Abstract
Cinnamon (
Introduction
Ovarian cancer (OC) is a malignant tumor of the reproductive tract that poses a serious threat to women's lives. 1 In 2015, there were 52,100 new cases of ovarian cancer and 22,500 deaths in China. 2 Most OC patients are found to be advanced, and their susceptibility to extensive pelvic and abdominal metastases, multidrug resistance to primary or secondary chemotherapy, and recurrence has led to a high mortality rate, with the 5-year survival rate still hovering at 40%.3, 4 At present, the treatment of OC is usually a combination of surgery and chemotherapy,5, 6 but chemical drugs have a certain toxicity and sometimes resistance develops to them, and Chinese medicine has unique advantages in this regard. 7 The vast majority of recurrent OC is incurable and treatment is mostly palliative with the aim of improving symptoms and quality of life, and so prevention of OC recurrence and reversal of platinum resistance are current clinical concerns.
Cinnamon is the dried bark of
In Chinese medicine research, analysis of the mechanism of active components for the treatment of diseases is a hot spot, but a difficult area. Network pharmacology provides a good platform for the study and analysis of multi-component, multi-target, and multi-pathway drugs.18, 19 The Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP) is a unique TCM systemic pharmacology platform from which relationships between drugs, target proteins, and diseases can be obtained.
In this study, network pharmacology integrated bioinformatics and molecular docking were applied to explore the potential targets and signaling pathways of cinnamon for the treatment of OC, and to predict the targets of the signaling pathways where cinnamon acts, with the aim of obtaining more detailed information on its mechanism for the treatment of OC.
Materials and Methods
Screening and Target Prediction of the Main Active Components of Cinnamon
The chemical composition of cinnamon was collected using the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP, https://old.tcmsp-e.com/tcmsp.php).
20
The TCSMP database was used to obtain all the main active components of cinnamon, and the database was used until 16th May, 2021. Two indicators, oral bioavailability (OB) ≥ 30% and drug-like index (DL) ≥ 0.08,
21
were used as screening thresholds to determine the final drug active components. The potential target proteins corresponding to the active components were then collected one by one, and the protein names were normalized on the Uniprot database (https://www.uniprot.org/), and the species “
OC Disease-Related Target Acquisition
OC-related genes were retrieved from GeneCards (https://www.genecards.org/) 22 and DisGeNET databases (https://www.disgenet.org/) using the search term “ovarian cancer”, and OC-related targets were obtained by processing and removing duplicates from the collected disease targets.
Construction of a Protein–Protein Network
The potential targets of cinnamon in the treatment of OC were analyzed using STRING database (https://string-db.org/), and the corresponding species was set to “
Biofunctional and Pathway Enrichment Analysis
The potential targets of cinnamon in the treatment of OC were enriched by Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis using the human genome annotation database DAVID database (https://david.ncifcr f.gov/) to investigate the biological processes and signaling pathways involved in the treatment of OC with cinnamon. The identifier “GENE OFFICIAL SYMBOL” and the species “
Network Construction and Analysis for OC
Venn diagrams were used to analyze “ component-target” and “disease-target” and identify shared targets. The Venn diagram was drawn to obtain the targets of cinnamon in the treatment of OC by using the diagram to map “component-target” to “disease-target”. Cytoscape 3.7.1 software was used to construct a “drug active component-target” visual network.
Molecular Docking
Molecular docking relies on the “lock-and-key principle” of ligand-receptor interactions, simulating the interaction between small molecule ligands and receptor biomolecules. 23 The binding pattern and affinity between components and proteins can be predicted computationally.24, 25 All target proteins were obtained from the RCSB PDB database (http://www.rcsb.org/) as 3D structures. The entire molecular docking process was performed with the Molecular Operating Environment (MOE) (v2015.10), including 6 steps: 1. input protein PDB file; 2. protein structure protonation; 3. removal of water molecules; 4. protein structure energy minimization; 5. protein binding site analysis; 6. input ligand molecules, and molecular docking.
Results
the Main Active Components of Cinnamon
One hundred chemical components of
Screening Results of Active Components of Cinnamon.

