Abstract
Keywords
Introduction
Alcoholic liver disease (ALD) is a condition arising from chronic heavy alcohol consumption. It typically begins with simple fatty liver, potentially progressing to alcoholic hepatitis, liver fibrosis, and cirrhosis. 1 Chronic severe alcohol abuse can lead to extensive hepatocyte necrosis and may even result in liver failure, which are primary causes of cirrhosis. 2 The clinical classification of ALD comprises five types: mild ALD, alcoholic fatty liver disease, alcoholic hepatitis, alcoholic liver fibrosis, and alcoholic cirrhosis. 3 Mild ALD is typically asymptomatic, with liver biochemical markers, imaging, and histopathological findings generally being normal or only mildly abnormal. 4 Serum total bile acids (TBA) serve as a sensitive marker of liver clearance function, particularly in patients with suspected liver disease where other biochemical markers are normal or only mildly abnormal. 5 If appropriate preventive and therapeutic measures are implemented during the early stages of ALD, specifically mild ALD, effective treatment for ALD can be achieved, thereby preventing disease progression.
Sesamin (SES) is the highest content of lignamin in sesame and sesame oil. Domestic and international studies, including those from our research group, have demonstrated that SES possesses antioxidant,6,7 anti-inflammatory,8,9 antihypertensive,10,11 cholesterol-lowering12,13 and hepatoprotective 14 properties. Recently, our research has shown that SES can decrease serum TBA levels in rats with mild ALD; however, the underlying mechanism remains unclear, with no correlations reported in the domestic and international literature. Consequently, this study employed mRNA-seq technology to identify differentially expressed genes associated with bile acid metabolism, followed by a series of validation experiments aimed at elucidating the specific mechanism by which SES reduces serum TBA levels in rats with mild ALD, and at establishing an experimental and theoretical foundation for the development of effective ALD treatments.
Materials and Methods
Drugs and Reagents
SES was obtained from Nanjing Guangrun Biological Products Co., Ltd. HE and Masson staining kits, as well as reagents for Western blot and q-PCR, were sourced from Biyun Biotechnology. The design and synthesis of primers (Slco1b2, Abcc3, Abcb4, Slc27a2, Baat, and Nr1d1) were carried out by Beijing Tsingke Biotech Co., Ltd. Antibodies (Oatp1b2, MRP3, and REV-ERBα) were acquired from Thermo Fisher Scientific Inc. Details of the primers used in this study are provided in Table 1.
Primers in This Study.

Experimental Animals and Group Administration
Twenty-one SD rats, obtained from Beijing Victoria Experimental Animal Technology Co., Ltd, were randomly assigned to three groups: normal control (Normal), mild ALD model (Model), and SES at 160 mg/kg (Model + SES). The sample size was determined based on a power analysis, which ensured that the study had a power of at least 0.80 to detect a significant difference at the α = 0.05 level between the groups. The SES 160 mg/kg group was administered the SES solution once daily at 12:00 PM. The normal control group was administered an equivalent volume of distilled water. Rats were weighed prior to each administration to adjust dosages accordingly. The experimental period was set for 8 days.
Establishment of the Mild ALD Rat Model
To establish the mild ALD rat model, we followed a previously validated protocol with slight modifications. Male Sprague-Dawley rats (n = 7) weighing 200-250 g were used. The rats were administered ethanol (EtOH) by gavage at a dose of 7 mL/kg body weight, 2 times per day, for a period of 1 week. The EtOH was diluted in a vehicle solution of distilled water. Control rats received the same volume of the vehicle solution without EtOH. The model was considered successfully established when the rats displayed mild liver enzyme elevation without significant liver inflammation or fibrosis, as confirmed by histopathological analysis.
Housing and Maintenance of Rats
All rats were housed in a specific pathogen-free environment within the animal facility at Wannan Medical College. The animals were maintained under controlled conditions with a 12-h light/dark cycle, a temperature of 22 ± 2 °C, and a relative humidity of 40%-70%. Each cage contained up to five rats and was equipped with bedding material for comfort. The rats had free access to a standard laboratory diet and tap water, ad libitum. The health and well-being of the animals were monitored daily by the animal care staff, and any signs of illness or distress were promptly addressed by a veterinarian. All procedures were conducted in accordance with the guidelines for the care and use of laboratory animals and were approved by the Wannan Medical College Animal Care and Use Committee.
Weight and Food Intake
To evaluate the effects of the study on the body weight and food intake of rats, we performed a series of assessments before and after the experimental period. At the commencement of the study, all rats were weighed to establish their initial body weight. We measured food intake over a 24-h period for each group by supplying a predetermined quantity of food and recording the residual after 24 h. This process was repeated at the end of the 7-day study period. The collected data on body weight and food intake were subsequently compiled and subjected to statistical analysis.
Ultrasonic Examination
Rats were anesthetized using isoflurane, followed by preparation of the abdominal skin. An ultrasound examination device was utilized to assess the uniformity of the liver parenchymal echo, the integrity of the liver capsule, and the blood flow within the liver portal vein.
