Abstract
Glioma, originated from the neuroectoderm, is one of the prevalent tumors. It is reported that ginsenoside Rh2, a compound extracted from Panax ginseng, shows pharmacological activity in inhibiting the proliferation and metastasis of glioma cells. However, the precise underlying mechanism has not been completely clarified. In this study, the molecular mechanism of ginsenoside Rh2 in glioma treatment was investigated using network pharmacology, through which the target of ginsenoside Rh2 in glioma treatment was screened, and further the biological functions were analyzed by gene ontology biological annotation and Kyoto Encyclopedia of Genes and Genomes pathway enrichment. In total, 80 potential targets of ginsenoside Rh2 for glioma treatment were screened. Biological functions analysis revealed that ginsenoside Rh2 was associated with several cancer-related signaling pathways such as VEGF signaling pathway, thereby regulating cell processes, including cell migration and cell proliferation, which suggests that ginsenoside Rh2 targets multiple genes, further mediating indicated signaling network in anti-glioma and providing a potential mechanism of ginsenoside Rh2 in glioma treatment.
Introduction
Glioma originates from the neuroectoderm and is a common intracranial malignancy, with the characteristics of malignancy and metastasis.1,2 The clinical treatment of glioma mainly involves surgery, combined with radiotherapy and chemotherapy as an adjuvant therapy. 3 Compounds derived from natural products, especially traditional herbs, show multiple activities such as anti-inflammatory, immune stimulating, and anti-cancer. 4
Ginsenoside Rh2, one of the active ingredients of Panax ginseng, has been identified as a bioactive effect in inhibiting the proliferation (in vivo and in vitro) as well as metastasis of glioma.5‐7 In detail, ginsenoside Rh2 suppresses the viability of U251 glioma cells via targeting microRNA-128, 5 arresting the cell cycle of A172 glioma cells at G1 phase by inhibiting the expression of cyclin-dependent kinase (CDK4) and cyclin D. 8 Moreover, ginsenoside Rh2 inhibits the expression of matrix metallopeptidase-13 (MMP13), which, mediated by the phosphatidylinositol-3-kinase (PI3K)/Akt (PI3k/Akt) signaling pathway, further blocks U251 glioma cells migration. 5 However, the mechanism of ginsenoside Rh2 inhibiting glioma progression has not been completely clarified.
In this study, via network pharmacology, the targets of glioma and ginsenoside Rh2 were screened respectively, and further the potential molecular mechanisms of ginsenoside Rh2 in glioma treatment were analyzed. Through this, we propose to provide thoughts and foundation for the future ginsenoside Rh2–glioma research.
Materials and Methods
Prediction of Target of Ginsenoside Rh2
The term “ginsenoside Rh2” was searched for in the PubChem database; the SMILES number for ginsenoside Rh2 (CC( = CCCC(C)(C1CCC2(C1C(CC3C2(CCC4C3(CCC(C4(C)C)OC5C(C(C(C(O5)CO)O)O)O)C)C)O)C)O)C) was obtained, which was used in a SwissTargetPrediction database search to obtain the candidate target of the active ingredient.
Screening of Glioma Target Proteins
Glioma target proteins were obtained by searching for the keywords “glioma” and “nervglioma” in GeneCards, Therapeutic Target Database (TTD), and Online Mendelian Inheritance in Man (OMIM); the species Homo sapiens (human) was selected in the UniProt database. Gene names and UniProt numbers of target proteins were obtained.
Construction of the Component–Target Network
The obtained components and disease targets were sorted, deduplicated, and mapped to obtain the intersection genes and draw the Venn diagram. Cytoscape 3.2.1 was used to visualize the network of the intersection proteins, and a “component-target” network of ginsenoside Rh2 and the target for glioma treatment were constructed.
Construction of the Protein–Protein Interaction Network
The “Multiple Proteins” function of the STRING database (https://string-db.org) was used, the species was set as human (Homo sapiens), the protein target of ginsenoside Rh2 intervention in glioma was imported into the system, and the score value was set to > 0.900 to search for and draw a protein–protein interaction network.
Analysis of Gene Ontology Biological Function and KEGG Pathway Enrichment
The ginsenoside Rh2 intervention gene for glioma was input into the Database for Annotation, Visualization and Integrated Discovery (DAVID). The species was set as human (Homo sapiens), and the biological process, molecular function, and cell component categories were selected for gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses.
Results
Prediction and Interaction of Ginsenoside Rh2–-Glioma Targets
Through searching indicated databases, combined with sorting and deduplicating, 110 candidate targets of ginsenoside Rh2 were obtained, and 4805 glioma-related genes were collected. As shown in Figure 1A, 80 overlapping genes were obtained, including epidermal growth factor receptor (EFGR), MMP2, CKD4, histone deacetylase 2 (HDAC2), etc. (Figure 1B); the details are listed in Table 1. To identify the interaction between the crossed gene, the STRING database was used to establish a PPI network, including 80 nodes and 128 connections (confidence score > 0.95) (Figure 1C).

