Abstract
Introduction
Lupus nephritis (LN) represents one of the serious manifestations of systemic lupus erythematosus (SLE), an autoimmune disease affecting the kidneys. 1 The clinical manifestations of the LN are mostly similar to nephrotic syndrome or chronic glomerulonephritis and mainly include edema, proteinuria, and hematuria, which can cause renal failure, uremia, and even death. 2 Glucocorticoids and immunosuppressants are commonly used in treatments management to prevent the development of LN; however, there is a high probability of disease recurrence, the occurrence of secondary infections, and other complications that could inversely impact patients life quality and threaten patients’ lives. 3
Nowadays, Traditional Chinese medicine (TCM) is increasingly attracting the interests from investigators and clinicians globally because it has been verified with efficacy owing to its multi-component, multi-target, and multi-pathway mechanism of action.4,5 It is believed that the pathogenesis of LN is based on the deficiency of the spleen and kidney, and on this basis, it is caused by exogenous heat toxins, dampness, and turbidity, and its remission period is mostly seen in liver and kidney Yin deficiency syndrome. The TCM Erzhi Pill consists of equal amounts (1:1) of Ecliptae Herba (aerial parts of Eclipta prostrata L. (Asteraceae)) and Fructus Ligustri Lucidi (dried mature fruit of Ligustrum lucidum Ait. (Oleaceae)), both of them can be used for treating and nourishing liver and kidneys, and is thus a good agent to tonic Yin, nourish kidneys, nourish qi, and soothe liver. 6 Various preclinical studies have confirmed multiple pharmacological effects of the Erzhi Pill, including body's immune function regulation, hypoglycemia, lipid-lowering, anti-oxidant, improving blood microcirculation, and protecting the liver, and it is quite effective on treating cardiovascular, hepatobiliary, nephropathy, and gynecological disorders.7,8 Although Erzhi Pill has shown a protective effect on the kidneys, its mechanism is still unclear.
Network pharmacology (NP) integrates bioinformatics, pharmacology, as well as some additional subjects with network analysis. It is capable of describing the drug action mechanism involving multiple components as well as multiple targets by analyzing interaction between gene phenotypes, molecular functions, together with pathways based on the constructed disease-phenotype-gene-drug network. 9 The present work focused on exploring Erzhi Pill's mechanism and possible active ingredients against LN by using NP together with molecular docking (MD) for providing the basis in future experiments and clinical application of the Erzhi Pill.
Materials and Methods
Bioactive Ingredient Selection for Erzhi Pill
This work applied the TCM systems pharmacology database and analysis platform (TCMSP, https://old.tcmsp-e.com/tcmsp.php) and high-throughput experiment and reference-guided database of TCM(HERB,http://herb.ac.cn/) for identifying chemical constituents in Erzhi Pill. The roots were screened in line with ADME factors (eg, distribution, absorption, excretion, and metabolism. Only components with drug-likeness (DL) > 0.18 and oral bioavailability (OB) > 30% were retained. 10
Screening for Possible Targets for LN and Erzhi Pill
This work obtained predicted targets in Erzhi Pill, including those of Eclipta Herba and Fructus Ligustri Lucidi in the TCMSP and DrugBank (https://www.drugbank.ca/) database. The GeneCards database is an integrated database, which offers concise gene-centric data, including human genomics together with clinical information. 11 This work obtained LN-related targets in GeneCards database (https://www.genecards.org/), Online Mendelian Inheritance in Man (OMIM, https://omim.org/), DrugBank (https://www.drugbank.ca/), and the Pharmacogenomics Knowledgebase (PharmGKB, https://www.pharmgkb.org/) using “lupus nephritis” as the keyword.
