Abstract
Objective
The aim of this research was to explore the effects on echinococcosis of harmine, an active ingredient of
Methods
The gene targets were screened out by use of an online database. STRING was exploited to build a protein interaction network model, DAVID to carry out gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses, and AutoDockTools for molecular docking. All mice were randomly divided into 3 groups: a model group, a normal group, and a harmine group. Blood indicators were detected and histological analysis was carried out. NCTC-1469 was treated with either harmine or control. Gene expression was evaluated by Western blot.
Results
Network pharmacological analysis concluded that 6 gene targets (HLA-B, HLA-DQB1, IL-6, tumor necrosis factor [TNF], HLA-A, and F3) might act as the major targets in the treatment of echinococcosis with harmine. GO function and KEGG pathway enrichment analyses showed that harmine might treat echinococcosis mainly by regulating the immune response. Molecular docking confirmed that harmine docked well to the targets and that the interactions were reliable. Harmine could regulate the content of inflammatory factors to modulate the immune response of mice, but caused liver damage. Pathological histology analysis confirmed that harmine had great suppressive efficacy against echinococcosis infection. However, the use of harmine would have a certain negative effect on the spleen of mice. The contents of HLA-B, HLA-DQB1, TNF, HLA-A, and F3 were observably enhanced, while Interleukin-6 was reduced in mouse liver tissue and cells after harmine treatment. As forecast by network pharmacology, harmine exerted antiechinococcosis effects by multiple targets and pathways. Finally, harmine treatment might regulate the Th17/Treg balance in mice with echinococcosis infection.
Conclusions
Harmine displayed great suppressive efficacy against echinococcosis infection by regulating the immune response. These findings suggest that harmine could be a potential therapeutic agent for echinococcosis management by targeting specific gene targets.
Introduction
Echinococcosis is an anthropozoonosis attributed to
Harmine (HM) is a β-carboline alkaloid extracted from the seeds of
In this study, the mechanism of HM on echinococcosis was investigated by network pharmacology.10-12 At the same time, we established a mouse model of echinococcosis and used this for the study of HM. After that, we examined blood indicators, performed pathological histology analysis, and measured the level of these targets. In this paper, we aim to investigate the regulatory mechanisms involved in HM's treatment of echinococcosis and the potential gene targets for its regulation. This study provides a reference for further pharmacological research and clinical management of echinococcosis.
Results
Protein–Protein Interaction Network of HM Targets in Echinococcosis
With the intention of revealing the association between HM (Figure 1A) and echinococcosis, we carried out a network pharmacological analysis. Forty-six gene targets of echinococcosis were obtained from GeneCards. These gene targets were imported into STRING to construct an interaction relationship (Figure 1B). A total of 1673 gene targets of HM were forecast by HERB. The protein–protein interaction (PPI) network relationship is displayed in Figure 1C. Among these, 6 genes were gene targets of both echinococcosis and HM (Figure 1D). The interaction relationship of these common gene targets (HLA-B, HLA-DQB1, interleukin [IL]-6, tumor necrosis factor [TNF], HLA-A, F3) is presented in Figure 1E. Therefore, we speculated that the 6 gene targets (HLA-B, HLA-DQB1, IL-6, TNF, HLA-A, and F3) refer to the major targets in the treatment of echinococcosis with HM.
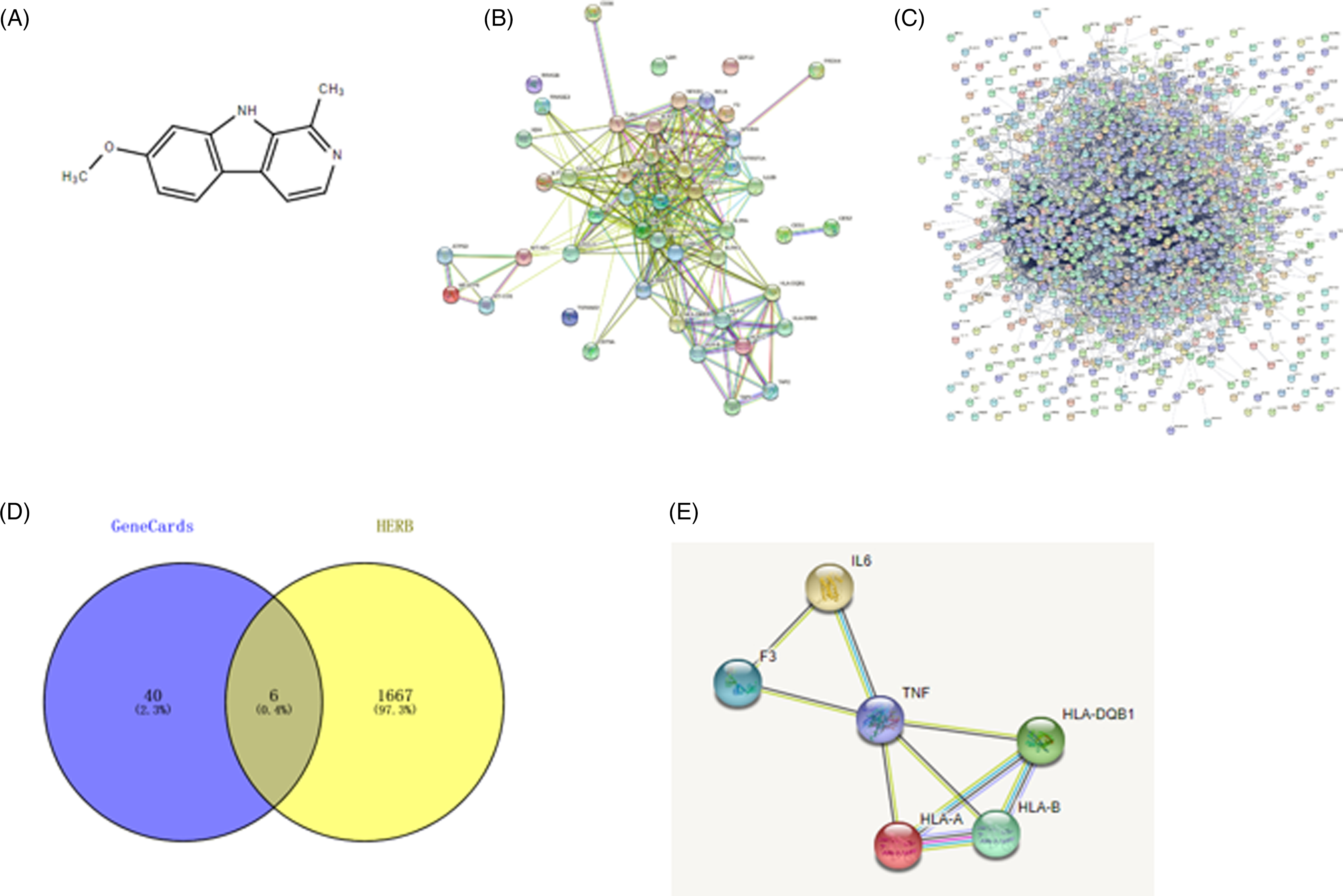
Protein–protein interaction (PPI) network of harmine targets in echinococcosis: (A) the chemical formula of harmine; (B) the echinococcosis targets PPI network; (C) the harmine targets PPI network; (D) Venn diagram of common targets in harmine and echinococcosis; and (E) the common targets PPI network.
