Abstract
Diabetic foot ulceration is a common complication of an uncontrolled diabetic regimen and is considered a serious type of wound. Matrix metalloprteinases (MMPs) are the common key enzymes in wound management, overexpression of MMPs can lead to chronic wounds and ulcers.
Introduction
Wound healing is a highly complicated multifactorial process, consisting of 4 consecutive phases: hemostasis, inflammation, proliferation, and tissue remodeling. 1 Interruption or irregularity in any of these phases will result in a delay in healing or even nonhealing. Patients with chronic wounds or ulcers especially those who are suffering from persistent conditions, such as diabetes mellitus (DM), ischemic heart disease, and irregular blood pressure will have more complicated and severe conditions.2,3 The uncontrolled glycemia in DM over prolonged periods causes peripheral neuropathy and loss of sensation.4,5 The combination of this neuropathy with ischemia in the foot whether due to macrovascular or microvascular diseases results in what is commonly known as “Diabetic Foot” (DF). 6 The DF impairs the healing cascade of wounds or ulcers and often results in amputation in up to 15% of the population especially in developing countries that suffer from lack of awareness, low foot care services and delay in professional help. 7 Furthermore, these ulcers are frequently colonized with pathogens such as bacteria whether they are the cause or the consequence, which further delays ulcer healing.8,9 As a result, the management of wounds and DF ulcers primarily involves the eradication of infection using antibiotics, improving peripheral blood flow and wound management.10,11
At the cellular level, chronic ulcers are characterized by the overactivity of various proteinases which cause a defect in collagen turnover and tissue remodeling. 12 The zinc-dependent matrix metalloproteinases (MMPs) are a family of 23 metalloproteins manipulating extracellular matrix in constructive and destructive manners. 13 Normally, matrix metalloprteinase-8 (MMP-8) (collagenase-2) and matrix metalloprteinase-9 (MMP-9) (gelatinase B) are expressed by neutrophils, macrophages, and fibroblasts14,15 during wounds, however, their levels increase in DM. 16 While their normal levels are beneficial in the wound healing process, their high level in chronic cases impairs their wound-healing capabilities due to their potentially destructive effects, hence their inhibition is proposed to allow normal healing.17,18
The consequences of DF ulcers on mortality and morbidity are getting more serious by the day, especially with current antibiotic use.11,19 The health care systems tend to empirically use antibiotics to avoid infection exacerbation thus participating in the antibiotic resistance crisis worldwide. 20 Although this can cause a short-term decrease in mortality and morbidity, the emergence of more resistant strains every year indicates an imminent rise in both markers as evidenced by the 171 resistant strains found in 2010. 21
Based on this evidence, it is imperative to search for new natural supportive remedies to be used alone or as adjunctive treatment to decrease our antibiotic reliance, especially with the occurrence of multidrug-resistant bacteria in 18% of cases.
22
Our study tackles this problem using an in silico screening evaluation of potential plant-derived compounds useful in targeting MMPs to be useful in the healing of wounds and DF ulcers.

Overview of Calendula plant, some constituents, and reported pharmacological activities.

Phytochemical constituents of
Material and Methods
Molecular Docking
Initial compounds screening was conducted using MOE 2019 software. 39 Both the receptor and the ligand were prepared according to the standard structure optimization protocol of MOE software using the same protein data bank files (PDBID: 2OVX and 4QKZ for MMP-9 and MMP-8, respectively), and energy minimization was performed under AMBER12: EHT force field. The active sites were set as the bound positions of the co-crystallized ligand using the general protocol of MOE DOCKTITE Wizard. The validation of docking experiments was achieved through the re-docking of the co-crystallized ligands into their corresponding active sites and then the root-mean-square deviation (RMSD) was calculated. The docking results were visualized and analyzed by a DS visualizer available from Biovia Inc, and the obtained results are represented in Table 1.
Screening Scores for the Docked Compounds Using MOE.

