Abstract
Background
Bats, critical to controlling insect populations, harbor unique immune systems that can provide insight into preventing and treating diseases. Unfortunately, white-nose syndrome (WNS), caused by the fungus Pseudogymnoascus destructans, has devastated North American bat populations, killing millions since the winter of 2006 in one of the greatest wildlife disease epidemics in U.S. history. Several reports have found evidence that bat microbiomes play a role in protecting their host against WNS. However, investigations into the antifungal metabolites produced by these symbionts are extremely limited.
Objective
The aim of this study was to investigate the antifungal constituents of Streptomyces buecherae, a novel bat-associated bacterium isolated from bats in New Mexico.
Methods
To identify bioactive compounds from the complex bacterial extract, we combined untargeted mass spectrometry metabolomics with in vitro antifungal activity data against P. destructans, identifying a suite of antifungal metabolites for targeted study using biochemometric partial least squares analyses combined with molecular networking.
Results
The metabolomics analyses revealed that nigericin—a known polyether antibiotic—was the major bioactive compound in the extract, and molecular networking revealed additional nigericin analogs present in the bioactive extract. Nigericin showed moderate antifungal activity against P. destructans in a disc diffusion assay down to a concentration of 6.25 μg/disc.
Conclusions
Nigericin is the primary bioactive constituent present in S. buecherae extracts that may be responsible for the antifungal effect this bacterium shows against P. destructans in vitro. This study underscores the potential of bat-associated Streptomyces species in combatting fungal pathogens and provides insight into the chemical mechanisms involved in bat microbiome defenses against WNS.
Introduction
Bats are incredibly important organisms promoting global biodiversity and supporting the health of diverse ecosystems across the planet. Accounting for one-fifth of all mammals on Earth, bats consume copious insects that would otherwise destroy populations of many important crops. Published estimates of the economic value of bats’ pest-control activities range from $3.7 to $53 billion annually in the U.S. alone. 1 Unfortunately, in the last decade, millions of bats have been killed across the U.S. by the invasive clonal fungal pathogen Pseudogymnoascus destructans causing white-nose syndrome (WNS). 2 In response, farmers have had to increase the usage of pesticides which has in turn been linked to human infant mortality. 3 In hopes to protect precious bat populations, scientists have begun investigating the interactions between this pathogen and bats’ microbial defenses against them.
Surveys have shown that bats’ fur and skin surfaces have a high microbial diversity including some members that have antagonistic properties against P. destructans.4–8 Several studies have focused on bacterial community dynamics of bats in the Palearctic, where P. destructans originates.9–12 Palearctic bats, including Murina leucogaster, Rhinolophus ferrumequinum, and Myotis petax, can tolerate high fungal loads without developing symptoms of WNS, highlighting their immune responses developed through millennia of coevolution with P. destructans.9,11,12 The prevalence of antifungal microbes established on the skin of Palearctic bats provides further evidence that bats have achieved tolerance to P. destructans in part through their microbiome defenses.9–12 Even in North America, where bat species have varying levels of disease susceptibility, host microbial defenses may protect some bats against P. destructans colonization or development of severe disease.6,8,13,14
Bats across North America are struggling to survive the WNS epidemic posed by P. destructans, and numerous tactics have been taken to prevent infection and mitigate the spread of P. destructans including biological, chemical, immunological, and mechanical treatments.15–19 While these approaches show promise, they do require continual treatments that can be disruptive to bat populations and delicate cave ecosystems.20,21 Probiotic treatments that seek to expose bats to antifungal bacterial strains that are already naturally part of the microbiome of some bats could minimize disturbance to these cave systems, allowing beneficial bacteria that are already part of natural ecosystem to persist and propagate on their own. Probiotics are increasingly accepted as tools for wildlife infectious disease mitigation. 22 In amphibians, members of the natural microbiome have been leveraged to protect against the lethal fungus Batrachochytrium dendrobatidis causing chytridiomycosis, 23 although critiques have noted that probiotic treatments only protect amphibians for a short duration and may only be effective for certain amphibian species.24,25 In bats, results from probiotic trials have also been variable. Hooper and Amelon (2023) found that treatment with volatile organic compounds from Rhodococcus rhodochrous DAP96253, which showed in vitro antifungal activity, did not improve survival rates of little brown bats (M. lucifugus) in their field trials. 26 Conversely, Hoyt and colleagues (2019) found that using a probiotic cocktail containing four Pseudomonas fluorescens strains increased the little brown bats’ survival rate from 8.4% to 46.2%. 17 While promising, the authors failed to confirm that the bacteria were persistent after P. destructans was introduced or if the P. destructans loads in the treatment group differed from controls. It is also important to note that these preliminary probiotic studies have only focused on little brown bat populations.