Drug-Disease Network Construction
The intersected gene targets between Erzhi Pill and LN were determined by R.4.1.0 software, which were later adopted for mapping targets between drug and disease. In order to obtain important active molecules, the common target genes of Erzhi Pill and LN were introduced into Cytoscape3.7.0, and the relation of effective ingredients with intersection gene targets was plotted. In that generated target network, nodes represent active components of the Erzhi Pill and potential action targets, while the edge is used to connect active components as well as putative targets for showing association of compounds with targets. The degrees represent the line numbers between nodes and can be assessed by using “Network Analyzer,” which is an additional function in the Cytoscape software.
Analysis of Network Topology
This work uploaded the targets intersected between effective ingredients and disease in STRING database (https://string-db.org/) for constructing a protein-protein interaction (PPI) network. We set biological species and minimal required interaction scores as “Homo sapiens” and 0.9, separately. The results were then input in the Cytoscape 3.7.0, while topological analysis of PPI network was carried out using “cytoNAC” package. Hub targets were screened through centralities, including Closeness Centrality (CC), Betweenness Centrality (BC), Eigenvector Centrality (EC), Degree Centrality (DC), Network Centrality (NC), and local average connectivity (LAC), and their values were twice of medians, respectively. These targets were key global geometric quantities in the network.
Analysis Based on GEO Database
GEO database (https://www.ncbi.nlm.nih.gov/geo/) has been developed as the free repository for genome data. 12 Genetic data were selected by searching for “Lupus Nephritis” within the GEO database, followed by downloading the gene matrix and platform data file. Perl's script was used to map the downloaded probes ids to gene ids. This work identified differentially expressed genes with up-regulation or down-regulation in LN group and normal group by R software “Limma” package, upon the thresholds of |logFC > 1| and P < .05.
MD of Effective Components with Targets
We chose three top components from those hub components in Erzhi Pill for MD with two proteins chosen based on the GEO database, for the sake of validating whether our selected components were accurate. Chemical structures of ligands were retrieved based on PubChem database (https://pubchem.ncbi.nlm.nim.nih.gov). Meanwhile, the target 3D structural information was obtained in RCSB protein data bank (http://www.rcsb.org). Binding affinity was predicted using AutoDock Vina 1.1.2, which was simulated by binding energy between two molecules. Typically, the binding energy <0 indicating spontaneous binding of two molecules, with a lower value indicating the more stable structure. At last, we input these findings in Discovery Studio 2019 client to conduct MD to obtain the 2D structure.
Gene Ontology Enrichment as Well as Kyoto Encyclopedia of Genes and Genomes Analysis
Biological function characteristics enriched by key genes were analyzed by Gene Ontology (GO), while specific pathways were analyzed through Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis. GO is mainly completed using “clusterProfiler” package of R software, which includes the Biological Process (BP), Molecular Function (MF), and Cellular Component (CC). 13 The R 4.1.0 software packages, including “clusterProfiler,” “GGploT2,” and “enrichPlot,” etc, were to identify the names of metabolic pathways in which the target participates. The top 30 pathways were then selected to draw the bubble map. We imported the above results of KEGG pathways in Cytoscape. The corresponding relationship between pathway and gene was displayed by Degree, and the network diagram was drawn.
Results and Discussion
Effective Component and Target Gene Selection for Erzhi Pill
Through TCMSP database, with as the limiting conditions, we obtained 21 effective chemical compounds, including 11 of Fructus Ligustri Lucidi, 8 of Eclipta Herba, and 2 (luteolin, quercetin) of both Fructus Ligustri Lucidi and Eclipta Herba (Table 1). We acquired altogether 646 drug targets, finally, 189 target genes were obtained after gene annotation and deleting duplicates.
Available Compounds in the Erzhi Pill.

DL, drug-likeness; OB, oral bioavailability.
Screening for LN Disease Targets
This work screen targets related to LN by searching against OMIM, GeneCards, DrugBank, and PharmGKB databases. This work obtained altogether 1147 LN-related targets after integrating data and removing duplicated data (Figure 1).

Overlapping LN-related target genes. LN, lupus nephritis.