Biochemical Testing
Rats were anesthetized with isoflurane. Euthanasia was performed via cervical dislocation, which is in accordance with the protocols approved by the Institutional Animal Care and Use Committee (IACUC). This method ensures a humane and rapid death for the animals. Following anesthetization with isoflurane, a midline laparotomy was conducted to expose the abdominal aorta. Blood samples were drawn from the abdominal aorta of the anesthetized rats. An approximate volume of 5 mL of blood was withdrawn using a 23-gauge needle and a 10-mL syringe. The collected blood was promptly transferred to serum separator tubes and incubated at room temperature for 30 min to allow clotting prior to centrifugation. Blood samples from rats were centrifuged at 2000 rpm for 10 min at 4 °C to isolate the supernatant. The serum concentrations of thiobarbituric acid-reactive substances (TBA), alanine aminotransferase (ALT), and aspartate aminotransferase (AST) were quantified in rat serum with a fully automated biochemical analyzer.
HE Staining and Masson Staining
Liver tissues were fixed in paraformaldehyde, embedded in paraffin, and sectioned into 5-μm slices. The sections were then stained using HE and Masson staining kits, and images were captured under a microscope.
Transcriptome Sequencing
Liver tissues were stored in pre-cooled RNase-free frozen storage tubes, rapidly frozen in liquid nitrogen, and subsequently sent to Novogene Co., Ltd for transcriptome sequencing analysis. Samples that passed quality control were sequenced on the Illumina NovaSeq 6000 platform, and differential genes were identified using DESeq2 software. The differential genes were analyzed for Gene Ontology (GO) functional categorization and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment.
Quantitative Polymerase Chain Reaction
Total RNA was extracted from liver tissue, and mRNA was reverse transcribed into cDNA using a reverse transcription kit. The synthesized cDNA was mixed with SYBR Green reagent and primers, and then analyzed by q-PCR. Relative mRNA expression levels were calculated using the 2^−ΔΔCT method, with actin as the internal reference gene. The gene names and sequences of the primers are shown in Table 1.
Western Blotting
Total proteins were extracted from each group, and their concentrations were determined using the BCA assay. The proteins were separated by 10% SDS-PAGE, transferred to a PVDF membrane, blocked in 5% non-fat milk for 2 h, incubated overnight at 4 °C with the primary antibody, washed in PBST, and then incubated with the secondary antibody for 30 min at room temperature. The bands were visualized by chemiluminescence and quantified using Image-Pro Plus software.
Statistical Analysis
Statistical Analysis Measurement data are expressed as mean ± standard deviation (SD). Statistical analysis was performed using one-way analysis of variance (ANOVA). Prior to ANOVA, the normality of the data was assessed using the Shapiro-Wilk test, and the homogeneity of variances was evaluated using Levene's test. If the data were normally distributed and variances were homogeneous, a Tukey's post hoc test was applied to determine the significance of differences between groups. In cases where the assumption of homogeneity of variances was violated, a Welch's ANOVA was used, followed by a Games-Howell post hoc test. The significance level for all tests was set at P < .05. Differential mRNA expression analysis of two conditions/groups (two biological replicates per condition) was performed using the DESeq2 R package (1.20.0). DESeq2 provides statistical routines for determining differential expression in digital gene expression data using a model based on the negative binomial distribution, with a significance level set at P < .05.
Statement of Compliance with Guidelines
The reporting of this study conforms to ARRIVE 2.0 guidelines. 15
Results
Effect of Sesamin on Body Weight and Food Intake in Mild ALD Rats
Data on changes in body weight and food intake for each group of rats are presented in Table 2. At the commencement of the study (Day 0), no significant differences were observed in the average body weight among the Normal, Model, and Model + SES groups. By Day 7, the Model group exhibited a significant increase in body weight relative to the initial weight (P < .05). Conversely, the Model + SES group manifested a less pronounced increase in body weight, which was significantly different from that of the Model group (P < .05), suggesting that the treatment may have attenuated the weight gain. As for food intake, at Day 0, no significant differences were observed among the groups. After 7 days, the Model group displayed an increase in food intake compared to the initial intake. The Model + SES group also evidenced an increase in food intake, although the increase was less marked than in the Model group, implying that the treatment may have impacted the dietary patterns of the rats. These findings suggest that the development of mild ALD in the Model group was associated with an increase in both body weight and food intake. The SES treatment in the Model + SES group seemed to lessen these effects, as evidenced by the diminished body weight gain and food intake, aligning with the observed improvement in ALD symptoms within this group.
Effect of Sesamin on Body Weight and Food Intake in Mild ALD Rats.
