Abstract
Background
This study used a holistic network pharmacological approach to clarify the mechanism of Compound Zeqi Granule (CZG) in treating psoriasis vulgaris (PV).
Methods
The potential bioactive compounds and corresponding targets of CZG were collected and searched via the TCMSP database. Disease targets of PV were obtained from the GeneCards database. The targets of CZG and PV were standardized by the UniProt database. GO and KEGG enrichment analyses were performed using the Metascape database. Then, molecular docking verification was conducted using the CB-Dock database. Animal experiments were conducted to validate the effects of CZG on PV.
Results
This approach identified 99 bioactive compounds, 579 potential drug targets, 822 PV-related targets, and 108 potential target proteins. The biological processes were primarily related to regulation of defense response, regulation of inflammatory responses, and cellular response to lipid. The targets of CZG for treating PV were significantly associated with 20 pathways including JAK-STAT signaling pathway. Animal experiments confirmed that CZG exerted its therapeutic effects on PV by promoting the expression of IL-10 and JUN, and inhibiting the activation of STAT1 and STAT3.
Conclusion
The present study provides evidence supporting the efficacy of CZG for the treatment of PV through a multi-compound, multi-target and multi-pathway approach and lays a theoretical foundation for further experimental research in this field.
Keywords
Introduction
Psoriasis vulgaris (PV) is a common chronic inflammatory skin disease that affects nearly 60 million people worldwide. 1 It is characterized by the presence of recurrent, red, scaly plaques and papules, and typically observed on the elbow, knee and scalp. 2 The development of PV is influenced by multi-gene inheritance and can be stimulated by multiple environmental factors.3,4 Despite significant progress in understanding the underlying pathophysiology, the specific mechanisms by which these factors trigger and perpetuate PV remain elusive, and specific targeted therapies are lacking. Thus, it is of great significance to actively explore the pathogenesis of PV and formulate effective drugs with clear targets.
As an effective and safe Traditional Chinese Medicine (TCM) prescription for the clinical treatment of PV, Compound Zeqi Granule (CZG) have clearing heat and detoxifying effects, thereby promoting blood circulation and removing blood stasis. It is composed of seven kinds of herbs, including Herba Euphorbia Helioscopia (HEH), Herba Hedyotis Diffusae (HHD), Folium Clerodendri Cyrtophylli (FCC), Radix Scutellariae (RS), Radix Gentianae (RG), Herba Scutellariae Barbatae (HSB) and Rhizoma Dryopteris Crassirhizomae (RDC). Some clinical studies have confirmed the efficacy of CZG in improving clinical symptoms, increasing the dermatology life quality index (DLQI) and decreasing the psoriasis area and severity index (PASI) of PV patients, without causing adverse reactions.5,6 However, there is still a lack of relevant investigations on the molecular mechanism of CZG in treating PV.
In this investigation, we utilized the pharmacological components of CZG to pinpoint its target and then combined it with PV target in order to identify a potential treatment target for PV. By conducting enrichment analysis of functions and pathways, we were able to determine the signal pathways associated with treating PV and validate them through experiments. This offers fresh perspectives and guidance for comprehensively and systematically explaining the mechanism of action of CZG in the treatment of PV. Figure 1 shows the detailed protocol of this present study.

The detailed schema of this study.
Materials and Methods
Collection of Bioactive Compounds and Drug Targets for CZG
Natural compounds possess distinct pharmacokinetic properties that determine their efficacy based on their absorption, distribution, metabolism and excretion (ADME) in the human body, which includes oral bioavailability (OB), drug-likeness (DL), blood-brain-barrier (BBB), drug-half-life (HL), internal affinity performance (Caco-2) and etc The TCMSP database (https://tcmspw.com/tcmsp.php), a comprehensive platform for TCM analysis, comprises drug compounds, pharmacokinetic data of compounds, targets, related diseases and the network relationship among drugs, targets and diseases. In this study, we used the TCMSP database to screen for bioactive ingredients and their drug targets in CZG, with OB ≥ 30% and DL ≥ 0.18 as the screening criteria. Then, all the targets obtained were converted into standard gene names and confirmed to be of human origin through the UniProt KB 7 database (https://www.uniprot.org/).
Construction of “Drug-Compound-Target” Network for CZG
The bioactive compounds of CZG and drug targets obtained from the TCMSP database were imported into the Cytoscape v3.7.2 software (https://www.cytoscape.org/) with the drugs of CZG to build a “Drug-Compound-Target” (D-T-C) network to determine the relationship among the drugs, bioactive ingredients and drug targets. The nodes represent the imported terms, and the edges represent the interaction between terms.
Acquisition of Disease Targets for PV
The keyword “psoriasis vulgaris” was queried in the GeneCards database (https://www.genecards.org/) to retrieve the disease targets, which were then converted to standard gene names and confirmed to be of human origin using the UniProt KB database. After data integration, null and duplicate values were removed to obtain the disease targets of PV.
