Abstract
Mdm2 is an essential regulator of the p53 tumor suppressor. Mdm2 is modified at transcriptional, post-transcriptional, and post-translational levels to control p53 activity in normal versus stressed cells. Importantly, errors in these regulatory mechanisms can result in aberrant Mdm2 expression and failure to initiate programmed cell death in response to DNA damage. Such errors can have severe consequences as evidenced by tumor phenotypes resulting from amplification at the
Introduction
Murine double minute 2 (Mdm2) is a RING finger containing E3 ubiquitin ligase that negatively regulates p53 by inhibiting p53-mediated transactivation of target genes and targeting it p53 for proteosomal degradation (reviewed in Marine
The mouse models described above demonstrate that Mdm2 and p53 expression levels must be tightly regulated for normal embryonic development and homeostasis. Errors in this signaling network can lead to aberrant expression of effectors of the p53 pathway, resulting in tumorigenesis.10,11 For example, approximately 30% of all sarcomas bear
Taken together, these
Proteins That Interact with Mdm2
Yeast 2 hybrid screens, mass spectrometry, and immunoprecipitation experiments have identified a large number of proteins that physically interact with
Effectors That Interact with MDM2
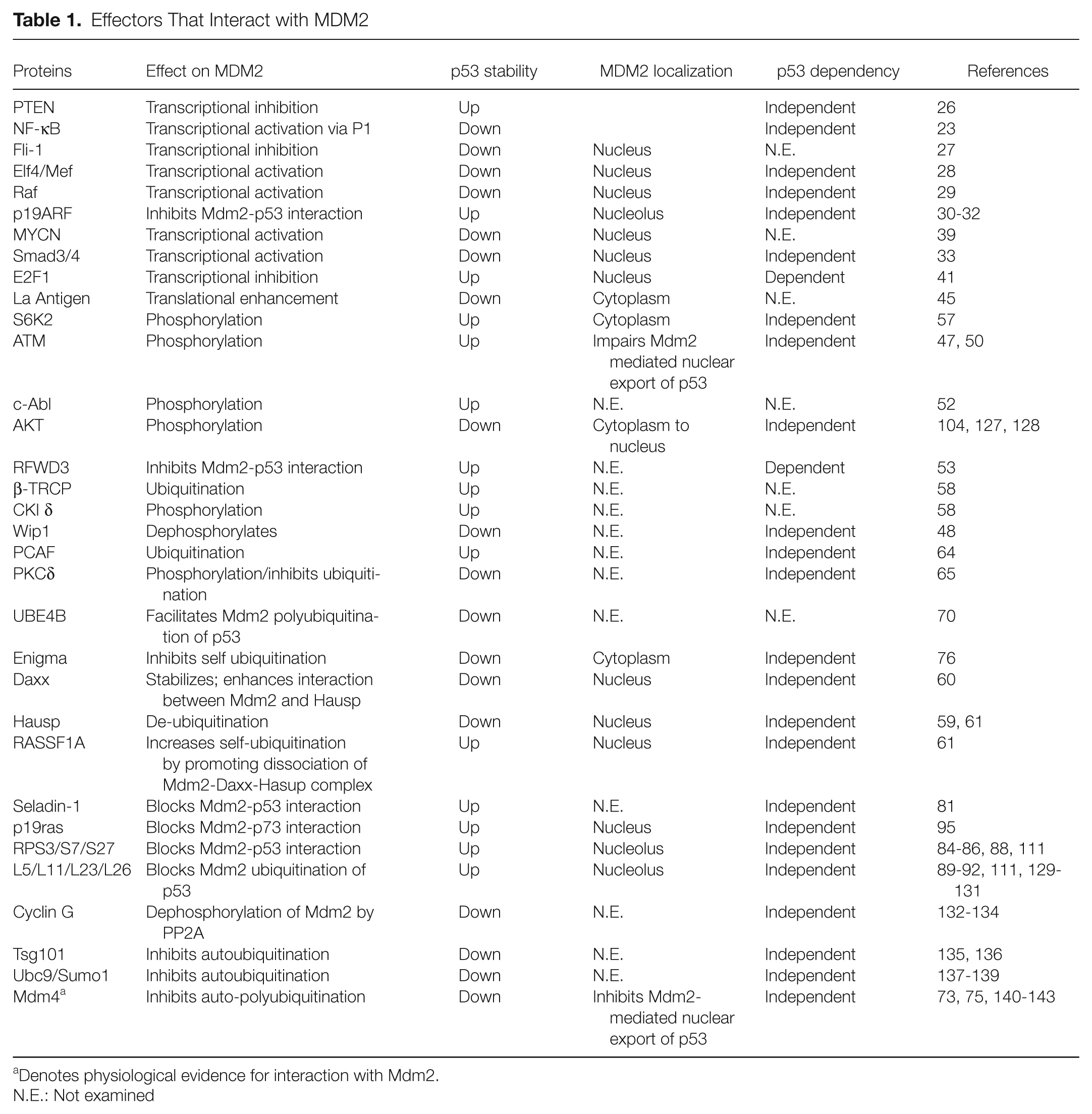
Denotes physiological evidence for interaction with Mdm2.