Drug-Target-Disease Network Establishment
Those core active target components of Erzhi Pill were matched with the disease targets of LN, and 87 common targets of the Erzhi Pill and LN were identified (Figure 2). In the relationship network of the Erzhi Pill for the treatment of LN (Figure 3), there were 100 nodes and 191 edges. The elliptic nodes represented the active components of the drugs, among which green nodes represent Eclipta Herba, blue nodes represent Fructus Ligustri Lucidi, and yellow nodes represent both. The red nodes correspond to common targets of Erzhi Pill and LN. The node size reflects the degree value, which is the most direct measurement index. 14 The higher degree value indicates the higher node importance within the network. Every component showed interaction with average 14.69 targets in our network, and every target showed interaction with average 2.2 compounds. Thereby indicating that one compound can interact with multiple targets of the Erzhi Pill, and different compounds also have action on one individual target. This indicates integration and connection of the multi-component and multi-target interaction in TCM. As for compounds, the top three compounds were MOL000098-quercetin, MOL00006-luteolin, and MOL000422-kaempferol, which, respectively, can interact with 72, 30, and 28 target proteins. The average degree value of these three compounds in Erzhi Pill was 43.3, about three times the average degree value of the 13 active components, indicating their efficacy against most of the targets in LN, as well as that these three compounds were very likely to account for most of the clinical effects of Erzhi Pill. These were the main therapeutically active components of the Erzhi Pill useful for treating LN. Quercetin is one of the main active components of the Erzhi Pill, which is found in a variety of other plants and has demonstrated antibacterial, anti-inflammatory, anti-tumor, anti-allergic, hypolipidemic, hypoglycemic, and other pharmacological effects.15,16 Quercetin can significantly inhibit renal tubule interstitial damage, as well as can reduce inflammatory factor production by regulating macrophage polarization, thus, improving renal injury. 17 Besides, quercetin alleviate renal tubular epithelial cells (RTEC) aging while alleviating renal fibrosis via SIRT1/PINK1/mitochondrial-autophagy axis. 18 Quercetin has been shown to ameliorate renal injury and pyroptosis in LN in vitro and in vivo through inhibiting IL-33/ST2 pathway. 19 Luteolin, another active ingredient of the Erzhi Pill, has anti-inflammatory, anti-oxidant, immunomodulatory, and other effects. 20 Luteolin has shown a significant protective effect on renal damage caused by PbAc poisoning by activating the anti-oxidation, anti-inflammation, as well as anti-apoptosis of Nrf2/ARE pathway. 21 Also, luteolin is suggested to resist renal ischemia-reperfusion (I/R) injury by decreasing OS, up-regulating anti-oxidant enzymes, while down-regulating miR320 and Nrf2 levels. 22 Tao Ding et al have demonstrated that luteolin can improve renal oxidative stress and reduce urinary protein levels in mice with LN, which may act by inhibiting the expression of HIF-1α in macrophages. 23 Kaempferol is the main active ingredient of Fructus Ligustri Lucidi in the Erzhi Pill, which has been observed to exhibit pharmacological activities like anti-bacterium, anti-inflammation, anti-oxidation, and anti-tumor, has been widely confirmed worldwide.24,25 In addition, Kaempferol has been found to increase FOXP3 expression in regulatory T cells (Tregs) and reduce its phosphorylation in position S422 induced by PIM1 and is thus useful for preventing and treating inflammatory disorders, like SLE, ankylosing spondylitis, and rheumatoid arthritis. 26 Kaempferol has also been found to promote insulin and GLP-1 production and inhibit RhoA/Rho Kinase to ameliorate renal fibrosis and damage, thereby delaying the chronic progression of renal pathology. 27 Based on NP, the present study demonstrated that the above-discussed compounds present in the Erzhi Pill may be helpful in the treatment of LN.

The intersection of Erzhi Pill and LN targets, with a total of 87 targets. LN, lupus nephritis.