Constructing a Cinnamon Active Component Target Network
A total of 166 drug targets (no corresponding targets for hexadec-11-enal) were obtained by searching the 15 actives of cinnamon using TCMSP, and 74 active targets were obtained after de-duplication and standardization in the Uniprot database. The active components and active targets were imported into Cytoscape network visualization software to construct the active component-target network of cinnamon. As shown in Figure 1, there were 88 nodes (14 active components and 74 active targets) and 166 edges in the composition-target network, in which the red nodes represented the active components of cinnamon and the blue nodes represented the drug targets. As shown in Table S1, oleic acid in cinnamon can act on 48 targets. The highest degree of target action was found for muscarinic acetylcholine receptor M1 (CHRM1), which was associated with 11 chemical components, followed by γ-aminobutyric acid receptor subunit α-1 (GABRA1), which was associated with 10 chemical components.

Component-target network of cinnamon.
Potential Targets of Cinnamon in the Treatment of OC
7255 OC-related targets were obtained from the Gene Cards database, as shown in Table S2. The DisGeNET database was used to obtain 2563 targets related to OC, as shown in Table S3. The 74 targets obtained from the drug prediction were mapped to the corresponding targets of the disease using Venn diagrams, resulting in 26 potential targets of cinnamon in OC, including ACHE, ADH1B, ADRA1A, ADRB2, BDNF, CAT, CCK, CRP, DPP4, EDN1, ERBB2, MPO, NR3C1, PGR, PLAU, PLG, PON1, PRSS3, PTGS1, PTGS2, SCD, SERPINE1, SLC2A1, SOAT1, SOD1, and TEP1, as shown in Figure 2.
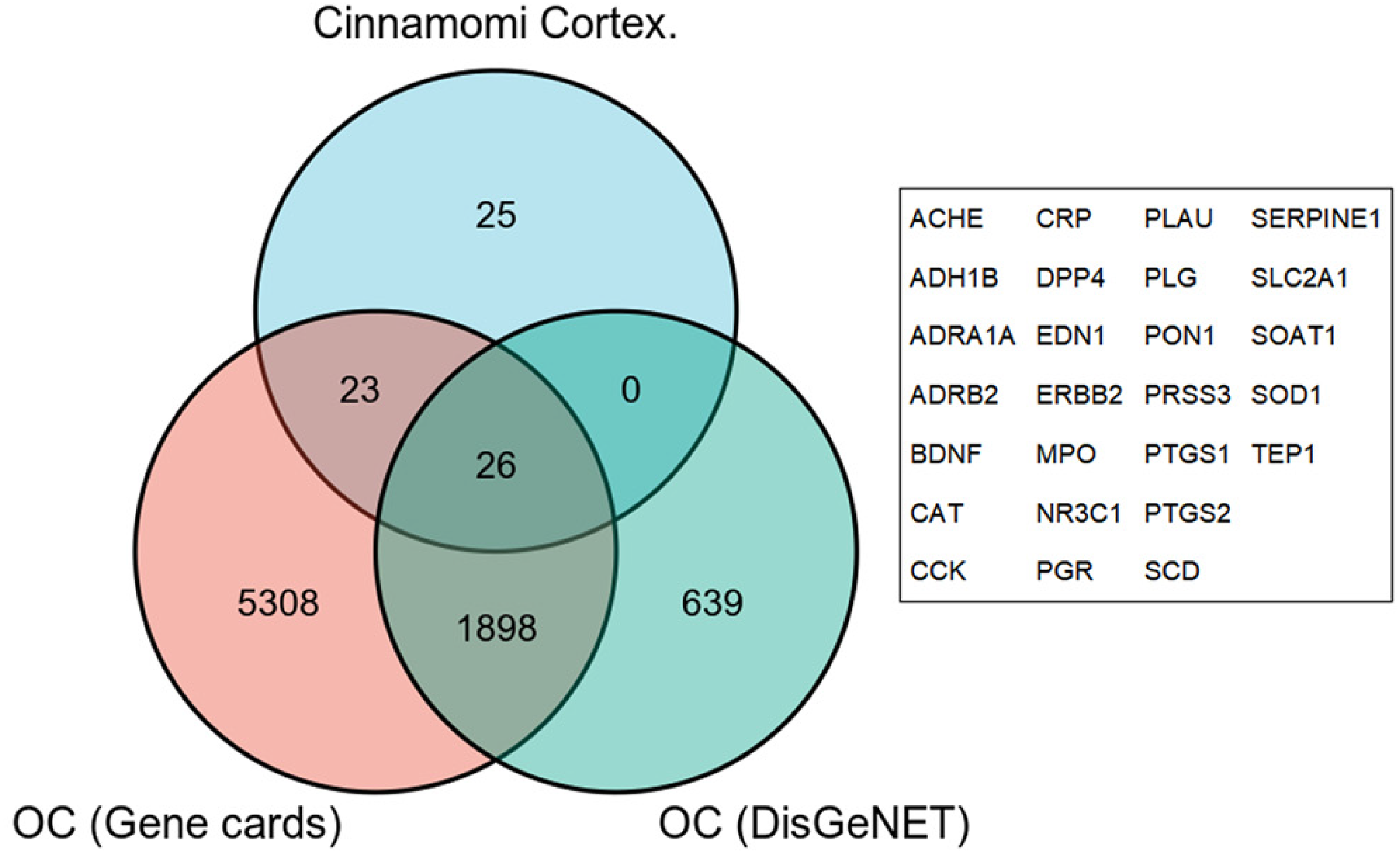
Screening of targets for OC treated by cinnamon.
Target PPI Network Analysis
As shown in Figure 3, the PPI network consisted of 26 targets of cinnamon for OC contained 26 nodes and 85 edges, with an average node degree of 6.54, an average local clustering coefficient of 0.534, an expected number of edges of 16, and a PPI concentration

Protein-protein interaction network of potential targets.
GO Functional Enrichment of key Targets of Cinnamon in the Treatment of OC Analysis
As shown in Table S4, 368 GO functional enrichment analyses were listed based on

Go and enrichment analysis of cinnamon for treating OC. The 26 potential targets of cinnamon in OC were imported into the DAVID database, and the key targets were analyzed for different biological processes at three levels: biological process (BP), cellular component (CC), and molecular function (MF).
KEGG Pathway Analysis of Cinnamon as a Target for OC Treatment