N.E.: Not examined
Downstream Targets of MDM2
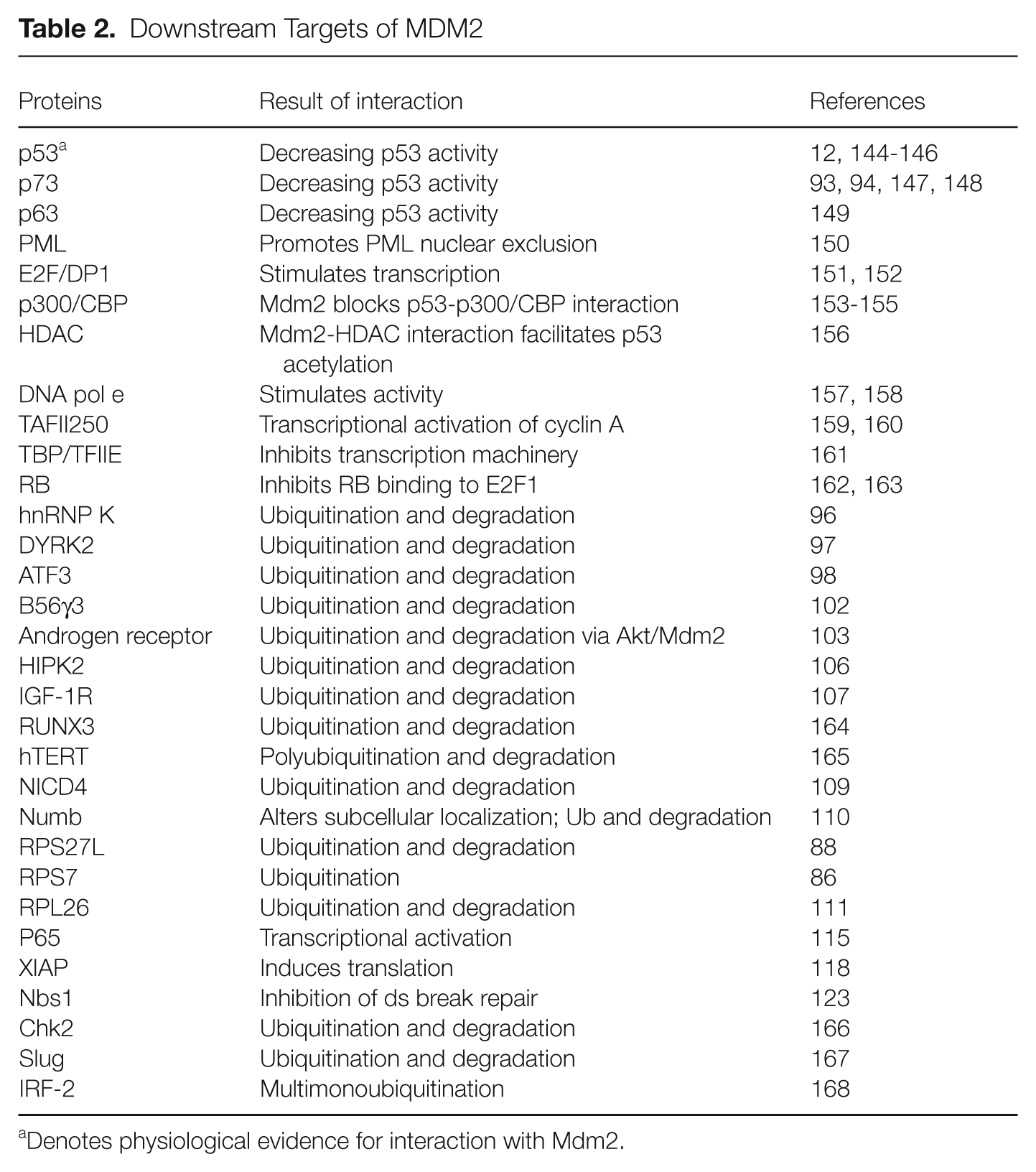
Denotes physiological evidence for interaction with Mdm2.