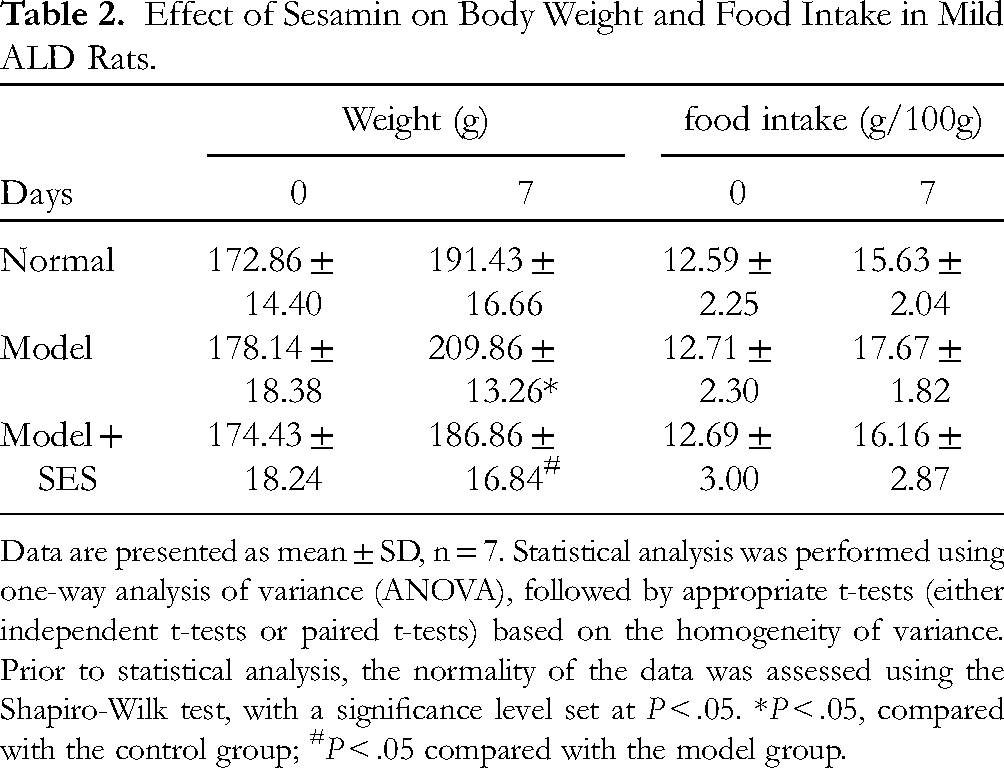
Data are presented as mean ± SD, n = 7. Statistical analysis was performed using one-way analysis of variance (ANOVA), followed by appropriate t-tests (either independent t-tests or paired t-tests) based on the homogeneity of variance. Prior to statistical analysis, the normality of the data was assessed using the Shapiro-Wilk test, with a significance level set at P < .05. *P < .05, compared with the control group; #P < .05 compared with the model group.
Effect of SES on Pathological Changes in the Liver of Rats with Mild ALD
Liver ultrasound examination revealed normal liver morphology and size, uniform parenchymal echo, unenlarged portal veins, no biliary dilation, and absence of space-occupying lesions (Figure 1A). HE staining demonstrated normal cellular architecture, without evident inflammatory cell infiltration, steatosis, pseudolobule formation, or fibrosis (Figure 1B). Masson staining showed no evidence of collagen deposition in the liver tissues across all groups (Figure 1C). These observations indicate that the gross morphological and structural integrity of the liver was preserved.

Effect of sesamin on pathological changes in the liver of mild ALD rats (n = 7). A, ultrasonic examination, B, HE staining, C, Masson staining. Scale bar: 50 μm. Magnification: 200×. SES, sesamin; ALD, alcoholic liver disease.
Effect of SES on serum TBA, ALT and AST in Rats with Mild ALD
As illustrated in Figure 2, serum TBA levels were significantly higher in the mild ALD model group compared to the normal controls (P < .05). Following SES treatment, serum TBA levels showed a significant reduction (P < .05). In this study, our observations revealed that serum TBA levels in rats were significantly higher than those in the normal control group, whereas AST and ALT levels did not change significantly. These results suggest that, despite the absence of overt pathological changes, liver function, as indicated by TBA levels, was impaired in the mild ALD model. The subsequent reduction in TBA levels after SES treatment suggests a potential therapeutic effect.

Effect of sesamin on serum TBA, ALT, and AST in rats with mild ALD. Data are presented as mean ± SD, n = 7. Statistical analysis was performed using one-way analysis of variance (ANOVA), followed by a t-test for homogeneity of variance and a t’test for heterogeneous variance. Prior to statistical analysis, the normality of the data was assessed using the Shapiro-Wilk test, with a significance level set at P < .05. *P < .05, compared with the control group; #P < .05 compared with the model group. TBA, thiobarbituric acid reactive substances; ALT, alanine aminotransferase; AST, aspartate aminotransferase; ALD, alcoholic liver disease.
Effect of SES on Differential mRNA in the Liver of Mild ALD Rats
In comparison with the normal control group, there were 2010 mRNAs with significant differences in the livers of rats in the mild ALD model group, of which 910 were upregulated and 1100 were downregulated. In comparison with the model group, there were 1933 mRNAs with significant differences in the livers of rats in the SES treatment group, of which 1133 were upregulated and 800 were downregulated, as illustrated in Figure 3A.

Effect of SES on differential mRNA expression in the liver of mild ALD rats (n = 7). A. Bar graph illustrating the combined statistics of differential gene numbers. B. Venn diagram showing the commonality of differential mRNA quantities in the liver among the normal control group, mild ALD model group, and SES administration group. C. Among the 442 genes shared by the three groups, relative to the normal control group, there were 286 upregulated and 157 downregulated mRNAs with significant differences in the liver of the mild ALD model group, determined by RNA-Seq analysis and a false discovery rate (FDR) < .05. D. Among the 442 genes shared by the three groups, in the livers of rats administered SES, there were 179 upregulated and 263 downregulated mRNAs with significant differences compared to the model group, as assessed by RNA-Seq analysis and a false discovery rate (FDR) < 0.05. SES, sesamin; ALD, alcoholic liver disease; FDR, false discovery rate.