Collection of Intersection Targets Related to CZG and PV
The drug targets and disease targets obtained from 1.1 and 1.3, respectively, were imported into the Draw Venn Diagram online platform (http://bioinformatics.psb.ugent.be/webtools/Venn/) to identify the “drug-disease” intersecting gene targets, namely the potential targets of CZG in the treatment of PV, and to construct Venn diagrams to visualize the intersection targets.
Establishment and Analysis of “Compound-Target” Network for Bioactive Compounds and Intersecting Targets
According to the intersection targets of drugs and disease, the effective bioactive compounds of CZG in the treatment of PV were reversely screened to identify the bioactive compounds associated with the intersection targets. The effective bioactive compounds were then imported into the Cytoscape 3.7.2 software, along with the intersection targets, to establish the “Compound-Target” (C-T) network for data visualization. Using the “Network Analyzer” plugin, the top 10 effective bioactive ingredients were screened based on their degree parameter.
Construction and Topology of Protein-Protein Interaction (PPI) Network
Based on the human multiple proteins analysis platform of the STRING database (https://cn.string-db.org/), a protein-protein interaction (PPI) network of intersection targets was generated, with a confidence level set at 0.950 for the minimum required interaction score, and the disconnected nodes in the network were hidden. The analysis results of the PPI network in “tsv” format were imported into the Cytoscape 3.7.2 software, and the “Network Analyzer” plugin was used to analyze the topological properties using parameters medians of degree centrality (DC), betweenness centrality (BC) and closeness centrality (CC). A higher parameter indicated a more important node. After two rounds of screening, the top 10 core intersection targets were obtained.
Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) Enrichment Analysis
GO function (including molecular function, MF; biological process, BP; and cellular component, CC) and KEGG pathway enrichment analysis of intersection targets were conducted using the Metascape database (https://metascape.org/) with a P value <.01, Min Overlap = 3, and Min Enrichment > 1.5 as the cut-off criteria. The results were visualized using RStudio.
Molecular Docking Validation
The CB-Dock website (http://clab.labshare.cn/cb-dock/php/) is based on the detection of curvature cavities, thereby enabling the automatic and precise identification of the binding site, calculation center and size, and the customized docking box dimensions, which seamlessly integrates with AutoDock Vina for molecular docking. This approach has a high hit rate and accuracy for blind docking. Based on the docking mode of AutoDock Vina, the binding tightness was evaluated by calculating the binding energy of the ligand and receptor, with a lower binding energy indicating a higher binding affinity. It is generally accepted that a binding energy <−4.25 kcal/mol represents a certain degree of binding tightness, while <−5.5 kcal/mol denotes a favorable binding combination.
The results of the above steps were analyzed to further confirm whether the gene targets STAT1 and IL-10 could play an important role in the treatment of PV with CZG. We also used the RCSB PDB database (https://www1.rcsb.org/) to retrieve and download the “pdb” formats of the targets and used the PubChem 8 database (https://pubchem.ncbi.nlm.nih.gov/) to obtain the “sdf” structures of the bioactive compounds that were screened from the top 10 bioactive ingredients with the highest degree value in the “C-T” network and corresponding to the targets. Then, we imported the “pdb” formats and “sdf” structures into the CB-Dock online platform for molecular docking and visualization. Citing Methotrexate was used as the positive control drug.
Reagents, Experimental Drugs and Animals
White vaseline (Jiangmen Hengjian Pharmaceutical Co., Ltd, Lot: F20110023); 5% Imiquimod cream (Sichuan Med-Shine Pharmaceutical Co., Ltd, Lot: H20030128); RIPA lysis buffer (Beyotime, Lot: 051418180626); STAT1 antibody (Wanleibio, Lot: WL02273); IL-10 antibody (Wanleibio, Lot: WL03088); Tubulin antibody (Proteintech; Lot: 10094-1-AP); Goat anti-rabbit IgG (Proteintech, Lot: B900210); SuperSignal™ West Pico PLUS Chemiluminescent Substrate (Thermo, Lot: 34577); Trizol (Life, Lot: 350508); Novostart SYBR qPCR SuperMix Plus (Novoprotein, Lot: 05229413); PrimeScriptTM RT reagent Kit with gDNA Eraser (TaKaRa, Lot: AL21115A); Electrophoresis tank, Electrophoresis apparatus (Tanon, Type: VE-180, EPS300 respectively); Electronic balance (Shanghai Jinghai Instrument Co., Ltd, Type: FA2004N); Chemiluminescence gel imager (Bio-Rad, Type: ChemiDoc XRS+); Real-time fluorescent quantitative PCR instrument (Thermo, Type: PikoReal96). CZG (batch number: Z20210055000) was purchased from the First Affiliated Hospital of Anhui University of Chinese Medicine (Hefei, China). Methotrexate tablets (MTX) were bought from Shanghai Sine Pharmaceutical Laboratories Co., Ltd (Shanghai, China). Thirty-two SPF male BALB/c mice (certificate number: SCXK(沪) 2018-0006; 6-8 weeks old, weighing 20-22 g) were purchased from Hangzhou Ziyuan Laboratory Animal Technology Co., Ltd (Certificate No. SCXK (Zhejiang) 2019-0004).