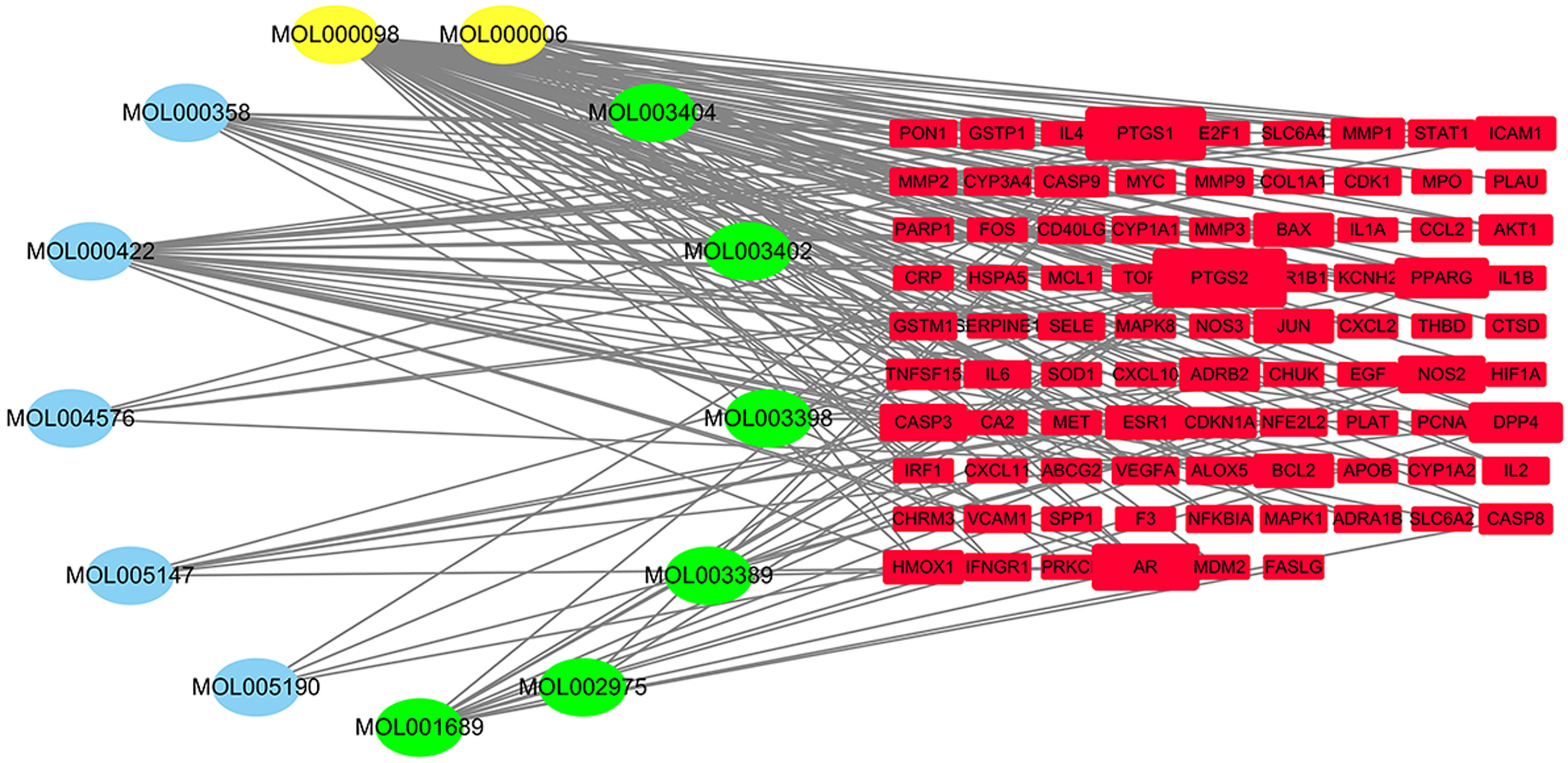
Network diagram of Erzhi Pill-LN-active components-targets. LN, lupus nephritis.