Prediction and interaction of ginsenoside Rh2-glioma targets. (A) Wayne diagram of ginsenoside Rh2 and glioma candidate targets. (B) Ginsenoside Rh2-targeting glioma network. The blue rhombus represents the potential target of ginsenoside Rh2 in the treatment of glioma. (C) The protein-protein interaction (PPI) network. The "node" in the network represents the target protein, and the "edge" represents the interaction between the target proteins. The more the number of edges, the more important the role of the target protein corresponding to the node in the network.
The Therapeutic Targets of Ginsenoside Rh2 in Glioma Treatment.

GO Enrichment of Putative Targets
Eighty screened ginsenoside Rh2-glioma crossed genes (Table 1) were subjected to the DAVID database, and the functions of the putative targets were further analyzed. With the limit of P value < .05, the putative genes were enriched in indicated processes, of which the top 20 are listed in Figure 2A. In detail, the genes involved in biological process were functions in negative regulation of apoptotic process, cell proliferation, growth hormone receptor signaling pathway, positive regulation of transcription form, RNA polymerase II promoter, and so on (Figure 2B), whereas the putative targets involved in cellular component (CC) participated in regulating centrosome, membrane, spindle microtubule, cytoplasm, nucleus, midbody, and cytosol (Figure 2C). Moreover, the overlapping genes involved in molecular function (MF) were associated with ATP binding, non-membrane spanning protein tyrosine kinase activity, and heme binding (Figure 2D).
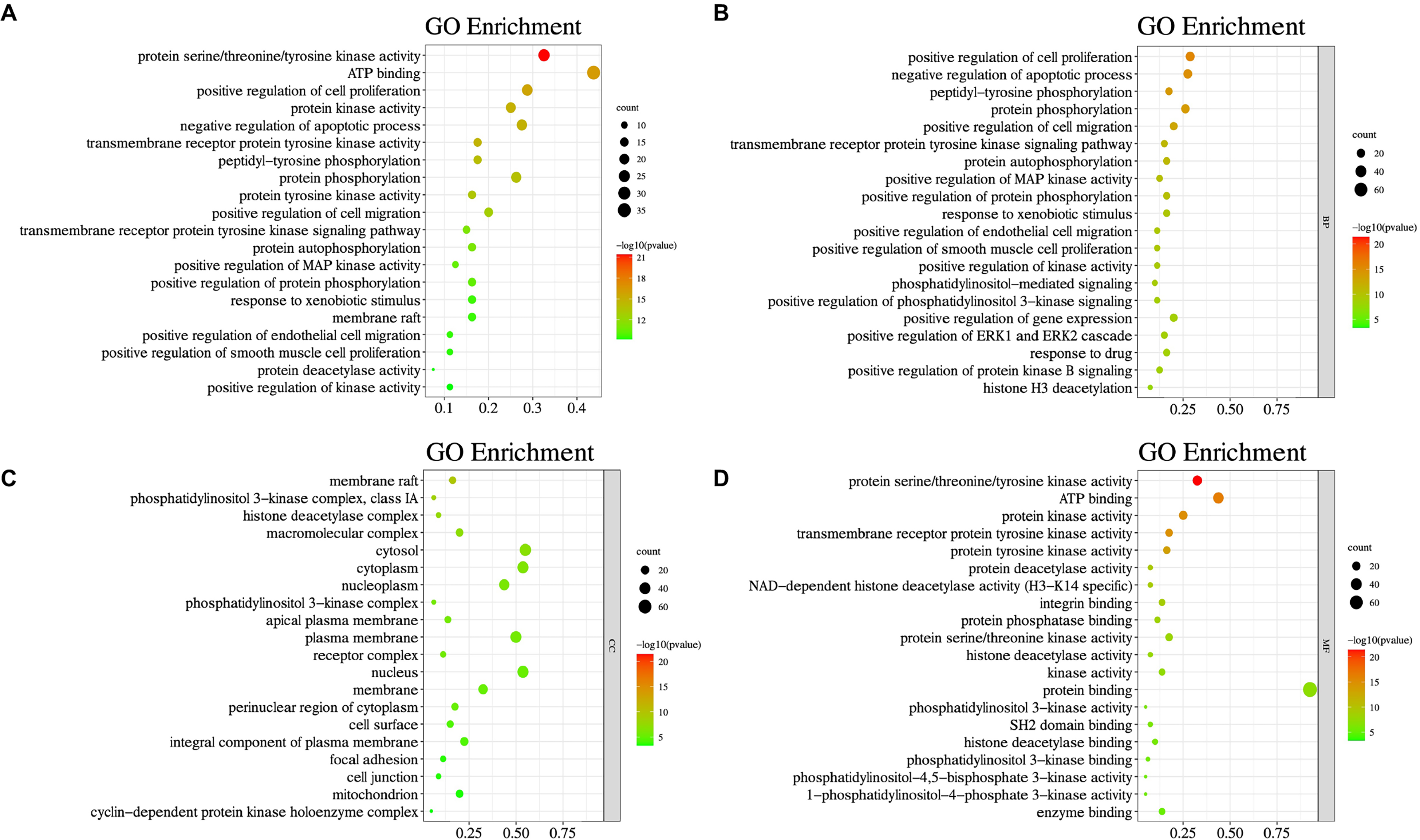
Gene ontology (GO) enrichment of putative targets. GO analysis of the identified targets of the top 20 enrichment (A), and in terms of biological process (B), cellular component (C), as well as molecular function (D).