Preparation of CZG Decoction
First, we added 10 times amount of water to the Chinese herbal medicines mentioned above to soak for 30 min and then boiled to decoct for 1.5 h. The herbal juice was obtained via filtration. Second, 8 times amount of water was added to herbal medicines and repeated the above steps. The two filtrates were combined. After sterilization and cooling, the extracts were stored at 4 °C for succeeding experiments.
Experimental Protocol
This experiment was executed in compliance with the NIH Guide for the Care and Use of Laboratory Animals, abided by The ARRIVE guidelines (Animal Research: Reporting of In Vivo Experiments) and approved by the Laboratory Animal Ethics Committee of Anhui University of Chinese Medicine (approval ID: AHUCM-mouse-2021113). All the mice were raised in the Laboratory Animal Room of the First Affiliated Hospital of Anhui University of Chinese Medicine. The temperature was maintained at 24 ± 1 °C with a regular light/dark cycle. Water and food supply was made readily available to mice during the course of the study. According to the equivalent dose conversion of the body surface area between mice and adults, CZG and MTX were mixed with normal saline as a suspension at a final concentration of 2.925 g/ml and 0.1 mg/ml, respectively. BALB/c mice were fed adaptively for three days before being divided into different random groups, including the control group, model group, treatment group and positive drug group. Each group consisted of 8 mice (n = 8). As previously described,9–11 the back of mice in each group was shaved with a mouse shaver to expose a skin area of approximately 2 cm × 3 cm, and residual hairs were removed with 10% Na2S. On the first day, the appearance of abnormal reactions such as redness, swelling and ulceration on the surface areas was assessed. On subsequent days, white vaseline was uniformly applied on the back hair removal area of the control group, while the other 3 groups were evenly smeared with 5% imiquimod ointment of dose 62.5 mg/mouse for 7 days. Concurrently, mice in the treatment group received CZG (29.25 g/kg/d), mice in the positive drug group received MTX (1 mg/kg/d), and an equal amount of normal saline was administered to mice in the control and model groups by gavage of 10 ml/kg/d for 7 days.
Twenty-four hours after the last administration, the mice from each group were sacrificed via cervical dislocation, and the skin lesions in the back modeling area were resected for subsequent hematoxylin & eosin (HE) staining, western blotting and Real-Time Quantitative PCR (qRT-PCR) detection.
Evaluation of Skin Lesion Index
After modeling, the changes in skin lesions in the back modeling area were recorded daily. The Psoriasis Area and Severity Index (PASI) score 12 was used to evaluate the skin lesion index of psoriasis modeling mice. It mainly comprises the following three indicators: erythema, scales and thickening degree. The score of each indicator was 0, 1, 2, 3 and 4 according to the severity of the skin lesions in the back modeling area. The total score was obtained by adding the score of the three indicators. The PASI scoring criteria was as follows: no = 0; mild = 1; moderate = 2; severe = 3; extremely severe = 4; total score = erythema + scales + thickening degree (0-12).
HE Staining
Fresh skin lesions were fixed with 4% paraformaldehyde solution, dehydrated, embedded, sliced and dewaxed with a dehydrator. HE staining was performed, followed by staining in eosin solution for 3 min and sealing. The pathological changes of the skin lesions were observed under a microscope, and the thickness of the skin lesions was measured using the CaseViewer software.
Western Blotting
The protein expressions of STAT1 and IL-10 of skin tissues in the back modeling area were determined via WB. The proteins were extricated on ice, and approximately 0.1 g modeling skin tissue of mice was dissolved in 0.6 ml RIPA lysis buffer, followed by centrifugation at 12,000 rpm for 20 min at 4°C to obtain the protein sample. Protein concentration was quantified using the BCA method. Protein samples of the same amount were subjected to SDS-PAGE before they were transferred onto PVDF membranes. After blocking at room temperature with 5% skimmed milk for 2 h, the membranes were incubated with STAT1 (1:500), IL-10 (1:500) and Tubulin (1:5000) antibodies overnight at 4 °C. The following day, the membranes were washed thrice with TBST buffer, 10 min each, and incubated with a secondary antibody (1:20000) at room temperature for 2 h. After washing the membranes with the above method, SuperSignal™ West Pico PLUS chemiluminescent substrate and ChemiDoc XRS + were used to detect and display proteins. Quantity One software was used for the gray-scale analysis of protein bands.
qRT-PCR
Total RNA was extracted from modeling skin tissue homogenate using the RNA extraction kit according to the manufacturer's instructions. The PrimeScriptTM RT reagent Kit was used to reversely transcribe the total RNA into cDNA. The target cDNA fragment was amplified with the Real-time fluorescent quantitative PCR instrument. Gene primer sequences were designed by Primer Premier 5.0 software and synthesized in Sangon Biotech (Shanghai, China), as shown in Table 1. β-actin was used as the internal reference. The Bio-Rad Iqtm5 Optical System software and the 2−ΔΔCt method were used to analyze gene expressions.