Gene Ontology Function and KEGG Pathway Enrichment Analyses
To elucidate the function of the appraised targets, gene ontology (GO) function and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis were implemented. The 6 common targets of HM-echinococcosis were enriched in molecular function (MF), cellular component (CC), and biological process (BP); the GO terms are displayed in Figure 2. As shown from the results, the targets of HM-echinococcosis mainly participated in antigen binding, cytokine activity, antigen processing and presentation, and humoral immune response, which are closely associated with the body's immune response. The DAVID web server was utilized to implement KEGG pathway enrichment analysis; the signal pathways are presented in Figure 3. HM might play management effects on echinococcosis by modulating signaling pathways, which mainly involve graft-versus-host disease, allograft rejection, antigen processing, and presentation. According to the above results, we speculated that HM might treat echinococcosis mainly by regulating the immune response.
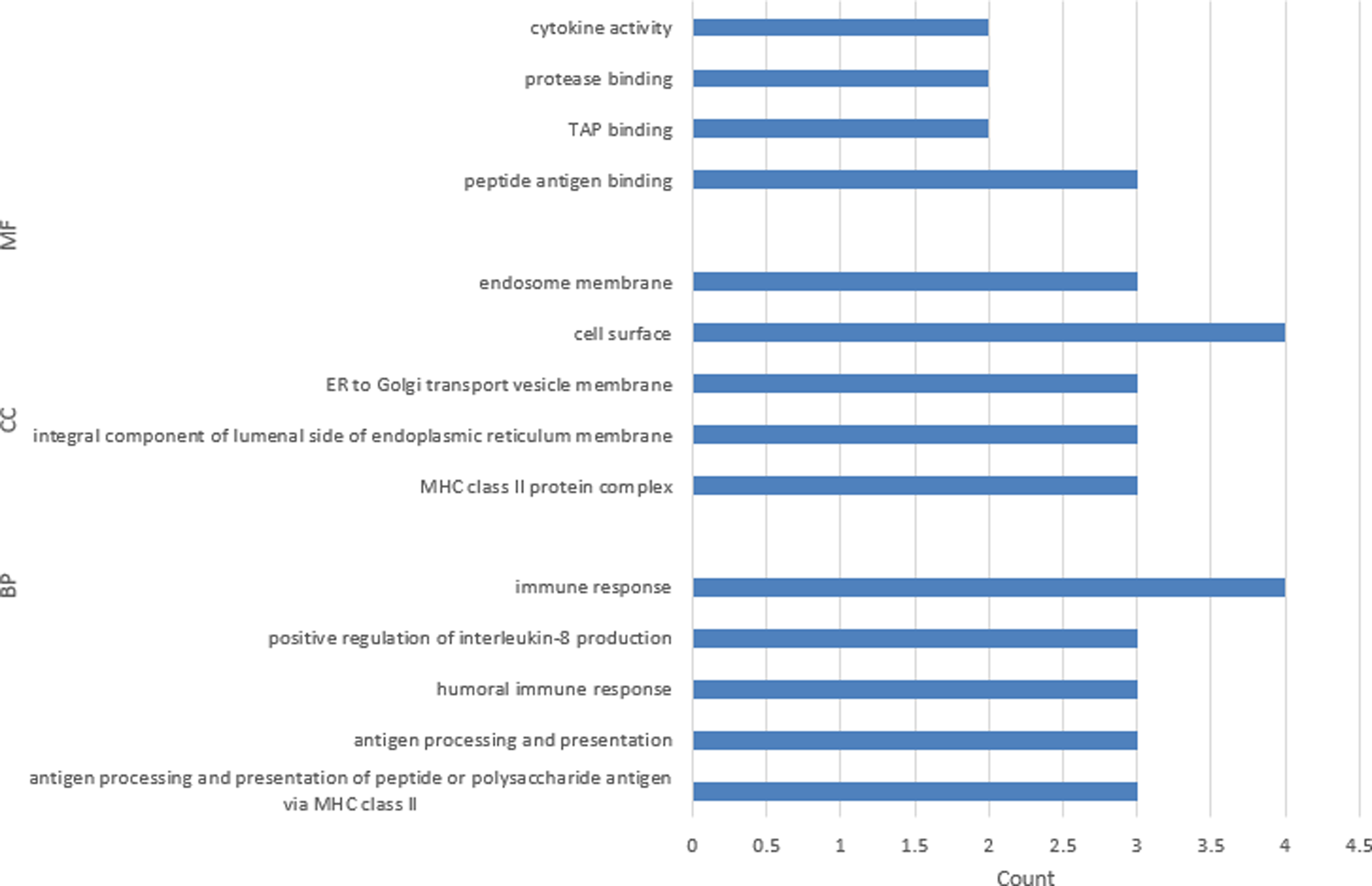
Gene ontology (GO) analysis for the common targets of harmine and echinococcosis.

Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis for the common targets of harmine and echinococcosis.
Molecular Docking Analysis
We further scrutinized the interactions between HM and targets. The results of molecular docking confirmed that HM docked well to HLA-B, HLA-DQB1, IL-6, TNF, HLA-A, and F3 proteins, and the interactions were dependable. The specific binding affinity is displayed in Table 1. The conformations of HM and targets are presented in Figure 4.
