Abstract
Berberine (BBR), an isoquinoline alkaloid, possesses multiply pharmacological effects as a potential therapeutic drug for antipsychotic-induced metabolic syndrome (MetS). However, the underlying therapeutic mechanisms have not been fully elucidated. In this study, we aim to investigate the possible mechanisms by identifying the key targets and biological pathways through network pharmacology and molecular docking analysis. A total of 23 overlapping targets in the intersection set among BBR, atypical antipsychotic drugs (AADs), and MetS were determined. PPI network analysis showed that 22 out of the 23 overlapping targets closely interacted with the others. The following pathway enrichment analysis and molecular docking indicated a central role of peroxisome proliferator-activated receptor-γ (PPARG) as the key target of BBR against AAD-induced MetS by acting on the PPAR signaling pathway, lipid and atherosclerosis, and AMP activated protein kinase (AMPK) signaling pathway. In addition, cytochrome P-450 2D6 (CYP2D6) could be considered as another target of BBR in ameliorating antipsychotic-induced metabolic side effects. Collectively, this study investigated the central targets and biological pathways of BBR against AAD-induced MetS from a systematic perspective, and thus brings novel insights into further understanding of the protective effects of BBR.
Keywords
Introduction
Antipsychotics, widely used for treating schizophrenia, include “typical” and “atypical” antipsychotics. Although the atypical antipsychotic drugs (AADs) are often preferred to typical antipsychotics due to the reduced risk of extrapyramidal symptoms, some AADs are associated with metabolic adverse effects, such as weight gain, impaired glucose metabolism, dyslipidemia, and cardiovascular complications.1–4 Clinical studies found that patients with schizophrenia and related psychotic disorders were more susceptible to metabolic syndrome (MetS), which could be further aggravated by some atypical antipsychotics. 5 Among AADs, clozapine and olanzapine are associated with the highest risk of treatment-related metabolic dysfunction,6,7 with a mean weight gain of 4.45 kg after 10-week clozapine use and 4.15 kg weight increase, as well as a doubled risk for MetS, with olanzapine treatment.8,9
Therapeutic guidelines for the management of antipsychotic-induced MetS include: (a) lifestyle interventions, such as diet control and exercise, (b) substitute antipsychotics with alternative drugs with lower metabolic risk, and (c) supplying additional pharmacological interventions in metabolism, such as metformin. 10 However, the disappointing outcome of lifestyle interventions limited the clinical application in patients with schizophrenia-spectrum disorders.11,12 In light of its favorable therapeutic effects, clozapine as one of the high metabolic risk AADs, plays an indispensable role in the treatment-resistant schizophrenia. 13 Therefore, pharmacological interventions are considered to be a promising therapy. Although metformin presented prospective effectiveness in ameliorating antipsychotic-induced weight gain and insulin resistance,14,15 there was insufficient evidence to recommend a broad promotion to clinical usage, probably considering its fatal side effect of lactic acidosis and gastrointestinal damage.16,17 Thus, it is urgently warranted to explore more safe and effective treatments to ameliorate AAD-induced MetS and to promote life expectancy in schizophrenia.
Traditional Chinese Medicine (TCM) has recently attracted worldwide attention, with lines of evidence suggesting its modulatory effects and treatment advantages for chronic complex diseases. 18 Berberine (BBR), an isoquinoline alkaloid of the protoberberine type, is isolated from several plants and possesses the advantages of low-cost and negligible side effects. Recent research has demonstrated that treatment with BBR could reduce the negative symptoms in patients with schizophrenia. 19 In addition, evidence from animal and clinical studies has revealed the therapeutic potential of BBR for antipsychotic-induced MetS.20,21 However, the mechanisms responsible for the therapeutic effect of BBR against AAD-induced MetS have not yet been fully illuminated and need to be further explored. Therefore, the present study aims to investigate the possible mechanisms by identifying the key targets and biological pathways through network pharmacology and molecular docking analysis.