Abbreviations: MMP-8, matrixmetalloproteinase-8; MMP-9, matrixmetalloproteinase-9.
Docking refinement was performed with AutoDock Vina software.
40
The receptor and the ligand were prepared using MGL tools. The active sites were determined using MGL tools as well. The docking was performed using the molecular database of
Binding Interactions of the Docked Compounds with the Target Proteins Using AutoDock Vina.
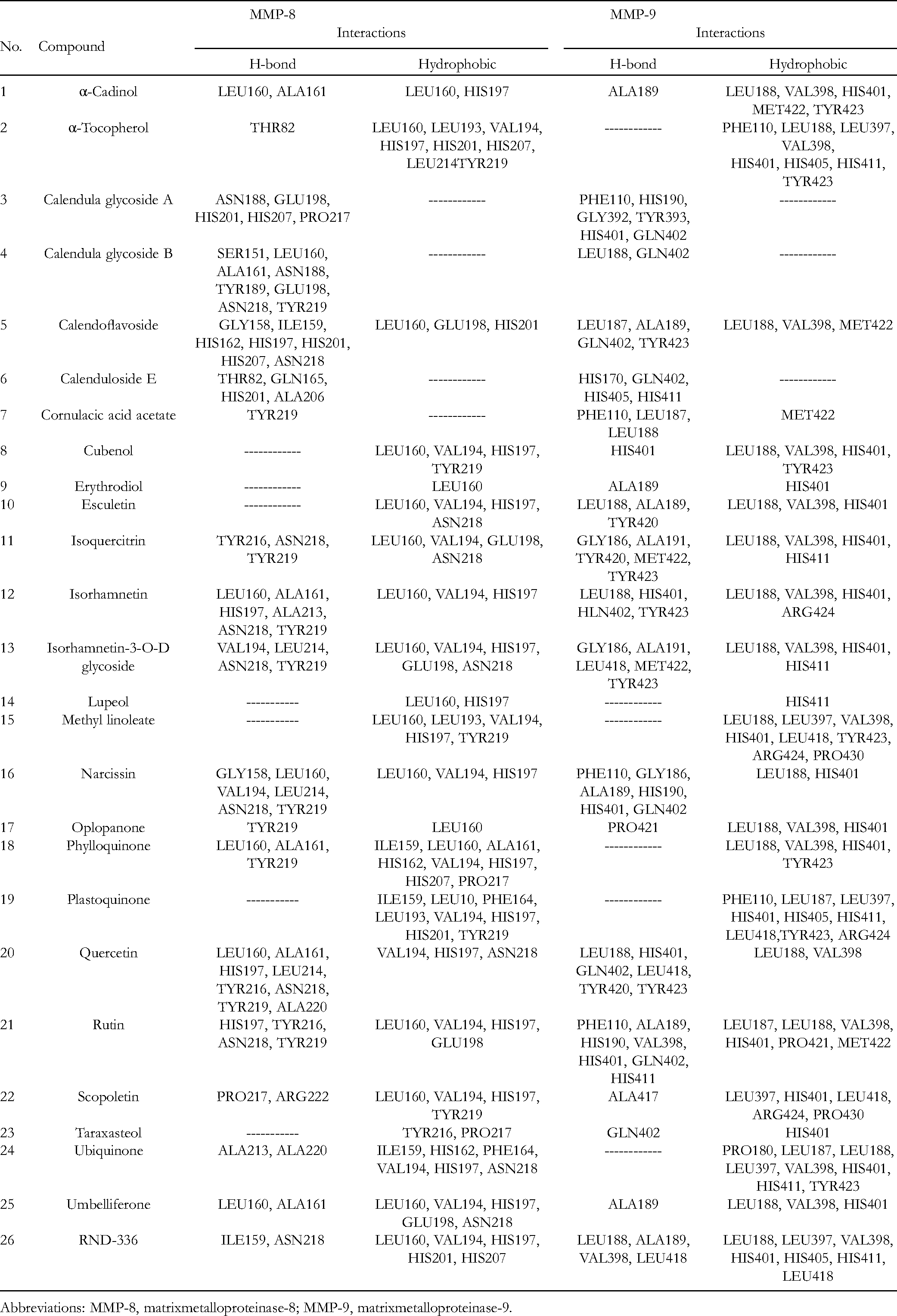
Abbreviations: MMP-8, matrixmetalloproteinase-8; MMP-9, matrixmetalloproteinase-9.
Molecular Dynamics
Isorhamnetin-3-O-D glycoside and quercetin, the most promising molecules in binding with MMP-9 and MMP-8 enzymes, respectively, were selected for further investigation. All molecular dynamics (MD) simulations were performed using GROningen MAchine for Chemical Simulations (GROMACS 2020.4) software. 38 Four MD simulation experiments were conducted to validate the results retrieved from the docking steps. Two experiments were performed on the ligand-free protein for MMP-9 and MMP-8, while the other 2 were conducted on the MMP-9 and MMP-8 enzyme in complex with isorhamnetin-3-O-D glycoside and quercetin, respectively. The enzyme and the ligand topologies were generated and joined using the standard published protocol. 39 The typical workflow of GROMACS enzyme–ligand simulations was applied as reported in the literature. The energy minimization step for the unbound and the complex enzyme structures was achieved using the steepest descent minimization algorithm with a maximum of 50,000 steps and < 10.0 KJ/mol force under GROMOS96 43a1 force field. 40 The energy minimized structures were equilibrated for 10 ns. The equilibrated systems entered the production stage without any restrains for 50 ns with a time step of 2 fs, and the structural coordinates were saved every 5 ps. The RMSD of the entire system residues was calculated from the generated trajectories from the production step.