A recent study evaluated antifungal activity of over 600 bacteria from 12 bat species in WNS-free New Mexico and Arizona caves against P. destructans. Thirty-six of the isolates, including 15 novel Streptomyces species, inhibited or completely stopped P. destructans growth, suggesting that bats in the American West possess specialized microbiota that may protect them against P. destructans infection. 27 Chemical investigations into the natural product(s) responsible for these inhibitory activities were not completed at that time. To address this knowledge gap, we sought to identify the chemical constituents responsible for the biological activity of one of the most potent antifungal strains from the 2017 study, the novel bat-associated species Streptomyces buecherae. 28 Given the propensity for natural products discovery efforts to reisolate known compounds, we utilized a biochemometrics approach combining chemical fingerprints obtained from mass spectrometry analyses with biological activity data into a statistical model that explains how changes in bioactivity correlate to chemical changes across samples (Figure 1) to identify expressed antifungal compounds from S. buecherae that inhibit P. destructans.

Integrated Workflow Combining Antifungal Activity Screening Results and Mass Spectral Profiling of Bioactive Bacterial Extracts to Predict Antifungal Compounds for Targeted Study. Created in BioRender. Caesar, L. (2025) https://BioRender.com/z16g949.
Experimental
General Experimental Procedures
Ultra-high performance liquid chromatography-mass spectrometry (UHPLC-MS) analysis was completed in two stages using methodologies standardized in our laboratory. Preliminary analyses of crude extracts were completed using a Thermo-Fisher Q-Exactive Plus Orbitrap mass spectrometer (ThermoFisher Scientific, MA, USA) connected to an Acquity UPLC system (Waters, Milford, MA, USA) with reversed phase UPLC column (BEH C18, 1.7 μm, 2.1 × 50 mm, Waters Corporation, Milford, MA, USA) or an Agilent 6546 q-TOF coupled to a 1290 Infinity II UHPLC with reversed phase ZORBAX RRHD Eclipse Plus C18 column (1.8 µm. 2.1 × 150 mm) (Agilent Technologies, CA, USA). All fractions were analyzed at 0.1 mg mL−1 in methanol (mass of sample per volume of solvent), with 3 μL injections. All studies utilized gradients comprised of solvent A (water with 0.1% formic acid) and solvent B (acetonitrile with 0.1% formic acid).
Preliminary studies for biochemometrics analyses were conducted on the Q-Exactive with a 0.3 mL min−1 flow rate and utilized a gradient beginning with 65:35 (A:B) from 0–0.5 min, and increased to 0:100 (A:B) from 0.5–8.5 min after which it was held at 100% B for 0.5 min, before returning to starting conditions over 0.5 min and held from 9.0–10.0 min. Analysis was performed in both positive and negative modes over a m/z range of 150–1500 with the spray voltage at 3.7 kV, capillary temperature at 257 °C, S-lens RF level at 50.00, sheath gas flow at 47.50, and auxiliary gas flow at 11.25. The four most intense ions were fragmented with HCD of 30.0. Analyses conducted during compound purification efforts were completed on the 6546 q-TOF with a 0.4 mL min−1 flow rate and utilized a gradient increasing from 95:5 (A:B) to 0:100 (A:B) over 12 min, after which it was held at 100% B for three minutes. The column temperature was kept at 50 °C. Analysis was completed in positive mode over a m/z range of 100–1700 with a capillary voltage of 3500 V and a fixed collision energy of 30 eV.
Flash chromatography was conducted with a CombiFlash NextGen 300 + system (Teledyne-Isco, NE, USA) with a photo-diode array (PDA) detector. All chemicals were acquired through Sigma-Aldrich and were spectroscopic or microbiological grade.