The Gene Primer Sequences of qRT-PCR.

Statistical Analysis
In vivo experiments were performed using 8 mice per group. The variations among the data were gauged by applying the one-way ANOVA accordingly. Data analysis was performed in SPSS software v22.0. The data were indicated as the mean ± standard deviation. P < .05 was counted as statistically significant.
Results
Collection of Bioactive Compounds and Drug Targets for CZG
Using the TCMSP database, we obtained 123 bioactive compounds of CZG, consisting of 15 compounds from HEH, 7 from HHD, 15 from FCC, 39 from RS, 10 from RG, 30 from HSB, and 7 from RDC. In addition, 17 duplicate bioactive compounds were identified among the drugs. After excluding the duplicate components, 99 bioactive ingredients were obtained. The corresponding results are detailed in Table 2. After screening the targets of the 99 bioactive compounds from the TCMSP database, performing protein-gene named transformations and confirming that they were of human origin in the UniProt KB database, we obtained 1950 drug targets. Then, after removing the duplicate drug targets, a total of 579 targets were obtained.
The Information of CZG Bioactive Compounds.

Construction of Drug-Compound-Target (D-C-T) Network for CZG
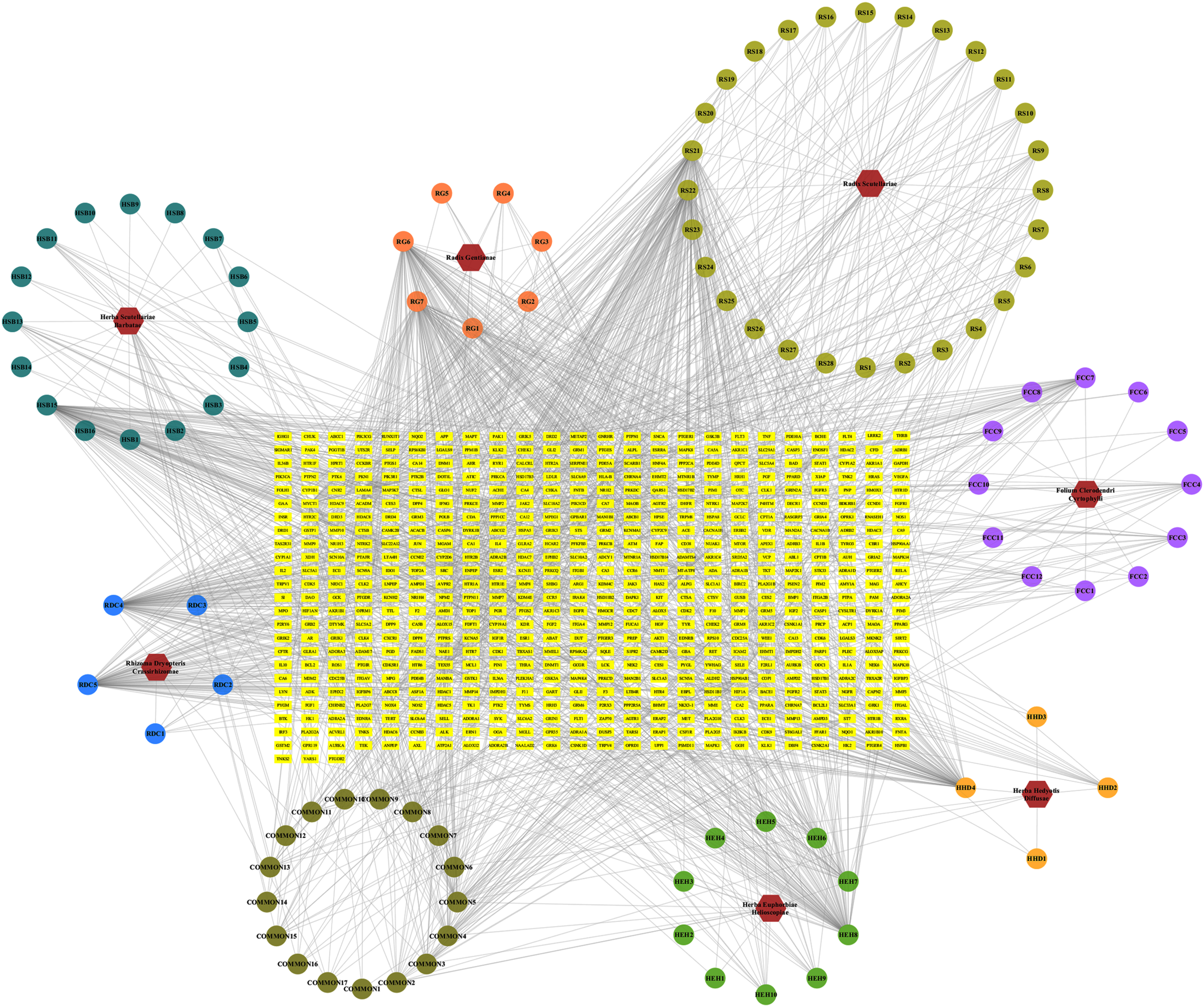
To visualize the relationship among the drugs, bioactive ingredients and drug targets, we imported the files containing them into Cytoscape 3.7.2 software to successfully construct the “D-C-T” network. As shown in Figure 2, there are 685 nodes and 1758 edges in the “D-C-T” network. The hexagon represents the drug, the circles represent the bioactive compound, and the rectangles represent the target.
