Abstract
Introduction
Non-Hodgkin's lymphoma (NHL) is the most common hematologic malignancy in the world, accounting for nearly 3% of cancer diagnoses and deaths. In 2020, the number of new cases of NHL worldwide was 544,352, ranking 13th among all new cases of malignant tumors. Simultaneously, the number of deaths from NHL worldwide reached 259,793, placing it 12th in the ranking of deaths from malignant tumors. In regions such as Europe and North America, the incidence and mortality of NHL are relatively high. Despite generally having better access to healthcare, people in these areas may also face higher levels of stress and risk factors, including environmental pollution. Conversely, in some parts of Asia and Africa, the incidence and mortality of NHL may be lower. However, the relative scarcity of medical resources and less advanced cancer screening and diagnostic technology in these regions can lead to late diagnosis and non-standard treatment, thereby increasing the mortality rate.1,2 Currently, the incidence of NHL is increasing year by year, and the age of these patient trends to become young. 3 At present, the drugs used to treat NHL primarily include chemotherapy drugs (such as cyclophosphamide, doxorubicin hydrochloride, and etoposide), targeted therapies (such as rituximab and golicitinib), and immunomodulators (such as cyclosporine and lenalidomide). 4 However, radiotherapy can lead to decreased immune function, cutaneous and mucosal reactions, and radiation pneumonia. Chemotherapy, on the other hand, can cause bone marrow suppression, immunosuppression, nausea, and vomiting. 5 Therefore, it is necessary to find safe drugs with small side effects to treat NHL.
With a long history, Traditional Chinese Medicine (TCM) is one of the most enduring and continuously practiced herbal medicine systems. More than two millennia have passed since the documented application of TCM in cancer treatment in Chinese medical texts and publications. TCM has attracted significant attention recently in the field of lymphoma treatment for its crucial role in cancer prevention and treatment. 6 Euscaphic acid (EA) is a natural triterpenoid saponin, which was first found in the peel of Eriobotrya japonica. 7 The EA exhibits anti-HIV, 8 anti-inflammatory9,10 and anti-tumor effects,11-13 while demonstrating almost no cytotoxicity on non-tumor cells. 10 Our previous research found for the first time that EA has anti-tumor effects in NHL cells. 14 However, the specific mechanism remains unclear. Network pharmacology can analyze the chemical structure and action targets of drugs, and carry out systematic network analysis of the relevant targets of drugs and diseases, so as to predict the key targets and molecular mechanisms of drug treatment of diseases.15,16
In this study, we analyzed the targets and underlying mechanisms of EA in the treatment of NHL through network pharmacology, molecular docking and cell experiment (Figure 1). This study will provide a theoretical basis for the clinical research and application of EA in the treatment of NHL.

The approach of this manuscript and the flow chart.
Results
Network Pharmacology Analysis
Acquisition of EA and NHL Related Targets
We obtained 121 EA action targets and 2341 targets of NHL from the mentioned databases above. Venny 2.1.0 was used to map the intersection of EA-related targets and NHL-related targets, and 51 EA anti-NHL targets were obtained (Figure 2).

Common target analysis of EA and NHL.
Identified 8 Target Genes for EA Treatment of NHL
The PPI network shows that 51 proteins (nodes) and 208 interactions (edges) may interact between EA and NHL (Figure 3). Figure 3A illustrates the PPI network diagram sourced from the String database. In this network, nodes denote target proteins, while lines between nodes indicate interactions, with more lines signifying stronger correlations. To visually represent the relationships and significance among targets, PPI data were imported into CytoScape in TSV format, producing the concentric PPI network diagram shown in Figure 3B. The node color intensity and radius reflect the degree, with darker colors and larger radii indicating higher degrees. Additionally, the MCC and degree algorithms were employed using Cytohubba in CytoScape to identify the top 10 core targets, and the results were largely consistent. (Figure 4A, 4B).

PPI network diagram. A: PPI network diagram by String database; B: PPI network diagram by CytoScape software.

Core target genes screened. A: The top 10 hub gene networks of the target proteins in the Cytohubba pluginby MCC algorithm; B: The PPI network bar plot; C: MCODE cluster analysis was performed on 51 targets; D: Venn diagram of core target genes.