Bacterial Growth and Extraction
Streptomyces buecherae AC541T was isolated from the skin of a female cave myotis bat (Myotis velifer) in the Carlsbad Caverns National Park as part of a previous study, and details on the isolation process are outlined by Hamm and colleagues (2017). 27 To optimize culture conditions, we grew isolates on Reasoner's 2 Agar (R2A; Research Products International, Mount Prospect, IL), ½ strength R2A, International Streptomyces Project medium 2 (ISP2), 29 ½ strength ISP2, and potato dextrose agar (PDA; Sigma Aldrich, Milwaukee, WI). In addition to evaluating the medium itself, we also evaluated three incubation temperatures (4 °C, 20 °C, and 37 °C), two light conditions (complete darkness and a 16:8 h light:dark cycle), and three incubation durations (1 week, 2 weeks, and 4 weeks). Using this information, isolates were grown for scale up studies on ISP2 and incubated at room temperature for 14 days in darkness as this medium maximized yield of chemical extracts. After 15 days, plates were frozen, then thawed and pressed to release spent liquid media. Media was filtered and extracted using 30 mg Supel-Select HLB SPE cartridges (Supelco, PA, USA). Three separate extracts (SB-1, SB-2, and SB-3) were prepared as described using 900 plates each of S. buecherae AC541T. Given that these extracts had slightly different chemical profiles (Figure S1), we chose not to combine them for flash chromatography efforts.
Chromatographic Separation
Each extract (1004 mg, 1624 mg, and 1176 mg for SB-1, SB-2, and SB-3 respectively) was separated with normal-stage flash chromatography (24 g silica gel column) with a 50 min hexane/EtOAc/MeOH gradient at a 35 mL min−1 flow rate using methodology standardized in the laboratory. SB-1 was separated into 13 simplified fractions (SB-1-1 through SB-1-13), SB-2 into 18 simplified fractions (SB-2-1 through SB-2-18), and SB-3 into 17 simplified fractions (SB-3-1 through SB-3-17), each of which was subjected to LC-MS analysis. LC-MS analysis revealed that several fractions from the different extracts had similar chemical profiles which were combined into a single sample prior to bioactivity testing (Figure S2). These fractions were combined into new fractions SB-4-1 through SB-4-7 (see Table S1). Following combination of chemically similar samples, fractions were subjected to antifungal susceptibility testing and biochemometrics analysis.
Antifungal Activity Assays
Antifungal assays were completed using an adapted version of the Clinical and Laboratory Standards Institute (CLSI) standardized disk diffusion protocol for nondermatophyte filamentous fungi susceptibility testing. 30 Eighteen extracts and their simplified fractions were tested in duplicate using a disc diffusion assay against P. destructans strain 20631-21 (ATCC® MYA4855™) stored in −80 °C. Each extract was dissolved in methanol at a concentration of 10 mg mL−1, and 20 µL were delivered onto sterile 6 mm discs. Cavicide was used as positive control, and methanol was used as negative control. P. destructans was activated from glycerol stock on Yeast Extract Peptone Dextrose (YPD; 1% yeast extract, 2% peptone, 2% glucose) media by incubating at 11 °C-15 °C for 10 days, following which spores were harvested using 0.01% tween water and filtered using a 0.40 µm strainer. YPD agar media was prepared with 5 mL of molten agar inoculated with 106 spores, allowed to solidify, and incubated at 15 °C for 4 days. Prepared discs were added to P. destructans plates (6 discs per plate, including controls) under sterile conditions and returned to the incubator for three days, after which zones of inhibition were measured. After this point, plates were returned to the incubator and ZOIs were measured periodically for two weeks. The measurements from day 3 held consistent, and those measurements were used for biochemometrics analysis.
Biochemometrics Analysis
Triplicate UHPLC-MS data sets for each sample (including crude extracts SB-1 through SB-3 and fractions from their chromatographic separation) were individually analyzed, aligned, and filtered with MZmine 3.4.16 software (http://mzmine.sourceforge.net/) in both positive and negative ion modes. 31 First, m/z values were detected within each spectrum above a defined baseline following which a chromatogram was produced for each m/z value that spanned longer than 0.1 min. Chromatograms were then deconvoluted to identify individual chromatographic peaks. Specific peak list parameters can be found in Table S2. The resulting data matrices (which consisted of m/z, retention time, and peak area values for each peak) was imported to Excel (Microsoft, Redmond, WA, USA) and filtered further. First, all ions detected in blanks with an average peak area equal to or higher than the selected baselines for each ion were removed from the peak list, as were all ions detected before 0.5 and after 9.5 min. Finally, relative standard deviation (RSD) filtering was utilized for all triplicate analyses. Ions detected within the analytical replicates for which peak area differed by more than 35% were designated as instrumental artifacts and excluded from the final metabolomics analysis. 32 Remaining ions were averaged across replicate injections to produce a final peak list. Filtered peak lists in positive and negative modes were combined into a single list and merged with the bioactivity data set (zones of inhibition) to form a final biochemometric data matrix.