Methods
Screening Potential Targets of BBR
The potential targets of BBR were screened out through the database analysis of Traditional Chinese medicine system pharmacology platform (TCMSP) (https://old.tcmsp-e.com/tcmsp.php), Encyclopedia of Traditional Chinese Medicine (ETCM) (http://www.tcmip.cn/ETCM/), PharmMapper (http://lilab-ecust.cn/pharmmapper/index.html), and Swiss Target Prediction (STP) (http://swisstargetprediction.ch/).
Screening Targets of AADs
GeneCards database (https://www.genecards.org/) was used to explore the targets of AADs. Given that clozapine and olanzapine are associated with the highest risk of treatment-related metabolic dysfunction, quetiapine and risperidone with intermediate risk, and ziprasidone and aripiprazole with the least risk, we performed a thorough screening with the keywords of “Clozapine,” “Olanzapine,” “Quetiapine,” and “Risperidone.” Optimal gene targets with relevance scores exceeding the median value were screened.
Screening MetS-Associated Targets
We searched the database GeneCards, DisGeNET, CTD, and PharmGKB to identify targets associated with MetS by using “metabolic syndrome” as a keyword.
GeneCards (https://www.genecards.org/): the selection filter was set as relevance score >= 10.
DisGeNET (https://www.disgenet.org/): data with an EI score of 1 were selected for further analysis.
Comparative Toxicogenomics Database (CTD) (https://ctdbase.org/): only genes with direct evidence (marker, mechanism or therapeutic) in CTD were defined as targets.
PharmGKB (https://www.pharmgkb.org/): the query data in PharmGKB were all included.
PPI Network Construction and Visualization
Overlapping targets among BBR, AADs, and MetS were identified and visualized by Venn diagrams (http://bioinformatics.psb.ugent.be/webtools/Venn/). After excluding disconnected nodes, we constructed the protein-protein interaction (PPI) network based on the STRING database (https://string-db.org/) by setting medium confidence at 0.400. The results were visualized using Cytoscape software (version 3.9.1). Cytohubba was used to calculate the degree of each node in the network.
GO and KEGG Enrichment Analysis
ClusterProfiler package in R studio software was used to perform Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis. GO analysis included categories of biological processes (BP), cellular component (CC), and molecular function (MF). The ggplot2 package was used to visualize the data.
BBR-Target-Pathway Network
The top 20 KEGG pathways were selected to construct the “BBR-targets pathway” network, and then the origin software was applied to draw the network diagram.
Conducting Molecular Docking
The molecular docking simulation was performed to validate the BBR-target associations. First, the spatial structure of BBR (Compound ID: 2353) was downloaded from Pubchem database (https://pubchem.ncbi.nlm.nih.gov/) and saved in SDF format. The file was then converted from SDF to PDBQT format via AutoDock Tools. Next, the protein structures of candidate targets were downloaded from the PDB database (https://www.rcsb.org/) and addressed by AutoDock Tools. In this study, the top eight targets in the interaction network were selected for molecular docking. Water molecules in each file were deleted and polar hydrogen atoms were added, and the results were exported in PDBQT format. Then, the open-source molecular docking program AutoDock Vina was used to explore the affinity and binding modes of BBR with the key targets. PyMOL was used to visualize the docking results.
Results
Overlapping Targets among BBR, AADs, and MetS
To determine the molecular mechanisms underlying the therapeutic effect of BBR against AAD-induced MetS, a total of 208 potential targets of BBR, 466 targets of AADs, and 2855 MetS associated targets were eventually detected (Supplementary Tables 1-3). Figure 1 showed the mutual overlapping target proteins between BBR, AADs, and MetS. In this study, 23 overlapping targets in the intersection set among BBR, AADs, and MetS were selected for further analysis (Supplementary Table 4).

Venn diagram analysis of overlapping targets among BBR, AADs, and MetS.