Screening the Key Targets by Network Topology Analysis
The mentioned 87 interaction targets of the Erzhi Pill for LN treatment were imported into the STRING database for building the PPI network, and it contained 1325 edges and 87 nodes. The “cytoNAC” software package was used for the topological analysis twice. Finally, 15 key targets, such as FOS, JUN, MMP9, EGF, CASP3, AKT1, PTGS2, IL6, HIF1A, NFKBIA, IL1B, MMP2, VEGFA, PPARG, and HMOX1, were obtained (Figure 4), all of which are likely to be considered as Erzhi Pill's critical targets for treating LN and worth studying further.
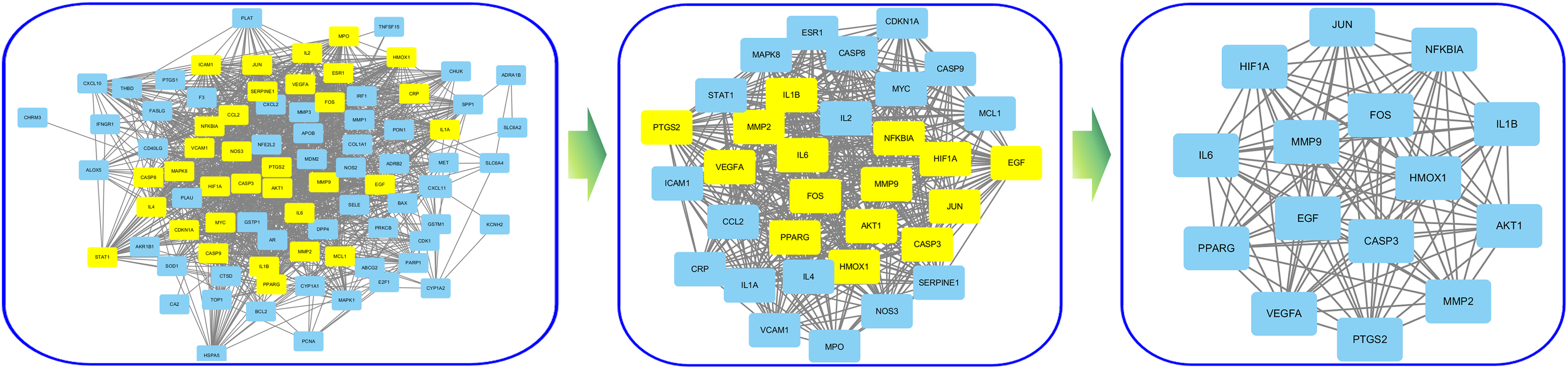
Topological network of Erzhi Pill for the treatment of LN. LN, lupus nephritis.
Validation and Analysis of Key Targets
A set of GSE99967 chips related to LN was found in the GEO database that contained 59 chips, including 17 normal samples and 42 LN samples. The volcanic map (Figure 5a) showed the overall gene expression, where the abscissa represented log2 (Fold Change), and the ordinate represented -log10 (adjusted P-value). One dot stands for one gene, with red dot representing upregulation of differential gene expression, the green dot representing the down-regulation of differential gene expression, and the black dot representing genes with no significant differential expression. There were 576 genes with significant differences, including 429 upregulated genes and 147 downregulated genes. Through the verification of 15 key targets, CASP3 and MMP9 showed significant differences. A box diagram (Figure 5b) was made, with the horizontal coordinate representing the differential genes and the vertical coordinate representing the gene expression level. As shown in the Figure 5b, both the two differential genes were highly expressed in LN. These results suggested that CASP3 and MMP9 may promote the development of LN. Further, CASP3 and MMP9 might serve as the critical core targets in the Erzhi Pill for treating LN. The mesangial cell matrix proliferation and renal basement membrane thickening are significant morphological changes in LN. Mesangial matrix and glomerular basement membrane destruction induced by matrix metalloproteinase (MMP) enzymes is the huge chromatin fragment deposit within renal tissue. 28 The enhanced MMP9 activity is observed in glomeruli within LN patients as well as nephritic mice. 29 As suggested in a study, for mice with lupus-susceptibility (NZBxNZW F1), proteolytic activities markedly elevate during proteinuria, which is possibly associated with MMP9 level. 28 CASP3 is an apoptotic factor that directly participates in the apoptosis of renal cells. 30 Studies have established the pivotal role of CASP3 for modulating renal fibrosis along with apoptosis of microvascular endothelial cells following ischemia-reperfusion injury. 31 Expression of caspase-3 within interstitial inflammatory cells, RTECs, and glomerular parenchymal cells have been reported to be higher in LN tissues compared with controls, suggesting the role of protein metabolism pathway in inducing apoptosis with regard to disease pathogenic mechanism, which has a certain effect on LN development via caspase-3 and MMP. 32

(a) Volcanic map of differential gene expression. (b) Analysis of key target genes CASP3 and MMP9 (**P < .01, ***P < .001).