Furthermore, to verify the accuracy of the identified core target genes and analyze the clustering of key targets, the authors employed the MCODE plug-in for functional module analysis on the PPI network. This analysis revealed three clusters, with Cluster1 achieving the highest score and aligning closely with previous findings, indicating that Cluster1 represents the core gene cluster (Figure 4C). The intersection of these results with prior findings confirms the core target of our prediction. (Figure 4D): tumor necrosis factor (TNF), peroxisome proliferator-activated receptor gamma (PPARG), matrix metalloproteinase-9 (MMP9), heat shock protein 90 alpha family class A member 1 (HSP90AA1), prostaglandin-endoperoxide synthase 2 (PTGS2), insulin like growth factor 1 receptor (IGF1R), androgen receptor (AR), estrogen receptor beta 2 (ESR2).
GO and KEGG Pathway Enrichment Analysis
GO and KEGG enrichment analysis of 51 potential target proteins reveal the potential mechanism of EA treatment of NHL. A total of 181 items were obtained from GO enrichment analysis with P < 0.05, including 119 biological processes (BP), 27 cellular components (CC) and 35 molecular functions (MF). The top 10 GO items are shown in Figure 5A. In BP category, target proteins are mainly concentrated in cellular response to UV-A, extracellular matrix organization, positive regulation of protein kinase B signaling, positive regulation of apoptosis process, etc. In CC category, target proteins are mainly concentrated in cytoplasm, nucleus, cytosol, nucleoplasm, etc. In MF category, target proteins are mainly concentrated in zinc ion binding, steroid binding, endopeptidase activity, etc.
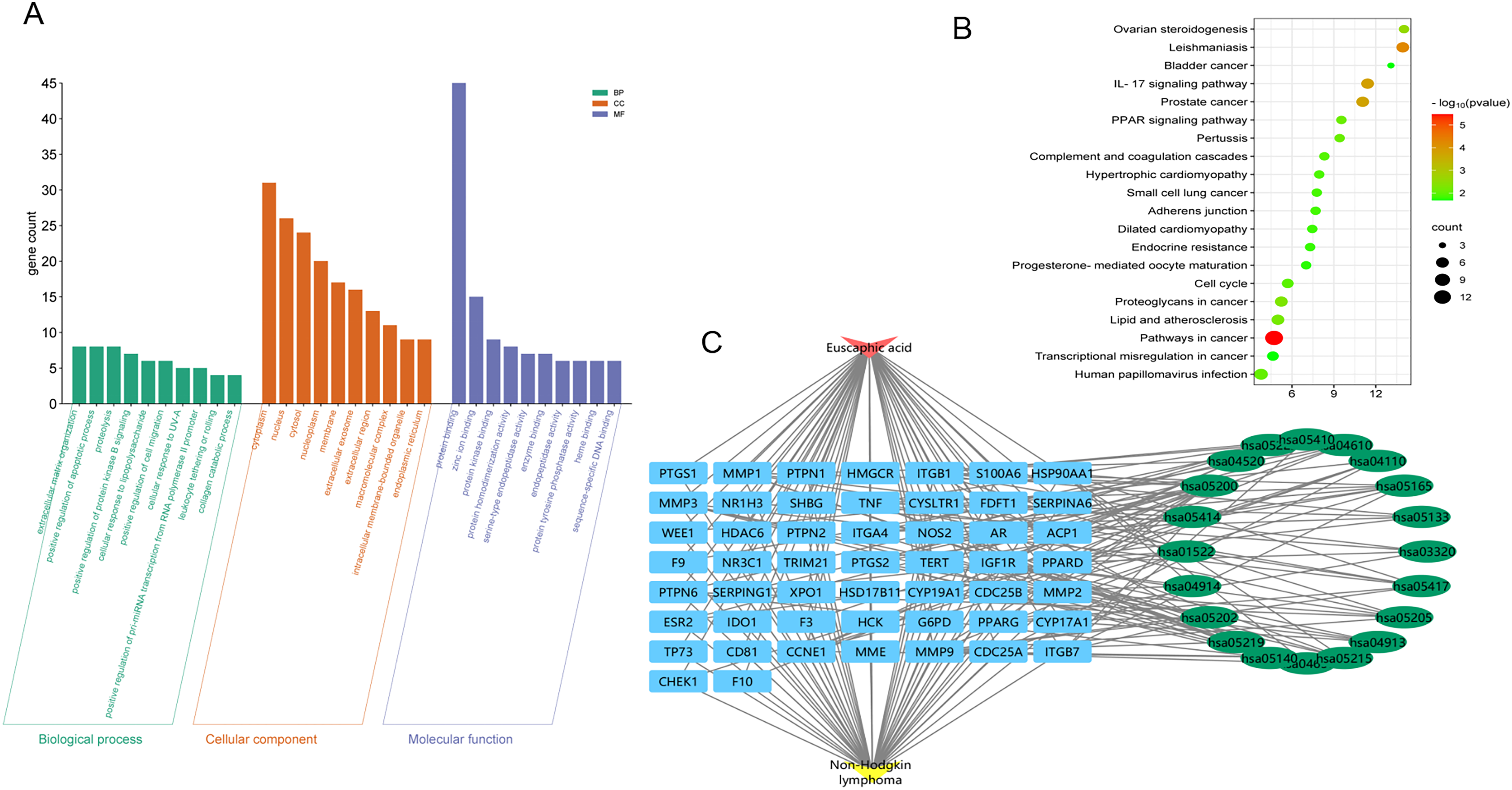
Enrichment analysis. A: GO enrichment maps of common targets of EA and NHL, including biological processes, cell components, and molecular functions; B: KEGG pathway analysis of common target of EA and NHL; C: Drug-target-pathway-disease network.
A total of 34 KEGG enrichment projects were selected with P < 0.05. The top 20 projects are shown in Figure 5B, mainly including pathways in cancer, Leishmaniasis, IL-17 signaling pathway, PPAR signaling pathway, etc. To understand the relationship among drug, target, pathway and disease, the top 20 pathways of KEGG items were integrated to construct drug-target-pathway-disease network (Figure 5C). EA is closely related to NHL through multiple targets and pathways. This interaction reflects the “multi-target point and multi-pathway” characteristics of EA treatment in TCM, which is also suitable for the holistic treatment strategy of TCM.
