Abstract
Objective
This study applied ultra-high performance liquid chromatography-Q Exactive-mass spectrometry (UHPLC-QE-MS) to identify the primary components of Panax notoginseng and, in combination with network pharmacology, molecular docking, and molecular dynamics (MD) simulations, explored its mechanisms in treating steroid-associated necrosis of femoral head (SANFH).
Methods
After identifying Panax notoginseng components using UHPLC-QE-MS, effective components were selected through the TCMSP database. Corresponding targets were obtained from TCMSP and Uniprot, while SANFH-related targets were retrieved from disease databases. Intersection targets between drug and disease targets were identified, and a protein-protein interaction (PPI) network was constructed using STRING and Cytoscape. An “active components-targets” network was built, and data for key active components were compared with PubChem reference standards. The “effective component-target” network was created for key components, followed by enrichment analysis. Molecular docking was used to explore interactions between key components and targets, and MD simulations were conducted on the top candidates with the lowest binding energy.
Results
Fifty-seven effective components of Panax notoginseng were identified, with 50 intersecting targets between disease and component targets. Four key effective components and four key targets were identified. Comparison with PubChem LC-MS data confirmed similar retention times for key components. Gene Ontology (GO) annotation yielded 683 entries, while KEGG enrichment identified 39 pathways. Molecular docking revealed strong binding of key components to targets, and MD simulations demonstrated that quercetin interacted stably with PTGS2.
Conclusion
This comprehensive study on Panax notoginseng components suggests it may exert anti-inflammatory, cartilage-protective, and anti-osteoporotic effects in treating SANFH through multiple components, targets, and pathways.
Keywords
Introduction
Steroid-associated necrosis of the femoral head (SANFH) is the progressive skeletal disorder resulting from improper or excessive steroid application, causing insufficient femoral head blood supply, osteocyte apoptosis, and subsequent femoral head structural damage. This leads to local collapse or dysfunction of hip joint, severely impacting patient's life quality. 1 Due to the high doses and long-term use of glucocorticoids in the clinical treatment of rheumatic diseases, immune-related diseases, spinal cord shock, and other conditions, the incidence of SANFH is increasing every year. According to an epidemiological survey, the number of femoral head necrosis patients in Japan has reached 25,000, while in China, the number of patients aged 15 and above has exceeded 8 million. The disability rate of this disease is extremely high. Without early intervention for SANFH, the collapse rate of the femoral head structure can exceed 80% within two years, placing a heavy burden on patients and their families.2,3 The effective components of natural products can enhance endothelial cell activity, inhibit steroid-induced apoptosis and functional loss, and improve pathological changes in the early and middle stages of SANFH.4,5 Panax notoginseng is the dried rhizome and root of Panax notoginseng plant from the Araliaceae family, native to regions including Yunnan and Guangxi in China. It displays different pharmacological activities, including anti-inflammation, antioxidation, analgesia, hemostasis, osteogenesis and angiogenesis. It has been commonly used to treat cardiovascular disorders, pain, inflammation, and both internal and external bleeding.6,7
Panax notoginseng treatment can promote angiogenesis because it activates the VEGF-KDR and PI3K-Akt-eNOS pathways, thereby enhancing angiogenesis. 8 Additionally, Panax notoginseng induces the production of VEGF-A, VEGFR-1, and VEGFR-2 in bone marrow mesenchymal stem cells (BMSCs). 9 Relevant studies have found that Panax notoginseng has the potential to treat SANFH by up-regulating the expression of osteogenic markers while down-regulating that of adipogenic markers, therefore stimulating mouse bone marrow stromal cell growth while enhancing their osteogenic differentiation.10,11 In this study, ultra-high performance liquid chromatography-Q Exactive-mass spectrometry (UHPLC-QE-MS) was employed to identify Panax notoginseng components. Subsequently, network pharmacology was performed with the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP), as well as molecular docking and molecular dynamics (MD) simulations for validation.12,13 According to mass spectrometry identification, network pharmacology predictions, molecular docking, together with MD simulations, this study aimed to explore the mechanisms through which Panax notoginseng intervened with SANFH, providing further scientific evidence for its clinical use.
Materials and Methods
Reagents and Chemicals
Panax notoginseng (Product Batch No.: 20230922, purchased from Guangxi University of Chinese Medicine Affiliated Ruikang Hospital); Methanol (CAS: 67-56-1, Purity: LC-MS grade, CNW Technologies); Acetonitrile (CAS: 75-05-8, Purity: LC-MS grade, CNW Technologies); Formic acid (CAS: 64-18-6, Purity: LC-MS grade, SIGMA); L-2-Chlorophenylalanine (CAS: 103616-89-3, Purity: ≥ 98%, purchased from Shanghai Hengbo Biotechnology Co., Ltd).
Experimental Instruments
Vanquish Ultra High Performance Liquid Chromatograph (Thermo Fisher Scientific, USA); Orbitrap Exploris 120 High-Resolution Mass Spectrometer (Thermo Fisher Scientific, USA); JXFSTPRP-24 Grinder (Shanghai Jingxin Technology Co., Ltd, China); Heraeus Fresco17 Centrifuge (Thermo Fisher Scientific, USA); Mingche D24 UV Pure Water System (Merck Millipore, Germany); YM-080S Ultrasonic Device (Shenzhen Fangao Microelectronics Co., Ltd, China); BSA124S-CW Balance (Sartorius, Germany); UPLC BEH C18 Chromatography Column (Waters, USA).
Extraction of Active Ingredients from Panax notoginseng
The components of Panax notoginseng were detected using UHPLC-QE-MS technology. A 100 mg sample of Panax notoginseng was ground at a frequency of 60 Hz for 120 s. The ground sample was then added to 500 μL of extraction solvent (methanol:water = 4:1, internal standard concentration: 10 μg/mL), vortexed for 30 s, and homogenized at 45 Hz for 4 min. The sample was then subjected to ultrasound treatment in an ice-water bath for 1 h. Afterward, the sample was left to stand at −40 °C for 1 h, followed by centrifugation at 12,000 rpm for 15 min at 4 °C (centrifugal force: 13,800×g, radius: 8.6 cm), and the supernatant was collected. Finally, the supernatant was filtered through a 0.22 μm microporous filter membrane to obtain the Panax notoginseng sample solution for subsequent qualitative analysis. 14
Chromatographic Conditions
① A UPLC BEH C18 column (1.7 μm × 2.1 mm × 100 mm) was used, with an injection volume of 5 μL.
② For the liquid chromatography analysis of Panax notoginseng samples, the following gradient conditions were applied: From 0–11 min, the flow rate was 500 μL/min, with 85% water as phase A and 15% acetonitrile as phase B. At 11 min, phase B was gradually increased to 75%, and phase A was reduced to 25%, maintaining this condition until 12 min. Over the next 2 min, phase B increased to 98%, while phase A decreased to 2%, lasting until 14 min. At 14.1 min, the gradient returned to the initial conditions, with phase A restored to 85% and phase B reduced to 15%, maintaining this until the end of the 16-min period.
③ Both phase A and phase B contained 0.1% formic acid.