In the livers of rats from the normal control group, the mild ALD model group, and the SES treatment group, 442 mRNAs were commonly differentially expressed, as illustrated in Figure 3B. Among these 442 commonly differentially expressed genes, relative to the normal control group, 286 were upregulated and 157 were downregulated in the livers of rats in the mild ALD model group, as shown in Figure 3C. Among the 442 commonly differentially expressed genes, in comparison with the model group, 179 were upregulated and 263 were downregulated in the livers of rats in the SES treatment group, as depicted in Figure 3D.
To accentuate the most pronounced transcriptomic disparities, we have constructed a table enumerating the top ten differentially expressed genes, ranked by their log2FoldChange and statistical significance. Table 3 furnishes an exhaustive summary of these genes, the alterations in their expression levels, and their respective functional annotations.
Top Ten Differentially Expressed Genes in the Liver of Mild ALD Rats.
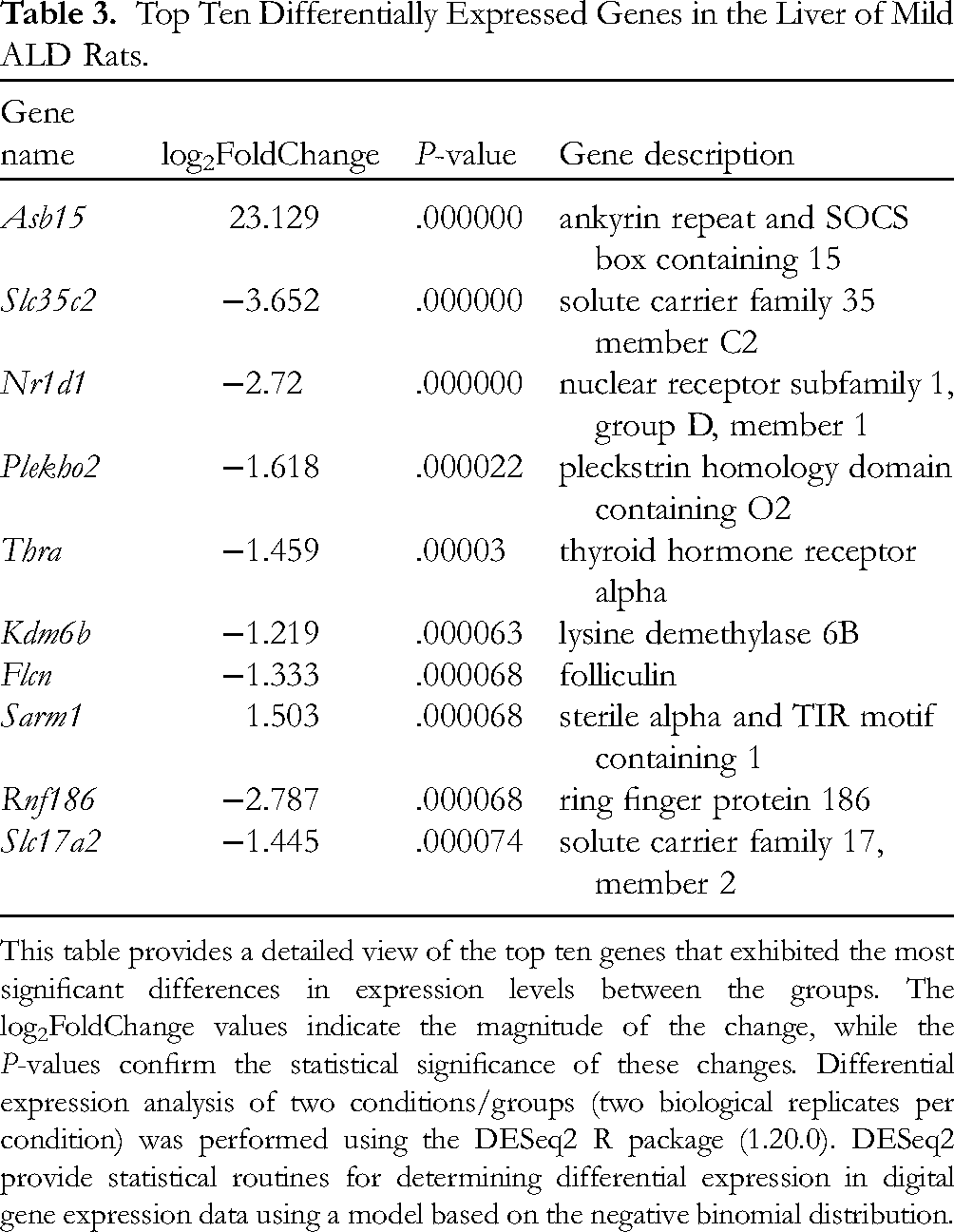
This table provides a detailed view of the top ten genes that exhibited the most significant differences in expression levels between the groups. The log2FoldChange values indicate the magnitude of the change, while the P-values confirm the statistical significance of these changes. Differential expression analysis of two conditions/groups (two biological replicates per condition) was performed using the DESeq2 R package (1.20.0). DESeq2 provide statistical routines for determining differential expression in digital gene expression data using a model based on the negative binomial distribution.