Acquisition of Disease Targets for PV and Intersection Targets Related to CZG and PV
Using the GeneCards database, we obtained 940 PV targets. After converting all of them into standard gene names and confirming that they were of human origin using the UniProt KB database, the null and duplicate targets were removed. Lastly, we obtained 822 PV targets. By importing the drug targets and the disease targets into Draw Venn Diagram online platform, 108 “drug-disease” intersecting gene targets were identified, including the potential targets of CZG for treating PV. Figure 3 showed the intersection targets related to CZG and PV.

The “drug-disease” intersecting targets through Venn diagram.
Establishment and Analysis of “C-T” Network for Bioactive Compounds and Intersecting Targets
By entering 108 intersection targets and their corresponding bioactive compounds into the Cytoscape 3.7.2 software, we established the “Compound-Target” (C-T) network. As shown in Figure 4, the network had 192 nodes and 339 edges. We identified the top 10 bioactive ingredients using the degree as the parameter and listed them in Table 3.

The top 10 Bioactive Compounds in the “C-T” Network.

Construction and Topology of Protein-Protein Interaction (PPI) Network
By inputting the 108 intersection targets into the STRING database, we acquired their protein-protein interaction (PPI) network (Figure 5A). Then, we imported the PPI network results in “tsv” format from the STRING database into the Cytoscape 3.7.2 software to analyze and visualize it based on DC, BC, and CC parameters using the “NetworkAnalyzer” plugin. The PPI network analysis revealed a total of 65 nodes and 221 edges. Using initial screening thresholds set as Degree ≥ 5, BC ≥ 0.005, and CC ≥ 0.421, they were reduced to 24 nodes and 92 edges. The second screening criteria were Degree ≥ 13, BC ≥ 0.044, and CC ≥ 0.471, based on which we obtained 10 nodes and 27 edges. Figure 5B illustrates the network topology. The top 10 core intersection targets were STAT3, JUN, SRC, HRAS, HSP90AA1, STAT1, MAPK1, ESR1, EGFR and IL-10, as summarized in Table 4.

The top 10 Targets in the PPI Network.

Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) Enrichment Analysis
GO enrichment analysis revealed 1768 entries, of which 1551 were BP entries, 77 were CC entries, and 140 were MF entries. The analysis of the BP category showed that the targets were mostly within the regulation of defense response, response to lipopolysaccharide, regulation of inflammatory response, positive regulation of cell migration, response to molecule of bacterial origin, inflammatory response, positive regulation of cell motility, positive regulation of locomotion, cellular response to lipid, and positive regulation of cytokine production. The CC of these targets included membrane raft, membrane microdomain, receptor complex, side of the membrane, plasma membrane raft, caveola, focal adhesion, cell-substrate junction, perinuclear region of cytoplasm, and lytic vacuole. MF analysis showed that the targets were principally related to protein homodimerization activity, protein kinase activity, protein tyrosine kinase activity, growth factor receptor binding, phosphatase binding, phosphotransferase activity, alcohol group as acceptor, cytokine receptor binding, kinase activity, protein domain specific binding, and nuclear receptor activity. Figure 6A shows the results of the GO analysis.

Gene ontology (GO) (A) and Kyoto encyclopedia of genes and genomes (KEGG) (B) enrichment analysis.
KEGG enrichment analysis revealed 178 entries, including the PI3K-Akt signaling pathway, MAPK signaling pathway, IL-17 signaling pathway, JAK-STAT signaling pathway, etc Figure 6B shows that the top 20 filtered pathways were most closely related to CZG in the treatment of PV.
Molecular Docking Validation
According to the above results, STAT1, IL-10, JUN, and STAT3 were identified as the important core targets of CZG for treating PV, which are also the key targets in the JAK-STAT signaling pathway, coinciding with the KEGG enrichment analysis. To further verify the effects of CZG on PV, we selected STAT1, IL-10, JUN, and STAT3 for molecular docking verification. The CB-Dock website was used, with STAT1, IL-10, JUN, and STAT3 proteins serving as receptors, while the corresponding bioactive compounds in the top 10 of the “C-T” network and Methotrexate were used as ligands for molecular docking. The results showed that when only STAT1 and IL-10 serve as receptors, the binding energy with ligands is significantly lower than −5.5 kcal/mol. It is generally accepted that a binding energy < -4.25 kcal/mol represents a certain degree of binding tightness, while < -5.5 kcal/mol denotes a good binding affinity. Therefore, both STAT1 and IL-10 may be as the key targets of CZG for treating PV. Table 5 shows the results of molecular docking, and Figure 7A-D shows the visualization of the molecular docking results.

