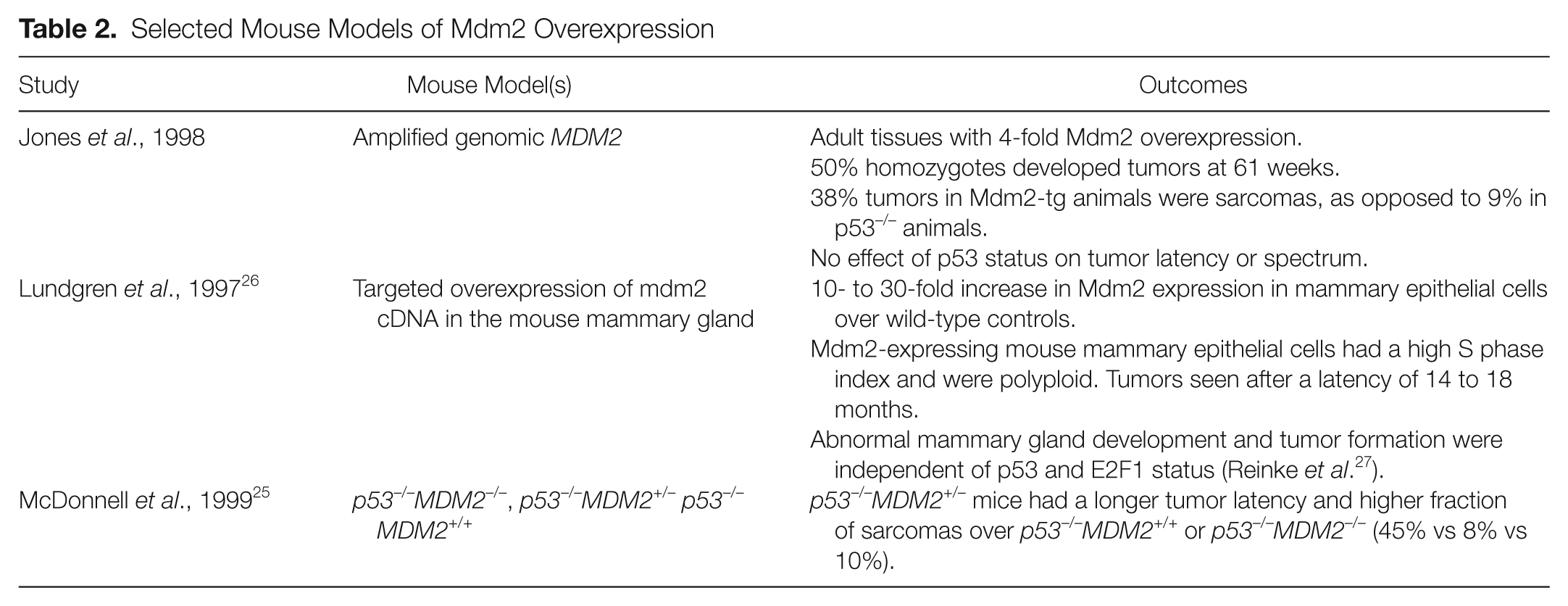
Mice Overexpressing Mdm2 Develop Spontaneous Tumors along a Histopathologic Spectrum That Is Distinct from p53 −/ − Mice and Display Phenotypes Independent of p53 Status
Jones
The Mdm2-transgenic mice in this study developed tumors at an accelerated rate compared with wild-type mice. Mice homozygous for the transgene (
A transgenic mouse with targeted overexpression of Mdm2 to the mammary gland has also been described.
26
To generate this mouse, investigators employed an
An Expanding Set of p53-Independent Functions Suggests Unique Contributions of Mdm2 Overexpression to Tumorigenesis
The ability to interact with the pRb-E2F1 complex and drive E2F1-dependent transcription and cell cycle progression was among the first p53-independent functions of Mdm2 to be proposed. An initial study showed that Mdm2 was able to directly bind E2F1 and its heterodimerization partner DP-1.
28
The interaction between Mdm2 and E2F1, but not Mdm2 and DP-1, was shown to stimulate E2F1 transcriptional activation by reporter assay, although a mechanism was not clear. These results, which were generated in a
More recent efforts have attempted to elucidate the mechanism by which Mdm2 stimulates a G1-S transition through its interaction with either pRb or E2F1. Mdm2 preferentially binds hypophosphorylated pRb and is able to decrease recruitment of pRb-E2F1 complexes to DNA. 33 Failure to recruit pRb, a well-known transcriptional repressor, to promoters containing E2F sites may permit inappropriate upregulation of factors governing S phase entry.34,35 Several groups have also shown that Mdm2 promotes the proteasomal degradation of pRb, although whether this outcome is dependent on the ability of Mdm2 to ubiquitinate pRb is not clear.32,36,37 Other studies have suggested that Mdm2 controls the intracellular level of the E2F1-DP1 complex, perhaps by regulating binding of E2F1 to the SCFSkp2 E3 ubiquitin ligase, so as to maintain levels below threshold that would otherwise trigger E2F1-dependent apoptosis in p53-null cells yet high enough to promote S phase entry.30,31 Mdm2 may thus perturb the cellular response to oncogenic stress, for example, by optimizing levels of E2F1 and promoting a pro-survival, antiapoptotic response to this protein.
Mdm2 has since been shown to influence the level or function of several other proteins involved in cell cycle control and apoptosis independent of its effects on p53. For example, Mdm2 promotes the ubiquitin-dependent proteasomal degradation of the FOXO family of transcription factors, which regulate p27, cyclin D1, TRAIL, and other genes involved in cell proliferation and survival. 38 In addition, the ability of Mdm2 to selectively ubiqutinate Erk-phosphorylated FOXO3a suggests that Mdm2 overexpression may cooperate with activated mitogenic signaling cascades to promote tumorigenesis. Mdm2 can also interact with the IRES on the 5′-UTR of the mRNA encoding the antiapoptotic protein XIAP. 39 By enhancing the translation of XIAP, Mdm2 overexpression may blunt the apoptotic response to DNA damage and confer radioresistance. 39 Finally, Mdm2 can bind to and impair the function of Nbs1, a critical mediator of the DNA double-strand break repair response, delaying the resolution of DNA lesions and promoting genomic instability. 40
Mdm2 and Cancer: A Prognostic and Therapeutic Opportunity Leaps from Past to Present
Although several efforts have been made to correlate Mdm2 status with clinical outcomes in human cancers, the role of Mdm2 in sarcomas has been of particular interest.
Studies of Mdm2 in human tumors have been extended to other malignancies, but these reports have undercut the mutually exclusive relationship between
Taken together, studies addressing Mdm2 in human tumors present contradictory clinical implications of its overexpression across a broad spectrum of malignancies.
48
One concern is the use of IHC to detect either Mdm2 or p53, particularly due to the use of different antibodies to Mdm2. Total Mdm2 protein levels assessed in this manner may not account for mutant or alternatively or aberrantly spliced Mdm2. Attempts to assign a prognostic value to Mdm2 overexpression alone and in the setting of
It has been proposed that therapies targeting the interaction between Mdm2 and p53 represent a possible means of pharmacologically reactivating the p53 pathway, and a number of such agents are currently in clinical trials.50,51 A more thorough understanding of the consequences of Mdm2 overexpression, particularly those that are independent of its roles as a regulator of p53, will greatly inform this therapeutic strategy.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