As shown in Figure 5 and Table S4, the KEGG pathway enrichment analysis of 26 key targets of cinnamon in OC identified four major signaling pathways, mainly related to neuroactive ligand-receptor interaction, HIF-1 signaling pathway, regulation of lipolysis in adipocytes, and complement and coagulation cascades.

KEGG signaling pathway analysis of cinnamon for treating OC.
Construction of an “Active Component - Target - Pathway” Network for Cinnamon
Cytoscape software was used to model the “active ingredient - target - pathway” network of Cinnamon. The correlations between 9 components, 13 targets, and 4 pathways are shown in Figure 6. Oleic acid, a component of Cinnamon, regulates genes including CCK, EDN1, PLG, PRSS3, ERBB2, SERPINE1, SLC2A1, PTGS1, PTGS2, and PLAU of pathways including neuroactive ligand-receptor interaction, HIF-1 signaling pathway, regulation of lipolysis in adipocytes, and complement and coagulation cascades. PTGS2 was associated with EIC, zoomaric acid, (Z)-caryophyllene, β-cubebene, ()-sativene, (-)-caryophyllene oxide, (-)-α-cedrene, and oleic acid.

Cinnamon active component-target-pathway network diagram. Cinnamon contains a complex network of effector components and targets, and between targets and pathways of action, where blue represents Cinnamon effector components, red represents component targets, and green represents pathways of action.
Molecular Docking Analysis
In our study, molecular docking was further applied to assess the interactions between components and targets, reducing the complexity and improving the accuracy of the composed target network. A virtual screen was performed using MOE to determine the binding affinity between protein models and 6 potentially active components obtained from component- component (including α-cedrene, caryophylene oxide, cubebene, DIBP, oleic acid, and sativene) target network. Twelve targets (ADRB2, CCK, EDN1, ERBB2, NR3C1, PLAU, PLG, PRSS3, PTGS1, PTGS2, SERPINE1, and SLC2A1) and 2 components (DIBP and oleic acid) were analyzed by molecular docking (Figure 7). There was a stable interaction between DIBP and ADRB2 and NR3C1. There is a stable interaction between oleic acid and C2K, EDN1, ERBB2, PLAU, PLG, PRSS3, PTGS1, PTGS2, SERPINE1 and SLC2A1. In conclusion, docking simulation studies showed that different binding models between components and proteins have different binding abilities. The detailed energy docking scores are in Table 2. The lower the score, the more stable the ligand-receptor binding. The calculated results show that representative components in cinnamon can bind well to the target genes.