Molecular Docking Verifcation
To verify drug-target interactions, we performed molecular docking on 8 screened targets. CB-DOCK2 was used to analyze the docking potential of the 8 screened targets. The lower the binding energy score, the more stable the binding of the ligand-target complex, indicating a stronger interaction between the ligand and protein. Among them, 6 targets (TNF, PPARG, HSP90AA1, PTGS2, IGF1R, and AR) have binding energy scores less than −7 kcal/mol, indicating that EA had good binding activity with the screened targets. The binding pose, 3D and 2D interaction were shown in Figure 6. Docking Score, cavity volume, docking center and docking size between selected targets and EA were provided in Table 1. Ranked of binding scores, the molecular docking results of top 4 were visualized.

Molecular docking showed strong interaction between the core targets and the EA. The 3D and 2D binding poses of EA with (A) TNF, (B) PTGS2, (C) PPARG, (D) IGF1R were presented.
Docking Score, Cavity Volume, Docking Center, and Docking Size Between Selected Targets.

Molecular Dynamic Simulation
To confirm the ligand–receptor binding stability, we analyzed the mobility of the receptor-ligand complex using the root-mean-square deviation (RMSD). The RMSD curve provides insight into fluctuations in protein conformation. Supplementary Figure 7 displays the RMSD of the IGF1R-EA complex. The entire structure of this complex was evaluated through RMSD analysis. The RMSD curve for the IGF1R-EA complex remains relatively stable between 50 and 300 ns.
Experimental Validation
EA Inhibited the Proliferation of Non-Hodgkin's Lymphoma Cells
To investigate the effect of EA on NHL cell proliferation, RAJI and OCI-LY3 cells were treated with different concentrations of EA for 48 h. The CCK8 experiment showed that the proliferation of RAJI and OCI-LY3 cells could be inhibited by EA in a dose-dependent manner (Figure 7, Supplemental Figure 6). EA inhibited the growth of NHL cells.

Effect of EA on survival rate of RAJI cells in vitro, representative CCK-8 results of EA on RAJI cell viability.
EA Induced the Apoptosis of Non-Hodgkin Lymphoma Cells
RAJI and OCI-LY3 cells were incubated with different concentrations of EA for 48 h. The cells apoptosis were detected using flow cytometry. EA induced apoptosis of cells in a dose-dependent manner (Figure 8, Supplemental Figure 8).