Mass Spectrometry Conditions
The Orbitrap Exploris 120 mass spectrometer was controlled using Xcalibur software to collect both FullScan-ddMS2 data for primary and secondary mass spectrometry. The specific parameter settings were as follows: ①Sheath gas flow rate: 35 Arb; ②Auxiliary gas flow rate: 15 Arb; ③Ion transfer tube temperature and evaporator temperature: 350 °C; ④Full scan resolution: 60,000; ⑤MS/MS resolution: 15,000; ⑥Collision energy: 16/32/48 in NCE mode; ⑦Spray voltage: 5.5 kV (positive) / −4 kV (negative).
Data Analysis
Following the conversion of raw data to mzXML format using proteowizard software, peaks were recognized, extracted, aligned and integrated with self-written R package (XCMS as kernel). During the identification process, the primary mass spectrometry data were compared (relative error < 10 ppm). On this basis, the effective components were further confirmed by secondary mass spectrometry analysis and comparison.
Target Acquisition of Effective Components in Panax notoginseng
The identified effective components in Panax notoginseng were imported into the “Chemical name” module of TCMSP database to discover target proteins associated with the effective components, and subsequently gene name associated with target protein was queried with UniProt database. After removing duplicates, those obtained results were effective component targets of Panax notoginseng.
Collection of SANFH Related Targets
With “SANFH” and “Gene expression” as the keywords, and “Expre-ssion profiling by array”, “Homo sapiens” and “Series” as the limiting standards, the relevant chips of SANFH were retrieved from GEO public database to acquire GSE123568 and GPL15207 matrix files. The data were reannotated using Perl language, and the R language was adopted for correction and classification. The limma package was used to analyze gene difference, and |logFC|≥0.5 and P < 0.05 were thresholds for screening differential genes. Moreover, pheatmap package was used to draw the differential gene heatmap, and findings represented SANFH targets.This study was approved by the Animal Ethics Committee of Guangxi University of Traditional Chinese Medicine (Approval No.: No.DW20231220-284).
Target Forecasting and “Effective Components-Targets” Network Establishment
By mapping and intersecting effective component targets of Panax notoginseng with associated SANFH targets into Venny platform, potential Panax notoginseng targets for the treatment of SANFH were acquired. Then, Cytoscape software was used for drawing the “effective components-targets” network diagram, with key effective components being selected based on the degree value. We referred to the analysis and comparison method of Cui Y, 15 the liquid chromatography-mass spectrometry (LC-MS) experimental data and chemical structures of the key active components of Panax notoginseng were obtained from the PubChem database and compared with the UHPLC-QE-MS detection data.
Protein Interaction Network Establishment
To clarify the mechanism by which Panax notoginseng treated SANFH, we input potential therapeutic targets in STRING database, restricting the search to “Homo sapiens” as research species, and setting connection score threshold at > 0.5 to retrieve protein interactions. Subsequently, the acquired results were visualized with Cytoscape software, and “NetworkAnalyzer” tool was used to construct a protein-protein interaction (PPI) network. Through “CytoHubba” approach, Panax notoginseng key targets in SANFH treatment were identified based on their degree centrality.
Gene Enrichment
Gene Ontology (GO) functional annotation was conducted on key targets with clusterprofiler package in R language to investigate the biological functions of Panax notoginseng in the treatment of SANFH. Key targets were subject to KEGG enrichment to explore pathways related to Panax notoginseng for treating SANFH. p < 0.05 indicated significant enrichment results. Finally, R package ggplot2 was used for visualization.
Construction of the KEGG Relationship Network
Based on the signaling pathways acquired in step 1.9, key targets of the principal effective components and corresponding pathways were examined to construct the “key effective components - key targets - key pathways” relationship network, which was later imported in Cytoscape. Moreover, those top 5 significant signaling pathways were chosen to create the KEGG relationship network diagram.
Molecular Docking Validation of Interactions Between Key Effective Components and Targets
The 2D structures representing key effective components were obtained based on PubChem database, imported into ChemOffice for the generation of 3D structural diagrams. In addition, 3D structures were obtained for crucial targets based on Protein Data Bank (PDB) (https://www.rcsb.org/). Subsequently, protein preprocessings, including dehydration and separation of original ligands, were performed using PyMOL. Next, the processed files were transformed into the pdbqt format with AutoDock Tools 1.5.6, and active pockets were identified. Finally, AutoDock Vina software was employed for molecular docking and visualization to validate predictions of this experiment.
MD Simulations
Effective components and targets with the lowest binding energy identified from molecular docking results were subjected to MD simulations to explore protein-small molecule binding and assess their interactions. The experimental system consisted of a protein-small molecule complex, comprising a protein molecule with a target site and an effective component small molecule. Simulations were performed using Gromacs software, and the specific steps were outlined as follows. Initially, we AmberTools22 was employed to apply a GAFF force field into the small molecule, and then Gaussian16 W was used for hydrogenation and calculation of the RESP potential. Subsequently, structure files were input for protein molecules and small molecules in Gromacs software. Simulation boxes and topology files were generated using pdb2gmx and gmxeditconf commands. Simulations were performed at the 300 K static temperature and air pressure (1 Bar), based on the Amber99sb-ildn force field. With water molecules (Tip3p water model) as the solvent, an appropriate quantity of Na + was introduced for neutralizing total charge in this simulation system. Subsequently, the steepest descent approach was used in energy minimization for protein and small molecule structures. Next, gmxgrompp and gmxmdrun commands were used to conduct 100,000 steps of isothermal-isobaric and isothermal-isochoric ensemble equilibration, respectively, at the simulation period and coupling constant of 100 ps and 0.1 ps, respectively. Finally, free MD simulations were executed, which included totally 5,000,000 steps at the 2 fs time step, and thus the overall simulation duration was 100 ns. After completing the simulations, the gmxrms command was utilized to compute the root mean square fluctuation (RMSF), root mean square deviation (RMSD), solvent accessible surface area (SASA), radius of gyration (Rg), hydrogen bonds (HBond), and the free energy landscape (FEL) of protein with the small molecule.
Results
Effective Components of Panax notoginseng
Figure 1 shows the mass spectrometry total ion chromatograms in positive or negative ion modes of Panax notoginseng solution. Based on the MS detection results, mass spectrometry fragments were obtained, and the theoretical values were compared with the observed values by calculating the relative error. Totally 2042 effective components could be obtained in positive or negative ion modes.