MM-PBSA Calculation
Binding free energy calculations were performed using the 
Results and Discussion
The screening results in Table 1 through the usage of MOE software showed that all compounds were placed into the active site of both enzymes using the GBVI/WSA dG scoring function. The validation of the protocol was performed by re-docking the co-crystallized ligands using the same parameters which results in a calculated RMSD between the docked and co-crystallized poses of 0.21 and 0.89 A° for MMP-9 and MMP-8, respectively. Generally, the docking results showed promising activity against both MMP-8 and MMP-9, demonstrated by the ability of all the compounds to achieve a negative kcal/mol score with both the receptors.
In MMP-8 docking, 4 compounds, namely quercetin, isorhamnetin, methyl linoleate, and ubiquinone achieved a higher score than the reference RND-336 that achieved a score of −6.73 kcal/mol. Quercetin was the best compound achieving a score of −8.01 kcal/mol, while isorhamnetin, methyl linoleate, and ubiquinone achieved scores of −7.99, −7.77, and −7.42 kcal/mol, respectively. The 2D interaction diagrams of the 4 compounds are shown in Figure 4. Similarly, the MMP-9 binding score for RND-336 was −6.63, whereas isorhamnetin-3-O-D glycoside, plastoquinone, and calenduloside achieved scores higher than the reference (RND-336). Isorhamnetin-3-O-D glycoside was able to strongly bind the receptor as indicated by its score of −8.33 kcal/mol, while plastoquinone and calenduloside achieved scores of −8.09 and −7.88, respectively. The 2D interaction diagrams of the three compounds are shown in Figure 5.