EA induced apoptosis of RAJI cells in a dose-dependent manner. Representative cellular results of EA on apoptosis of RAJI cells.
Effect of EA on Target Gene IGF1R
Since EA has a high molecular docking score with NHL, we used QPCR to detect the effect of EA on IGF1R mRNA expression in RAJI cells. RAJI cells were incubated with different concentrations of EA (0 μM,50μM, 100μM) for 48 h, and the mRNA expression of IGF1R in RAJI and OCI-LY3 cells were detected by QPCR. The results showed that IGF1R induced by EA decreased significantly compared with the control group (Figure 9, Supplemental Figure 9).

EA decreased IGF1R expression in RAJI.
Discussion
Our previous research demonstrated that EA can induces the apoptosis of RAJI cells by downregulating the expression of JAK2/STAT3 and c-Myc protein. 14 However, whether EA has the potential to treat NHL and the possible target mechanism are not clear. In this study, we investigated the molecular mechanism of EA in the treatment of NHL based on network pharmacology. EA and NHL share 51 intersection targets. The results of PPI network analysis predicted 8 core targets that played an important role in the treatment of EA in NHL probably. Among these, 6 core genes exhibited strong binding efficiency with EA. Meanwhile, this was confirmed by molecular dynamics simulations, which demonstrated that the structure of the IGF1R and EA complex was stable and compact. Therefore, IGF1R may be a potential target for EA's anti-NHL effect. GO and KEGG enrichment analysis showed that EA acted on NHL through multiple pathways, especially pathways in cancer, inducing apoptosis and inhibiting cell cycle. Whats more, TNF, PTGS2, PPARG, IGF1R, AR, HSP90AA1, MMP9 and ESR2 are enriched in these pathways.
Studies have shown that serum levels of TNF and its receptor are elevated in patients with NHL, and higher levels of TNF are associated with a poorer prognosis for NHL. 17 PTGS2, also known as COX-2, is a key enzyme in prostaglandin biosynthesis. PTGS2 is one of the inducers of lymphangiogenesis in NHL, and its expression level is negatively correlated with prognosis.18,19 PPARγ, encoded by PPARG gene, is a member of peroxisome proliferator-activated receptors. Activation of PPARγ in tumor cell growth can inhibit tumor angiogenesis, proliferation and differentiation of cancer cells, and induce apoptosis.20,21 In NHL, activation of PPARγ was also found to have a dose-dependent effect against cell proliferation. 22 Generally, insulinlike growth factors (IGFs) as potent mitogens are involved in the invasion and metastasis of many tumor cells, and overexpression of these factors and their receptors leads to malignancies like NHL.23,24 Studies have found that knock down IGF1R genes inhibited the proliferation of RAJI. 24 The proportion of male patients is significantly higher than female patients in NHL. This gender biased incidence reflect that AR signaling with gender differences may play a role in the pathogenesis of NHL. 25 HSP90α, encoded by HSP90AA1, is a kind of cancer -activation factor,with the role of anti-apoptosis, which is considered to be necessary to the malignant transformation and progression. 26 Studies have found that the level of HSP90α in DLBCL patients is statistically correlated with prognostic indicators such as Lugano staging, IPI score and the number of extranodal lesions, suggesting that HSP90α can be used as a plasma tumor marker for efficacy judgment and prognosis evaluation of DLBCL. 27 High levels of MMP9 play a crucial role in the aggressiveness of NHL by promoting ECM degradation, metastasis establishment and tissue destruction, and angiogenesis. 28 Studies have found that ESR2 is underexpressed in various tumor types such as prostate cancer, breast cancer and thyroid cancer, while overexpression of ESR2 gene can inhibit the proliferation of NHL cells, suggesting that ESR2 may become an important target for related treatment of NHL. 29
Consistent with previous research, the targets above are associated with the process of disease occurrence and progress of the NHL, suggesting that EA may treat non-Hodgkin lymphoma by targeting these targets. Most of the core targets are involved in the regulation of apoptosis. Apoptosis is one of the important keywords in the study of TCM treatment of lymphoma. 6 We further studied the effect of EA on apoptosis of RAJI. The results showed that EA could significantly promote apoptosis of RAJI cells, which was consistent with the prediction of network pharmacology. In addition, we found that EA inhibited the expression of IGF1R targets. It is reported that IGF1R can stimulate cell proliferation,survival, invasion antiapoptotic, playing a prominent role in the development and transformation of lymphoma. 30 The results indicate that EA may target IGF1R to inhibit the proliferation and induce apoptosis in NHL cells.