Effectors of Mdm2
The majority of proteins in this group regulate
Transcriptional regulation of Mdm2
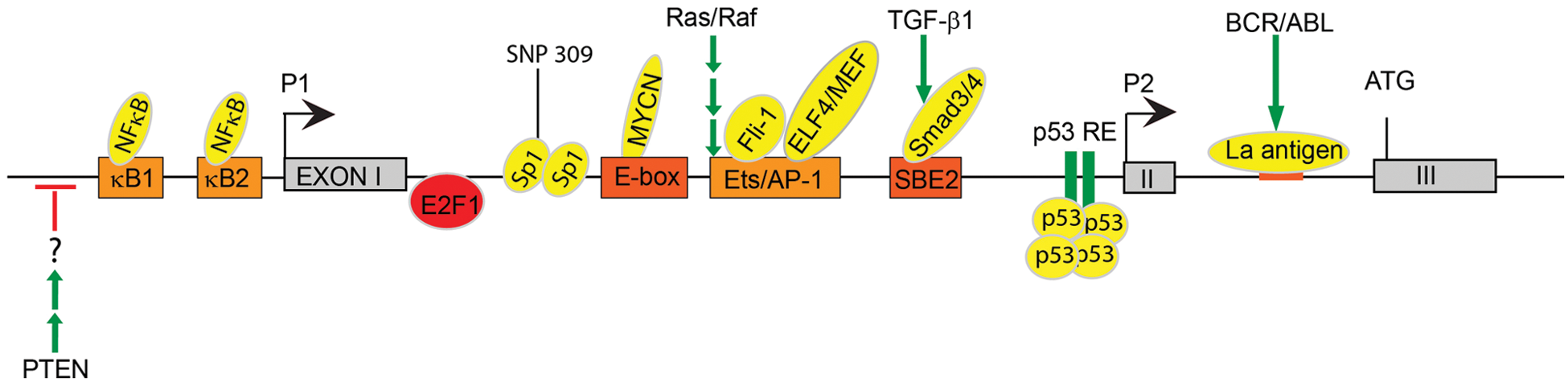
Transcriptional and post-transcriptional regulation of Mdm2.
Unlike the P1 promoter, the inducible P2 promoter contains several known response elements for transcription factors that either inhibit or activate
Mdm2 transcription is also induced by activated Ras and Raf via Ets/AP-1 motifs in the Mdm2-P2 promoter.
29
Expression of a constitutively active Ras or Raf allele in
TGF-β1 signaling also induces
The MYCN oncogene is a member of the MYC family of transcription factors and is amplified in a variety of tumors, including neuroblastomas. 34 MYCN induces both cell proliferation and apoptosis, which are opposing cellular processes. MYCN shortens the G1-S transition and increases cell proliferation35-37 while it simultaneously induces p53-mediated apoptosis (reviewed in van Noesel & Versteeg 38 ). MYCN functions as an oncogene when its pro-proliferative effects outweigh its pro-apoptotic effects, resulting in escape from p53-mediated apoptosis. A ChIP screen using an MYCN-amplified neuroblastoma cell line revealed that MYCN binds to a consensus E-box within the human MDM2-P2 promoter (Fig. 1). Accordingly, induction of MYCN resulted in increased endogenous MDM2 mRNA and MDM2 protein, and inhibition of MYCN resulted in decreased MDM2 expression. These data indicate that the MYCN oncogene inhibits p53 mediated apoptosis in neuroblastoma cells by direct binding to and activation of the MDM2 promoter. 39
Although most reported Mdm2 effectors positively regulate
Last, Mdm2 itself is transcriptionally regulated by wild-type p53 via p53 response elements located in the
Given that Mdm2 expression must be tightly regulated to ensure proper p53 expression and activity, deregulation of any of these bindings protein can affect Mdm2 levels and potentially lead to tumorigenesis. For example, a T-to-G single nucleotide polymorphism (SNP) in the P2 promoter was identified in humans that increases the binding affinity of the Sp1 transcriptional activator for the Mdm2 promoter (Fig. 1). This leads to increased Mdm2 protein levels and, consequently, decreased protein stability. Importantly, the MDM2SNP309G allele is correlated with increased cancer risk in humans. To directly examine the effect of this SNP in tumorigenesis, Post
Post-transcriptional regulation of Mdm2
Mdm2 is also regulated at the post-transcriptional level by an RNA binding protein named La antigen. The BCR/ABL oncoproteins are responsible for the transformation of the hematopoietic stem cells resulting in chronic myelogenous leukemia (CML). Recent data show that BCR/ABL upregulates
Post-translational regulation of Mdm2
During the DNA damage response (DDR), p53 is stabilized through post-translational modification of Mdm2 (Fig. 2). For example, after DNA damage, Mdm2 is phosphorylated by ATM at serine 395 (Fig. 2). This phosphorylated Mdm2 species cannot efficiently promote translocation of p53 from the nucleus to the cytoplasm, 47 resulting in increased p53 stability. Wild-type p53-induced phosphatase (Wip1), which is a p53 target gene, can dephosphorylate this same Mdm2 residue. This results in stabilization of Mdm2 and subsequent ubiquitination and degradation of p53. 48 Thus, the phosphorylation status of Mdm2-S395 regulates the p53 response after DNA damage and implicates Wip1 as a potential oncogene. Interestingly, Wip1 is amplified and overexpressed in a number of human cancers (reviewed in Le Guezennec & Bulavin 49 ).