Mass spectrometry total ion chromatograms for Panax notoginseng solution in positive/negative ion scan modes. Note: Positive (A) and negative (B) ion modes, 1:Quercetin, 2:Kaempferol, 3:Ursodeoxycholic acid, 4:Kainic Acid.
Targets of Effective Components in Panax notoginseng
Totally 2042 effective components were input into TCMSP database. After excluding 1985 components that did not satisfy the criteria, 57 effective components were selected, as shown in Table 1. After integrating target proteins associated with these 57 effective components, gene names corresponding to target proteins were retrieved using Uniprot database. This process identified 329 targets associated with the effective components.
Information on Effective Components of Panax notoginseng.

Note: NameEN indicates the English name of the substance, CompositeScore indicates the spectral similarity score, Formula stands for the molecular formula, mzmed refers to the median mass-to-charge ratio (m/z), rtmed suggests the median retention time, ppm indicates deviation between measured and theoretical values, while ms2Adduct is secondary fragment information.
SANFH-related Targets
Using R language and other software to analyze SANFH-related microarray data, 1328 SANFH-related targets were obtained, including 1088 with up-regulation, whereas 748 with down-regulation. Those 10 most significantly differential genes were selected to create a heatmap of differential genes, as shown in Figure 2.

Heatmap showing differential genes. Note: Those initial 10 columns represent genes from healthy control group, whereas those final 30 columns represent genes from SANFH group. The colors indicate gene expression levels across different samples, with green, black, and red representing low, medium and high expression, respectively.
“Effective Components-Targets” Network of Panax notoginseng for the Treatment of SANFH
Using Venny platform, we mapped the target proteins of effective components in Panax notoginseng against targets associated with SANFH, resulting in an intersection of 60 potential therapeutic targets, as shown in Figure 3. Then, interactions of effective components with intersection targets were input in Cytoscape software to construct an “Effective components-Targets” network of Panax notoginseng for treating SANFH, as depicted in Figure 4. This network comprised totally 104 nodes (including 43 effective component nodes, 60 target nodes, and 1 Panax notoginseng node) along with 177 edges. Those four effective components with the highest degree centrality in this network included Quercetin, Ursodeoxycholic acid, Kainic Acid, and Kaempferol, and they were identified as vital effective components in the network, exerting a vital role in treating SANFH. Their data are provided in Table 2. The liquid chromatography-mass spectrometry (LC-MS) experimental data and chemical structures of the key active components of Panax notoginseng standards in the PubChem database are shown in Supplementary Figure 1.16–19