However, there are some limitations to this study. First, we only verified the identified targets through computer simulation and in vitro experiments, and did not conduct in vivo experiments. Second, we did not study the effect of EA on the expression of IGF1R protein in NHL, but only the changes in the mRNA transcription level of IGF1R. Third, the results of this study did not clarify the changes of downstream protein and mRNA levels of IGF1R regulated by EA in NHL. Therefore, further in vitro and in vivo experiments are needed to confirm the mechanism and effect of EA anti-NHL, and provide scientific research theories for the clinical use of EA.
Conclusion
EA has the potential for clinical to treat NHL. TNF, PTGS2, PPARG, IGF1R, AR, and HSP90AA1 may play crucial roles in the anti-NHL of EA. Among then, EA may induce NHL cells apoptosis by acting on IGF1R.
Materials and Methods
Network Pharmacology Analysis
EA and NHL-Related Target Acquisition
The 2D structure and Canonical SMILES ID number of EA were obtained by PubChem database (https://pubchem.ncbi.nlm.nih.gov/). Subsequently, the Canonical SMILES ID number was imported into the Swiss Target Prediction database (https://SwissTargetPrediction.cn) to screen the targets of EA. The downloaded 2D structure of EA was imported into PharmMapper database (https://lilab-ecust.cn/pharmmapper/index.html) to predict the target proteins of EA. The target Uniplot number of the drug target was converted to the official gene symbol through the UniProt database (http://www.uniprot.org). NHL targets were searched in Genecard database (https://www.GeneCards.org) and OMIM database (https://www.OMIM.org), with the keywords “Non-Hodgkin lymphoma” and “Homo sapiens”, respectively. Then the targets were de-duplicated and merged to screen the targets of NHL. Finally, we obtained the common targets which were EA anti-NHL targets by Venn software.
PPI Network Construct and Identification of the Key Target Genes
The selected common targets were imported into String database (https://cn.string-db.org), the species was selected as “homo sapiens”, the minimum interaction threshold was set to medium confidence of 0.4, and the Protein-protein interactions (PPI) network. Subsequently, the results of the PPI network analysis were imported into Cytoscape 3.7.0 software in the TSV format, and the Network Analyzer plug-in was used for topology analysis to predict the core targets.
GO and KEGG Pathway Enrichment Analysis
These core targets were subjected to enrichment analysis of Gene Ontology (GO) and the Kyoto Encyclopedia of Genes and Genomes (KEGG) using the DAVID database (https://DAVID.ncifcrf.gov). GO consists of three ontologies: process (BP), cellular component (CC), and molecular function (MF). Then the bar chart of GO and the bubble map of KEGG signal pathway were drawn by the online bioinformatics tools (http://www.bioinformatics.com.cn). Finally, pathway results were integrated to construct drug-target-pathway-disease network.
Molecular Docking
In order to verify the reliability of the interaction between core targets and compounds, this study selected the core targets and compounds for molecular docking. Downloading the crystal structure of the core targets from the PDB database (http://www.rcsb.org/pdb) and 3D structure of EA from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/), we use CB-DOCK2 server (https://cadd.labshare.cn/cb-dock2/) for molecular docking to determine the binding affinity between EA and receptor protein. The affinity value measures the confidence of interaction between the intersection target and the compound. By default, the CB-DOCK2 server verifies the subject protein structure of missing hydrogen atoms and side chain atoms. Similarly, the submitted ligands are improved by adding hydrogen atoms and partial charges. In addition, by estimating the center size and size of the top N (default N¼5) cavities, the binding site of the selected protein is predicted based on the curvature based on cavity detection method. The PDB file of the receptor-ligand complex of the docking combination with the best Vina score was imported into the software Discovery Studio 2019 to visualize the molecular docking results.