The docking model of components with 12 genes. Binding model of DIBP on the molecular surface of ADRB2 (A) and NR3C1 (E). The interaction model of DIBP with ADRB2 (a) and NR3C1 (e). Binding model of oleic acid on the molecular surface of (B) CCK, (C) EDN1, (D) ERBB2, (F) PLAU, (G) PLG, (H) PRSS3, (I) PTGS1, (J) PTGS2, (K) SERPINE1, and (L) SLC2A1. The interaction model of oleic acid with (b) CCK, (c) EDN1, (d) ERBB2, (f) PLAU, (g) PLG, (h) PRSS3, (i) PTGS1, (j) PTGS2, (k) SERPINE1, and (l) SLC2A1. The ligands in binding model and interaction model are colored in green and within the dashed line, respectively. The length of the bond was added to it.
Virtual Docking of 2 Representative Components of Cinnamon with OC Targets.

Discussion
OC has an insidious clinical manifestation in the early stage, which makes diagnosis difficult. In the later stages, the disease progresses rapidly and is prone to extensive abdominal metastasis and chemotherapy multidrug resistance, and is prone to recurrence. Although surgical techniques are improving, chemotherapy regimens are becoming more standardized, and the side effects of new chemotherapeutic agents are diminishing, but there has been no significant improvement in the overall survival rate from ovarian cancer, which remains a major reproductive tract malignancy that poses a serious threat to women's lives. 3
Herbs are gaining attention as effective and complementary anticancer therapeutic agents that can enhance the efficacy and reduce the toxicity of anticancer agents.
26
Oleic acid-induced apoptosis in tumor cells is associated with increased intracellular reactive oxygen species (ROS) production or cystein 3 activity.
11
Excess reactive species of oxygen and nitrogen may still cause oxidative damage to tissues and organs. Hesperidin, cinnamon, and
The analysis in this study identified four pathways through which cinnamon acts in relation to ovarian cancer, including neuroactive ligand-receptor interaction, HIF-1 signaling pathway, regulation of lipolysis in adipocytes, and complement and coagulation cascades. Dual drug-loaded micelles targeting the HIF and mTOR signaling pathways have been used in combination therapy for OC treatment. 33 Metformin prevents tumor-to-tumor crosstalk in OC by metabolically downregulating the expression of HIF1α in mesothelial cells. 34 Knockdown of HIF-1α promotes autophagy and inhibits the PI3K/AKT/mTOR signaling pathway in OC cells. 35 Pancreatic cancer (PC)-secreted exosomes cause lipolysis in subcutaneous adipose tissue. 36 There is a complex metabolic symbiosis between the adipocytes surrounding the tumor and the cancer cells, which stimulates their aggressiveness. 37 Complement and coagulation cascade pathways correlate with chemosensitivity and overall survival in patients with soft tissue sarcoma (STS). 38 The KEGG enrichment analysis showed that it was associated with neuroactive ligand-receptor interaction, regulation of lipolysis in adipocytes, and complement and coagulation cascades.