Effect of SES on Differential mRNA GO Functional Enrichment in the Liver of Mild ALD Rats
GO functional enrichment analysis was performed on the 442 commonly differentially expressed mRNAs identified in the livers of rats across the normal control group, the mild ALD model group, and the SES treatment group. The analysis revealed 507 molecular functions (MF), with 71 being statistically significant (P < .05); 3364 biological processes (BP), with 468 being statistically significant (P < .05); and 371 cellular components (CC), with 35 being statistically significant (P < .05).
As depicted in Figure 4, BP was primarily enriched in processes such as peptide cross-linking, chondroitin sulfate proteoglycan metabolic process, mitochondrial genome maintenance, liver development, hepaticobiliary system development, cellular response to steroid hormone stimulus, regulation of system process, chondroitin sulfate metabolic process, negative regulation of muscle tissue development, monocarboxylic acid metabolic process, etc.
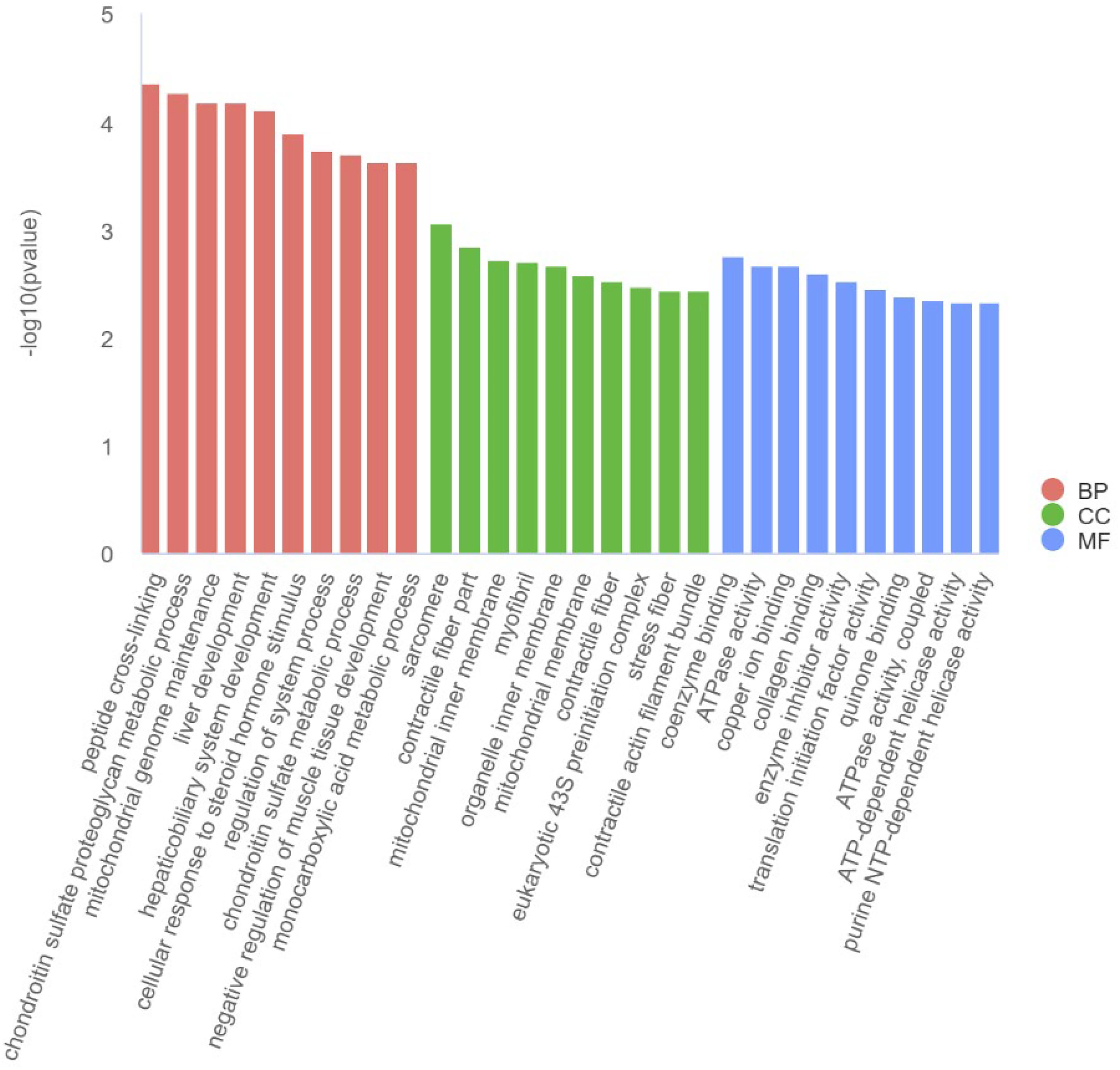
Effect of SES on differential mRNA GO functional enrichment in the liver of mild ALD rats, n = 7. GO functional enrichment analysis was performed using the Fisher's exact test, and the false discovery rate (FDR) was adjusted using the Benjamini-Hochberg procedure. Only GO terms with an FDR < 0.05 were considered significantly enriched. SES, sesamin treatment; ALD, alcoholic liver disease; GO, Gene Ontology; FDR, false discovery rate; BP, biological processes; CC, cellular components; MF, molecular functions.