Venn diagram showing possible targets of Panax notoginseng in SANFH treatment.
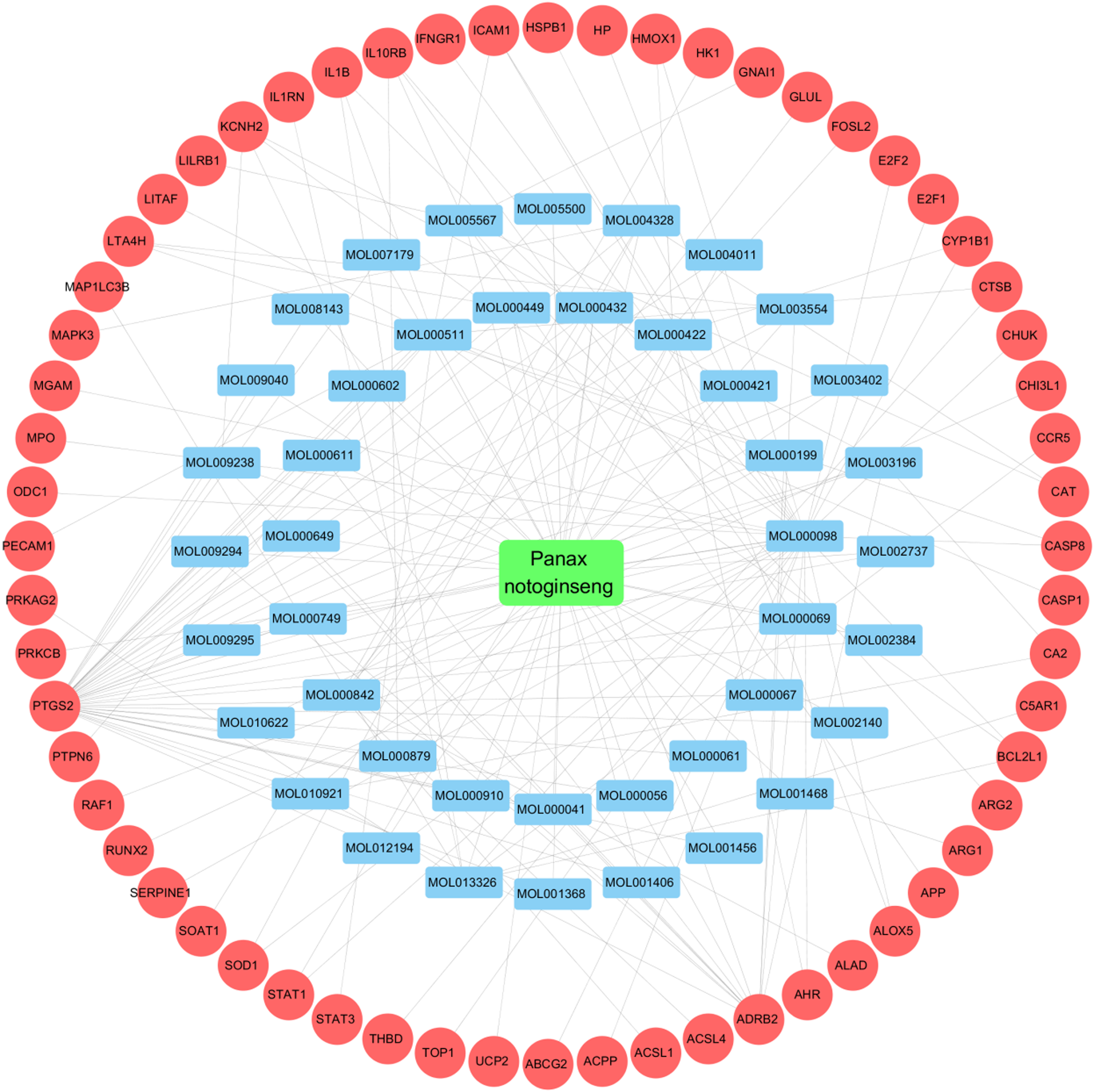
The “effective components-targets” network for Panax notoginseng in treating SANFH. Note: Red circles represent the target proteins, while blue rectangles stand for the effective components.
Baseline Data of Key Effective Components.

PPI Network
A PPI network was established using Cytoscape software and STRING database, as shown in Figure 5, including totally 203 edges and 50 nodes. Based on degree centrality, the top four protein genes identified were IL1B, STAT1, STAT3, and PTGS2. These protein genes were not only vital within this network, but also crucial for SANFH treatment using Panax notoginseng. They might be the critical targets for Panax notoginseng in the treatment of SANFH, and their basic information is provided in Table 3.

Protein interaction network. Note: Nodes stand for protein genes, whereas edges suggest interactions of these protein genes. Node size and color, as well as edge thickness, are topological parameters for this PPI network. The degree centrality, which reflects the connection number of a node with others, can intuitively assess a node's “influence” within this network. A larger connection number of a node indicates a greater degree centrality, and therefore the higher influence.
Basic Data of Key Targets.

Enrichment Analysis
In the process of GO annotation on the key targets, 683 terms were obtained. Among these, 646 terms corresponded to Biological Processes (BP), primarily involving response to peptide hormone, response to mechanical stimulus, positive regulation of cytokine production, and cellular response to peptide hormone stimulus. Two terms were related to Cellular Components (CC), focusing on nuclear outer membrane and RNA polymerase II transcription regulator complex. In addition, 35 terms were associated with Molecular Functions (MF), including protein phosphatase binding, cytokine receptor binding, phosphatase binding, interleukin-1 receptor binding, and RNA polymerase II-specific DNA-binding transcription factor binding. Based on GO annotation results, the BP, CC, and MF categories were closely associated with SANFH development and progression (Figure 6). KEGG analysis identified totally 39 relevant pathways for target proteins, primarily involving the AGE-RAGE, C-type lectin receptor, IL-17, NF-kappa B, and Toll-like receptor pathways (Figure 7).

Go annotation. Note: The top six items that meet the screening criteria for each category are displayed. The circular diagram from the outside to the inside represents the GO ID, with colors corresponding to the classification; gene number related to every GO term, with colors representing the degree of enrichment; and the number of key genes enriched in each GO term.