Results of the Molecular Docking.

The Effect of CZG on Psoriasis-Like Mice
Figure 8A shows the changes in skin lesions on the back of the mice in each group. Compared with the control group, the skin lesions in the modeling area were significantly thickened, and typical erythema, infiltration and scales were observed. Compared with the model group, the erythema and scales of skin lesions in the back modeling area of mice in the treatment group and the positive drug group were reduced, and the infiltration degree was reduced. Moreover, as shown in Figure 8B, the PASI score of skin lesions in the back modeling area of mice differed significantly (P < .05). As shown in Figure 8C, compared with the control group, the epidermal thickness in the skin lesion of the model group was significantly increased (P < .001). Compared with the model group, the thickness of skin lesions of the treatment group and the positive drug group were significantly decreased (P < .01). The pathological damage of skin in the back modeling area of mice in each group was also evaluated. As illustrated in Figure 8D, the control group had 1-3 stratum corneum, 3-5 stratum spinosum cells, intact basal layer cells, and no obvious inflammatory cell infiltration in the superficial dermis and perivascular. Compared with the control group, the pathological manifestations of the model group were hyperkeratosis with incomplete keratosis, spinous layer thickening, epidermal process extension, and superficial dermis lymphocyte infiltration. The pathological degree of mice in the treatment group and the positive drug group was consistent with the results of naked-eye observations.

Effects of CZG on the Levels of STAT1, IL-10, JUN, and STAT3 on the Skin Lesions of Psoriasis-Like Mice
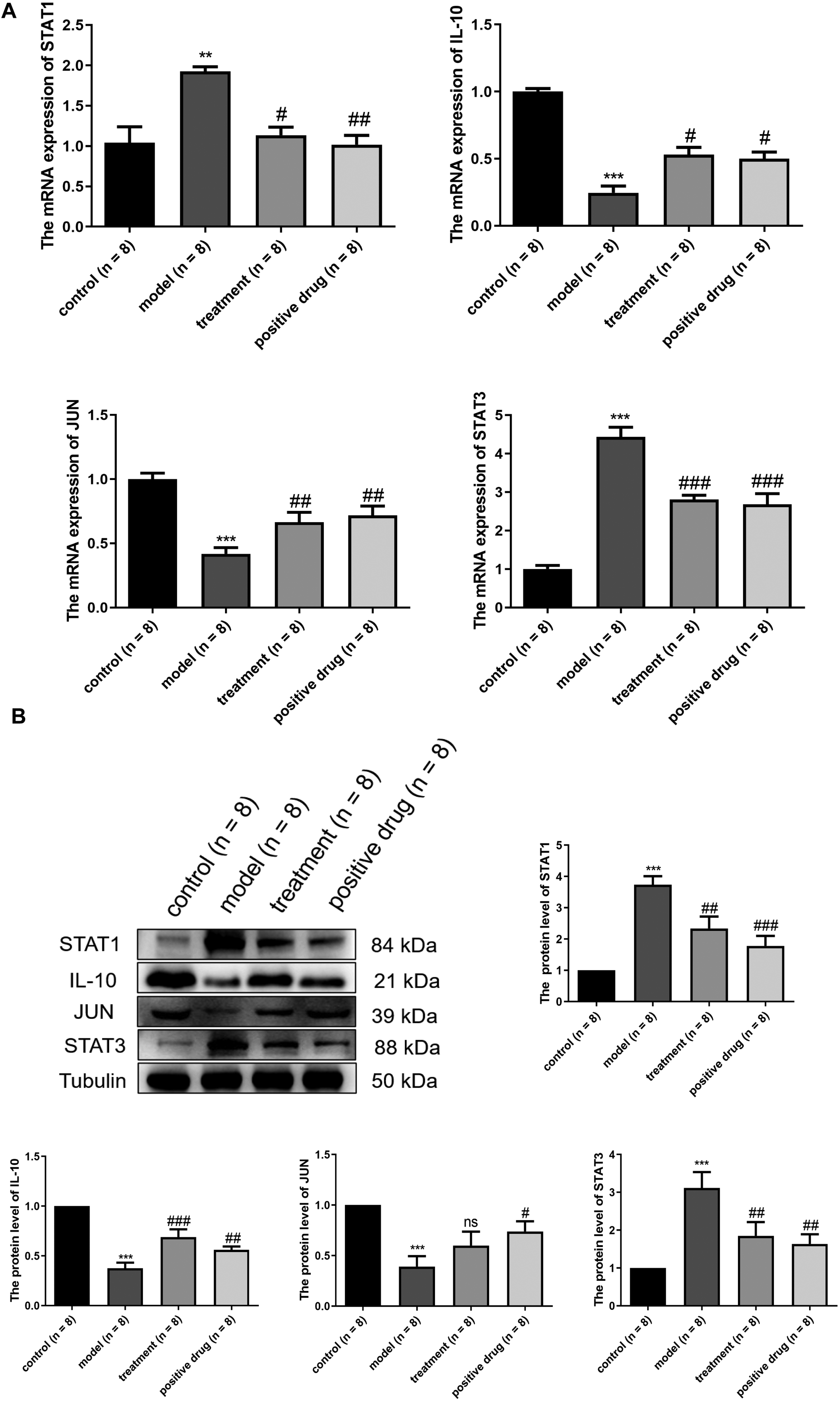
As STAT1, IL-10, JUN, and STAT3 are all core cytokines of the JAK-STAT signaling pathway, we conducted further analysis to investigate the influences of CZG on the expression of these molecules. As shown in Figure 9A, compared with the control group, the model group exhibited a significantly higher mRNA expression level of STAT1 and STAT3 (P < 0.01), and a significantly lower mRNA expression level of IL-10 and JUN (P < .001). Compared with the model group, the relative mRNA expression of STAT1 and STAT3 in the treatment group and the positive drug group was decreased significantly (P < .05), while the relative mRNA expression of IL-10 and JUN in the treatment group and the positive drug group was increased considerably (P < .05). As shown in Figure 9B, compared with the control group, the protein expression of STAT1 and STAT3 in the skin lesions of model mice was significantly enhanced (P < .001), and the protein expression of IL-10 and JUN was decreased remarkably (P < .001). Compared