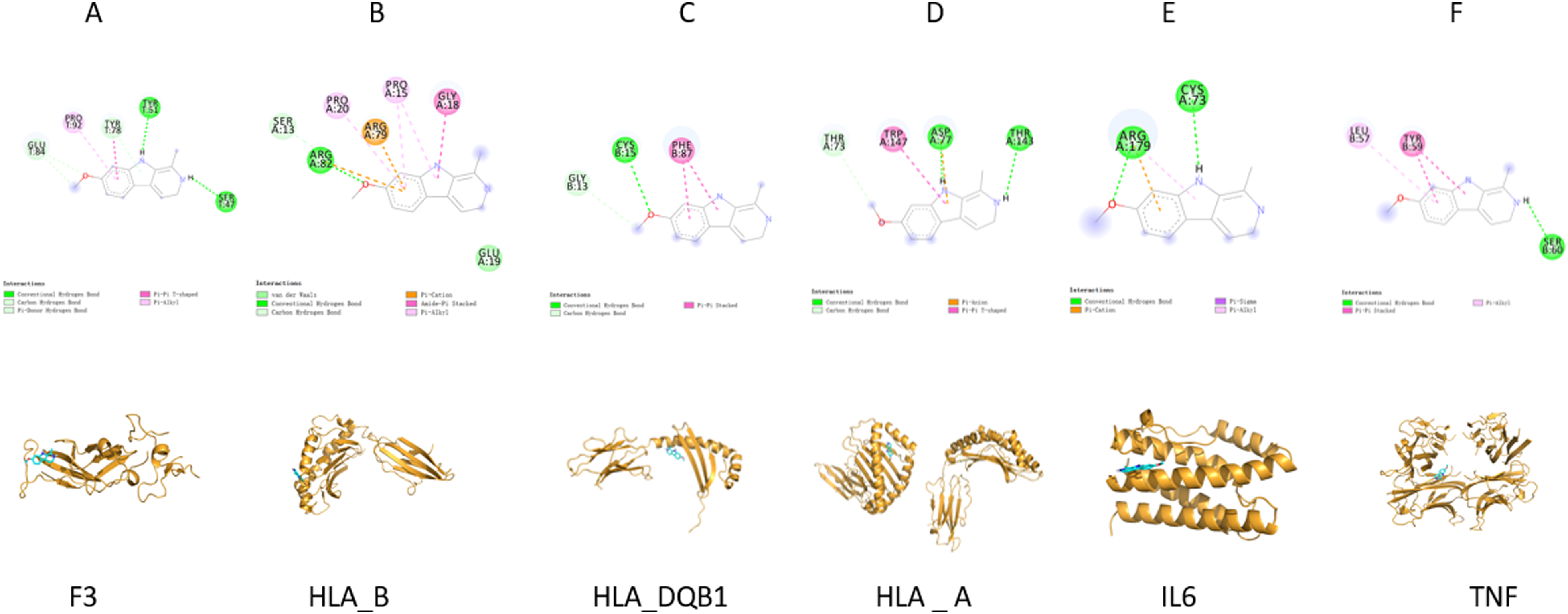
Conformations of harmine and targets.
Docking Results Between Harmine and Target Genes.
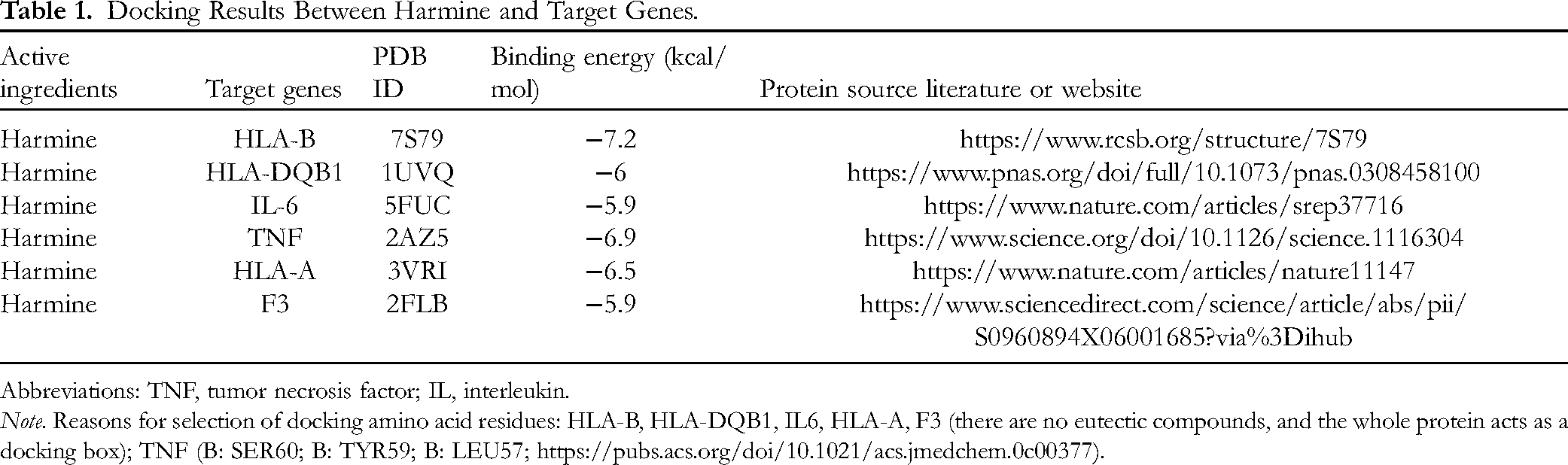

Abbreviations: TNF, tumor necrosis factor; IL, interleukin.
Blood Indicators Examination
The contents of IL-2 (Figure 5A) and IL-10 (Figure 5C) in the HM group were observably enhanced in comparison with those in the model group, whereas no apparent difference was found when compared with the normal group. IL-6 (Figure 5B) and immunoglobulin E (IgE; Figure 5D) levels in the HM group were apparently reduced compared to those in the model group, but there was no remarkable difference with respect to the normal group. Additionally, TNF-α abundance among the three groups presented no obvious difference (Figure 5E). The specific values are shown in Table 2. The aspartate transaminase (AST) content was improved in the HM and model groups versus the normal group (Figure 5F). It is clear that HM triggered hepatic damage. As for alanine transaminase (ALT) abundance, there was no difference between the 3 groups (Figure 5G). Hence, we speculated that HM could modulate the content of inflammatory factors to regulate the immune response of mice, but there was some damage to the liver.