KEGG pathway enrichment. Note: The horizontal axis stands for enrichment score. A greater bubble indicates the more genes enriched into that term. A redder bubble suggests the higher significant enrichment level.
KEGG Relationship Network
The “Key Effective Components-Key Targets-Key Pathways” relationship network for Panax notoginseng was visualized with Cytoscape, resulting in the construction of the “KEGG Relationship Network” (Figure 8). Therefore, Panax notoginseng treated SANFH via several targets and pathways. Table 4 displays more details.
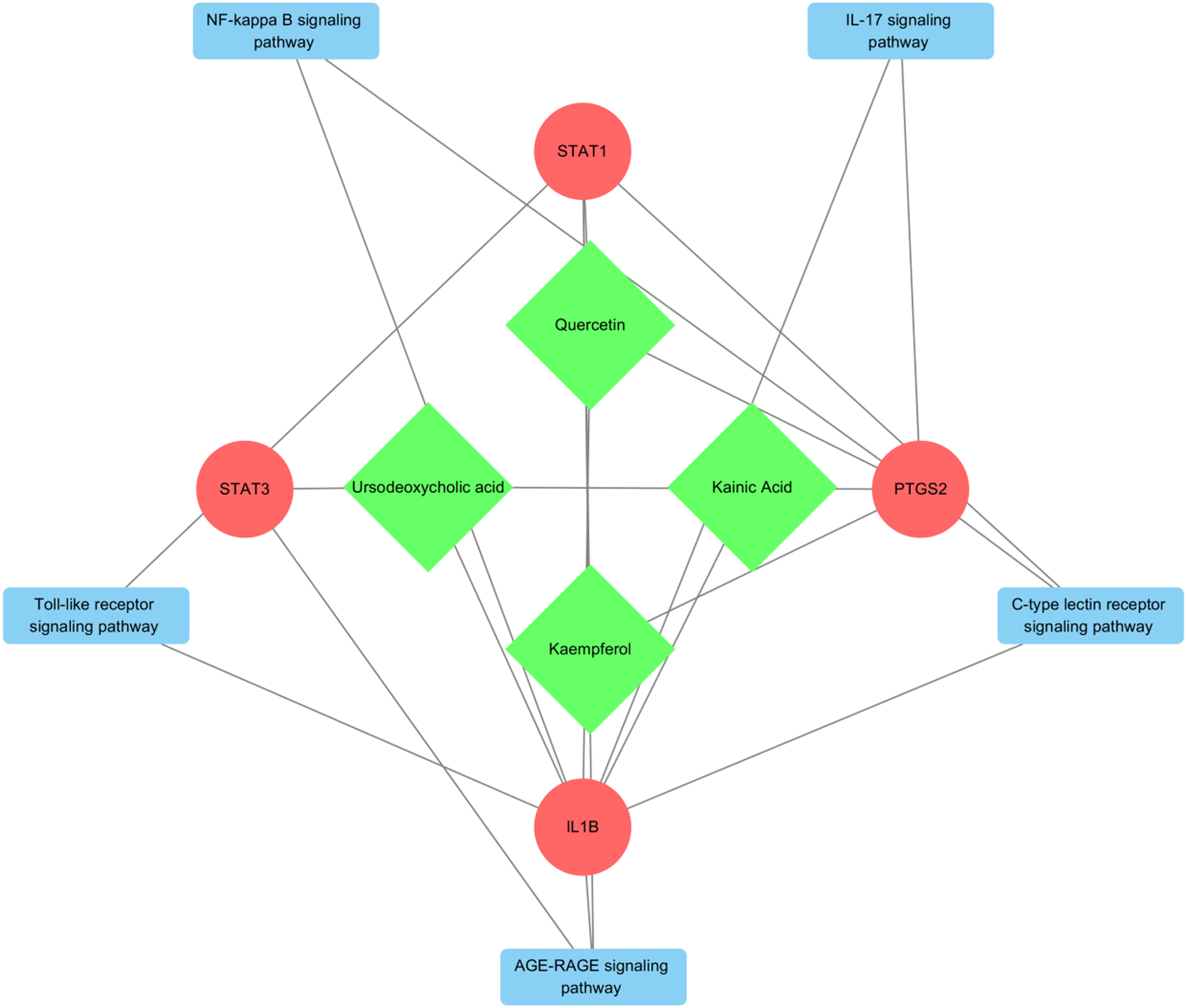
KEGG network diagram. Note: Inner ring diamonds represent key effective components, the middle ring circles represent key targets, and the outer ring rectangles stand for key signaling pathways.
KEGG Network.

Molecular Docking
Molecular docking was performed between key effective components and key targets with software including AutoDock. At binding energy <0, the ligand showed spontaneous binding to receptor, and lower binding energies suggested more stable binding conformations, with ≤ -4.25, ≤ -5.0 and ≤ -7.0 kcal/mol suggesting certain, good and strong binding activities, respectively. Our results showed binding energies < -7.0 kcal/mol, suggesting very strong binding activity of key effective components with key targets. This supported that our predictions were reliable. The results are presented in Table 5. The lowest affinity was observed between Quercetin and PTGS2, and their molecular docking can be observed from Figure 9.

Molecular docking.
Binding Energy Between Key Effective Components and Key Targets.

MD Simulations
MD simulations were performed to assess the stability and flexibility of the Quercetin-PTGS2 complex and to evaluate the interactions between Quercetin and PTGS2. RMSD can be used to evaluate changes in protein structure. RMSD between Quercetin and PTGS2 was determined (Figure 10). During the 100 ns simulation, the RMSD value between Quercetin and PTGS2 was initially high, which slightly became stable. In general, RMSD was in the small range (<0.25), suggesting that binding between Quercetin and PTGS2 remained relatively stable. RMSF can assess protein dynamics. RMSF between Quercetin and PTGS2 was determined (Figure 11). RMSF values of PTGS2 were lower in the binding regions and higher in the non-binding regions, suggesting that Quercetin increased the flexibility of PTGS2 domains. This made them more prone to conformational changes. Rg is an indicator for the evaluation of overall protein compactness. In this simulation, the Rg was calculated between Quercetin and PTGS2 (Figure 12). Therefore, Rg value declined following Quercetin binding to PTGS2, indicating that the binding of Quercetin made PTGS2 more compact. SASA is an indicator for assessing the the surface area of a protein. The SASA between Quercetin and PTGS2 was determined (Figure 13). PTGS2 had a larger SASA value before binding, which decreased after binding. This indicated that Quercetin binding reduced the surface area of PTGS2. Hydrogen Bonding (HBond) can evaluate the protein-small molecule hydrogen bonds. In the current simulation, the HBond between Quercetin and PTGS2 was calculated (Figure 14). The results demonstrated the formation of numerous hydrogen bonds between Quercetin and PTGS2, primarily between some key residues in PTGS2 and important groups in the small molecule. The FEL reflects energy distribution and interactions between molecules in this system, contributing to understanding the conformations, interactions and stability between Quercetin and PTGS2. The FEL showed a low-energy region (Figure 15), indicating that during the simulation, the molecular structure maintained a relatively stable state, mainly around 34 ns. In this low-energy region, the molecular structure probably exhibited a high similarity to the initial structure and was relatively compact, which indicates that the molecule maintained its stable status for a long time, indicating stability.

