Abstract
Objectives
Based on a report that Prunus mume fruits affect gut function and microflora, we hypothesized that mumefural (MF), one of the active compounds of processed Prunus mume fruits, could affect intestinal function or gut microbiome.
Methods
we investigated the effects of orally administered MF on intestinal function and the intestinal environment by analyzing the changes in the fecal, intestinal, and gut microbiome of a murine model.
Results
No changes in the body weight and fecal parameters of C57BL/6 mice were observed following MF administration. However, the quantity of residual feces in the large and small intestines and cecum was considerably reduced. The abundance of specific bacteria, ie, members of the Lactobacillus genus, was markedly increased without any change in the abundance or diversity of the gut microbiota, as determined by 16S rRNA gene-based microbiome taxonomic profiling.
Conclusions
Overall, our results suggest that oral MF administration could improve the gut microbiota in diseases related to specific bacterial changes because it increases the abundance of certain bacteria known to exhibit probiotic functions without any other obvious effects.
Introduction
The human microbiome is a consortium of various microorganisms that inhabit the gut and an important factor in human health and disease. Changes in the gut microbiome are induced by genetic and environmental factors, leading to changes in intestinal permeability, immune activity, metabolism, and neurological development. 1 Additionally, small molecules derived from the gut microbiome are associated with differences in physiological processes between individuals, such as immune system development, disease progression, and drug targets, via direct involvement in human cells and their receptors or indirect interactions with other microorganisms. 2 Gut dysbiosis caused by disruption of the gut microbiome has been implicated in diseases including cancer, 3 inflammatory bowel disease, asthma, obesity, metabolic syndrome, cardiovascular disease, immune-mediated diseases, and neurodevelopmental disorders, such as autism spectrum disorder. 4 The role of the gut microbiome in changing health and disease states in living organisms has previously been elucidated. 5 In practice, probiotic lactic acid bacteria are considered promising adjuvant therapeutics for inflammatory bowel diseases,6,7 including ulcerative colitis, Crohn's disease, 8 and irritable bowel syndrome. 9 Additionally, the potential mechanisms underlying the associations between intestinal microbiome imbalance and disease progression in various diseases, such as cancer,10,11 kidney disease, 12 obesity, 13 epilepsy,14,15 and cardiovascular disease, 16 are being considered. In this milieu, compounds with prebiotic potential are attracting attention for their applications in preventing and treating diseases.17–19
Prunus mume fruit extracts (PMFEs) are rich in fiber, free sugar, calcium, and iron and are considered alkaline foods containing several organic acids such as succinic, citric, malic, and tartaric acids. PMFEs have long been consumed as processed concentrates, tea, side dishes, alcoholic drinks, and vinegar. Moreover, they have been traditionally used to detoxify and treat fever, fatigue, sweating, cough, and indigestion in East Asia. PMFEs can regulate the intestinal microbiome 20 and have been shown to contribute to improvements in cognitive ability in animal models of Alzheimer's disease 21 and cognitive impairment. 22 Mumefural (MF), a major active compound obtained by processing the fruits of Prunus mume Sieb. et Zucc.23,24 MF helps improve blood circulation by inhibiting platelet aggregation25,26 and cognitive impairment caused by decreased cerebral blood flow. 27 Furthermore, MF improves recognition memory through extracellular signal-regulated kinase (ERK)-cyclic AMP-response element-binding protein (CREB)-brain-derived neurotrophic factor (BDNF) signaling 28 and exerts an antiviral effect. 23 Products containing MF are used as functional foods, particularly in Japan, acting as phosphodiesterase 4D inhibitors, 29 γ-amino butyric acid agonists, 30 and benzodiazepine agonists. 30
However, the effective ingredients that constitute PMFEs are poorly understood, especially in the context of their traditional application to treat intestinal disorders. Therefore, we hypothesized that MF, an effective compound in PMFEs, affects the intestinal environment and gut microbiota. Herein, we report the changes in the fecal, intestinal, and gut microbiota caused by MF administration in a murine model and suggest the potential of MF for use in maintaining a healthy intestinal microbial environment and preventing diseases associated with gut dysbiosis.
Results
Analysis of Collected Feces and Intestinal Content
No differences were observed in the change in body weight of mice between groups during oral MF administration (Figure 1A). The stool water content, recorded every 2 weeks, corresponded to the oral administration of MF after completing all experiments (Figure 1B). Furthermore, no difference was observed in the fecal number (Figure 1C), fecal weight per unit body weight (Figure 1D), or fecal weight per stool (Figure 1E) between the control and MF-administered groups.