MD
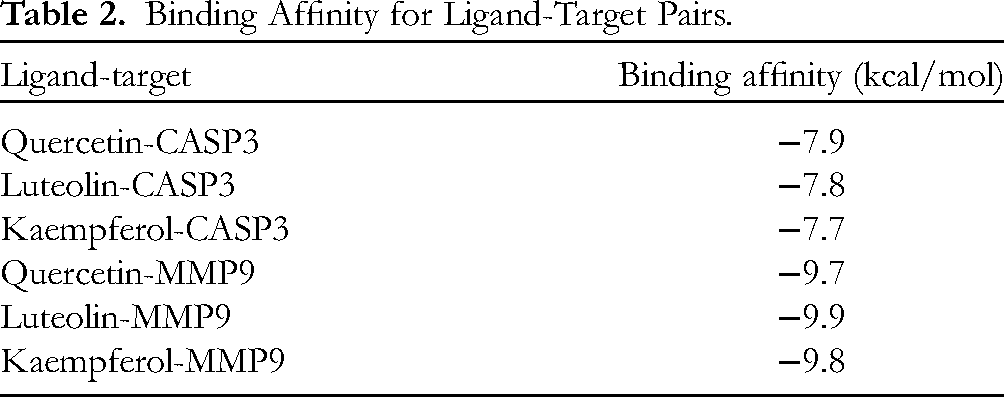
Based on the MD analysis, key compounds quercetin, luteolin, and kaempferol in the Erzhi Pill match well with CASP3 and MMP9 proteins. We predicted the ligand-target-binding affinity (Table 2), as a result, both Luteolin and MMP9 have the highest binding affinity.
Binding Affinity for Ligand-Target Pairs.

By adopting Discovery Studio, this work selected ligand-receptor interactions (Figure 6). We discovered the interaction between quercetin and ARG164 (CASP3 amino acids) via a π-cation; with TYR197 via a π-donor hydrogen bond; with LEU136, VAL266, PRO201, and VAL266 via a π-alkyl; and with GLY125, and ARG164 via a hydrogen bond. Luteolin showed no interaction with TYR195 (CASP3 amino acids) via a π-π stacked; with PRO201, LYS137, and VAL266 via a π-alkyl. Kaempferol interacted with ALA191, ALA244, and ILE187 (CASP3 amino acids) by a π-alkyl; as well as with GLY153, PRO188, and HIS185 via a carbon-hydrogen bond.

The protein-ligand of the 2D docking simulation. The three core compounds (Quercetin, Luteolin, and Kaempferol) of Erzhi Pill are docked with two targets (CASP3 and MMP9).
Quercetin interacted with THR426 (MMP9 amino acid) via a hydrogen bond; with LEU418 via hydrogen bond and π-alkyl; and with ARG424 via π-alkyl and unfavorable donor-donor. Luteolin was found to interact with MMP9 amino acids TYR423 via a π-sigma; with GLU402 via a π-anion; LEU188, and LEU418 via an alkyl bond; and PRO421, ALA417, and ARG424 via a hydrogen bond. Kaempferol interacted with LEU418, and ARG424 (MMP9 amino acids) through a π-alkyl; with THR426 and PRO415 through a hydrogen bond; as well as with TYR423, and HIS401 via a π-π stacked.