In this study, ADRA1A, ADRB2, CCK, EDN1, NR3C1, PLG, PRSS3, ERBB2, SERPINE1, SLC2A1, PTGS1, PTGS2, and PLAU were likely to be the therapeutic targets of cinnamon in the treatment of OC. Several factors related to stress and ovulation are differentially associated with ovarian tumors that respond to ADRB2 signaling. 39 The EDN1 axis promotes YAP-induced chemotherapy escape in OC. 40 Endothelin A receptor/β-arrestin signaling to the Wnt pathway makes ovarian cancer cells resistant to chemotherapy. 41 Elevated PLG expression represents a good prognostic biomarker for advanced (FIGO III/IV) high grade serous ovarian cancer (HGSOC). 42 Overexpression of PRSS3 can be used as a predictor of clinical prognosis in OC patients. 43 Silencing ERBB2 gene expression may inhibit activation of the MAPK1/MAPK3 signaling pathway, thereby suppressing the proliferation, invasion, and migration of OC cells. 44 FOXM1 and the EGFR/ERBB2 pathway are key points of treatment to disrupt the vulnerability of peritoneal spread and ovarian cancer cell adhesion. 45 ERBB2 reduces the drug sensitivity of OC cells by inducing stem cell-like properties. 46 Aberrant SERPINE1 DNA methylation is associated with carboplatin-induced epithelial-mesenchymal transition in epithelial ovarian cancer (EOC). 47 However, the role of genes including ADRA1A, CCK, NR3C1, SLC2A1, PTGS1, PTGS2, and PLAU in OC are not clear. The mechanism by which cinnamon acts through these targets in OC needs to be further investigated.
Modelling of the ‘active ingredient-target-pathway’ network of cinnamon using Cytoscape software showed correlations between 9 components, 13 targets, and 4 pathways (Figure 6). Subsequently, molecular docking simulations were performed for 9 components and 13 targets. A virtual screen using MOE determined the binding affinity between 13 proteins and 6 components including α-cedrene, caryophylene oxide, cubebene, DIBP, oleic acid, and sativene (Table 2). The molecular docking results for 12 target genes (ADRB2, CCK, EDN1, ERBB2, NR3C1, PLAU, PLG, PRSS3, PTGS1, PTGS2, SERPINE1, and SLC2A1) and two core components (DIBP and oleic acid) are shown in Figure 7. DIBP and oleic acid are expected to be potential drugs for the treatment of OC and will be the focus of future research.
Due to the limitations of network pharmacology and molecular docking, experiments need to be conducted in the future on the material basis, pharmacodynamics and pathway validation to provide a theoretical and experimental basis for the use of cinnamon in the treatment of OC.
Conclusion
This study showed that the main pathways involved in the treatment of OC by cinnamon are neuroactive ligand-receptor interaction, HIF-1 signaling pathway, regulation of lipolysis in adipocytes, and complement and coagulation cascades, and predicted that ADRA1A, ADRB2, CCK, EDN1, NR3C1, PLG, PRSS3, ERBB2, SERPINE1, SLC2A1, PTGS1, PTGS2, and PLAU may be potential targets of cinnamon for the treatment of OC.
Supplemental Material
sj-xlsx-1-npx-10.1177_1934578X221119118 - Supplemental material for Network Pharmacology, Integrated Bioinformatics, and Molecular Docking Reveals the Anti-Ovarian Cancer Molecular Mechanisms of Cinnamon (Cinnamomum cassia (L.) J. Presl)
Supplemental material, sj-xlsx-1-npx-10.1177_1934578X221119118 for Network Pharmacology, Integrated Bioinformatics, and Molecular Docking Reveals the Anti-Ovarian Cancer Molecular Mechanisms of Cinnamon (
Footnotes
Acknowledgements
This work was supported by Jiangsu Provincial Key Discipline of Maternal and Child Health (2017103033) and Xuzhou Key R&D Programme (ZYSB20210489).
Author Contributions
Buze Chen, Guilin Li, and Xiaoyuan Lu did the conception and design; Buze Chen, Guilin Li, and Xiaoyuan Lu did the acquisition; Buze Chen, Xin Jin, Haihong Wang, Qingmei Zhou, Guilin Li, and Xiaoyuan Lu did the analysis and interpretation of data; Buze Chen, Xin Jin, Guilin Li, and Xiaoyuan Lu did the drafting of the manuscript; Buze Chen, Xin Jin, Haihong Wang, Qingmei Zhou, Guilin Li, and Xiaoyuan Lu did the critical revision of the manuscript for important intellectual content; Buze Chen and Xiaoyuan Lu did the statistical analysis; Buze Chen and Xiaoyuan Lu did the obtaining funding; Buze Chen and Xiaoyuan Lu did the supervision.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Xuzhou Key R&D Programme, Jiangsu Provincial Key Discipline of Maternal and Child Health, (grant number ZYSB20210489, 2017103033).
Data Availability Statement
The data within the article are available from the corresponding author upon request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