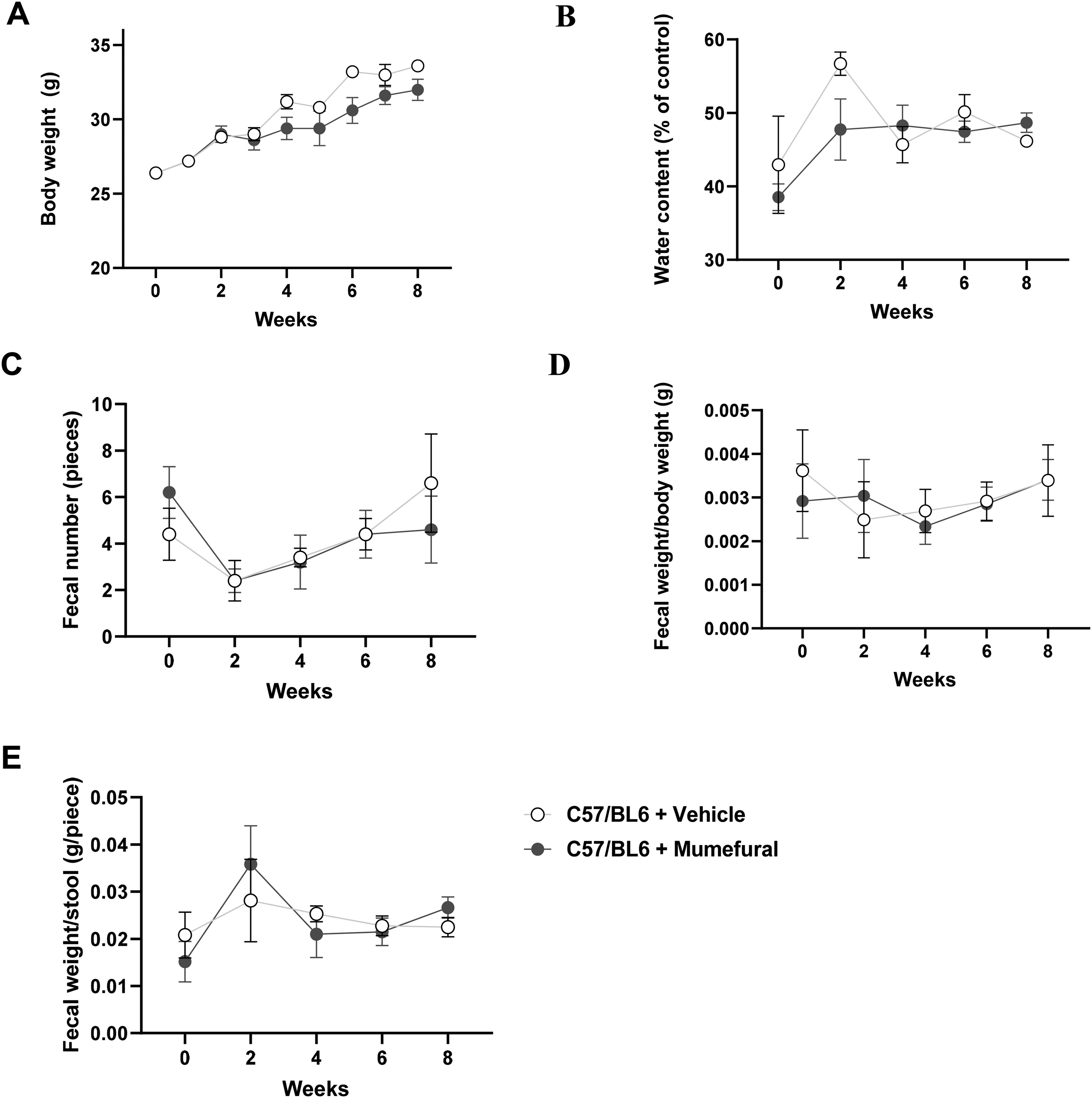
Effects of mumefural (MF) administration on the body weight and fecal parameters of C57BL/6 mice. (A) Body weight was measured once a week, and fecal samples were collected every 2 weeks during the administration of MF. The collected fecal samples were analyzed by measuring (B) water content, (C) fecal number, (D) fecal weight per unit body weight, and (E) fecal weight per stool. All values are presented as the mean ± SEM (n = 5).