Biochemometric analysis was completed using Sirius version 11.5 (Pattern Recognition Systems AS, Bergen, Norway). 33 Internally cross-validated PLS models were produced using 100 iterations and a significance level of 0.05, and data were visualized using PLS scores and loadings plots, S-plots, and selectivity ratio plots.34–36 Built-in statistical models internal to the Sirius software were used to calculate final selectivity ratios for biochemometric analyses.
Identification of Target Compounds and Molecular Networking Analysis
Biochemometric analysis revealed a suite of ions that were correlated to biological activity, all of which were different adducts of the same molecule. This molecule was tentatively identified as nigericin by searching the Natural Products Atlas (http://npatlas.org), 37 the Dictionary of Natural Products, 38 COCONUT, 39 and Lotus 40 for molecules of bacterial origin within 5 ppm of the exact mass of the neutral molecule. Rather than pursue isolation of this known molecule, a standard was purchased from Sigma Aldrich and MS/MS patterns of the standard compared to those detected in our bacterial extracts and fractions (Figure S3).
To determine if any relatives of nigericin existed in our extracts, molecular networks were created using the online workflow on the Global Natural Products Social Molecular Networking (GNPS) website (http://gnps.ucsd.edu) for both positive and negative modes. For each network, data were filtered by removing all MS/MS fragment ions within ±17 Da of the precursor m/z. MS/MS spectra were window filtered by choosing only the top six fragment ions in the ± 50 Da window throughout the spectrum. The precursor ion mass tolerance and MS/MS fragment ion tolerance were each set to 0.02 Da. A network was then created where edges were filtered to have a cosine score above 0.65 and more than three matched peaks. Further, edges between two nodes were kept in the network if and only if each of the nodes appeared in each other's respective top ten most similar nodes. Finally, the maximum size of a molecular family was set to 100, and the lowest scoring edges were removed from molecular families until the molecular family size was below this threshold. The spectra in the network were then searched against spectral libraries housed within GNPS, which were filtered in the same manner as previously described. All matches were required to have a cosine score above 0.7 and at least six matched peaks. 41 Positive and negative mode data can be downloaded from the GNPS database (dataset MSV000095771).
Confirmation of Biological Activity
To confirm biochemometrics predictions, nigericin (Sigma Aldrich, PA, USA) was subjected to another round of antifungal testing via disc diffusion assay. 30 P. destructans strain 20631-21 (ATCC® MYA4855™) was activated on malt extract agar (MEA, Difco, NJ, USA) at 12 °C for 14 days to induce sporulation. Spore suspensions were collected from MEA plates and resuspended in sterile distilled water at 105 spores/mL. Ten μL of the spore suspension was spread on MEA plates using a sterile cotton swab. Nigericin stock solutions were prepared in dimethyl sulfoxide (DMSO) at concentrations of 10 mg mL−1, 5 mg mL−1, 2.5 mg mL−1, 1.25 mg mL−1, 0.625 mg mL−1, and 0.312 mg mL−1. Ten μL of each stock solution was applied to 6-mm sterile paper discs, for final disc concentrations of 100, 50, 25, 12.5, 6.25, and 3.12 μg disc−1. Discs prepared using 10 μL of DMSO and untreated discs served as negative controls. Each MEA plate included four replicate disks of a single nigericin concentration, along with two negative controls. Discs were placed on the inoculated agar surface with sufficient spacing to prevent overlapping zones of inhibition. In parallel, antifungal activity testing was completed using the same protocol with amphotericin B as positive control, with concentrations of ranging from 0.312 to 10 μg disc−1. All plates were incubated at 12 °C for 14 days to allow sufficient growth of P. destructans and interaction with the antifungal-treated discs.
ZOIs were measured in centimeters following the 14-day incubation period for all tested concentrations of nigericin and amphotericin B. Statistical analysis was conducted using one-way analysis of variance (ANOVA) to evaluate differences in ZOI sizes across concentrations. To determine pairwise statistical significance between groups, a Tukey's post hoc test was performed. All analyses were carried out using Prism GraphPad software (GraphPad Software, Inc.), ensuring accurate computation and visualization of the data.