This is consistent with the synergistic effect of multiple components of TCM and thereby suggesting that CASP3 and MMP9 might serve as critical targets in Erzhi Pill for treating LN.
Mechanism Analysis of Erzhi Pill in Treating LN
Eight seven critical targets in Erzhi Pill in the treatment of LN were subject to GO analysis. The top five terms (BP, CC, and MF) with P < .05 are shown (Figure 7a). The horizontal axis stands for gene quantity enriched into every item, while longitudinal axis represents function of those enriched genes. Bar color shifting from red to blue is presented based on the increasing P-values, whereas bar length is displayed based on increasing gene quantity. According to GO analysis, for the BP category, the targets were mostly related to lipopolysaccharide (LPS) response, oxidative stress, and molecule of bacterial origin. In CC category, the targets were closely associated with membrane microdomain and membrane raft (MR). In the MF category, the core targets were enriched in signaling receptor activator activity, receptor-ligand activity, and cytokine receptor binding. Earlier studies have shown that NZB/W mice, after repeated LPS exposure, had increased activation of polyclonal B cells, with increased deposition of renal immune complexes, suggesting that LPS has an important effect on LN development. 33 MRs, which refer to the sphingolipid, sterol, as well as glycosphingolipid domains on cell membranes, have a close relation to OS activity. 34 Han et al first provided in vivo and in vitro evidence supporting that MRs-redox pathway had an important effect on fibrosis, inflammation, and renal epithelial-mesenchymal transition. 34 This suggests that LN has a complicated pathogenic mechanism, and the Erzhi Pill may have an effect on the regulation of these BPs.
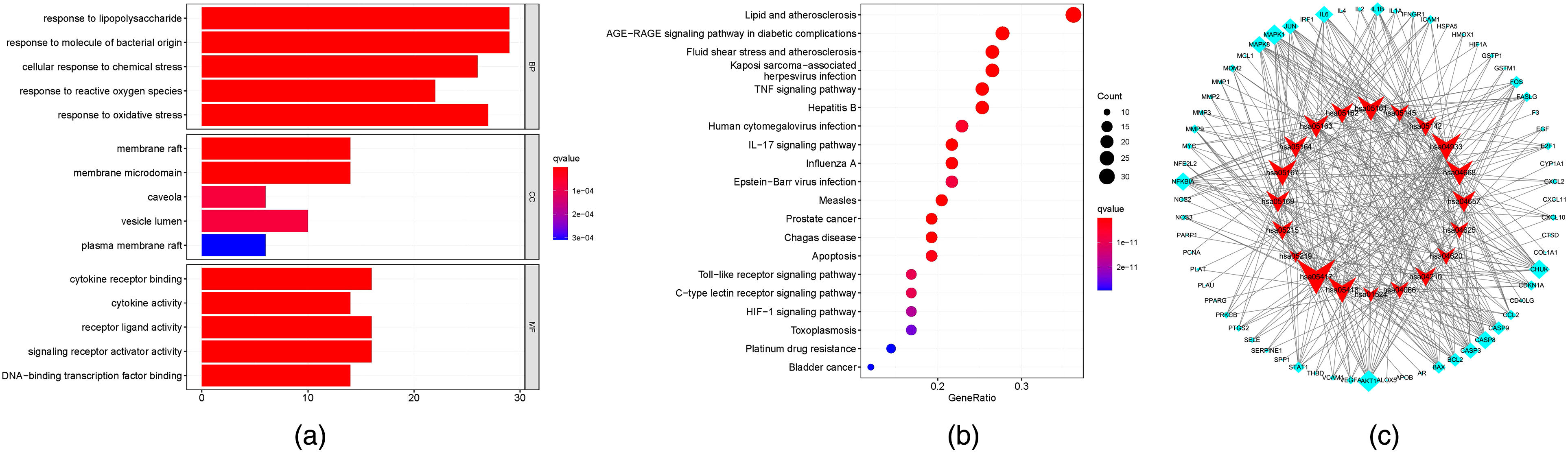
(a) GO enrichment analysis for BP, MF, and CC of Erzhi Pill targets for treating LN. (b) Erzhi Pill treatment of LN key signaling pathway. (c) The KEGG enrichment pathway of Erzhi Pill in LN treatment. The red nodes represent the pathways, and the blue nodes represent the targets involved in the enrichment pathways; the node size reflects the degree value. LN, lupus nephritis; GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes; BP, Biological Process; MF, Molecular Function; CC, Cellular Component.