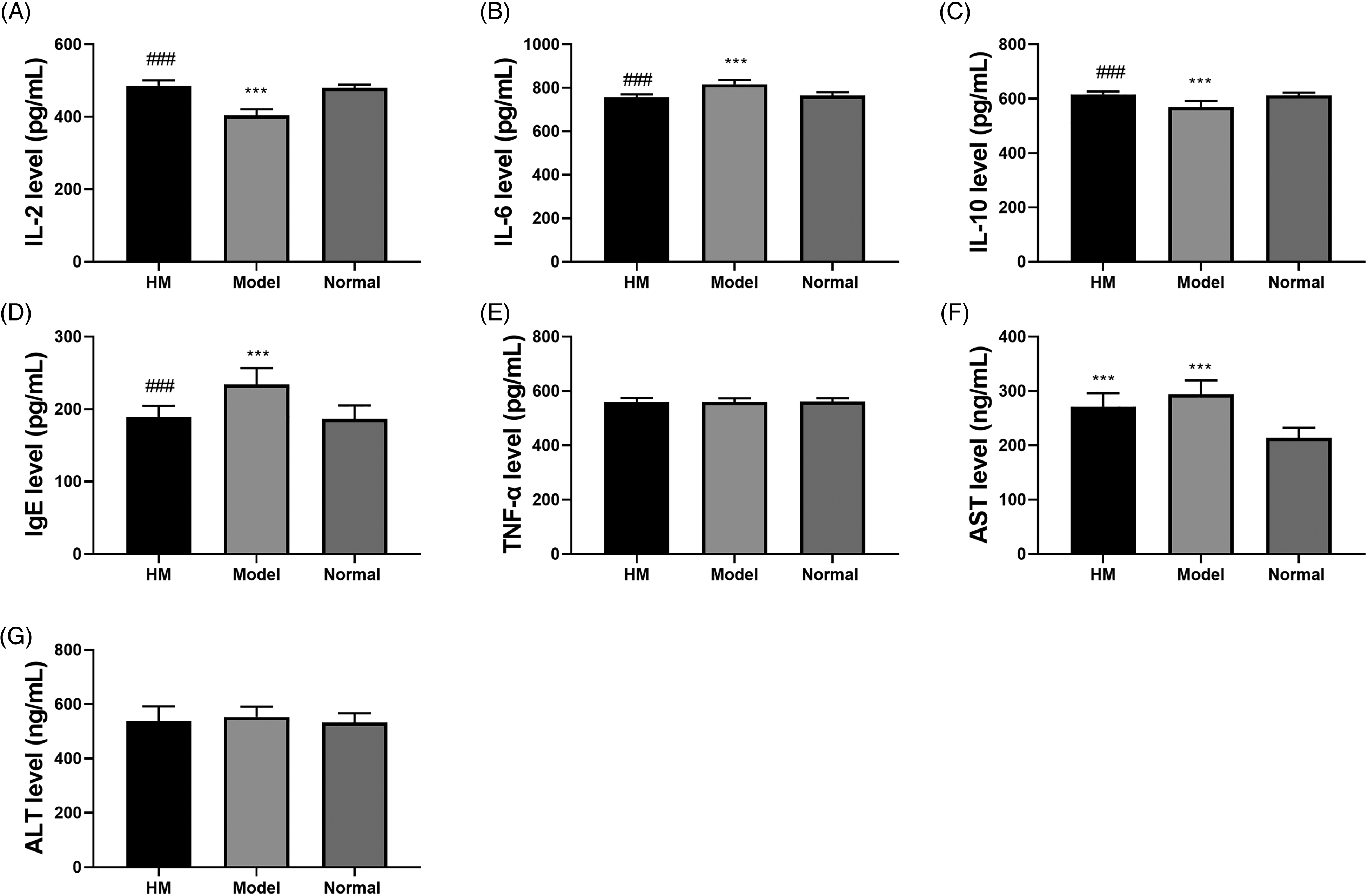
Examination of blood indicators. (A–G) The levels of IL-2, IL-6, IL-10, IgE, TNF-α, AST, and ALT in serum were detected by ELISA kits. ∗ Compared with the normal group; # compared with the model group; ∗∗∗
Level of Immunological Factors in mouse Serum.

Abbreviations: IL, interleukin; IgE, immunoglobulin E; TNF, tumor necrosis factor; HM, harmine.
∗Compared with normal group; #compared with model group; *
Pathological Histology Analysis
In this part, we inspected histological changes in hydatid cysts. As revealed in Figure 6B, the hydatid tissues in the model group were well developed. In the HM group, the structure of the brood capsule was shrinking and collapsing (Figure 6A), representing the management effect of HM. Besides, we scrutinized histological changes in liver and spleen tissues. A few depositions were found in both liver (Figure 6C) and spleen tissue (Figure 6F) of the HM group. In addition, in the model groups, lots of monocytes had filtered into the liver tissue (Figure 6D) and some polykaryocytes into the spleen tissue (Figure 6G). The liver and spleen tissues of the normal group are displayed in Figure 6E and H. The thymus index of the 3 groups presented no significant difference (Figure 6I) whereas the spleen index was boosted in the HM group versus the model and normal groups (Figure 6J). As displayed in Figure 6K, HM treatment markedly decreased cyst weight and enhanced the cyst inhibition rate versus the model group. The results of pathological histology analysis confirmed that HM had great suppressive efficacy against echinococcosis infection. However, the use of HM will have a certain deleterious effect on the spleen of mice.