with the model group, the protein expression of STAT1 and STAT3 in the treatment group and the positive drug group were dramatically reduced (P < .001), and the protein expression level of IL-10 in the treatment group and the positive drug group was increased (P < .001). Additionally, we found a significant increase of JUN protein level in positive drug group as compared to the model group (P < .05); however, there seemed no significant difference between the model and treatment groups in JUN protein level. These results implied that CZG mainly affects STAT1 and IL-10 in JAK-STAT signaling pathway, which is consistent with the results of molecular docking validation.
Discussion
Psoriasis vulgaris (PV) is a chronic inflammatory skin disease mediated by the immune system and has a prevalence rate of approximately 2%–3%. 13 Although CZG has been used as a hospital preparation with a definite therapeutic effect on PV in clinical settings, its underlying therapeutic mechanism remains unknown. TCM can treat many diseases because of the “multi-compound, multi-target, multi-pathway” treatment mode, most of which are clinically effective for symptomatic treatment. However, the complex biological mechanisms associated with TCM pose significant challenges to experimental verification of its efficacy. Network pharmacology, which combines systems biology and computer technology, can be used to establish a multi-level network that mimics the treatment mode of TCM. Therefore, in this study, we utilized this method to elucidate the potential intrinsic mechanisms of CZG in treating PV.
Through network pharmacology, this study identified 99 bioactive compounds, 579 CZG drug targets, 822 PV disease targets, and 108 overlapping targets of CZG for the therapeutic management of PV. By constructing the “C-T” network and conducting enrichment analysis, we identified 10 corresponding bioactive compounds with the highest degree value, namely 6-Hydroxynaringenin, 5,8,2'-Trihydroxy-7-methoxyflavone, Gentirigenic acid, Gentiopicroside tetraacetate, 1,3,4,5,6a,7,8,8a,10,11,12,13,14b-tetradecahydropicene-4a-carboxylic acid, Dihydroxylin A, Harmony, Albaspidin AA, 7-Angelyl-9-echimidinylheliotridine, and Quercetin. Additionally, the constructed PPI network of intersection targets and enrichment analysis results identified STAT3, JUN, SRC, HRAS, HSP90AA1, STAT1, MAPK1, ESR1, EGFR and IL-10 as potential core targets associated with treatment efficacy. The GO enrichment analysis of the intersection target identified 1768 entries, including 1551 BP entries, 77 CC entries and 140 MF entries. KEGG enrichment analysis of the intersection target revealed 178 pathways, mainly including the PI3K-Akt signaling pathway, MAPK signaling pathway, IL-17 signaling pathway, JAK-STAT signaling pathway, etc.