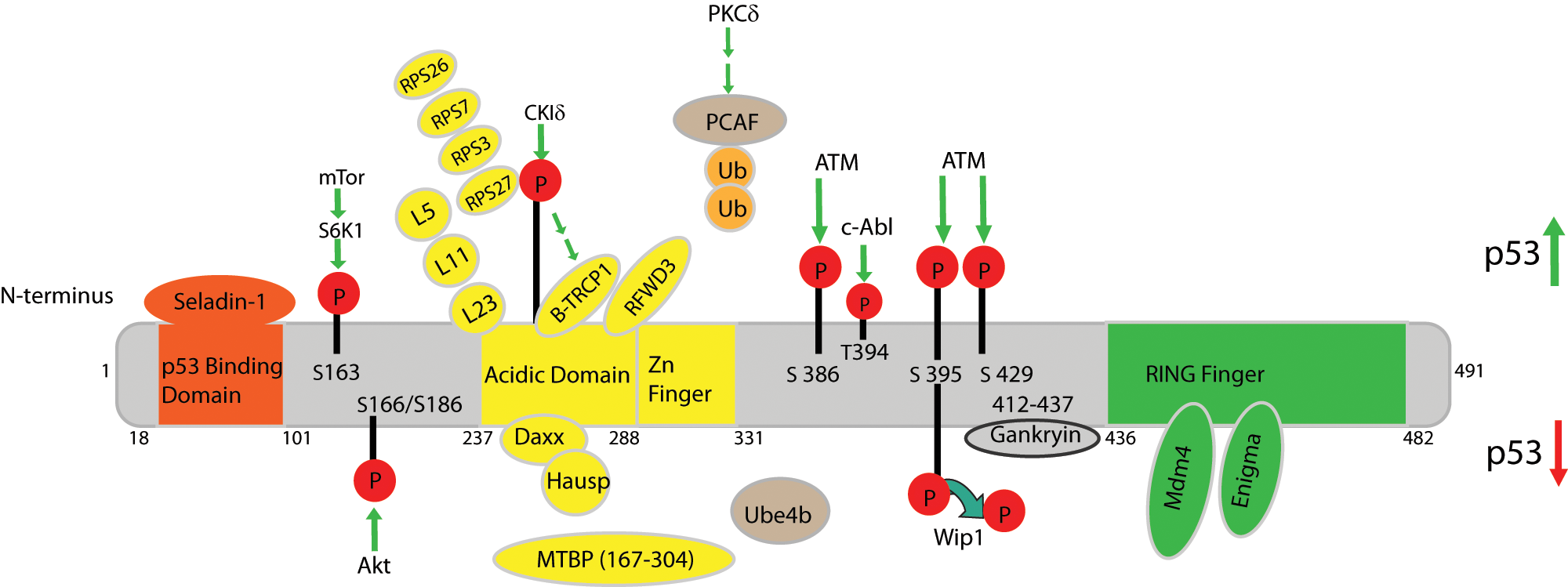
Effectors of MDM2. Schematic representing the characterized human MDM2 domains and known binding partners. Human MDM2 contains 491 amino acids and has 4 characterized domains: residues 18-101 comprise the p53 binding domain, residues 237-288 contain the acidic domain, which is adjacent to the Zinc finger domain, and the RING finger domain is within residues 436-482. MDM2 binding partners are depicted in orange, yellow, or green depending on the region of MDM2 to which they bind (orange = p53 binding domain, yellow = acidic domain/zinc finger, green = RING finger domain). Proteins whose binding locations on MDM2 are unknown are depicted in tan. Interactions that result in increased p53 stability and activity are depicted above the schematic, whereas interactions that result in decreased p53 stability and activity are shown below. P = phosphorylation; Ub = ubiquitination.