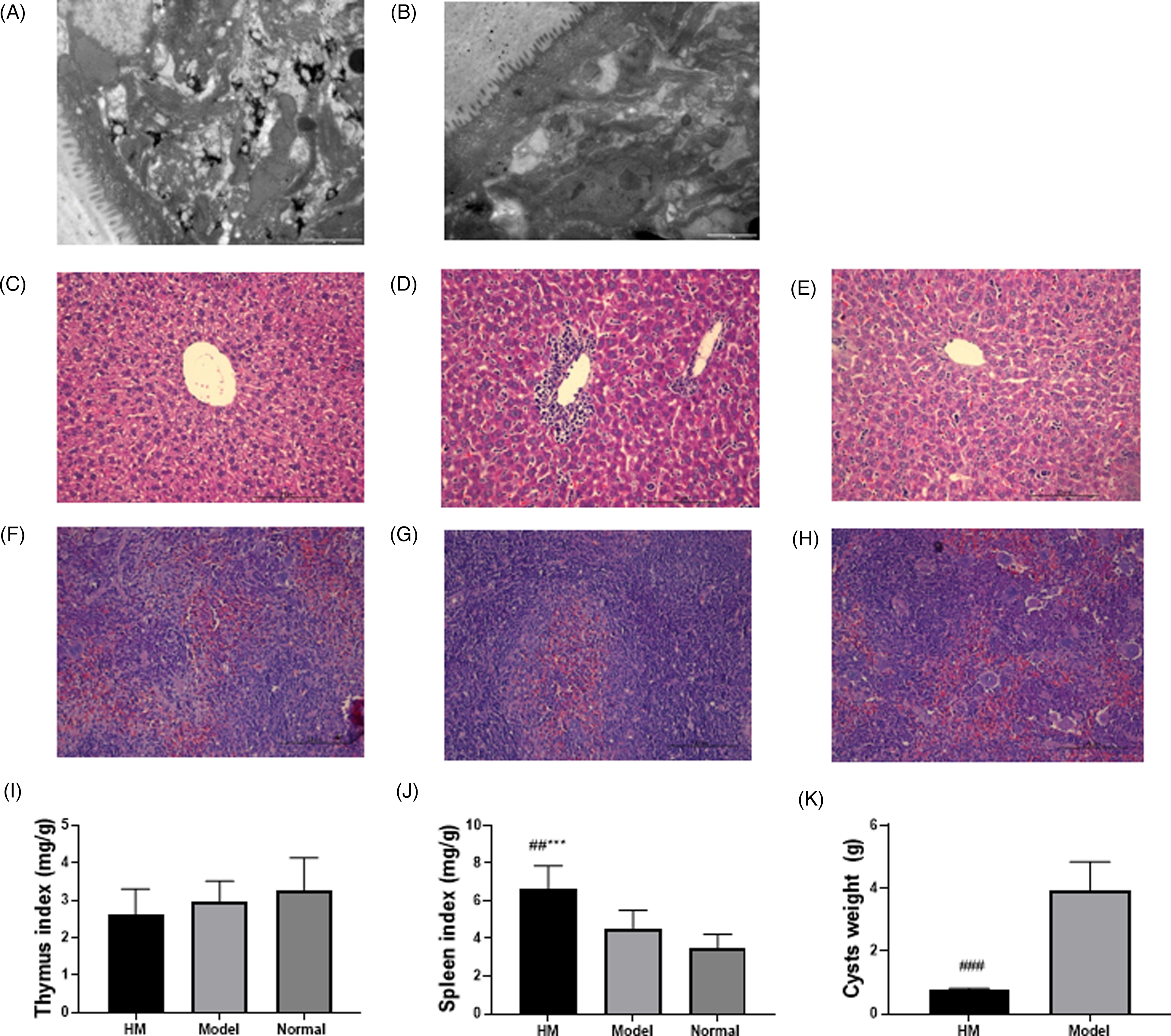
Pathologic histology analysis: (A) TEM images of metacestodes in the HM group; (B) TEM images of metacestodes in the model group. The scale bar was 2 μm. (C–E) HE stained sections of liver tissue in HM, model, and normal groups. (F–H) HE stained sections of spleen tissue in the HM, model, and normal groups. The scale bar was 10 μm. (I–K) The thymus index, spleen index, and cyst weights were detected. ∗ compared with normal group; # compared with model group; ***
Determination of Target Protein Levels in Rats
Lastly, we researched the effect of HM on target protein levels in mouse liver tissues. There was no obvious difference in the protein contents of HLA-B, HLA-DQB1, IL-6, TNF, HLA-A, and F3 between the model and normal groups (Figure 7). The contents of HLA-B (Figure 7A and B), HLA-DQB1 (Figure 7A and C), TNF (Figure 7A and E), HLA-A (Figure 7A and F), and F3 (Figure 7A and G) were observably enhanced, while IL-6 (Figure 7A and D) was reduced in the HM group in comparison with the model and normal groups. Then, we assessed the target protein levels in NCTC-1469 cells. The levels of HLA-B (Figure 8A and B), HLA-DQB1 (Figure 8A and C), TNF (Figure 8A and E), HLA-A (Figure 8A and F), and F3 (Figure 8A and G) were apparently boosted, but IL-6 (Figure 8A and D) was reduced in the HM group in contrast to the control group.
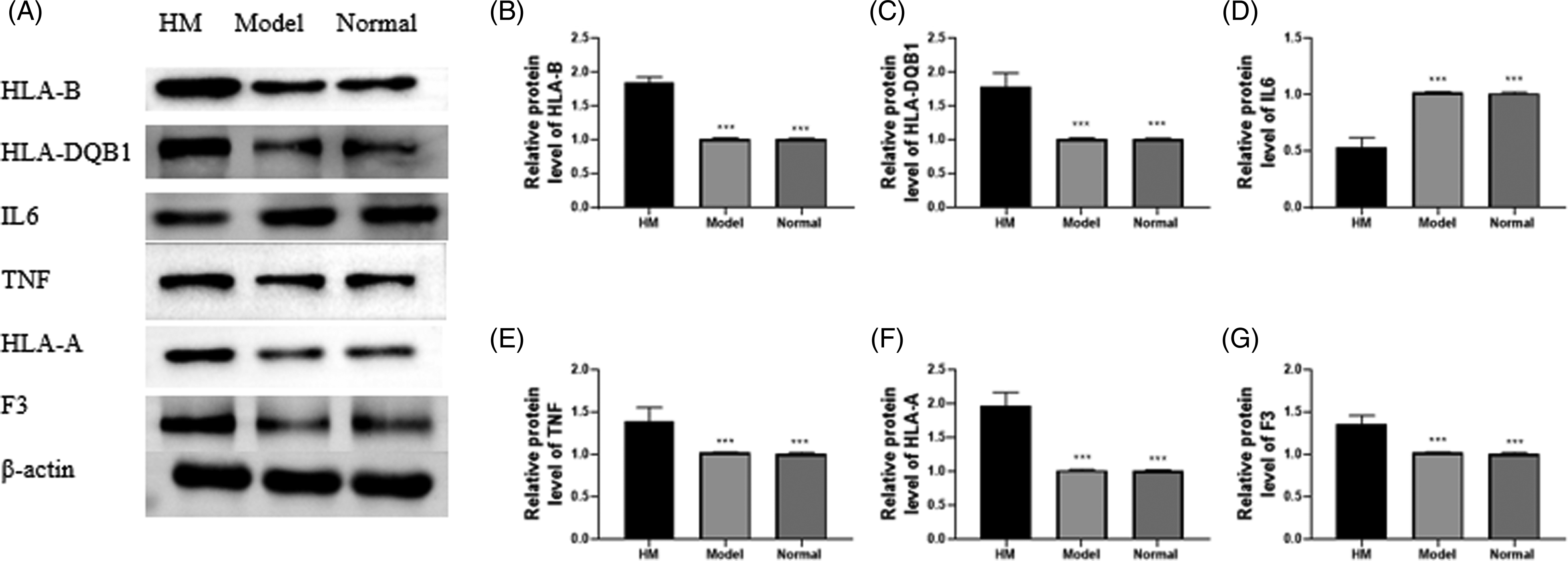