PPI Network of BBR Targets Involved in AAD-Induced MetS
A PPI network of the 23 overlapping targets was constructed based on the STRING database and visualized by Cytoscape. After excluding the disconnected node, we found that 22 out of the 23 overlapping targets were closely linked to each other with 22 nodes and 38 edges, as shown in Figure 2. Color depth of a node represents the degree value of a node; the darker the color, the greater the degree value. These targets include SRC, PPARG, NR3C1, ADRB2, DRD4, MAOB, CYP2D6, HSD11B1, GSR, HTR3A, GABRA1, IL2, ACHE, F2, KCNH2, NR1I2, NPY5R, GABRG2, SOD2, NR3C2, PIK3CG, and SCD, sorted according to the degree value (Supplementary Table 5).

Protein-protein interaction (PPI) network of BBR targets involved in AAD-induced MetS. The nodes represent the major hub targets, and edges between nodes signify the interactions.
GO and KEGG Enrichment Analysis
GO and KEGG enrichment analysis were carried out after submitting the 23 candidate targets of BBR associated with AAD-induced MetS to ClusterProfiler package. A total of 268 GO terms were obtained, including categories of biological processes (BP), cellular component (CC), and molecular function (MF) (Supplementary Table 6). The top 10 GO terms of BP-, CC-, and MF-based count are shown in Figure 3. For BP, those common DEGs are significantly enriched in regulation of membrane potential, response to histamine, response to ethanol, positive regulation of transmembrane transport, response to alcohol, response to xenobiotic stimulus, intracellular receptor signaling pathway, positive regulation of kinase activity, and positive regulation of ion transport. For CC, those common DEGs are significantly enriched in GABA-A receptor complex, GABA receptor complex, ion channel complex, transmembrane transporter complex, transporter complex, dendrite membrane, leading edge membrane, chloride channel complex, neuron projection membrane, and GABA-ergic synapse. For MF, those common DEGs are mainly enriched in nuclear receptor activity, ligand-activated transcription factor activity, steroid binding, neurotransmitter receptor activity, ligand-gated ion channel activity, ligand-gated channel activity, transmitter-gated ion channel activity, transmitter-gated channel activity, postsynaptic neurotransmitter receptor activity, and benzodiazepine receptor activity.

GO enrichment analysis of the targets. The top 10 GO terms of BP, CC, and MF based count are shown.
Figure 4 shows the top 20 pathways of KEGG enrichment (Supplementary Table 7). Strikingly, the selected targets are associated with the PPAR signaling pathway (hsa03320), lipid and atherosclerosis (hsa05417), and AMPK signaling pathway (hsa04152), which are closely associated with metabolic disturbance and indicated the important therapeutic function of BBR on AAD-induced MetS.

Bubble diagram of the top 20 KEGG enrichment.
BBR-Target-Pathway Network
The “BBR-target-pathway” network was then constructed based on the top 20 KEGG pathways. Figure 5 reveals for BBR 18 target proteins and 20 signaling pathway networks. The network elucidates the function of potential candidate targets of BBR against AAD-induced MetS in the KEGG biological pathway. Among these targets, PPARG was identified to be the key regulator of the PPAR, lipid and atherosclerosis, and AMPK signaling pathways. Subsequently, SCD was also found to be involved in the PPAR and AMPK signaling pathways, which might represent the key therapeutic efficacy of BBR.

BBR-targets-pathway network.