Other groups confirmed S395 as a direct substrate of ATM and revealed 2 additional phosphorylation sites (S386/429) adjacent to the RING domain (Fig. 2). Treatment with IR revealed that ATM phosphorylation of Mdm2 at these 2 sites prevents Mdm2-mediated polyubiquitination of p53 and is necessary for p53 stabilization.
50
ATM also activates the c-Abl kinase in response to DNA damage.
51
C-Abl in turn binds to and phosphorylates human MDM2 at Tyr 394, resulting in increased p53 activity.
52
In addition, the novel protein, E3 ligase RING finger and WD repeat domain 3 (RFWD3), which is another substrate of the checkpoint kinases ATM/ATR, was recently shown to interact with Mdm2 and p53, forming an RFWD3-Mdm2-p53 ternary complex.
53
Deletion analyses indicate that the Mdm2 acidic domain is required for its interaction with RFWD3. Consistent with its role as a negative regulator of Mdm2, overexpression of RFWD3 protects p53 from degradation in the presence of high Mdm2 levels after DNA damage. RFWD3 restricts polyubiquitination of p53 by Mdm2, resulting in increased p53 stability, thus inhibiting Mdm2-p53 interactions.
53
Together, these data indicate that ATM-mediated phosphorylation of Mdm2 is important for regulating the p53 response after DNA damage. Further studies using mouse models that disrupt ATM/c-Abl mediated phosphorylation and Wip1-mediated dephosphorylation of Mdm2 will be needed to determine the
mTOR is an ATM/ATR-related protein kinase that functions to regulate cell proliferation, metabolism, and growth.
54
S6K1 is a downstream signaling molecule of mTOR
55
that promotes translation of various proteins through phosphorylation of the eukaryotic translation factor, eIF4B, and the ribosomal protein S6 (reviewed in Averous & Proud
56
). A recent study shows that genotoxic stress caused by treatment with doxorubicin (which causes single and double stranded DNA breaks) results in phosphorylation of Mdm2 S163 by S6K1 in primary MEFs and 293 cells. Furthermore, immunoprecipitated S6K1 phosphorylated Mdm2 S163
Mdm2 stability is also regulated by the F-box ubiquitin ligase, β-TRCP, in a phosphorylation dependent manner. In response to DNA damage, Mdm2 interacts with β-TRCP1 and β-TRCP2
HAUSP, a ubiquitin-specific protease, and the death domain-associated protein (Daxx) also regulate Mdm2 stability. Disruption of
These data elucidate several mechanisms governing the cis- and trans-E3 activities of Mdm2 in stressed versus normal cells and demonstrate that Daxx plays a pivotal role in regulating Mdm2 self-ubiquitination.
P300-CBP-associated factor (PCAF), which is a histone acetyltransferase, exhibits intrinsic E3 ligase activity toward Mdm2. Ubiquitination of Mdm2 by PCAF is necessary for Mdm2 degradation and p53 activation in U2OS cells after DNA damage. 64 PCAF mediated ubiquitination appears to be regulated at least in part by the protein kinase, PKCδ. Overexpression of PKCδ is capable of inhibiting PCAF-mediated Mdm2 ubiquitination. Although the mechanism has not been fully elucidated, depletion of PKCδ results in down-regulation of Mdm2 protein expression independent of p53. 65 Taken together, these results suggest that PKCδ may regulate PCAF-mediated ubiquitination of Mdm2 during times of cellular stress.
As mentioned above, polyubiquitination of p53 by Mdm2 is necessary for proteosomal degradation. However, Mdm2 catalyzes p53 mono- or multiple monoubiquitinations
Since tight regulation of the antiproliferative and proapoptotic function of p53 is essential for cell viability, cells use several mechanisms to ensure proper Mdm2 expression and p53 activity during embryonic development and homeostasis. One mechanism involves Mdm4, a homolog of Mdm2 that negatively regulates p53 by binding.