Determination of target protein levels in mice. (A–G) The protein levels of HLA-B, HLA-DQB1, IL-6, TNF, HLA-A, and F3 in liver tissues were detected by Western blot. *Compared with HM group; ***
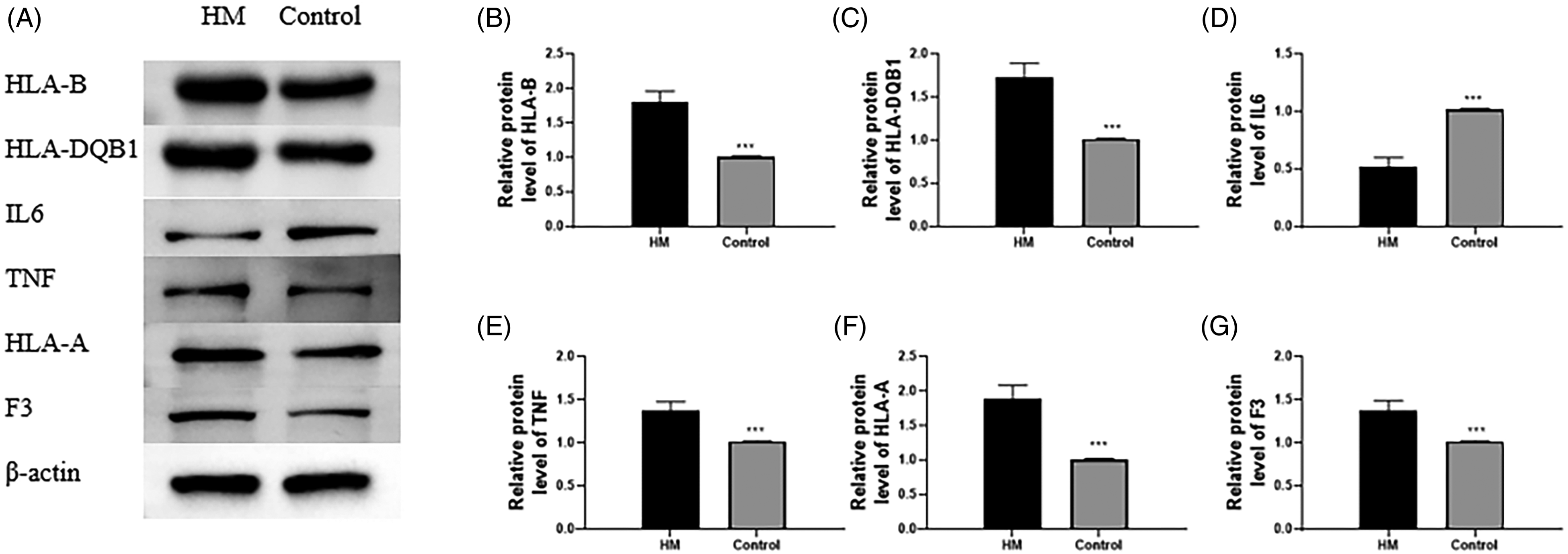
Determination of target protein levels in cells. (A–G) The protein levels of HLA-B, HLA-DQB1, IL-6, TNF, HLA-A, and F3 in liver cells were assessed by Western blot. *Compared with the HM group; ***
HM Treatment Regulated Th17/Treg Balance
The levels of Th17 cell markers (IL-17 and RAR-related orphan receptor alpha [RORα]) and Treg cell markers (Foxp3 and IL-10) in the liver tissues of rats in each group were detected by enzyme-linked immunosorbent assay (ELISA). We found that the IL-17 and RORα levels were enhanced in the model group, but reduced in the HM group (Figure 9A and B). However, the Foxp3 and IL-10 levels declined in the model group, but were boosted in the HM group (Figure 9A and B). Therefore, we speculated that HM treatment might promote the differentiation of Treg cells and inhibit the differentiation of Th17 cells.
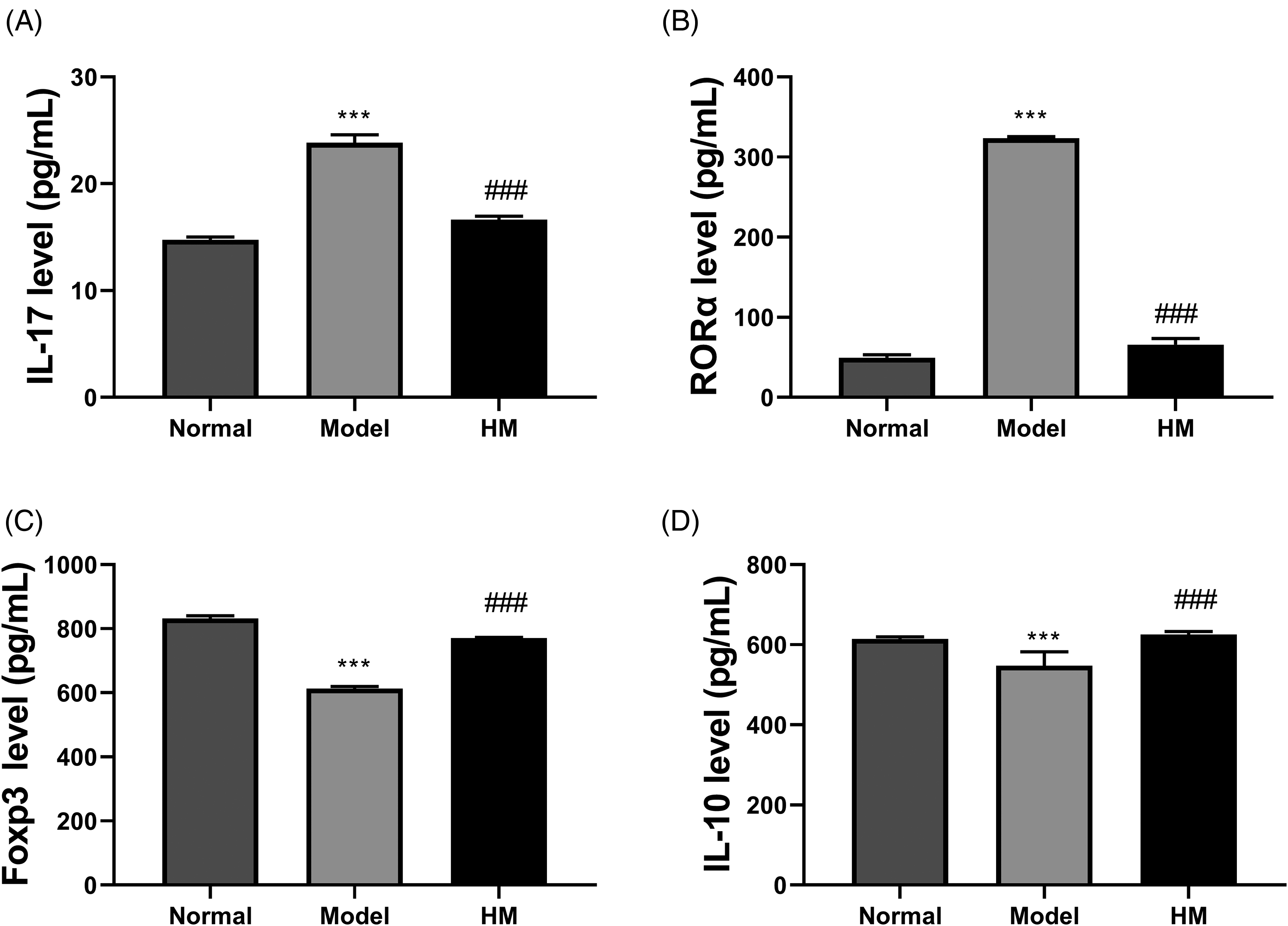
Examination of Th17/Treg indicators. (A–D) The levels of IL-17, RORα, Foxp3, and IL-10 in liver tissues were detected by ELISA kits. *Compared with normal group; #compared with model group; ***
Discussion
Echinococcosis is a serious zoonotic parasitic disease caused by
In current clinical practice, albendazole and mebendazole are the first choices for the management of echinococcosis, but they are relatively toxic and can cause serious liver damage to patients.