Figure 4 also illustrates that CC was primarily enriched in sarcomere, contractile fiber part, mitochondrial inner membrane, myofibril, organelle inner membrane, mitochondrial membrane, contractile fiber, eukaryotic 43S preinitiation complex, stress fiber, contractile actin filament bundle, etc.
Furthermore, Figure 4 shows that MF was primarily enriched in coenzyme binding, ATPase activity, copper ion binding, collagen binding, enzyme inhibitor activity, translation initiation factor activity, quinone binding, ATPase activity, coupled, ATP-dependent helicase activity, purine NTP-dependent helicase activity, etc.
Effect of SES on Hepatic Differential mRNA KEGG Signaling Pathway Enrichment in Mild ALD Rats
KEGG signaling pathway enrichment analysis was performed on 442 commonly differentially expressed mRNAs from the normal control group, mild ALD model group, and SES administration group. The analysis revealed 257 enriched signaling pathways, 45 of which were statistically significant (P < .05).
As shown in Figure 5, The top 20 most significantly enriched pathways included the Glucagon signaling pathway, Bladder cancer, Non-small cell lung cancer, Circadian rhythm, ABC transporters, Human papillomavirus infection, EGFR tyrosine kinase inhibitor resistance, Thermogenesis, Endometrial cancer, JAK-STAT signaling pathway.

Effect of SES on hepatic differential mRNA KEGG signaling pathway enrichment in mild ALD rats, n = 7. The KEGG signaling pathway enrichment analysis was conducted using the hypergeometric test, and the significance of enrichment was determined by a corrected P-value (FDR) < 0.05. SES, sesamin; ALD, alcoholic liver disease; KEGG, Kyoto Encyclopedia of Genes and Genomes; FDR, false discovery rate.
Effect of SES on Differential mRNA Associated to Bile Acid Metabolism in Liver of Mild ALD Rats
From the common differentially expressed mRNAs across the three groups, six were identified that are associated with bile acid metabolism. These include ATP binding cassette subfamily B member 4 (Abcb4), ATP binding cassette subfamily C member 3 (Abcc3), bile acid CoA: amino acid N-acyltransferase (Baat), nuclear receptor subfamily 1 group D member 1 (Nr1d1), solute carrier family 27 member 2 (Slc27a2), and solute carrier organic anion transporter family member 1B2 (Slco1b2).
Effect of SES on the Levels of Abcb 4, Abcc 3, Baat, Nr1d1, Slc27a2, and Slco1b2 mRNA in the Liver of Rats with Mild ALD
As shown in Figure 6, in comparison to the normal control group, the mRNA levels of Slco1b2 and Nr1d1 in the liver of the mild ALD model group were significantly reduced (P < .01), whereas the mRNA levels of Abcb4, Slc27a2, and Baat were significantly elevated (P < .01). In comparison to the model group, the mRNA levels of Slco1b2 and Nr1d1 in the liver of the SES administration group were significantly upregulated (P < .05 or P < .01), and the mRNA level of Abcc3 was significantly downregulated (P < .05). These findings suggest that SES may increase the mRNA levels of Slco1b2 and Nr1d1 and decrease the mRNA levels of Abcc3.

Effect of SES on the hepatic expression of Slco1b2 (A), Abcc3 (B), Nr1d1 (C), Abcb4 (D), Slc27a2 (E) and Baat (F) mRNA in rats with mild ALD. Data are presented as mean ± SD, n = 7. mRNA expression levels were quantified by qPCR and analyzed using one-way ANOVA followed by t-test. *P < .05, compared with the control; #P < .05, ##P < .01 compared with the model. SES, sesamin; ALD, alcoholic liver disease; qPCR, quantitative polymerase chain reaction; ANOVA, analysis of variance.
Effect of SES on the Expression of Oatp1b2 (Slco1b2), MRP 3 (Abcc3), and REV-ERB α (Nr1d1) in the Liver of Rats with Mild ALD
As shown in Figure 7, in comparison to the normal control group, hepatic Oatp1b2 (Slco1b2) and REV-ERBα (Nr1d1) protein expressions were significantly reduced (P < .01), and MRP3 (Abcc3) expression was significantly elevated (P < .01). In comparison to the Model group, Oatp1b2 (Slco1b2) and REV-ERBα (Nr1d1) protein expressions in the Model + SES group were significantly upregulated (P < .05 or P < .01), whereas MRP3 (Abcc3) expression was significantly downregulated (P < .05). These findings suggest that SES increased Oatp1b2 (Slco1b2) and REV-ERBα (Nr1d1) protein expressions and decreased MRP3 (Abcc3) expression.

Effect of SES on the protein expression of Oatp1b2 (A), MRP3 (B), and REV-ERBα (C) in the liver of mild ALD rats. Data are presented as mean ± SD, n = 7. Protein expression was measured by Western blotting and analyzed using one-way ANOVA with t-test. *P < .05, compared with the control; #P < .05, ##P < .01 compared with the model. SES, sesamin; ALD, alcoholic liver disease; Oatp1b2, organic anion transporting polypeptide 1b2; MRP3, multidrug resistance-associated protein 3; REV-ERBα, nuclear receptor subfamily 1 group D member 1.