RMSD between quercetin and PTGS2.

RMSF between quercetin and PTGS2.

Rg between quercetin and PTGS2.
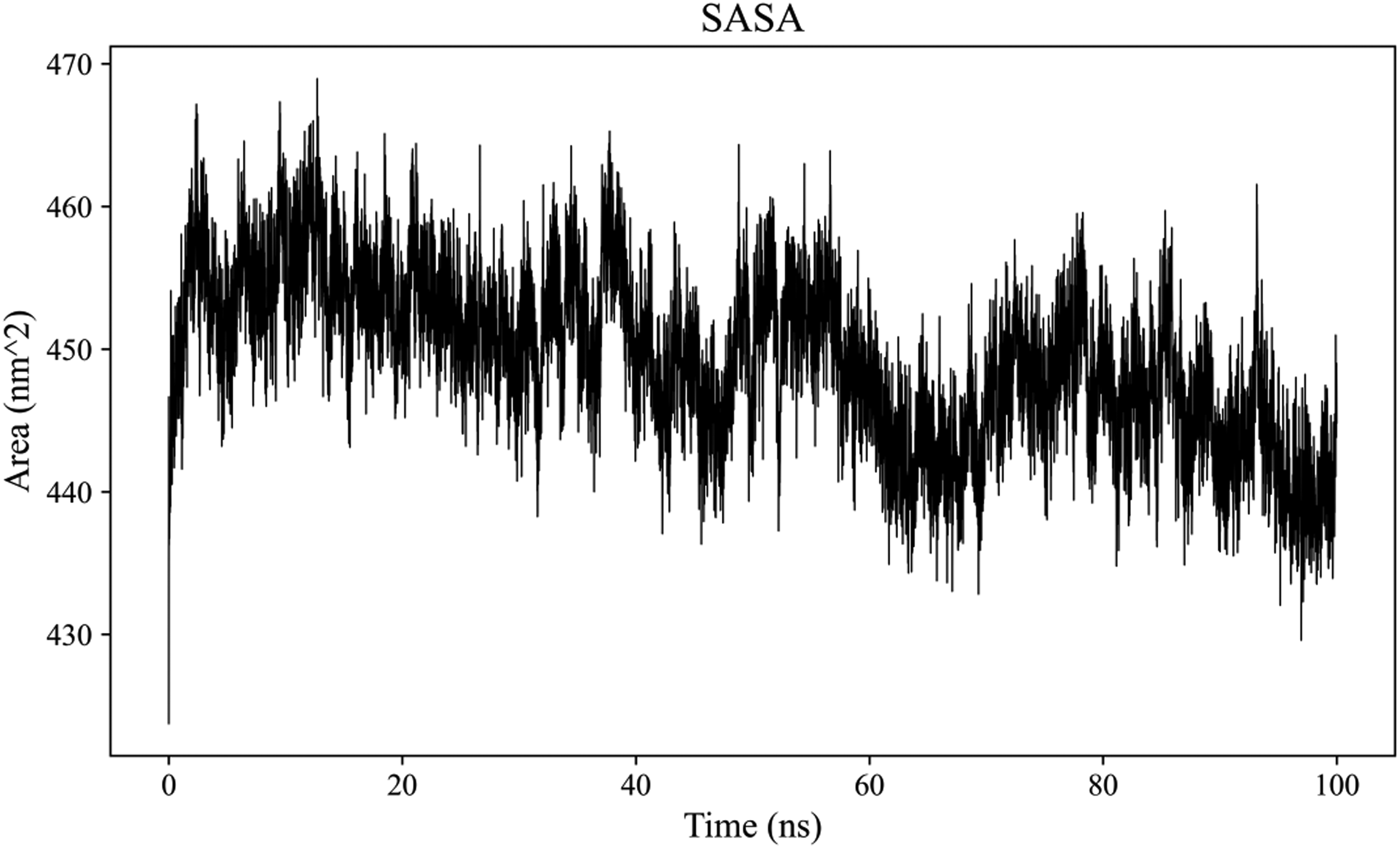
SASA between quercetin and PTGS2.

Hbond between quercetin and PTGS2.