71
Accordingly, genetic ablation of
Mdm2 also binds the LIM domain protein, Enigma, to regulate p53 stability and activity in unstressed cells. Endogenous Enigma co-immunoprecipitates with endogenous Mdm2 in the presence or absence of p53, forming a ternary complex. The C terminus of Mdm2 was sufficient for its binding to Enigma (Fig. 2). This interaction functions to inhibit Mdm2 self-ubiquitination and enhances Mdm2-mediated ubiquitination of p53. Interestingly, Enigma was co-expressed with Mdm2 in 10 cases of human liver and stomach tumors where p53 was undetectable and in 7 cases where p53 was detected. Tissue arrays revealed that Enigma co-localized with Mdm2 in the cytoplasm of human stomach and colorectal tumor cells. 76 The Enigma-Mdm2-p53 ternary complex was detected in both the cytosolic and nuclear fractions, suggesting that Enigma regulation of Mdm2 may function to decrease p53 activity mostly in the cytosol in proliferating cells but may also suppress p53 transactivation in the nucleus. 76 This is interesting given the finding that p53 activates the proapoptotic protein, BAX, in the cytoplasm.77-79 Taken together, these results indicate that Enigma may inappropriately stabilize Mdm2 in some human tumors, resulting in destruction of p53.
Gankyrin is another protein that regulates Mdm2 mediated ubiquitination and degradation of p53. Gankyrin, which is an ankyrin repeat oncoprotein overexpressed in hepatocellular carcinomas, interacts with the S6 proteasomal ATPase to increase degradation of the tumor suppressor Rb. Similarly, U2OS cells stably overexpressing Gankyrin exhibit decreased stability of endogenous p53 protein only in the presence of Mdm2, indicating that gankyrin enhanced Mdm2 mediated degradation of p53. Co-immunoprecipitation experiments demonstrate that endogenous gankyrin physically interacts with Mdm2 but not with p53, and ubiquitination assays show that gankyrin increases Mdm2 mediated mono- and polyubiquitination of p53. 80
A genetic screen revealed that following oncogenic and oxidative stress, Seladin-1, which is activated by Ras pathway signaling, directly binds p53 and Mdm2 to regulate p53 expression.
81
Co-immunoprecipitation assays reveal binding between Seladin-1 and the N-terminus of p53, which displaces Mdm2 from p53, resulting in p53 accumulation. These results indicate that Seladin-1 regulates p53 stability by binding to Mdm2 and interfering with Mdm2-mediated p53 ubiquitination. In addition, Seladin-1 binds to Mdm2 in
The majority of Mdm2 binding partners mentioned thus far regulate the MDM2-p53 feedback loop during homeostasis and in response to DNA damage. This feedback loop is also regulated in response to ribosomal stress or nucleolar stress.
82
Ribosome biogenesis involves expression of ribosomal RNA and ribosomal proteins (RPs), processing of rRNA, and subsequent assembly of the ribosome subunits in the nucleolus. The subunits then translocate to the cytoplasm, where they undergo further assembly and ultimately catalyze protein synthesis.
83
Ribosome biogenesis is the most energy demanding cellular process, and disruption of any step results in “nucleolar stress.” This triggers binding of several RPs (RPS3,
84
RPS7,85,86 RPS14,
87
RPS27,
88
RPL5,
89
RPL11,
90
RPL23,
91
RPL26
92
) to MDM2 (Fig. 2), inhibiting its binding to and degradation of p53, resulting in p53 stabilization and activation (Table 1). Given that these Mdm2-interacting RPs all regulate the p53 response to nucleolar stress in a similar manner, it is possible that each RP may regulate different steps of the nucleolar stress response. In addition, it is possible that a combinatorial effect of RP binding to Mdm2 is required for a strong stress response. These ideas can be tested
As mentioned above, Mdm2 regulates the p53 family member, p73. Mdm2 binds to p73 and suppresses transcription without targeting it for degradation.93,94 Jeong
Downstream Targets of Mdm2
Ubiquitin-Dependent Proteasomal Degradation
Since Mdm2 is an E3 ubiquitin ligase, the majority of Mdm2 targets, including p53, are ubiquitinated by Mdm2 and targeted for proteasomal degradation. A complete list of these proteins is provided in Table 2. Below we discuss in more detail the nature and significance of the reported interactions.