Molecular Docking Investigation of BBR-Target Associations
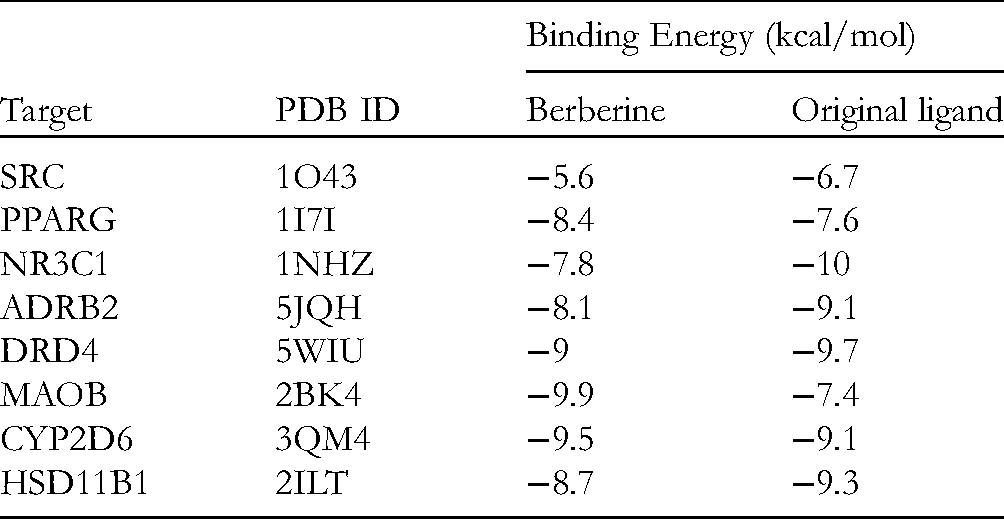
The molecular docking simulation was performed to confirm the valid bonding effects between BBR and its predicted targets. In this study, we selected the top eight targets in the interaction network for further analysis. As shown in Table 1, the promising binding energy of BBR with the potential targets SRC (PDB ID: 1O43), PPARG (PDB ID: 1I7I), NR3C1 (PDB ID: 1NHZ), ADRB2 (PDB ID: 5JQH), DRD4 (PDB ID: 5WIU), MAOB (PDB ID: 2BK4), CYP2D6 (PDB ID: 3QM4), and HSD11B1 (PDB ID: 2ILT) were −5.6, −8.4, −7.8, −8.1, −9, −9.9, −9.5, and −8.7 kcal/mol, respectively. As shown in Figure 6, analysis of the binding pattern between BBR and target proteins revealed that numerous interactive processes existed, such as hydrogen bonding and hydrophobic interaction. BBR interacts with four amino acid residues of SRC through hydrophobic interaction. These amino acid residues are Ile-73, Thr-74, Asp-94, and Leu-96. BBR binds with Ile-281, Arg-288, and Ile-341 of PPARG through hydrophobic interaction. Moreover, BBR interacts with four amino acid residues of NR3C1. The interactions between BBR and Leu-563 and Phe-623 were hydrophobic interactions, whereas interactions with Asn-564 and Gln-570 were hydrogen bonding. BBR interacts through hydrophobic interaction with three amino acid residues of CYP2D6, Phe-120, Leu-213, and Ala-305. These interactions have been proved essential in evaluating the affinity between the receptors and ligands, which made the BBR-target structure tend to be stable.

Molecular docking interactions between BBR and potential targets: (A) BBR-SRC; (B) BBR-PPARG; (C) BBR-NR3C1; (D) BBR-CYP2D6. The blue solid line represents hydrogen bonding and the gray dotted line represents hydrophobic interaction.
The Results of Molecular Docking Between BBR and the Top Eight Targets in the Interaction Network.