At the end of the experiment, images of the small intestine, cecum, and large intestine were obtained (Figure 2A), and the length of the residual feces in the colon was measured. Compared with that in the control group (4.56 cm), the quantity of residual feces was significantly reduced in the MF-administered group (2.6 cm, P = .016) (Figure 2B). The weight of the remaining contents in both the small intestine and cecum of the MF-administered group (large intestine: 45.7, small intestine: 24.8, cecum, 579.2 mg/cm, P < .05) was significantly reduced compared with that of the control group (large intestine: 51 mg/cm, small intestine: 38.78 mg/cm, cecum: 734.2 mg/cm) (Figure 2C).

Effects of mumefural (MF) administration on C57BL/6 mouse intestinal weight and residual feces. (A) Representative intestinal images captured after completing all experiments. (B) Residual feces were measured as the length of the feces in the colon. (C) Intestinal weight with residual feces was measured by separating the large intestine, small intestine, and cecum. All values are presented as the mean ± SEM (n = 5). *P < .05 compared with the control group.
Gut Microbiome Analysis
In the alpha-diversity analysis, species richness was analyzed using ACE, CHAO, and Jackknife indices, whereas diversity was analyzed using NPShannon, Shannon, and Simpson indices. No difference was observed in the number of species (Figure 3A) and species diversity (Figure 3B) between the control and MF-administered groups. Additionally, the structural similarity of the microbiome between the control and MF-administered groups was compared via beta-diversity analysis using the UniFrac Principal Coordinate (PC) analysis plot following Jensen-Shannon's method. The control and MF-administered groups showed separate clustering in the microbiome structure (first PC: 46.639%, second PC: 21.659%) (Figure 3C).

Alpha- and beta-diversity analyses of the gut microbiome of C57BL/6 mice administered mumefural (MF). Alpha-diversity analysis of (A) species richness (number of species) and (B) species diversity using the indicated indices. (C) Beta-diversity analysis of microbiome structural similarity presented as the principal component plot (PC1 and PC2) (n = 5). PC, principal coordinate.
The taxonomic composition was analyzed at the phylum, class, and genus levels. At the phylum level, the abundance of Bacteroidetes (32.16) and Firmicutes (66.45) in the MF-administered group did not differ compared with that in the control group (Bacteroidetes: 34.28, Firmicutes: 64.61) (Figure 4A). At the class level, the abundance of Bacilli (9.69) in the MF-administered group increased by 2.4-fold compared with that in the control group (P = .032) (Figure 4B). At the genus level, the abundance of Lactobacillus (9.64) in the MF-administered group increased by 2.4-fold compared with that in the control group (4.05) (P = .032). The abundance of AF349416_g, Eubacterium_g17, KE159600_g (1.8; 1.3; 1.5) in the MF-administered group increased by 2.2-, 2.2-, and 3.6-fold compared with that in the control group (0.8; 0.6; 0.4) (P = .016; .032; .008), respectively. The abundance of PAC000664_g (1.1) in the MF-administered group decreased by 0.5-fold compared with that in the control group (2.3) (P = .016) (Figure 4C). All results are expressed as a percentage.

Taxonomic composition via gut microbiome analysis in C57BL/6 mice administered mumefural (MF). The taxonomic composition was analyzed at the (A) phylum level, (B) class level, and (C) genus level. The taxonomic relative abundance is also presented to compare the changes in taxa between the control and MF-administered groups. All values are presented as the mean ± SEM (n = 5). *P < .05 compared with the control group.
In the order of class, order, family, and genus levels, the composition of Bacilli, Lactobacillales, Lactobacillaceae, and Lactobacillus, respectively, increased more than two-fold after MF administration. These results were confirmed through linear discriminant analysis (LDA) scores (Figure 5A) and cladograms (Figure 5B) with linear discriminant analysis effect size (LEfSe).

Taxonomic differences using linear discriminant analysis (LDA) effect size (LEfSe) analysis in mumefural (MF)-administered C57BL/6 mice. Enriched bacterial taxa are represented by LEfSe plots following the (A) LDA score and (B) cladogram (n = 5).
Discussion
The movement of feces in the intestine can determine the function of the large intestine. Residual feces affect the intestinal environment due to the production of various toxins derived from the gut microbiota and are associated with disease susceptibility and progression.31,32 Colonic transit times are closely related to the diversity of gut microbiota, microbial fermentation, 33 and exposure to toxins produced by the gut microbiota through protein or amino acid metabolism. Pathogenic bacteria interfere with signaling pathways related to the induction of cellular DNA damage or cell proliferation, apoptosis, and inflammation by producing enzyme-activated protein toxins that may potentially act as carcinogens. 34 In addition, uremic toxins produced during amino acid metabolism of intestinal microbes 35 can cause chronic kidney disease 36 or heart disease. 37 In the present study, we confirmed that oral administration of MF could reduce the quantity of residual feces in the intestine, which is expected to contribute to the improvement of the intestinal environment.