Immunoprecipitation experiments in U2OS cells reveal that in the absence of cellular stress, Mdm2 binds to and ubiquitinates the heterogeneous nuclear ribonucleoprotein K (hnRNP K), resulting in its proteasomal degradation. In response to cellular stress, however, Mdm2 is inhibited in an ATM dependent manner, leading to the rapid accumulation of hnRNP K. Interestingly, hnRNP K also functions as an important co-factor for p53 in response to DNA damage, as evidenced by decreased activation of p53 target genes after hnRNP K depletion. 96
Under normal growth conditions, Mdm2 also interacts with and ubiquitinates the p53 inducer, dual specificity tyrosine-phosphorylation-regulated kinase 2 (DYRK2), resulting in its degradation. However, under genotoxic stress, ATM phosphorylates DYRK2, enabling it to escape Mdm2-mediated degradation, resulting in its translocation to the nucleus, where it phosphorylates p53, inducing apoptosis. 97
The activating transcription factor 3 (ATF3) is a stress sensor whose induction is important for cellular responses to DNA damage. During times of normal growth, Mdm2 binds to the ATF3 leucine-zipper domain and ubiquitinates ATF3, resulting in its degradation. Importantly, deregulation of ATF3 is documented to contribute to tumorigenesis, presumably through lack of appropriate DNA damage responses. 98
Protein phosphatase 2A (PP2A) is a serine-threonine phosphatase involved in the DNA damage response. PP2A is a heterotrimer comprised of a catalytic C subunit, a structural A subunit, and several regulatory B subunits. Recent work has demonstrated that PP2A may function as a tumor suppressor (reviewed in Wurzenberger & Gerlich 99 ). Specifically, the PP2A B56γ3 isoform inhibits cell proliferation in human lung cancer cell lines. 100 Under normal growth conditions, p53 is phosphorylated by TAF1 at Thr55, rendering it inactive, but after DNA damage, B56γ-PPA dephosphorylates p53 at this residue, leading to its activation. However, phosphorylation of p53 at Ser15 by ATM is required for B56-p53 interaction and subsequent dephosphorylation at Thr55, indicating that ATM phosphorylation at Ser15 primes p53 for its interaction with B56-PPA. 101 Since B56γ3 can activate p53 during times of normal cell growth, B56γ3 must be negatively regulated, and this is achieved through Mdm2 mediated ubiquitination and proteasomal degradation. After DNA damage, B56γ3 is phosphorylated by ATM, which blocks Mdm2-mediated B56γ3 ubiquitination. ATM also phosphorylates Mdm2 at serine 395 after DNA damage, preventing Mdm2 polyubiquitination of p53.50,102 Thus, multiple mechanisms ensure p53 activity after DNA damage.
The androgen receptor (AR) is a phosphoprotein that regulates a variety of biological functions including prostate cell growth and apoptosis via the p53 pathway. Therefore, AR levels must be tightly regulated, and defects in this regulation are implicated in prostate tumorigenesis. One mechanism for regulating AR levels involves the Mdm2 and Akt pathway.
103
Mdm2, AR, and Akt form a complex
The homeodomain-interacting protein kinase, HIPK, is a serine-threonine kinase that functions to repress homeodomain transcription factors. As mentioned, after DNA damage, p53 induces cell cycle arrest, senescence, or apoptosis. p53 phosphorylates HIPK2 only after severe, unrepairable DNA damage, such as after UV radiation, resulting in apoptosis. However, during times of moderate DNA damage using sublethal doses of doxorubicin, the HIPK2 apoptotic activator must be silenced, and this is achieved through Mdm2 mediated ubiquitination and degradation of HIPK2. 106
The insulin-like growth factor 1 receptor (IGF-1R) is a receptor tyrosine kinase that is implicated in several cancers as a result of its antiapoptotic properties. Mdm2 physically associates with and ubiquitinates IGF-1R
Notch receptors and ligands function during development and are implicated in tumorigenesis. The Notch4 receptor, which promotes tumorigenesis in mouse mammary epithelium,
108
was recently shown to be a target of Mdm2 ubiquitination. Co-immunoprecipitation experiments in
As discussed in the previous section on Mdm2 effectors, several ribosomal proteins (RPs) bind to Mdm2, inhibiting its binding to p53. This ultimately results in p53 stabilization and activation. However, the interaction between Mdm2 and RPL26 performs a different function. Mdm2 acts as a ubiquitin ligase for RPL26, resulting in its degradation. 111 Since RPL26 binds to the 5′UTR of p53 to increase its translation, 112 in this scenario, Mdm2 inhibits p53 translation via RPL26 degradation. These data indicate that under nonstressed conditions, Mdm2 binds to RPL26 to keep p53 levels low. However, in response to stress, Mdm2 ubiquitination of RPL26 is inhibited by unknown mechanisms, resulting in increased p53 expression. These results reveal an additional mechanism used by cells to regulate p53 expression in stressed versus normal cells.