FEL diagram.
Discussion
UHPLC-QE-MS technology is known for its advantages such as high resolution, high separation ability, and high sensitivity. Using this technology, it is possible to accurately characterize the compounds contained in traditional Chinese medicine, including precise mass numbers, elemental composition, and mass spectrometry fragmentation information. It has now been widely applied in the identification of chemical components in traditional Chinese medicine. 20 UHPLC-QE-MS technology was used for identifying chemical constituents in Panax notoginseng. Additionally, mass spectrometry profiles in positive/negative ion full-scan modes were established for the Panax notoginseng solution. Through the integration of network pharmacology, key effective components in Panax notoginseng for the treatment of SANFH were identified as Quercetin, Ursodeoxycholic acid, Kainic Acid, and Kaempferol. The identified key targets were IL1B, STAT3, PTGS2 (also known as COX2), and STAT1. It was found in this study that Quercetin significantly attenuated the production of inflammatory factors, and reduced the NOD-like receptor thermal protein domain associated protein 3 (NLRP3) inflammasome level, as well as pyroptosis-associated proteins including NLRP3, cleaved IL-1β and active caspase-1. It plays a regulatory role in osteoclasts, osteoblasts, and BMSCs, simultaneously promoting metabolic balance and bone health through antioxidant and anti-inflammatory pathways.21,22 In the meanwhile, studies have indicated that Quercetin can up-regulate STAT1 expression, therefore inhibiting lipopolysaccharide (LPS)/interferon-γ (IFN-γ)-induced macrophage activation and promoting IL-4-induced macrophage M2 polarization. 23 In the cartilage of patients with SANFH, tumor necrosis factor-α (TNF-α), COX-2, and IL-1β mRNA expression increase. 24 Based on Papakonstantinou's study, 25 ursodeoxycholic acid significantly decreased COX-2 and TNF-α expression. In addition, ursodeoxycholic acid decreased STAT3 phosphorylation as well as peroxiredoxin 2 (Prx2) expression, reducing reactive oxygen species (ROS)-mediated inflammation and oxidative stress. 26 Kaempferol has been found to improve osteogenic activity suppressed by dexamethasone, inhibit glucocorticoid-induced bone loss, and exert anti-osteoporotic effects. 27 The research conducted by Xu 28 suggested that Kaempferol mitigated BMSC apoptosis, enhanced their growth, angiogenesis, and migration, and reduced endothelial damage and dysfunction in SANFH. Many other studies have suggested that Kaempferol exerts bone-protection through suppressing adipogenesis, inflammation, osteoblast apoptosis, osteoclast autophagy, and oxidative stress, whereas simultaneously activates osteoblast autophagy. Kaempferol exerts its impact against osteoporosis by regulating the Estrogen Receptor (ER), Bone Morphogenetic Protein-2 (BMP-2), NF-κB, mammalian target of rapamycin (mTOR), and Mitogen-Activated Protein Kinase (MAPK) pathways.29,30 Based on the above studies, effective components in Panax notoginseng possess significant activities of anti-inflammation, bone protection, and immunomodulation in treating SANFH. Moreover, extensive literature demonstrates that these key targets are critical for the SANFH pathogenic mechanism and treatment.31,32
The PPI network revealed the interconnections among 50 targets, indicating that these targets did not function in isolation but exhibited close interactions. Among them, IL1B, STAT3, PTGS2, and STAT1 played crucial roles within the entire network. In our constructed compound-target network, numerous target genes were modulated through several effective components in Panax notoginseng, highlighting the multi-component and multi-target characteristics in traditional Chinese medicine therapies. Further functional annotation of key targets in the PPI network identified AGE-RAGE, NF-κB, IL-17, Toll-like receptor, and C-type lectin receptor as the main pathways associated with the core network of SANFH. Consistent with previous studies, these signaling pathways are shown to significantly influence the development and progression of SANFH.33–35
The AGE-RAGE signaling pathway is an upstream pathway of the NF-κB signaling pathway. Upon activation, both pathways lead to the increased expression of inflammatory factors, which in turn reduces cartilage elasticity and accelerates cartilage wear. Liu 36 indicated that the AGEs/RAGE/NF-κB pathway exerted a vital impact on mitigating oxidative stress, improving bone quality and providing bone protection. Furthermore, compared with normal tissues, NF-κB pathway can be significantly activated within bone tissue in SANFH patients. Inhibiting NF-κB pathway significantly attenuates TNF-α-mediated autophagy. The increase in TNF-α within SANFH regulates osteoblast apoptosis and autophagy through the NF-κB pathway. Blockage of this pathway with inhibitors exacerbates TNF-α-mediated apoptosis by damaging autophagic flux. 35 Zou's research 37 indicated that IL-17 levels increased in SANFH and were positively correlated with the severity of pain. Th17 cells can be recruited into the inflamed synovium, while the inflammatory cytokine IL-17 exhibits increased levels within hip joint synovium in SANFH patients, facilitating the development of clinical symptoms. Based on Cheng, 38 inflammatory factors IL-1β, IL-33, and IL-17A were induced within articular cartilage in a rat model of SANFH. Therefore, blocking IL-17 pathway may slow SANFH progression and alleviate associated pain symptoms. In osteoclastogenesis, RANKL stimulation promotes the secretion of pro-inflammatory proteins, enhancing RANKL-induced osteoclastogenesis. The C-type lectin receptor can regulate the expression of RANKL-induced osteoclast marker genes and intracellular MAPK/NF-κB signal transduction, improving osteoporosis in rats. 39 Thus, targeting the C-type lectin receptor may represent an effective approach for the treatment of osteoporotic symptoms associated with SANFH. In recent studies, Toll-like receptor 4 (TLR4), which is expressed on osteoblast and osteoclast surfaces, exerts an important effect on osteoporosis development. The aberrant TLR4 expression can trigger osteoclast and osteoblast dysregulation, which is a primary molecular mechanism in osteoporosis pathology, potentially involving interactions of TLR4 with the NF-κB pathway, pro-inflammatory effects, RUNX2 and ncRNAs. 40 Research has shown that TNF-α can increase the expression of TLR4, thereby promoting osteocyte necroptosis in osteoporosis. In osteoporosis patients, TLR4 is highly expressed in skeletal sites where osteocyte necroptosis occurs, and high TLR4 expression is associated with increased TNF-α expression. Osteocytes exposed to necroptotic osteocytes in vivo and in vitro show strong TLR4 expression. TNF-α upregulates TLR4 expression in vitro, which may further promote osteocyte necroptosis. Moreover, TAK-242 inhibits TLR4 in vitro, effectively blocking TNF-α-induced osteocyte necroptosis. 41 In addition, research has indicated that corticosteroids may interfere with immune responses via the TLR4 signaling pathway, inducing femoral head necrosis. 42 Moreover, the findings of this study suggested that the effective components of Panax notoginseng exerted therapeutic effects on SANFH in various biological processes or pathways, providing anti-inflammatory, antioxidant, analgesic, and bone-improving benefits.