This work obtained altogether 154 KEGG pathways based on P ≤ .05, of which 20 most significant pathways were screened according to their enrichment number and significance, and a table (Table 3) and bubble map (Figure 7b) were drawn. Signaling pathways included lipid and atherosclerosis (AS), TNF pathway, as well as AGE-RAGE pathway within diabetic complications. The enrichment results of the KEGG signal pathway were then imported into Cytoscape3.7.0, and the network diagram was drawn to show the degree of enrichment of pathways and genes (Figure 7c). The network could indicate the relationship between genes and pathways. The line indicated that the gene belongs to the pathway, while graph size reflecting connection point quantity. Hsa05417 was the pathway that had most genes. Premature AS is identified to be the main concurrent disease of SLE. Inflammation and cholesterol metabolic disturbance are key components related to AS pathological mechanism. 35 Systemic inflammation has been increasingly suggested to disrupt cholesterol homeostasis, which facilitates cholesterol deposition within the arterial wall cells, like endothelium and macrophages. 36 In SLE, OS participates in forming proinflammatory mediators and advanced glycation end-products (AGEs). AGEs can trigger the signaling cascades via their corresponding receptors called RAGE. RAGE polymorphisms are related to the risk of SLE and LN. 37 Based on the unexpected results obtained from AGEs within LN, AGEs possibly have a certain effect on glomerular damage in the context of acute inflammatory glomerulonephritis, which may be achieved through the oxidation role in the glomerular matrix proteins. 38 TNF-α is a key adverse factor that predicts ihypertension as well as renal injury and may enhance OS within renal cortex by activating the NF-κB-mediated pathway. 39 In some in vivo models, treatment with anti-TNF-α drugs has been observed to delay the progression of LN. 40 These discussions further support that the Erzhi Pill works through multiple signaling pathways for the treatment of LN. However, subsequent experiments in vitro and in vivo should be conducted to confirm relevant targets and determine whether these targets are involved in LN.
Pathway of Erzhi Pill for the Treatment of LN.

LN, lupus nephritis.
Conclusion
To sum up, through scientific means integrating NP, preliminary discussion on Erzhi Pill's mechanism in treating LN has been carried on and found that the Erzhi Pill for the treatment of LN involves multiple components that act through multiple targets, and multiple pathways, such as lipid metabolic pathway, oxidative stress, anti-inflammatory pathway, and cellular immune function regulatory pathway. In the present study, the GEO database and MD were used to analyze key targets, data validation and to confirm the feasibility of this study, which provided certain scientific foundation in further discussion of Erzhi Pill's mechanism and a new approach for the treatment of LN.
Footnotes
Author Contributions
Participated in research design: Xue Ru, Xiaolei Chen, and Tao Fan. Contributed analytic tools: Xue Ru, Xingqian Sun, Honglin Lv, and Yi Zhang. Performed data analysis: Xue Ru, Yi Zhang, and Tao Fan. Wrote or contributed to the writing of the manuscript: Xue Ru and Yi Zhang.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Ethical Approval is not applicable for this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by Special Science Research Foundation of Sichuan Administration of traditional Chinese Medicine of China (NO.2021MS373).
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