Discussion and Conclusions
Patients with mild ALD are typically asymptomatic, and liver biochemical markers, imaging, and histopathological examinations are generally normal or only mildly abnormal. 3 TBA measurement serves as a sensitive indicator of liver clearance function, particularly for diagnosing patients with suspected liver disease where other biochemical markers are normal or only mildly abnormal. 5 In this study, we observed that serum TBA levels in rats were significantly higher than those in the normal control group, whereas AST and ALT levels did not change significantly, and no damage was detected by ultrasound examination, HE, or Masson staining. This suggests that alcohol may cause an increase in serum TBA at the onset of ALD, consistent with a previous report. 16 Studies have suggested that disrupted bile acid homeostasis is a significant feature and a potent driver of alcohol-induced liver injury. 17 Thus, targeted interventions in bile acids and their metabolic pathways may represent a viable therapeutic alternative for ALD. 18
The liver is the sole organ responsible for synthesizing bile acids, which are derived from cholesterol in hepatocytes. 19 Bile acids are secreted into the gallbladder via the bile salt export pump (BSEP) and multidrug resistance protein 2 (MRP2), stored in the gallbladder, and then subsequently released into the duodenum. When cholestasis occurs, large amounts of bile acids are excreted into the systemic circulation through the hepatocyte multidrug resistance proteins 3 (MRP3) and 4 (MRP4), as well as the organic solute transporters α (OSTα) and β (OSTβ) transport system. 20 The bile acids in the systemic circulation are actively taken up by hepatocytes via the sodium-taurocholate cotransporting polypeptide (NTCP) and organic anion transporting polypeptide 1 (OATP1, rat Oatp1b2) transport systems. Long-term alcohol consumption can significantly increase the mRNA levels encoding the bile acid efflux proteins MRP4 and OSTα/β, and significantly decrease the expression of the bile acid uptake transport protein NTCP. This results in a significant increase in total bile acid levels and alterations in the bile acid profile, leading to bile acid accumulation in the liver. The accumulation of bile acids further leads to hepatocellular injury, inflammation, and fibrosis. 21
SES, the most abundant lignan in sesame and sesame oil, possesses antioxidant, anti-inflammatory, antihypertensive, cholesterol-lowering, and neuroprotective effects. 22 Our research group has been actively involved in researching SES. Previous studies have demonstrated that SES ameliorates endothelium-dependent vascular dysfunction and renal injury in hypertensive rats fed a high-fat, high-glucose diet. 23 SES enhances aortic endothelium-dependent relaxation in spontaneously hypertensive rats by increasing nitric oxide activity, upregulating endothelial nitric oxide synthase expression, and downregulating NADPH oxidase subunit expression.10,24 SES also prevents myocardial fibrosis in spontaneously hypertensive rats by inhibiting the TGF-β1/Smad signaling pathway. 25 Furthermore, SES suppresses STZ-induced apoptosis in INS-1 cells by inhibiting NF-κB activation and regulating the expression of Bcl-2 family proteins. 26 Recently, we unexpectedly discovered that SES can reduce serum TBA levels in mild ALD rats; however, the underlying mechanism remains unclear, with no previous reports documented domestically or internationally. Subsequently, we performed mRNA-seq analysis on liver tissue and identified six differentially expressed genes associated with bile acid metabolism: Slco1b2, Abcc3, Abcb4, Slc27a2, Baat, and Nr1d1. Following this, q-PCR and Western blot analyses were conducted for further verification, revealing the most significant differences in Slco1b2 (Oatp1b2), Abcc3 (MRP3), and Nr1d1 (REV-ERBα). Compared to the normal control group, the mRNA expression levels of Slco1b2 and Nr1d1 in the livers of mild ALD model rats were significantly lower, whereas the mRNA expression levels of Abcb4, Slc27a2, and Baat were significantly higher. Furthermore, the protein expression levels of Oatp1b2 (Slco1b2) and REV-ERBα (Nr1d1) were significantly reduced, and those of MRP3 (Abcc3) were significantly elevated. Compared to the mild ALD model group, the expression levels of Oatp1b2 (Slco1b2) and REV-ERBα (Nr1d1) in the livers of the sesamin-treated group were significantly higher, whereas the expression level of MRP3 (Abcc3) was significantly lower. The findings indicated that SES enhanced the active uptake of bile acids into hepatocytes by upregulating the expression of the bile acid transporter Oatp1b2 (Slco1b2), and simultaneously downregulated the expression of MRP3 (Abcc3), thereby decreasing the efflux of bile acids into the systemic circulation, which consequently led to a reduction in serum bile acid levels in mild ALD.
The Nr1d1 gene, a member of the nuclear receptor subfamily 1, encodes ligand-sensitive transcription factors that negatively regulate the expression of core clock genes, particularly the circadian clock transcription factor aryl hydrocarbon receptor nuclear translocator, involved in regulating genes that play roles in metabolism, inflammation, and cardiovascular processes. 27 Studies have indicated that Nr1d1 mediates cholesterol level elevation. 28 Nr1d1 regulates the expression of the serine protease inhibitor, group A member 3, thereby influencing lipid metabolism. 29 Deletion of exons 3 and 4 in the murine Nr1d1 gene intensifies hepatic steatosis induced by a high-fat diet. 30 Therefore, alcohol may modulate bile acid transporter activity via the Nr1d1 receptor, leading to increased serum TBA levels. Compared to the model group, hepatic Nr1d1 mRNA levels and REV-ERBα protein expression were significantly elevated in the sesamin-treated group.