Discussion
Berberine (BBR; [C20H18NO4]+), the major bioactive constituent of
We first identified the overlapping targets among BBR, AADs, and MetS, and then constructed the PPI networks. Results showed that 22 out of the 23 overlapping targets closely interacted with the others. GO and KEGG enrichment analysis were conducted in order to explore further the key biological pathways. Intriguingly, “regulation of membrane potential” ranked first in the significantly enriched GO biological process while KEGG enrichment analysis revealed that these selected targets participated in the pathways closely related with metabolic disturbance. In particular, we focused on the “PPAR signaling pathway,” “Lipid and atherosclerosis,” and “AMPK signaling pathway” because these pathways have emerged as central regulators of glucose and lipid homeostasis. Specifically speaking, PPARs are ligand dependent transcription factors which function in the PPAR signaling pathway, and engage in the regulation of glucose homeostasis, fatty acid and lipoprotein metabolism, cellular proliferation/differentiation and the immune response.27,28 Additionally, lipid-derived products seem to play a major role in the pathophysiology of atherosclerosis in relation to the MetS. 29 Furthermore, AMPK modulates glucose transport, lipid and protein synthesis, and food metabolism. The activation of AMPK improves insulin sensitivity and metabolic health, which sheds light on the future treatment of type 2 diabetes.30,31
To illustrate the drug-target-pathway networking, a “BBR-target-pathway” analysis was constructed, which reflected the 18 potential targets of BBR against AAD-induced MetS in 20 signaling pathway networks. Among these networks, PPARG represents the most important therapeutic effector of BBR, which is associated with the PPAR signaling pathway, lipid and atherosclerosis and AMPK signaling pathway. Finally, molecular docking simulation was performed to confirm the valid bonding effects between BBR and the top eight target proteins in the interaction network. Among the top eight targets, SRC, 32 PPARG, 28 NR3C1, 33 ADRB2, 34 MAOB, 35 and HSD11B1 36 have been reported to be correlated with insulin signaling in glucose metabolism. PPARG, 37 ADRB2, 38 DRD4, 39 and HSD11B1 40 participate in lipid metabolism, while CYP2D6 41 is associated with pharmacokinetics. These potential targets showed potent to moderate binding energy towards BBR, ranging from −5.6 to −9.9 kcal/mol. Results also revealed that BBR interacted with multiple residues of the potential target proteins through hydrogen bonding and hydrophobic interaction. It is worth noting that the binding energy between the key potential target PPARG and BBR (−8.7 kcal/mol) tended to be enhanced compared with the original ligand (−7.6 kcal/mol), indicating that BBR could bind potently to PPARG. This substantial function of BBR may be further supported by a previous study demonstrating that BBR has PPARs activator activities. 42
In addition, CYP2D6 was also on the list of overlapping target proteins, which attracted much attention for further surveys. It is well known that CYP2D6 is associated with antipsychotic pharmacokinetics and the propensity for adverse drug reactions.41,43 Previous research has already documented the relationship between BBR and CYP2D6. Notably, CYP2D6 plays a pivotal role in the metabolism of BBR, whereas administration of BBR alleviated the activity of CYP2D6.44,45 Our results suggested that the binding energy of BBR with CYP2D6 was −9.5 kcal/mol, and BBR interacts with Phe-120, Leu-213, and Ala-305 of CYP2D6 through hydrophobic interaction, indicating direct interactions between BBR and CYP2D6. Accordingly, it is speculated that BBR might ameliorate the metabolic side effects of atypical antipsychotics through interacting with CYP2D6 and inhibiting its activity.
Conclusion
In summary, the current study identifies the key targets and biological pathways of BBR against AAD-induced MetS through combining network pharmacology and molecular docking. “BBR-target-pathway” analysis revealed 18 potential targets of BBR against AAD-induced MetS in 20 signaling pathway networks, which elucidate the potential candidate targets and therapeutic mechanisms from a systematic perspective. In particular, PPARG is considered to be the critical target of BBR by acting on the PPAR signaling pathway, lipid and atherosclerosis, and AMPK signaling pathway to maintain glucose and lipid homeostasis. Furthermore, CYP2D6 could be considered as another candidate target of BBR that ameliorates the metabolic side effects of atypical antipsychotics. Although further experiments are needed to support our findings, this insight provides innovative avenues for further understanding of the therapeutic mechanisms of BBR in AAD-induced MetS.