Based on the results, we concluded that STAT1, IL-10, JUN, and STAT3 could be important targets in the treatment of PV by CZG, which coincides with the JAK-STAT signaling pathway obtained from KEGG enrichment analysis. The molecular docking results confirmed that the bioactive compounds with top-degree values associated with STAT1 and IL-10 had good binding activity corresponding to the target, with the binding energy significantly lower than −5.5 kcal/mol. Therefore, we speculated both STAT1 and IL-10, two cytokines on the JAK-STAT signaling pathway, may be the direct targets for CZG. As an important cytokine signaling pathway, the JAK-STAT signaling pathway is widely involved in cell proliferation, differentiation, apoptosis and immune regulation.14–16 Studies have shown that the JAK-STAT signaling pathway in PV patients is abnormally active and blocking it improved the efficacy, quality of life and immune function of PV patients.17–20 Studies have also reported that IL-10 can reduce epidermal mitosis, keratinization, inflammatory cell infiltration and Munro micro-abscess in psoriasis, thereby thinning the epidermis.21,22 The abnormal activation of STAT1 in the STAT family can mediate the occurrence and development of PV by promoting the excessive differentiation and proliferation of cells and inhibiting cell apoptosis. 23 Additionally, a comprehensive analysis of the results of PPI, KEGG enrichment analysis and molecular docking showed that the bioactive compounds of CZG played a role in the treatment of PV by targeting multiple proteins and multiple pathways, similar to JAK-STAT signaling pathway and its STAT1 and IL-10 cytokines.
After conducting animal experiments to verify our findings, we evaluated the severity of skin lesions in psoriasis-like mice by assessing the PASI score, observing the histopathological changes and epidermal thickness using HE. Our results showed that CZG could effectively reduce the PASI score of psoriasis-like mice, alleviate pathological damage and epidermal thickness of back skin lesions. Additionally, given that STAT1, IL-10, JUN, and STAT3 are pivotal cytokines in the JAK-STAT signaling pathway, we conducted further analysis to investigate the impact of CZG on the expression of these molecules. Our findings indicated that while CZG can effectively modulate the mRNA expression levels of four factors, it appears to have no impact on the protein expression of JUN. These results were consistent with the results of molecular docking validation and implied the potential therapeutic role for CZG in the prevention and treatment of PV through targeting STAT1 and IL-10.
From the perspective of network pharmacology, our present study can be considered reliable, normative and reasonable. However, some limitations should be acknowledged, including the incompleteness of the database, the subjective nature of the algorithms, the limitations of the analytic software used, and the fact that the efficacy of CZG in treating PV has only been validated through animal experiments. Therefore, we still need further experimental verification and exploration, such as preparing CZG drug-containing serum and mass spectrometry analysis of bioactive compounds of CZG, to further validate these findings.
There is a limitation of this study that should not be ignored. In fact, the significance of single herb is greater, and there are too many influencing factors of compound formula. Therefore, we will consider using single herb or active ingredient for further research in the future.
Conclusion
In summary, this present study utilized network pharmacology to elucidate the multifaceted mechanism of CZG in treating PV, taking into account its multi-compound, multi-target and multi-pathway effects. Furthermore, our animal experiments provided evidence showing the promising therapeutic effects of CZG on PV. These findings offer a foundation for extracting bioactive drug components from CZG for the treatment of PV and provide a theoretical framework for further experimental validations.
Supplemental Material
sj-docx-1-npx-10.1177_1934578X241300924 - Supplemental material for Study on the Mechanism of Compound Zeqi Granule in Treating Psoriasis vulgaris Based on Network Pharmacology and Experimental Verification
Supplemental material, sj-docx-1-npx-10.1177_1934578X241300924 for Study on the Mechanism of Compound Zeqi Granule in Treating Psoriasis vulgaris Based on Network Pharmacology and Experimental Verification by Min Wu, Tongtong Dong, Ting Zhang, Jiuxiang Wang, Taofeng Liu and Changzhong Wang in Natural Product Communications
Footnotes
List of Abbreviations
Acknowledgements
We acknowledge the strong support of Anhui University of Chinese Medicine and The First Clinical Medical College of Anhui University of Chinese Medicine.
Availability of Data and Materials
The data used to support the findings of this study are included within the article and are available from the first author upon request.
Authors’ Contributions
Changzhong Wang and Taofeng Liu made substantial contributions to the conception and design of the work. Min Wu, Tongtong Dong and Ting Zhang made substantial contributions to the acquisition, analysis and interpretation of data for the work and drafted the manuscript. Min Wu and Jiuxiang Wang revised the manuscript critically for important intellectual content. All authors agreed to be accountable for all aspects of the work in ensuring that questions related to the accuracy and agreed final approval of the version to be published.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
This study was approved by the Laboratory Animal Ethics Committee of Anhui University of Chinese Medicine (approval ID: AHUCM-mouse-2021113).
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the the Clinical Scientific Research Project of the First Affiliated Hospital of Anhui University of Chinese Medicine, (grant number 2020yfyzc18) and Key Project of Scientific Research in Higher Education Institutions in Anhui Province, (grant number 2024AH051042).
Statement of Human and Animal Rights
All of the experimental procedures involving animals were conducted in accordance with the NIH Guide for the Care and Use of Laboratory Animals and approved by the Laboratory Animal Ethics Committee of Anhui University of Chinese Medicine (approval ID: AHUCM-mouse-2021113).
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