KEGG Pathway Enrichment Analysis
KEGG pathway analysis revealed that the overlapping targets mediated signaling pathways; the details including the involved genes and indicated signaling pathways are listed in Table 2. Figure 3A shows the top 20 pathways, including cell cycle, cancer, PI3K-Akt, HIS-1, FoxO, and Jak-STAT signaling pathways, in which the bubbles are related with gene number (size) and P value (color) (Figure 3A, Table 2). Furthermore, targets such as EFGR, HDAC2, heat shock protein 90 alpha family class A member 1 (HSP90AA1), vascular endothelial growth factor (VEGFA), B-cell lymphoma-2 like 1 (BCL2L1), and PIK3R1 enriched in cancer pathway are shown in Figure 3B, marked with red stars, tightly related with tumorigenesis and cancer proliferation.

Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis. (A) KEGG pathways of the identified targets. (B) Rh2 potential targets enriched in cancer pathway.
Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathway Enrichment Analysis.

Conclusion
The current study screens the potential targets of ginsenoside Rh2 in glioma treatment, which are involved in cancer-related signaling pathways, thereby functioning in regulating cell process including cell apoptosis, proliferation, and gene transcription. The results demonstrate the potential mechanisms of ginsenoside Rh2 in glioma treatment.
Discussion
Although ginsenoside Rh2 has been reported as a therapeutic effect on various tumors, including glioma, 9 the mechanisms of ginsenoside Rh2 in glioma therapy have not been precisely clarified.
In this study, the targets of ginsenoside Rh2 in glioma treatment have been screened. For instance, cell cycle-related protein CDKs, including CDK1, CDK2, CDK4, and CDK6, are identified in the putative targets, suggesting the role of ginsenoside Rh2 in regulating glioma cell cycle as well as cell division. It is reported that ginsenoside Rh2 suppresses various types of human cancer cells’ proliferation and viability, including hepatocellular carcinoma cell lines, leukemia cell lines, colorectal cancer cell lines, breast cancer cell lines, and glioma cell lines. 10 Correspondingly, ginsenoside Rh2 drives glioma cell cycle arrest at G1 phase. 8 Consistent with this, ginsenoside Rh2 is associated with cell proliferation in the result of GO enrichment (Figure 2A).
Several studies confirmed that histone acetylation exhibits reduced modification level in various types of cancer and correlated with the pathological stages and prognosis. 11 In glioma, HDAC class I (HDAC1, 2, 3, 8) as well as HDAC6 are upregulated and corelated with more aggressive phonotype. 12 HDAC inhibitors (HDACi) targeting HDACs have been investigated in cancer therapy; more than 30 inhibitors have been applied in clinic trials, while five HDACis have been approved in clinical cancer treatment. 13 Our data demonstrated that ginsenoside Rh2 can regulate HDACs’ (HDAC1, 2, 3, 5, 6, 7) expression (Table 1) and functions in regulating histone deacetylase complex and histone deacetylase activity in CC and MF of GO analysis (Figure 2C and 2D), suggesting an HDACs inhibitor role of ginsenoside Rh2 in anti-glioma.
EGFR, a type of receptor tyrosine kinases, is involved in activating several downstream signaling pathways such as PI3K-AKT pathway. 14 EGFR is often overexpressed in various cancer cells, including glioma, and related with cancer cell proliferation, autophagy, and migration. 15 It is reported that ginsenoside Rh2 downregulates the expression level of EGFR and inhibits the proliferation of glioma. 6 In our result, EGFR is one of the overlapping genes of ginsenoside Rh2 and glioma, through targeting EGFR, ginsenoside Rh2 mediating PI3K-AKT signaling pathway, estrogen signaling pathway, Ras signaling pathway, and FOXO signaling pathway, indicating that ginsenoside Rh2 can target EGFR, function in regulating indicated signal transduction, thereby suppressing glioma progression.
VEGF-mediated angiogenesis plays a critical role in tumor growth, metastasis, and recurrence. 16 Several drugs targeting VEGF and VEGF signaling pathway have been approved by U.S. FDA in various types of cancer, including glioma.17,18 Study has shown that ginsenoside Rh2 inhibits the expression of VEGFA, thereby suppressing the proliferation of the cells causing oral squamous cell carcinoma. 19 Consistent with this, through the network pharmacology, VEGFA was screened, and further biological analysis revealed that ginsenoside Rh2 regulating VEGF signaling pathway, HIF signaling pathway, Ras signaling pathway, etc. is mediated by VEGFA. In summary, this study reveals ginsenoside Rh2 can inhibit the proliferation and metastasis of glioma by regulating multi-targets and related signaling network.
Footnotes
Author Contributions
D.H. and X.W. designed the experiments; X.W. and X.Q. performed the experiments; D.H. and X.W. contributed to manuscript preparation.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