MF administration distinctly altered the intestinal microbiota in terms of Lactobacillus abundance. No difference in the alpha-diversity was observed between the control and MF-administered groups, probably because MF was not administered together with a new bacterial strain. However, we observed a difference in the microbial community between the control and MF-administered groups in terms of beta diversity. The abundance of Lactobacillus can be increased by improving the environment for their survival using substances produced by MF metabolism, such as citric acid, in the intestine. 26 Citric acid increases the abundance of Lactobacillus and decreases bacterial diversity. A beneficial synergistic effect between Lactobacillus and citric acid has also been reported, involving a reduction in the abundance of Dysgonomonas and Pseudomonas, which are known to be harmful fermentation quality. 38 Additionally, Lactobacillus exhibits strong viability and reduces bacterial diversity in acidic environments.39,40 Furthermore, MF regulates the pH by increasing the expression SMCT1, which is involved in monocarboxylate absorption in the colon. 41 Colonic pH determines the structure and function of the gut microbial community and the production of short-chain fatty acids. 42 These findings provide a basis for the selective increase in Lactobacillus abundance following oral administration of MF.
Our study findings revealed that the oral administration of MF decreases residual feces in the murine intestine, regardless of fecal output and water content per unit time, and characteristically increases the abundance of Lactobacillus. Probiotics containing one or more validated microbial strains, such as Lactobacillus, Bifidobacterium, Lactococcus, Streptococcus, and Enterococcus,43–45 exert beneficial effects in maintaining the intestinal microbial system and regulating microbial activity via the immune system, antibacterial effects, and inhibition of microbe-derived toxin production.46–49
Conclusion
Our results demonstrated the efficacy of MF in reducing intestinal transit time and selectively increasing beneficial bacteria, ie, Lactobacillus members. Nevertheless, our study has some limitations. The interaction between Lactobacillus members and host metabolism could not be elucidated. Furthermore, we did not evaluate the efficacy of MF in models with intestinal abnormalities. Thus, follow-up studies are required to improve the reliability of the results.
Consequently, we demonstrate that MF can be expected to have a positive effect in terms of maintaining health and preventing disease by facilitating an appropriate balance between pathogens and bacteria.
Materials and Methods
Materials
MF (CAS No. 222973-44-6) was purchased from U CHEM (Gyeonggi-do, Republic of Korea). It was dissolved in saline and administered orally to mice once a day at 40 mg/kg after vortexing.
Animals
Eight-week-old male C57BL/6 mice (body weight, 22-23 g) were obtained from OrientBio (Seongnam, Republic of Korea). All mice were housed at 22 ± 1 °C and 55 ± 10% humidity under a 12-h light/dark cycle with ad libitum access to food and water. All experimental procedures were performed following the Guide for Animal Care and Use of Laboratory Animals of the Institutional Animal Care and Use Committee of the Korean Institute of Science and Technology (approval number: KIST-2020-023).
Fecal Weight, Number, and Water Content
The number and weight of fecal pellets collected for 1 h separately from individual cages at the beginning of the experiment were measured. All experiments were conducted under constant conditions and time. The weight of the fecal pellets dried in an 80 °C dry oven (JEIO TECH, Daejeon, Republic of Korea) after 24 h was recorded. This process was performed before the start of oral administration and at 2-week intervals after administration to measure fecal weight, number, and water content. Moisture content was calculated using the following equation: (fecal weight before drying − fecal weight after drying) / fecal weight before drying × 100.
Experimental Design and Sample Collection
The mice were randomly divided into two groups (n = 5) and administered saline (vehicle) or MF (40 mg/kg, dissolved in saline) via oral administration once a day for 2 months. The mice were anesthetized using inhalational isoflurane, intestinal images were captured, and the length of the separated intestines placed on ice was measured. Thereafter, the small intestine, cecum, large intestine, and feces from each mouse were collected. The collected samples were frozen on dry ice and stored at − 80 °C until microbiome analysis.