16
Traditional Chinese medicine is considered a safe and effective complementary and alternative therapy for various diseases. For instance, ginsenoside Rg1 ameliorated blood–brain barrier disruption and traumatic brain injury.
17
Schisandra has been linked to immune inflammation and may have therapeutic potential against atherosclerosis.
18
Besides, compound Fengshiding capsule and liquiritin have been found to be effective drugs for rheumatoid arthritis.19,20 Gong et al
9
showed that HM could be active against
In the current research, we carried out network pharmacology analysis to explore the link between HM, gene targets, and signaling pathways. In this study, the conclusions of network pharmacological analysis exposed that 6 gene targets (HLA-B, HLA-DQB1, IL-6, TNF, HLA-A, and F3) might act as the major targets in the HM treatment of echinococcosis. Some studies have shown that HLA-B, HLA-DQB1, and HLA-A were linked with the resistance and susceptibility to echinococcus in humans.23-25 Besides, IL-6 and TNF are related to the immune evasion of echinococcus infection. 26 Moreover, F3 is closely related to inflammatory bowel disease. 27 These conclusions indicate that our prediction results have a certain significance. The results of enrichment analyses suggest that HM may treat echinococcosis mainly by regulating the immune response. Studies have shown that a binding affinity lower than −5.0 kcal/mol signified that the validations have stable interactions. 28 The results of molecular docking confirmed that HM docked well to the targets and that the interactions were reliable. We speculate that HM may exert therapeutic effects on echinococcus by modulating these gene targets and signaling pathways.
In vivo experiments were implemented, thereby revealing the potential mechanisms of HM. HM regulated the content of inflammatory factors to regulate the immune response of mice, but it might also cause some damage to their livers. Jiang et al 29 revealed that HM could curb the inflammatory response and be used to treat inflammatory diseases, which was consistent with our findings. The results of pathological histology analysis confirmed that HM had great suppressive efficacy against echinococcosis infection. However, the use of HM had a certain effect on the spleen of the mice. After HM treatment, the contents of HLA-B, HLA-DQB1, TNF, HLA-A, and F3 in mouse liver tissue and cells were observably enhanced, while IL-6 was reduced. Among the body's immune cells, CD4 T-helper lymphocytes play an important role in regulating immune response and inflammatory diseases. 30 Th17 cells can stimulate a variety of cells, gather neutrophils to the site of infection, and mediate immune responses against extracellular bacteria and fungi. Treg cells suppress the immune response to maintain immune homeostasis. 31 We found that HM treatment might promote the differentiation of Treg cells and inhibit the differentiation of Th17 cells in mouse liver tissues. The result indicates that HM has a good anti-inflammatory effect. The next step of this study will be to continue to study how to reduce the damage to the liver and spleen caused by HM in animals. However, there are still some limitations to this paper. The study was only completed in a mouse model, which needs to be further verified in clinical work.
Conclusion
Here we exposed the antiechinococcosis function of HM and confirmed that it exerted antiechinococcosis effects by targeting multiple genes (HLA-B, HLA-DQB1, IL-6, TNF, HLA-A, and F3) and pathways. Our conclusions suggest that HM has great suppressive efficacy against echinococcosis infection by regulating the immune response. This research implies that HM could have a helpful influence on the management of echinococcosis and provides a theoretical basis for follow-up studies.
Materials and Methods
Target Screening
Gene targets for the management of echinococcosis were predicted by using “echinococcosis” as a keyword in GeneCards (http://www.genecards.org/). HERB (http://herb.ac.cn/) was employed to predict the pharmacological targets of HM. The above targets were investigated by drawing a Venn diagram to determine the overlapping targets. These overlapping gene targets generated a list of possible targets of HM for treating echinococcosis.
PPI Network Construction
The PPI network serves a vital role in uncovering cell function and biological processes.
32
The authentication of PPI is helpful for a good understanding of the molecular mechanism of HM in the therapy of echinococcosis. STRING (https://string-db.org) was utilized to build a PPI network model.
33
The specific steps were based on previous articles.
34
First of all,
Gene Ontology and Kyoto Encyclopedia of Genes and Genomes Enrichment Analyses
DAVID (https://david.ncifcrf.gov/summary.jsp) was employed to carry out GO and KEGG enrichment analyses for the overlapped gene targets.35,36 The overlapped gene targets of HM for treating echinococcosis were imported into the Metascape platform, followed by enrichment. The GO function, comprising BP, MF, and CC, was enriched by setting the
Molecular Docking Confirmation
Molecular docking technology was employed to confirm the reliability of the relationship between HM and related targets and to explore its binding mode. Protein crystal structures were downloaded from PDB (https://www.rcsb.org/), and AutoDock Vina software (http://vina.scripps.edu/) was used to prepare molecular docking ligands from PubChem (https://pubchem.ncbi.nlm.nih.gov/) and protein. 35 The crystal structure of the target protein was pretreated, including removal of hydrogenation, modification of amino acids, optimization of energy, adjustment of force field parameters, and then meeting the low energy conformation of the ligand structure. Finally, molecular docking was performed between the target structure and the active component structure using Vina in pyrx software. The affinity (kcal/mol) value represented the binding ability of the two. The lower the binding ability, the more stable the binding between the ligand and the receptor. Visualization analysis was performed using Pymol and Discovery Studio 2020 Client for 2-dimensional drawings.