Results
Preliminary Antifungal Analysis
S. buecherae AC541T extracts were grown under varying culturing conditions; however, media type, temperature, light exposure, and incubation duration did not have major impacts on chemical profiles (data not shown). Given that cultures grown on ISP2 at room temperature for two weeks in complete darkness produced the most material, we utilized these extracts for chromatographic separation and bioactivity screening experiments. A single stage of bioassay-guided fractionation was conducted to simplify the S. buecherae AC541T extracts and provide data for biochemometrics analysis. Crude extracts SB-1, SB-2, and SB-3 were subjected to flash chromatography and divided into 48 unique flash chromatography fractions. Prior to bioactivity testing, these fractions were subjected to LC-MS analysis and combined if their mass spectral profiles were similar (Figure S2). Using this process, 26 of the flash chromatography fractions were combined into 7 fractions (Table S1). Crude extracts and simplified fractions were then evaluated for antifungal activity against P. destructans using a disk diffusion assay. Results illustrated that bioactive constituents were concentrated during chromatographic separation, as multiple flash chromatography fractions (SB-1-5, SB-2-11, SB-3-12, SB-3-13, and SB-4-6) showed higher inhibition of P. destructans than the crude extracts SB-1 through SB-3. Zones of inhibition ranged from 0-2.0 cm, with the most potent fractions (SB-2-11, SB-3-12, SB-3-13, and SB-4-6) showing higher inhibition than the positive control, cavicide (Figure 2, Figure S4).
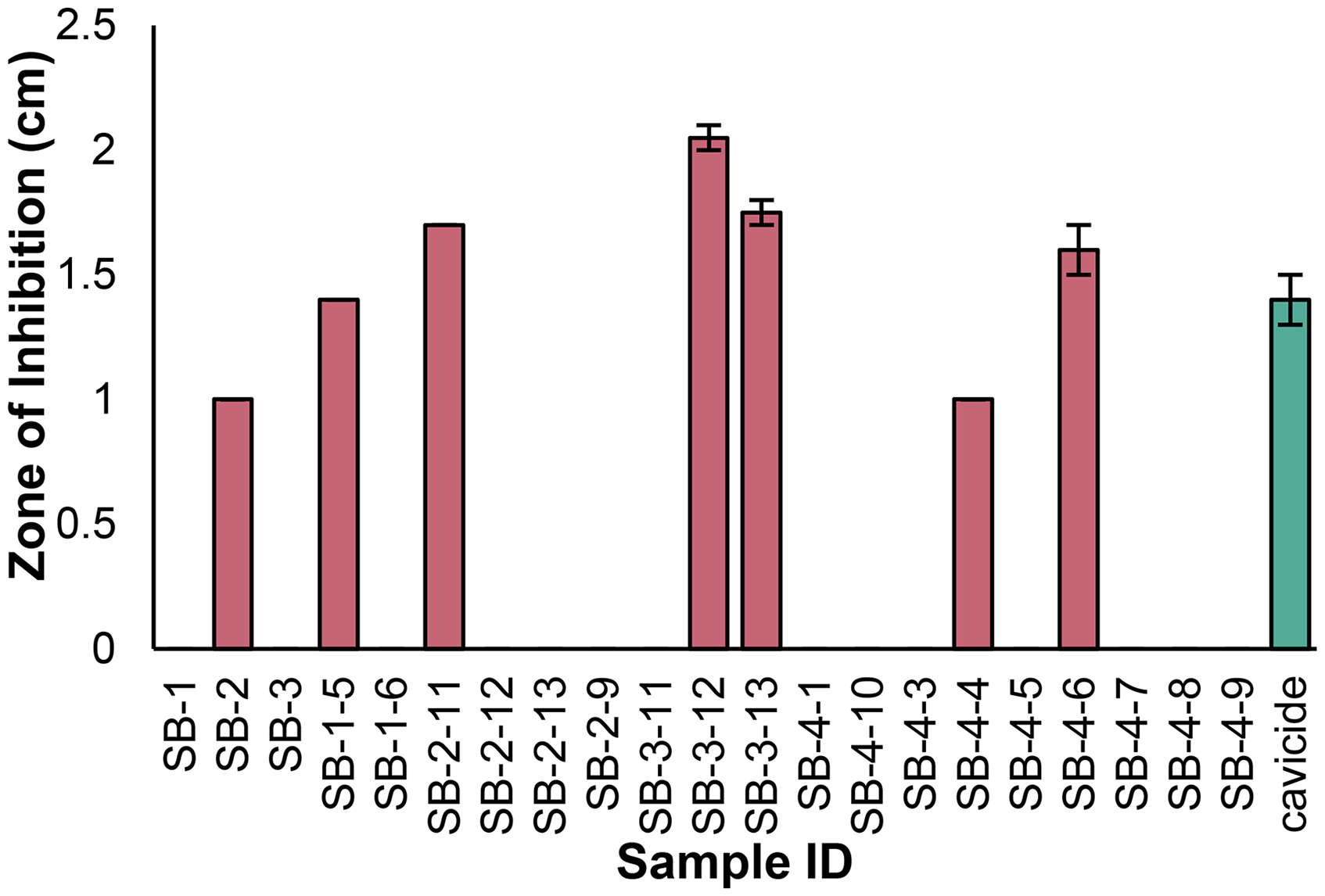
Zones of Inhibition of P. destructans by S. buecherae AC541T Extracts (SB-1 Through SB-3) and Their Associated Flash Chromatography Fractions. Seven of the 21 Samples Tested Inhibited of P. destructans. Strength of Inhibition was Defined in Reference to the Positive Control, Cavicide, with Samples Showing Slight (Inhibition Zones Less Than Positive Control), Intermediate (Inhibition Zone Equal to Positive Control), or Strong Inhibition (Inhibition Zone Greater Than Positive Control). Error Bars Represent Standard Error, Calculated Using Duplicate Bioactivity Tests per Sample, and Were Equal to Zero for Several Samples.
Biochemometrics Analysis
On pairing the antifungal screening with high-resolving power mass spectral data, biochemometric analyses revealed chemical differences between the S. buecherae fractions based on their bioactivity. The internally cross-validated PLS model yielded two components, accounting for 63.82% of the dependent variable variation (bioactivity) using just 57.58% of the independent variable variation (mass spectral profiles; component 1: 18.85% independent, 61.57% dependent; component 2: 38.73% independent, 2.25% dependent). The PLS scores plot (Figure 3A) showed distinct clustering of the most potent bioactive samples in the upper righthand quadrant of the plot. Examination of the corresponding PLS loadings plot (Figure 3B