Molecular Dynamics Simulation
Molecular dynamics (MD) simulations were conducted utilizing Groningen Machine for Chemical Simulations (GROMACS-2021) on the complexes derived from molecular docking procedures. The protein force field parameters were generated using the July 2022 release of Chemistry at Harvard Macromolecular Mechanics (CHARMM36), 31 whereas the ligand force field parameters were obtained from CGenFF (accessible at https://cgenff.com). The system was enclosed within a dodecahedral box that featured periodic boundary conditions, maintaining a distance of 10 Å from the protein's edge. The system was subsequently neutralized by adding Na+ ions to the solvated box using the three-point transferable water model (TIP3P). To mimic physiological conditions, an extra concentration of 0.15 M of Na+ and Cl− ions was introduced. A structural refinement process was implemented using the steepest descent algorithm, aimed at minimizing the system's energy until the maximum force (Fmax) did not exceed 1000 kJ mol-1 nm-1. This was followed by a 100 ps equilibration phase in both the NVT and NPT ensembles, employing the V-rescale thermostat and Berendsen barostat for temperature and pressure control, respectively. The Particle Mesh Ewald technique was adopted to compute long-range electrostatic interactions, with the van der Waals and electrostatic interaction cut-offs established at 1.2 nm. During the production run, the temperature and pressure were held constant at 310 K and 1 bar. 32 The Berendsen barostat regulated pressure with a time constant of 1 ps and a compressibility of 4.5 × 105 bar−1, while the V-rescale thermostat utilized a time constant of 0.1 ps. A trajectory snapshot was captured every 100 ps throughout entire simulation run that lasted 300 ns. Following this, the stability of the generated trajectories was evaluated using the root-mean-square deviation (RMSD) metric.
Experimental Validation
Materials
Euscaphic acid (No.Y-176) was purchased from Herbpurify, Chengdu. Annexin V-FITC/PI kit (No.70110100) was obtained from Biosharp, Shanghai. CCK8 was purchased from Meilun, Dalian (No. MA0218-2-Sep-08F).
Cell Culture
RAJI and OCI-LY3 cells were purchased from Procell CL-0189. The cells were cultured in 1640 culture medium with 10% fetal bovine serum, 100μg/mL penicillin and 100μg/mL streptomycin, placed at 37 °C in an incubator with 5% CO2.
Cell Proliferation Assays
Cell proliferation was detected with cell Counting Kit-8 according to instruction. 2 × 104 cells/well were inoculated in 96-well plates, and then final concentrations of 0, 12.5, 25, 50 and 100μM of EA were added to incubate for 48 h. At the certain time, add 10μl CCK-8 solution to each well and wait for another 2 h. The absorbance was measured at 450 nm with the Infinite 200PRO (Science&Technology, Männedorf, Swiss).
Apoptosis Assay
Cells at logarithmic growth stage were taken, centrifuged, collected, counted, and inoculated with 5 × 105 cells per well into a 6-well plate. At the same time, EA liquid with final concentrations of 0, 50 and 100μM were added, and the cells were centrifuged and collected after co-culture in a carbon dioxide incubator for 48 h. The cells were washed twice with pre-cooled PBS, and the cells were re-suspended with 1 × binding buffer. Appropriate amounts of cells were added with 5μL Annexin V-FITC, mixed, incubated at room temperature for 15 min in dark light, then gently mixed with 10μL PI dyeing solution and incubated for 5 min in dark light. Finally, 400μL 1 × binding buffer was added and gently mixed, and apoptosis detection was completed within 1 h.
RNA Extraction and qRT-PCR
Total RNA was extracted with Trizol reagent (TaKaRa, Tokyo, Japan), according to the manufacturer's instructions. RNA concentration was measured via Infinite 200PRO (Science&Technology, Männedorf, Swiss). Subsequently, cDNA synthesis and quantitative PCR (qPCR) were performed by a reverse transcriptase kit (Vazyme) and an SYBR Premix Ex Taq™ II kit (Vazyme), respectively, each according to the manufacturer's instructions. All of the primers used were synthesized by SangonBiotech (China, Guang zhou). Relative expression levels of IGF1R was quantified according to the 2−ΔΔCt.
Statistical Analysis
The data were analyzed by GraphPad Prism 9.0.2 (San Diego, USA) and expressed as mean ± SD. Statistical analysis was carried out by one-way ANOVA followed by the Tukey's multiple comparisons test. *p < 0.05, **p < 0.01, and ***p < 0.001 were considered statistically significant differences.