RPS7 is another substrate for Mdm2 ubiquitination. 86 Since RPS7 can also bind Mdm2, inhibiting its interaction with p53, RPS7 is both a substrate and effector of Mdm2. 86 Finally, RPS27L, an RPS27-like protein, is also a substrate of Mdm2 ubiquitination. 88 Although both family members (RPS27L and RPS27) bind to Mdm2, only RPS27L can be degraded by Mdm2 under physiological conditions. Since RPS27L competes with p53 for binding to Mdm2, overexpression of this protein results in accumulation of p53. Given that RPS27L is a direct p53 target,113,114 these data demonstrate a feedback loop wherein in response to stress, p53 activates RPS27L, which then competes with p53 for Mdm2 binding, resulting in p53 stabilization. However, p53 also induces Mdm2, which degrades both p53 and RPS27L. Together, these results reveal a complex relationship between RPS27L, Mdm2, and p53 that could have important implications for regulation of cell proliferation under stressed conditions.
Ubiquitin Ligase Independent Functions of Mdm2
Mdm2 has E3-ligase independent functions as observed with Mdm2 dependent transcriptional activation of the NFκB subunit p65. p65 is an antiapoptotic factor expressed in neoplastic cells, namely leukemic bone marrow cells. Chromatin immunoprecipitation and EMSA demonstrate that Mdm2 directly binds the Sp1-binding site of the p65 promoter. Interestingly, Mdm2 overexpression in B-cell precursor acute lymphoblastic leukemia (BCP-ALL) and an ALL cell line (EU-4) results in increased expression of p65 and resistance to doxorubicin. Together, these results implicate Mdm2 activation of p65 in chemotherapy resistance in ALL. 115
Mdm2 can also induce translation of the Inhibitor of Apoptosis (IAP) protein family member, XIAP. XIAP is a caspase inhibitor that is overexpressed in cancer cells and confers resistance to DNA damage induced by irradiation and chemotherapy (reviewed in Galban & Duckett
116
and Dubrez-Daloz
Nbs1 is a member of the Mre11/Rad50/Nbs1DNA repair complex that functions in double strand break repair, meiotic recombination, and telomere maintenance.
119
Mutations in Nbs1 result in genetic instability syndromes such as Nijmegen breakage syndrome (NBS) and have a very high incidence of cancer as a result of defective DNA repair.120-122 Mdm2 directly binds to Nbs1 and Mdm2-Nbs1 co-localized to DNA damage sites following gamma-radiation in HeLA and IMR90 cells, inhibiting DNA double strand break repair as measured by a comet assay. Moreover, the ubiquitin ligase domain of Mdm2 was dispensable for its binding to Nbs1, and this interaction was p53 independent as it was observed in
p53 Independent Functions of Mdm2
As demonstrated by the p53-independent function of Mdm2 in inhibiting DNA double strand break repair, Mdm2 has p53-independent functions. For example, in a mouse model of breast cancer, high Mdm2 protein levels resulted in chromosomal abnormalities in a p53-independent manner.
124
Interestingly, the tumor spectrum of Mdm2 transgenic mice differs from that observed in
Concluding Remarks
Although the most documented function of Mdm2 involves its negative regulation of p53, the discovery of new Mdm2 binding partners and substrates demonstrates that Mdm2 has many functions in addition to regulating p53. Analysis of the Mdm2 binding partners presented in Tables 1 and 2 demonstrates that transcriptional, post-transcriptional, and post-translational regulation of Mdm2 is important to facilitate appropriate Mdm2 levels during development, in homeostasis, and in response to DNA damage. Importantly, errors in the transcriptional regulation of Mdm2 or post-translational modifications of Mdm2 can contribute to tumorigenesis in a p53-independent manner. Mouse models harboring genetic alterations in Mdm2 binding proteins and/or amino acid substitutions in Mdm2 phosphorylation sites will reveal the physiological relevance of these reported interactions. Understanding how Mdm2 activities are regulated
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
NIH/NCI (T32 CA009299-32 and R01 CA47296-24) to G.L.