(A) Partial Least-squares (PLS) Scores Plot Showing the Grouping of Bioactive and Non-bioactive Fractions from Streptomyces buecherae AC541T. Each Fraction was Analyzed Via UHPLC-MS and Subjected to Antifungal Activity Testing Against Pseudogymnoascus destructans. The Most Potent Samples Clustered in the Upper Righthand Quadrant of the Plot. (B) Loadings Plot from the PLS Analysis of Biochemometric Data. Variables Located in the Same Region in (B) as the Bioactive Groups in (A) Have the Highest Correlation with Bioactivity. Several Ions Corresponding to the Same Molecule were Identified from the Loadings Plot.
Two other visualization tools were used to identify bioactive constituents from the S. buecherae fractions using the PLS model: the S-plot and the selectivity ratio (Figure 4). The S-plot displays the covariance and correlation of loading variables (individual mass spectral features) against the biological activity data as a scatterplot that forms a characteristic “S” shape. For the S. buecherae biochemometric model, ions located in the upper righthand quadrant of the S-plot contributed most to the differentiation of biologically active versus inactive fractions (Figure 4A). The same data can be visualized using a selectivity ratio plot, similar to a mass spectrum, in which the x-axis represents the m/z of the detected ion and the y-axis represents the strength of the association of that particular ion with the antifungal activity (Figure 4B). Notably, all ions identified by both the S-plot and selectivity ratio analyses correspond to different adducts and/or isotopes of the same compound

(A) S-Plot from PLS Model of Antifungal Activity of S. buecherae Extract and Fractions. the Upper Right Quadrant Contains Ions That Have the Highest Correlation to Bioactivity and Includes Several Ions Corresponding to Compound
Dereplication of Bioactive Molecules
To avoid isolating already known compounds, we dereplicated our dataset by searching the chemical formula of compound
Molecular Networking Analysis
To discern if there were other nigericin derivatives in our S. buecherae extracts, we conducted molecular networking analysis using the GNPS website (https://gnps.ucsd.edu). The positive mode molecular contained a molecular family containing two nigericin ions (the ammonium and sodium adducts), as well as a suite of 33 additional ions that were connected based on similarities in their MS/MS fragmentation patterns (Figure 5). Through inspection of S. buecherae LC-MS profiles, we were able to organize 22 of the 33 ions in the nigericin molecular family into six groups based on both retention time and scan number, suggesting that there are at least six yet unknown nigericin derivatives produced by S. buecherae. Due to material limitations, we were unable to pursue these low-abundance molecules for purification.