Supplemental Material
sj-xlsx-1-npx-10.1177_1934578X221129106 - Supplemental material for Network Pharmacology and Molecular Docking Analysis on Targets and Mechanisms of Berberine in Atypical Antipsychotic-Induced Metabolic Syndrome
Supplemental material, sj-xlsx-1-npx-10.1177_1934578X221129106 for Network Pharmacology and Molecular Docking Analysis on Targets and Mechanisms of Berberine in Atypical Antipsychotic-Induced Metabolic Syndrome by Zhuowei Huang and Xiaolan Liu in Natural Product Communications
Supplemental Material
sj-xlsx-2-npx-10.1177_1934578X221129106 - Supplemental material for Network Pharmacology and Molecular Docking Analysis on Targets and Mechanisms of Berberine in Atypical Antipsychotic-Induced Metabolic Syndrome
Supplemental material, sj-xlsx-2-npx-10.1177_1934578X221129106 for Network Pharmacology and Molecular Docking Analysis on Targets and Mechanisms of Berberine in Atypical Antipsychotic-Induced Metabolic Syndrome by Zhuowei Huang and Xiaolan Liu in Natural Product Communications
Supplemental Material
sj-xlsx-3-npx-10.1177_1934578X221129106 - Supplemental material for Network Pharmacology and Molecular Docking Analysis on Targets and Mechanisms of Berberine in Atypical Antipsychotic-Induced Metabolic Syndrome
Supplemental material, sj-xlsx-3-npx-10.1177_1934578X221129106 for Network Pharmacology and Molecular Docking Analysis on Targets and Mechanisms of Berberine in Atypical Antipsychotic-Induced Metabolic Syndrome by Zhuowei Huang and Xiaolan Liu in Natural Product Communications
Supplemental Material
sj-xlsx-4-npx-10.1177_1934578X221129106 - Supplemental material for Network Pharmacology and Molecular Docking Analysis on Targets and Mechanisms of Berberine in Atypical Antipsychotic-Induced Metabolic Syndrome
Supplemental material, sj-xlsx-4-npx-10.1177_1934578X221129106 for Network Pharmacology and Molecular Docking Analysis on Targets and Mechanisms of Berberine in Atypical Antipsychotic-Induced Metabolic Syndrome by Zhuowei Huang and Xiaolan Liu in Natural Product Communications
Supplemental Material
sj-xlsx-5-npx-10.1177_1934578X221129106 - Supplemental material for Network Pharmacology and Molecular Docking Analysis on Targets and Mechanisms of Berberine in Atypical Antipsychotic-Induced Metabolic Syndrome
Supplemental material, sj-xlsx-5-npx-10.1177_1934578X221129106 for Network Pharmacology and Molecular Docking Analysis on Targets and Mechanisms of Berberine in Atypical Antipsychotic-Induced Metabolic Syndrome by Zhuowei Huang and Xiaolan Liu in Natural Product Communications
Supplemental Material
sj-xlsx-6-npx-10.1177_1934578X221129106 - Supplemental material for Network Pharmacology and Molecular Docking Analysis on Targets and Mechanisms of Berberine in Atypical Antipsychotic-Induced Metabolic Syndrome
Supplemental material, sj-xlsx-6-npx-10.1177_1934578X221129106 for Network Pharmacology and Molecular Docking Analysis on Targets and Mechanisms of Berberine in Atypical Antipsychotic-Induced Metabolic Syndrome by Zhuowei Huang and Xiaolan Liu in Natural Product Communications
Supplemental Material
sj-xlsx-7-npx-10.1177_1934578X221129106 - Supplemental material for Network Pharmacology and Molecular Docking Analysis on Targets and Mechanisms of Berberine in Atypical Antipsychotic-Induced Metabolic Syndrome
Supplemental material, sj-xlsx-7-npx-10.1177_1934578X221129106 for Network Pharmacology and Molecular Docking Analysis on Targets and Mechanisms of Berberine in Atypical Antipsychotic-Induced Metabolic Syndrome by Zhuowei Huang and Xiaolan Liu in Natural Product Communications
Footnotes
Acknowledgements
This work was supported by grants WG13B08 and WZ18Z09 from the Medical Scientific Research Foundation of Wuhan Municipal Health Commission.
Authors' Contributions
Z. Huang and X. Liu conceived and drafted the manuscript. X. Liu analyzed the network pharmacology data. Z. Huang collected the candidate literature. All authors (Z. Huang and X. Liu) read and approved the final manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Medical Scientific Research Foundation of Wuhan Municipal Health Commission (grant numbers WG13B08 and WZ18Z09).
Ethical Approval
Ethical approval is not applicable for this article.
Statement of Human and Animal Rights
Statement of human and animal rights is not applicable for this article.
Informed Consent statements
There are no human subjects in this article and informed consent is not applicable.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