Supplemental Material
sj-pptx-1-npx-10.1177_1934578X251322669 - Supplemental material for Exploration of Euscaphic Acid on Non-Hodgkin Lymphoma Based on Network Pharmacology, Molecular Docking and Cell Experiment
Supplemental material, sj-pptx-1-npx-10.1177_1934578X251322669 for Exploration of Euscaphic Acid on Non-Hodgkin Lymphoma Based on Network Pharmacology, Molecular Docking and Cell Experiment by Guangru Li, Chunyuan Liang, Yan Liu, Yueyuan Pan, Zhiyuan Li, Guocai Wu and Ruiting Wen in Natural Product Communications
Supplemental Material
sj-tif-2-npx-10.1177_1934578X251322669 - Supplemental material for Exploration of Euscaphic Acid on Non-Hodgkin Lymphoma Based on Network Pharmacology, Molecular Docking and Cell Experiment
Supplemental material, sj-tif-2-npx-10.1177_1934578X251322669 for Exploration of Euscaphic Acid on Non-Hodgkin Lymphoma Based on Network Pharmacology, Molecular Docking and Cell Experiment by Guangru Li, Chunyuan Liang, Yan Liu, Yueyuan Pan, Zhiyuan Li, Guocai Wu and Ruiting Wen in Natural Product Communications
Supplemental Material
sj-tif-3-npx-10.1177_1934578X251322669 - Supplemental material for Exploration of Euscaphic Acid on Non-Hodgkin Lymphoma Based on Network Pharmacology, Molecular Docking and Cell Experiment
Supplemental material, sj-tif-3-npx-10.1177_1934578X251322669 for Exploration of Euscaphic Acid on Non-Hodgkin Lymphoma Based on Network Pharmacology, Molecular Docking and Cell Experiment by Guangru Li, Chunyuan Liang, Yan Liu, Yueyuan Pan, Zhiyuan Li, Guocai Wu and Ruiting Wen in Natural Product Communications
Supplemental Material
sj-tif-4-npx-10.1177_1934578X251322669 - Supplemental material for Exploration of Euscaphic Acid on Non-Hodgkin Lymphoma Based on Network Pharmacology, Molecular Docking and Cell Experiment
Supplemental material, sj-tif-4-npx-10.1177_1934578X251322669 for Exploration of Euscaphic Acid on Non-Hodgkin Lymphoma Based on Network Pharmacology, Molecular Docking and Cell Experiment by Guangru Li, Chunyuan Liang, Yan Liu, Yueyuan Pan, Zhiyuan Li, Guocai Wu and Ruiting Wen in Natural Product Communications
Supplemental Material
sj-tif-5-npx-10.1177_1934578X251322669 - Supplemental material for Exploration of Euscaphic Acid on Non-Hodgkin Lymphoma Based on Network Pharmacology, Molecular Docking and Cell Experiment
Supplemental material, sj-tif-5-npx-10.1177_1934578X251322669 for Exploration of Euscaphic Acid on Non-Hodgkin Lymphoma Based on Network Pharmacology, Molecular Docking and Cell Experiment by Guangru Li, Chunyuan Liang, Yan Liu, Yueyuan Pan, Zhiyuan Li, Guocai Wu and Ruiting Wen in Natural Product Communications
Footnotes
Acknowledgements
The first author would like to thank Dr Wen Ruiting, Ms Liang Chunyuan and Ms Liu Yan for their continued support. Without their wisdom, talent and kindness, progress here would not be possible.
Data Availability
For reasonable requirements, the data related to this study can be requested from the corresponding author.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Ethical Approval is not applicable for this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Medical Scientific Research Foundation of Guangdong Province (No. A2024784), National Natural Science Foundation of China (82200238), the Natural Science Foundation of Guangdong Province (2021A1515010708, and 2023A1515010594), Science and Technology Plan Project of Zhanjiang city (2020A0611, 2021A05153, 2021A05137, and 2021A05150), China zhongguancun Precision Medicine science and technology foundation (No. ZGC-YXKY-68 and No. ZGC-YXKY-67).
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Trial Registration
Not applicable, because this article does not contain any clinical trials.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