MS/MS-based Molecular Networking Revealed a Molecular Family Containing Nigericin Ions (Teal Nodes with Thick Black Borders), as Well as 33 Additional Ions. Nodes Have Been Organized Based on Retention Time and MS Scan Number and Color-coded into Six Groups which May Represent Different Ions Associated with Unique Nigericin Derivatives.
Confirmation of Nigericin's Antifungal Activity
The disc diffusion assay evaluating nigericin's antifungal capacity against P. destructans revealed significant concentration-dependent ZOIs with a notable paradoxical effect at intermediate concentrations (Figure 6). An ANOVA revealed that our bioactivity results were normally distributed with equal variances, and a Tukey's post-hoc test determined which groups differed from one another. The largest ZOIs were observed at a nigericin concentration of 25 μg disc−1, where ZOIs measured an average of 1.87 cm. At higher concentrations of 100 and 50 μg disc−1, ZOIs were significantly smaller, averaging only 0.85 cm (averages and standard errors for all treatment concentrations provided in Table S3). A progressive decline in ZOI size was observed as nigericin concentration decreased below 25 μg disc−1 (Figure 6). In comparison, the positive control amphotericin B disc diffusion assay demonstrated a consistent linear pattern, with higher concentrations producing larger ZOIs (Figure S6). Nigericin showed moderate antifungal activity (>5 mm inhibition) against P. destructans down to a concentration of 6.25 μg disc−1, though partial activity was witnessed at even lower concentrations (3.12 μg disc−1).

(A)
Discussion
Although P. destructans loads appear to be decreasing in bat hibernacula after 10 + years of introduction in eastern North America, 42 recent studies suggest that even recovered bat populations could be vulnerable to reintroduction by a foreign genotype of P. destructans. 43 Moreover, its steady movement southward and westward across the continent raises considerable concern. P. destructans was only recently (<5 years) detected in the southwestern U.S., and it is still unclear how bat populations native to the region will respond. Of the ∼45 bat species that reside in the continental U.S., some of the greatest diversity is found in New Mexico, Arizona, and California. 7 Identification of probiotic candidate strains from bats native to this region may assist in disease mitigation strategies that cause the least amount of disturbance. S. buecherae is one such candidate strain, given that it has been isolated from multiple bat species in southern New Mexico, including members of Carlsbad Caverns’ renowned Brazilian free-tailed bat (Tadarida brasiliensis) maternal colony. 44 The widespread nature of S. buecherae as a member of the bat microbiome, combined with its robust antifungal activities against P. destructans in vitro, demonstrates its potential ecological importance in the American Southwest, making it a valuable candidate for further study.
The study described herein provides insight into the chemical constituents responsible for the antifungal activity of bat-associated S. buecherae against P. destructans in vitro. Chemical extracts for this study were collected from S. buecherae cultures grown at 20 °C in darkness. Given that M. velifer and T. brasiliensis are nocturnal, temperate vespertilionid bats, these growth conditions are representative of conditions that the bat skin microbiome would occupy in the field during daily torpor (where bat skin temperatures drop to temperatures ranging from 17 °C-26 °C). 45 The resulting extracts from these cultures contained hundreds of individual constituents, and we utilized biochemometrics analysis to identify putative bioactive constituents contributing to the antifungal activity of S. buecherae in vitro. 34 The utilization of this method allowed us to identify the known polyether antibiotic nigericin 46 as a primarily bioactive constituent early in the fractionation process without the need for multiple arduous chromatographic purification steps. These findings are consistent with previous in silico antiSMASH and MiBIG analyses that predicted nigericin biosynthesis associated with S. buecherae. 28
Notably, nigericin showed a concentration-dependent antifungal effect, though activity began to decrease above concentrations of 25 µg/disc (Figure 6). This decrease in activity can be explained by nigericin's poor solubility in aqueous media, potentially leading to incomplete diffusion in the MEA medium at higher concentrations. 47 Nigericin is renowned for its potent antibiotic and cancer activities which are well-documented in the scientific literature.48–50 Other studies have identified additional biological activities, including antiviral, 51 antiparasitic, 52 and antifungal effects, 53 though these activities are less thoroughly explored. Nigericin acts as an ionophore that forms stable complexes with monovalent metal cations (eg, K+) across cellular membranes, disrupting intracellular gradients, increasing proton concentration, and acidifying intracellular pH levels. 48 Such cellular acidification can induce apoptosis in cancer cells 54 and impair ATP production and membrane integrity in Gram-positive bacteria. 55 Prior to this study, nigericin has exhibited only weak antifungal activity against two fungi—Candida albicans and Escovopsis sp.,53,56 and this work highlights the need for additional research to fully elucidate nigericin's therapeutic potential as an antifungal agent. Molecular networking analysis also suggests that S. buecherae biosynthesizes additional nigericin analogs, and future studies will be required to increase yield and elucidate the structures and biological activities of these molecules.