Baat is a key enzyme involved in bile acid metabolism, and its expression level is closely associated with bile acid homeostasis and liver health. We investigated the effect of SES on hepatic Baat expression. Our results showed no significant effect of SES on hepatic Baat expression. However, when analyzing the model + SES group data, we observed a correlation between SES and hepatic Baat expression. Specifically, SES treatment was associated with lower hepatic Baat expression. These findings suggest that SES may indirectly influence hepatic Baat expression through other pathways. Currently, there is limited research on the effect of SES on hepatic Baat expression. Prior research indicates an association between SES and the risk of liver diseases, such as non-alcoholic fatty liver disease (NAFLD) and non-alcoholic steatohepatitis (NASH). This evidence suggests that SES may influence liver health through lifestyle, diet, and metabolic pathways. However, none of these studies has directly assessed the effect of SES on hepatic Baat expression. Our findings further suggest that SES may indirectly influence hepatic Baat expression through other pathways, such as affecting gut microbiota composition, which in turn could influence hepatic Baat expression. Furthermore, SES may influence lipid metabolism pathways, leading to hepatic fat accumulation and subsequently affecting hepatic Baat expression. Our findings regarding the effect of SES on hepatic Baat expression may provide partial explanation for the association between SES and the risk of liver diseases. Further research is needed to explore the mechanisms by which SES influences hepatic Baat expression and the impact of this effect on liver health.
In summary, our study provides novel insights into the molecular mechanisms underlying the protective effects of sesamin against ALD by identifying Nr1d1 (REV-ERBα) as a key therapeutic target for modulating bile acid metabolism. The modulation of bile acid transporters by sesamin, specifically the upregulation of Slco1b2 (OATP1) and downregulation of Abcc3 (MRP3), enhances the active uptake of bile acids by hepatocytes and diminishes their efflux into the systemic circulation, thereby reducing serum bile acid levels—a pivotal aspect of ALD management. Our findings establish sesamin as a potentially safe and effective natural compound for ALD management and set the stage for the development of targeted therapies. As the first to propose sesamin as a therapeutic agent for mild ALD, our results contribute to the emerging evidence supporting the use of natural products in the treatment of liver disease. The translational potential of our findings, supported by preclinical models and molecular analyses, highlights the significance of our discovery to the medical community and underscores the importance of further research in this area.
Our research has two key limitations, even as it identifies sesamin's potential protective effects on mild ALD. Firstly, the study's exclusive reliance on a rodent model may not adequately capture the human ALD pathology, implying the need for clinical studies to validate the applicability of our findings in humans. Secondly, our molecular analysis, despite revealing changes in specific bile acid transporters, does not comprehensively cover the broader genetic and biochemical landscape of mild ALD. A more in-depth molecular profiling is essential to fully understand the disease's complexity and sesamin's therapeutic mechanisms. In light of these limitations, our findings should be viewed as a preliminary basis for subsequent research rather than a conclusive conclusion. Future studies, with an expanded molecular focus and rigorous clinical trials, are crucial to fully elucidate the therapeutic potential of sesamin in treating mild ALD.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X251327306 - Supplemental material for Exploring the Mechanism of Sesamin in Treating Mild Alcoholic Liver Disease Based on Transcriptome Sequencing
Supplemental material, sj-docx-1-npx-10.1177_1934578X251327306 for Exploring the Mechanism of Sesamin in Treating Mild Alcoholic Liver Disease Based on Transcriptome Sequencing by Moxuan Li, Yundong Wang, Yunqing Peng, Ya Zhang, Le Qin, Hao Rong, Huanhuan Jin, Ye Wang, Zongyuan Hong and Junxiu Zhang in Natural Product Communications
Footnotes
Acknowledgements
This work was supported by the National Natural Science Foundation of China (81671318), Anhui Provincial University scientific Research Plan Project (2023AH030105, 2024AH051882), Science and Technology Project of Wuhu City (2022jc311), Young and Middle-aged Research Foundation of Wannan Medical College (WK202204), National Innovation and Entrepreneurship Training Program for College Students (202210368037, 202310368051). The authors would like to express their gratitude to Dapeng Yan from Fudan University for providing resources.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Innovation and Entrepreneurship Training Program for College Students, Anhui Provincial University scientific Research Plan Project, Young and Middle-aged Research Foundation of Wannan Medical College, Science and Technology Project of Wuhu City, the National Natural Science Foundation of China, (grant number 202210368037, 202310368051, 2023AH030105, WK202204, 2022jc311, 81671318).
Ethical Approval
The study protocol was approved by the Ethics Committee of Wannan Medical College (LLSC-2022-206).
Statement of Human and Animal Rights
All procedures in this study were conducted in accordance with the the Ethics Committee of Wannan Medical College (LLSC-2022-206) approved protocols.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