Intestinal microbiota 16S rRNA Sequencing
DNA from the collected intestinal fecal samples was extracted following the experimental method employed by CJ Bioscience, Inc. (Seoul, Korea). For PCR amplification, fusion primers targeting the V3-V4 regions of the 16S rRNA gene were used. Fusion primers 341F (5′- AATGATACGGCGACCACCGAGATCTACAC-XXXXXXXX-TCGTCGGCAGCGTCAGATGTGTATAAGAGACAG-CCTACGGGNGGCWGCAG-3′, the underlined sequence indicates the target region primer) and 805R (5′- CAAGCAGAAGACGGCATACGAGAT-XXXXXXXXGTCTCGTGGGCTCGG-AGATGTGTATAAGAGACAG-GACTACHVGGGTATCTAATCC-3′) were used for bacterial genetic material amplification. The amplification conditions were as follows: initial denaturation at 95 °C for 3 min, followed by 25 cycles of denaturation at 95 °C for 30 s, primer annealing at 55 °C for 30 s, extension at 72 °C for 30 s, and final elongation at 72 °C for 5 min. Following amplification, the PCR product was electrophoresed using a 1% agarose gel and confirmed via visualization using the Gel Doc system (Bio-Rad, Hercules, CA, USA). After purification using CleanPCR (CleanNA, The Netherlands), all PCR products of the same concentration were pooled, and the short fragments (non-target products) were removed using CleanPCR (CleanNA). Quality and product size were assessed using the Bioanalyzer 2100 (Agilent, Palo Alto, CA, USA) with a DNA 7500 chip. Mixed amplicons were pooled, and sequencing was performed at CJ Bioscience, Inc. using an Illumina MiSeq Sequencing system (Illumina, San Diego, CA, USA).
Data Analysis
The processed raw data were checked and filtered for low-quality (<Q25) reads using Trimmomatic ver. 0.32. 50 After quality checking, paired-end sequence data were integrated using the fastq_mergepairs command of VSEARCH version 2.13.4. 51 Parameters and primers were trimmed using the alignment algorithm as stated by Myers and Miller 52 at a similarity cutoff of 0.8. Nonspecific amplicons not encoding 16S rRNA were detected using nhmmer 53 in the HMMER software package ver. 3.2.1 (EMBL-EBI, UK). The extracted unique reads were collected after processing redundant reads using the derep_fulllength command of VSEARCH. 51 Based on the EzBioCloud database, 54 taxonomic assignment was performed using the userarch_global command of VSEARCH, 51 followed by pairwise alignment, 52 and chimeric reads with <97% similarity were filtered using the UCHIME algorithm. 55 After filtering the chimeras and compiling unidentified reads using the cluster_fast command, 51 additional operational taxonomic units (OTUs) were generated. OTUs with single reads were omitted from further analysis. The alpha-diversity indices, including ACE, 56 Chao, 57 Jackknife, 58 Shannon, 59 NPShannon, 60 and Simpson 59 and beta-diversity distances with Jensen-Shannon 61 were computed using the bioinformatic cloud platform of CJ Bioscience and EzBioCloud 16S rRNA gene-based microbiome taxonomic profile.
Statistical Analysis
All statistical analyses were performed using SPSS (version 12.0, SPSS Inc., Chicago, IL, USA). Differences between the control and MF-administered groups were analyzed using the Student's t-test. The data did not follow an assumption of analysis and were analyzed using the Mann-Whitney U test. Data were presented as the mean ± SEM. Results with P < .05 were considered statistically significant.
Footnotes
Author Contributions
JK and WKJ were involved in conceptualization; JK and MK were involved in methodology; JK was involved in software; JK validated the manuscript; JK wasinvolved in formal analysis; JK and MK were involved in investigation; JK was involved in data curation; JK written the original draft preparation; JK and WKJ reviewed and edited the manuscript. All authors have read and agreed to the publication of the final version of the manuscript.
Availability of Data and Materials
Not applicable.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Ethical Approval is not applicable for this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Republic of Korea National Research Council of Science & Technology (NST), the Korea Institute of Planning and Evaluation for Technology in Food, Agriculture, Forestry and Fisheries (IPET) and the Ministry of Agriculture, Food and Rural Affairs (MAFRA), (grant number CRC-15-04-KIST, NSN1523380, KSN 1621850, 821023-03, and NAN2111120).
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Statement of Human and Animal Rights
All experimental procedures were performed following the Guide for Animal Care and Use of Laboratory Animals of the Institutional Animal Care and Use Committee of the Korean Institute of Science and Technology (approval number: KIST-2020-023).