Our preliminary studies highlight that S. buecherae, its chemical extracts, and its major constituent nigericin inhibit the growth of P. destructans in vitro on several growth media (R2A, MEA, and YPD). Of course, given the limited scale of this study, further research beyond our assessment in vitro needs to be conducted to confirm the efficacy of S. buecherae and/or nigericin as a probiotic to prevent symptoms of WNS. Moreover, it is likely that there are additional bioactive constituents produced by S. buecherae that material limitations prevented us from identifying, and further studies aimed at inducing formation of these constituents for bioactivity testing and structural elucidation are warranted. Finally, this study only investigated one bioactive strain from the bat microbiome, and host microbiome defenses are likely more complex given the antifungal activity of other bacterial strains isolated from resistant bats. 27 Given the complexity of host microbiome defenses, future studies geared towards investigating other bioactive strains, including fungi, alone and in mixed microbial consortia, may identify additional bioactive molecules and/or probiotic candidates for WNS disease management.
Conclusions
Herein, we demonstrate a biochemometrics-guided approach to identifying antifungal molecules active against the bat pathogen Pseudogymnoascus destructans. We identified nigericin (compound
Supplemental Material
sj-docx-1-npx-10.1177_1934578X251376175 - Supplemental material for Biochemometric Analysis of Bat Skin Microbiome-Associated Streptomyces buecherae Identifies Polyether Antibiotics Active Against the Bat Pathogen Pseudogymnoascus destructans
Supplemental material, sj-docx-1-npx-10.1177_1934578X251376175 for Biochemometric Analysis of Bat Skin Microbiome-Associated Streptomyces buecherae Identifies Polyether Antibiotics Active Against the Bat Pathogen Pseudogymnoascus destructans by Ashley E Clements, Alexa G Lee, Benjamin N Otoo, Roberto R Regalado, David C Dainko, Nancy P Keller, Paris S Salazar-Hamm and Lindsay K Caesar in Natural Product Communications
Footnotes
Acknowledgements
The authors would like to acknowledge Dr Michael Mullowney for initiating this collaboration and the University of North Carolina at Greensboro's Triad Mass Spectrometry Facility (specifically, Dr Warren Vidar) for their assistance with mass spectrometry analysis. Dr Diana Northup, Debbie Buecher, and the National Park Service, particularly Carlsbad Caverns, is acknowledged for providing Streptomyces buecherae strains for study.
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Author Contributions
AEC and AGL completed extract fractionation and conducted biochemometrics and molecular networking analysis to identify bioactive constituents under the supervision of LKC. BNO and RRR conducted biological activity assays under the supervision of NPK. DD grew bacterial extracts for analysis. PSSH provided expertise on S. buecherae and helped organize the manuscript. LKC wrote the manuscript and coordinated experiments between collaborating institutions. All authors read and edited the final manuscript.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was financially supported by the National Science Foundation (REU CHE-2150091 supporting AEC and BRC-BIO 2334070 to LKC and PSSH); Division of Biological Infrastructure; Division of Chemistry, and the American Society of Pharmacognosy's Audrey S. Bingel Fellowship for Female Scientists to LKC. RR was funded by an Advanced Opportunity Fellowship through SciMed Graduate Research Scholars at University of Wisconsin- Madison.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The authors have made available all positive and negative mode mass spectrometry data used for this analysis through the GNPS database under dataset #MSV0000957771.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