According to our molecular docking analysis, key effective components of Panax notoginseng exhibited favorable docking activity with their respective key targets, suggesting that these components might exert a vital role in treating SANFH. Our predictions lay a foundation for therapeutic action of effective components in Panax notoginseng on specific targets. Among all the docking results, the binding affinity between Quercetin and PTGS2 was the highest, implying its inherent high bioactivity. This prompted further MD simulations of Quercetin and PTGS2. MD simulation is the strong calculation method widely adopted for studying the motion, structure, and interactions of molecules in various environments. By simulating the movement of atoms, MD provides detailed insights into MD, offering perspectives which are hardly be obtained through conventional pathological or biochemical assays. For example, MD simulations can reveal more details concerning the roles of mutations in protein structure and protein-protein/ligand interactions at an atom scale, making it an invaluable tool in the biomedical field. 43 MD simulations can explore not only the static structures of molecules but also their dynamic changes under different conditions, revealing the characteristics of molecular interactions. This method provides critical support for comprehending biological molecular mechanisms and drug-receptor interactions. Therefore, in this study, MD simulations were employed to conduct an in-depth investigation into the binding between Quercetin and PTGS2. 44 In the MD simulation of Quercetin and PTGS2, the RMSD results indicated that their binding remained relatively stable. Quercetin was found to increase the flexibility of PTGS2 domains, making them more prone to conformational changes. The decrease in Rg value suggested that PTGS2 became more compact after binding. The reduction in SASA value suggested a reduction in the surface area of PTGS2 post-binding. HBond analysis proved that several hydrogen bonds were formed between Quercetin and PTGS2, while the FEL showed that the molecular structure was most stable and compact around 34 ns. Overall, these results suggested that the interaction of Quercetin with PTGS2 promoted the stability of the protein's overall structure, internal stability, compactness, surface characteristics, and interaction strength. This implies that Panax notoginseng may exert its therapeutic impacts on SANFH by binding to PTGS2, potentially inhibiting PTGS2 expression and/or phosphorylation. This also suppressed the C-type lectin receptor pathway, contributing the treatment of SANFH.
Although the study identified 57 active components using UHPLC-QE-MS technology, due to the limitations of mass spectrometry, it may not have fully identified all the components present, especially those with low concentrations or difficult-to-interpret compounds. This study relied on the TCMSP and Uniprot databases to predict the targets of active components, and while these databases provided useful clues for target screening, there were still errors in the target predictions. While molecular docking and molecular dynamics simulations validated the binding ability and stability of the active components with the targets, these were only computer simulation results, and the actual biological effects may be influenced by various factors, failing to fully reflect the complexity of the in vivo environment. The absence of animal models or clinical trials for validation prevents a comprehensive evaluation of the true efficacy and safety of Panax notoginseng in treating steroid-induced femoral head necrosis. Future research can employ more advanced mass spectrometry techniques or combine other modern analytical methods to further comprehensively identify active components in Panax notoginseng, especially those with low concentrations. Further cell and animal experiments can be conducted to validate the key targets identified in this study and explore the specific physiological mechanisms of these targets in steroid-induced femoral head necrosis, thereby improving the multi-target pharmacological mechanism of Panax notoginseng. Additionally, through high-throughput screening and other methods, other potential targets and pathways of Panax notoginseng can be further explored to investigate its therapeutic potential in other osteoporosis-related diseases.
This study applied UHPLC-QE-MS technology to rapidly and comprehensively identify the active components of Panax notoginseng, providing a basis for further research on the treatment of SANFH and the pharmacological mechanisms of Panax notoginseng. Meanwhile, based on network pharmacology, molecular docking, and molecular dynamics simulation results, it is suggested that the active components of Panax notoginseng act on multiple targets and pathways to exert anti-inflammatory, cartilage-protective, and anti-osteoporosis effects, offering theoretical support for experimental and clinical studies on the treatment of SANFH with Panax notoginseng.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X251340455 - Supplemental material for Comprehensive Analysis of the Active Components and Mechanisms of Panax notoginseng in Steroid-associated Necrosis of Femoral Head Using UHPLC-QE-MS, Network Pharmacology, and Molecular Dynamics Simulations
Supplemental material, sj-docx-1-npx-10.1177_1934578X251340455 for Comprehensive Analysis of the Active Components and Mechanisms of Panax notoginseng in Steroid-associated Necrosis of Femoral Head Using UHPLC-QE-MS, Network Pharmacology, and Molecular Dynamics Simulations by Jie Han, Kebin Shang, Qinglin Peng, Yu kun Wu, Yachao Wu and Bin Nong in Natural Product Communications
Footnotes
Acknowledgments
Everyone who contributed significantly to this study has been listed.
Ethical Considerations
This study was approved by the Medical Ethics Committee of Ruikang Hospital Affiliated to Guangxi University of Chinese Medicine (YJS2023-003)*, Guangxi Zhuang Autonomous Region, China.
Funding
National Natural Science Foundation of China Project (82260858); Guangxi Natural Science Foundation (2023JJA140414); Pilot project for the construction of high-level key disciplines of traditional Chinese medicine in Guangxi (Guizhong Medical Science and Education Development [2023] No. 13); The construction project of Guangxi Key Research Laboratory of Traditional Chinese Medicine (Guizhong Pharmaceutical Science and Education Development [2023] No. 9); Guangxi Youth Qihuang Scholar Training Program (Guizhong Medical Science and Education Development [2022] No. 13); Guangxi University of Traditional Chinese Medicine “Qihuang Project” High level Talent Team Cultivation Project (202413).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
Data sharing is not applicable to this article as no datasets were generated or analyzed during the current study.
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