Animals
Thirty kunming white mice (18-20 g, female, 10-week-old) were purchased from the Experimental Animal Center of Xinjiang Medical University. These mice were fed in standard cages (5 mice per cage) for 1 week under standard conditions (20 ± 2 °C; 50%–60% relative humidity; 12 h light/dark cycle), with free access to food and water before the experiment. This research was permitted by the Animal Ethics Committee of the First Affiliated Hospital of Xinjiang Medical University (Ethical approval No. K202202-08).
Animal Treatment
All 30 mice were randomly divided into 3 groups (
Blood Indicator Determinations
The treated mice were anesthetized and sacrificed, and their blood was collected to obtain serum after centrifugation. The liver tissues were also collected. ELISA kits were employed to assess the IL-2, IL-6, IL-10, IgE, TNF-α, IL-17, RORα, and Foxp3 contents in either serum or liver tissues, including IL-2 (ab221834; Abcam), IL-6 (ab178013; Abcam), IL-10 (ab214566; Abcam), IgE (ab157736; Abcam), TNF-α (ab181421; Abcam), IL-17 (ab214028; Abcam), RORα (RS-10076; MengWei), and Foxp3 (ab289645; Abcam). Besides, aspartate transaminase (AST) and ALT levels in serum were measured by AST (ab263883; Abcam) and ALT (ab234579; Abcam) ELISA kits.
Pathological Histology Analysis
The weight of the hydatid cysts, spleen, thymus, and mouse body was examined. The cysts' inhibition rate, spleen index, and thymus index were measured. The specific formulas were as follows: the cysts inhibition rate = (cysts weight in model group − cysts weight in HM group)/cysts weight in model group × 100%; the spleen index = (spleen weight/body weight) × 10; the thymus index = (thymus weight/body weight) × 10.
Hematoxylin-Eosin Staining Assay
The treated rats were anesthetized and sacrificed, and their liver and spleen were collected. The liver and spleen tissues were fixed with 10% paraformaldehyde (Sigma-Aldrich) for 48 h, then embedded in paraffin (Sigma-Aldrich), and 5 μm thick paraffin slices were prepared. These paraffin slices were treated with a hematoxylin–eosin (HE) staining kit (Sigma-Aldrich), and the morphological alterations were detected through a microscope (Leica).
Transmission Electron Microscopy Analysis
The treated mice were anesthetized, sacrificed, and the metacestode vesicles were removed and fixed in glutaraldehyde (2.5%, Sigma-Aldrich) for 2 h. Next, the samples were exposed to osmium tetroxide (2%, Sigma-Aldrich) for 2 h. After dehydration, the samples were embedded. After staining the sections, the samples were observed through a JEM-1400 plus TEM (JEOL).
Cell Cultures and Treatment
Mouse normal liver cell line NCTC-1469 (China Center for Type Culture Collection) was selected for this study. NCTC-1469 cells were grown in RPMI-1640 medium (Sigma-Aldrich) containing 15% fetal bovine serum (Invitrogen) and 2% penicillin/streptomycin (Invitrogen) at 37°C with 5% CO2. Then, the NCTC-1469 cells were divided into 2 groups, a control group and an HM group. The solubility of HM in water is very low, but it was dissolved in dimethyl sulfoxide (DMSO). 37 Therefore, DMSO was used as the solvent in this experiment. The cells of the HM group were treated with 40 μg/mL HM for 24 h, and the cells of the control group with an equivalent volume of DMSO (Sigma-Aldrich) for 24 h. These cells were used for subsequent experiments.
Western Blot
The tissues and cells were lysed by exploiting RIPA buffer (Sigma-Aldrich), and the quality of protein was scrutinized using a BCA kit (Sigma-Aldrich). Afterward, the proteins were separated on a 12% sodium dodecyl sulfate-polyacrylamide gel electrophoresis gel and moved to polyvinylidene fluoride membranes (Sigma-Aldrich). After blocking, these membranes were incubated overnight with primary antibodies at 4 °C. The antibodies used were as follow: anti-HLA-B (SAB1405937; 1:1000; Sigma-Aldrich), anti-HLA-DQB1 (ab200733; 1:1000; Abcam), anti-IL-6 (ab9324; 1:2000; Abcam), anti-TNF (ab1793; 1:1000; Abcam), anti-HLA-A (SAB1405936; 1:1000; Sigma-Aldrich), anti-F3 (SAB1410355; 1:1000; Sigma-Aldrich), and anti-β-actin (ab179467; 1:1000; Abcam). Successively, these membranes were treated with goat anti-rabbit immunoglobulin G (IgG; ab6733; 1:2500; Abcam) for 1 h. The protein blots were observed exploiting an ECL-Plus reagent (Invitrogen).
Statistical Assay
The statistical analysis utilized SPSS 22.0. Group differences were scrutinized by Student's
Supplemental Material
sj-rar-1-npx-10.1177_1934578X231182905 - Supplemental material for Effect and Mechanism of Harmine on Echinococcosis Based on Network Pharmacology
Supplemental material, sj-rar-1-npx-10.1177_1934578X231182905 for Effect and Mechanism of Harmine on Echinococcosis Based on Network Pharmacology by Bei Chen, Jun Zhao, Liang Teng, Yimaierjiang Buhuliqianmu, Zhaohui Xu, Yuehong Gong and Huijing Gao in Natural Product Communications
Footnotes
Acknowledgement
None
Supporting Information
The unclipped protein gel images in Figures 7 and ![]() , the Animal Ethics Committee approval document, and the Certificate of Editing are included in the supplementary file.
, the Animal Ethics Committee approval document, and the Certificate of Editing are included in the supplementary file.
Data Availability Statement
The data used to support the findings of this study are available from the corresponding author upon request.
Author Contributions
BC, JZ, and LT designed research studies. BC, JZ, LT, and YB conducted the experiments. ZX, YG, and HG carried out data analysis and provided reagents. All the authors took part in manuscript preparation and approved the final manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by The National Natural Science Foundation of China (no. 82260722), State Key Laboratory of Pathogenesis, Prevention, and Treatment of High Incidence Diseases in Central Asia Fund (Nos. SKL-HIDCA-2022-26, SKL-HIDCA-2022-BC3 and SKL-HIDCA-2023-YX1), The National Natural Science Foundation of China (No. 82160700) and Key R&D projects in Xinjiang Uygur Autonomous Region(2022B03013-4).
Ethical Approval
This research was permitted by the Animal Ethics Committee of the First Affiliated Hospital of Xinjiang Medical University (Ethical approval No. K202202-08).
Statement of Human and Animal Rights
All procedures in this study were conducted in accordance with the Animal Ethics Committee of the First Affiliated Hospital of Xinjiang Medical University (Ethical approval No. K202202-08) approved protocols.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