Phytochemical constituents of

2D interactions of matrixmetalloproteinase-8 (MMP-8) with (A) quercetin, (B) isorhamnetin, (C) methyl linoleate, and (D) ubiquinone.

2D interactions of matrixmetalloproteinase-9 (MMP-9) with (A) isorhamnetin-3-O-D glycoside, (B) calenduloside, and (C) plastoquinone.
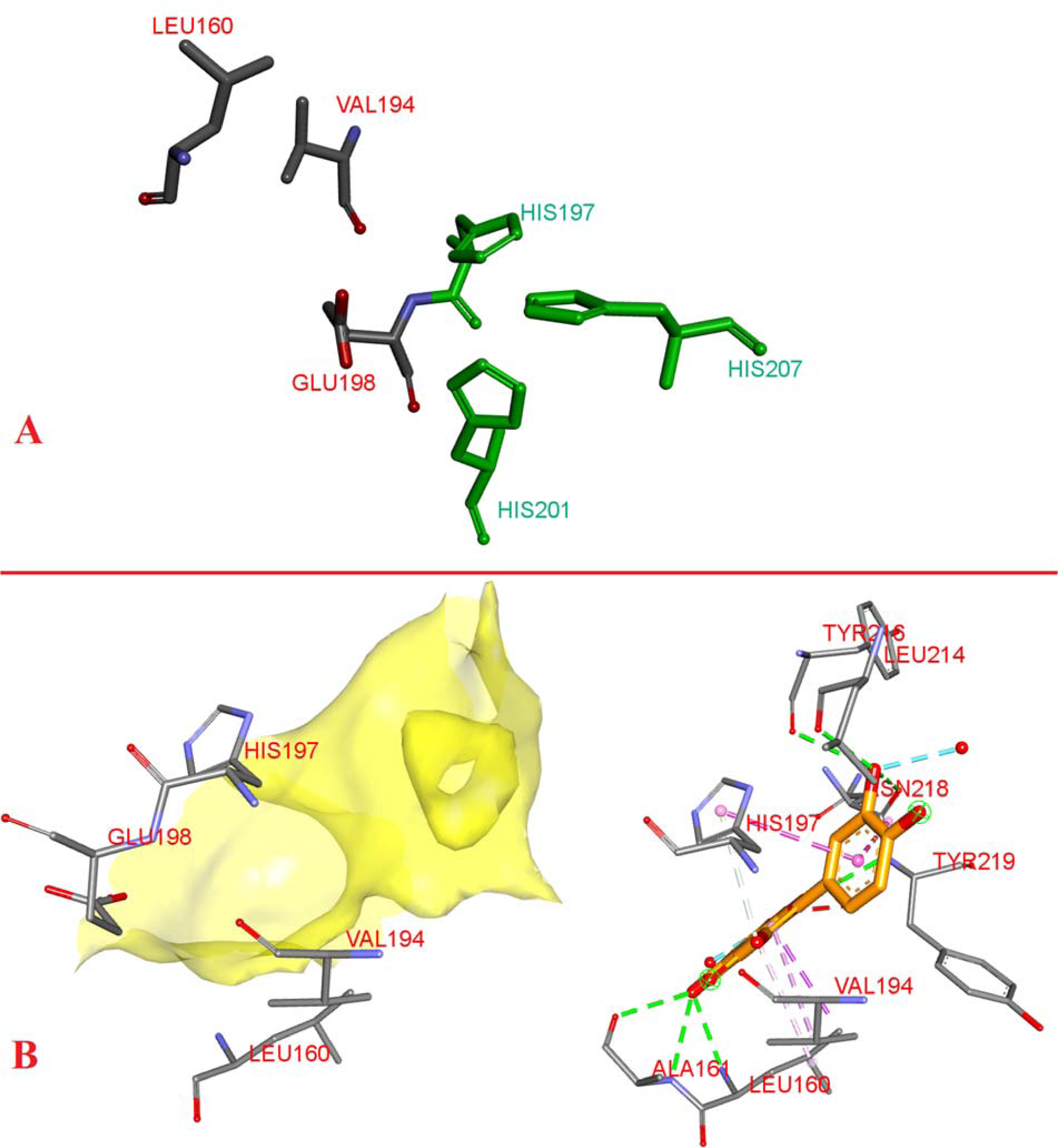
(A) 3D binding pocket of matrixmetalloproteinase-8 (MMP-8) with the histidine triangle in green. (B) 3D binding interactions of quercetin.
Moreover, we performed docking refinement using AutoDock Vina to attain more in-depth results pertaining to future selectivity. The obtained results of refinement showed that all the compounds were placed into the active site of both enzymes appropriately and validation of the protocol was performed by re-docking of the co-crystallized ligands using the same parameters which results in a calculated RMSD between the docked and co-crystallized poses of 1.50 and 0.58 A° for MMP-8and MMP-9, respectively (Figure 6).

Validation results of (A) matrixmetalloproteinase-8 (MMP-8) and (B) matrixmetalloproteinase-9 (MMP-9). (Pink is the internal ligand, Blue is the docked pose).
Consistent with previous screening, the docking results in Tables 2 and 3 showed promising activity against both enzymes as proven by the ability of all the compounds to achieve a negative kcal/mol score with both receptors with minor differences. In MMP-8 docking, over half of the compounds showed good bindings when compared to RND-336 which scored −8.2 kcal/mol. Quercetin, Calendula glycoside B, isorhamnetin, and isoquercetin scored the highest binding of −10.4, −10.2, −9.9, and −9.8, respectively (Figure 7). Studying the 3D structure of the interactions, 4 common moieties are present: LEU160, VAL197, GLU198, and HIS (197, 201, or 207). The3 histidine amino acids form what can be called a “Histidine Triangle,” nearly all poses generated especially among the strong binding derivatives interacted with at least one of those 3 amino acids as demonstrated by quercetin in Figure 8.

2D interactions of matrixmetalloproteinase-8 (MMP-8) with (A) quercetin, (B) calendula glycoside B, (C) isorhamnetin, and (D) isoquercetin.
Binding Scores for the Docked Compounds Using AutoDock Vina.

Abbreviations: MMP-8, matrixmetalloproteinase-8; MMP-9, matrixmetalloproteinase-9.
The refinement in MMP-9 showed the ability of most compounds to bind stronger than RND-336 which scored −8.7 kcal/mol. Quercetin and rutin achieved the highest binding potential with a score of −10.3 kcal/mol for each one of them. Isorhamnetin, calendoflaviside, Calendula glycoside B, and isorhamnetin 3-O glycoside scored good binding with scores of −10.1, −10.0, −9.8, and −9.7 kcal/mol, respectively (Figure 9). When focusing on the 3D interactions, a “Histidine Triangle” is also observed among all compounds especially those with the highest scores. Additionally, LEU188 and VAL398 take part in binding interactions as well (Figure 10).
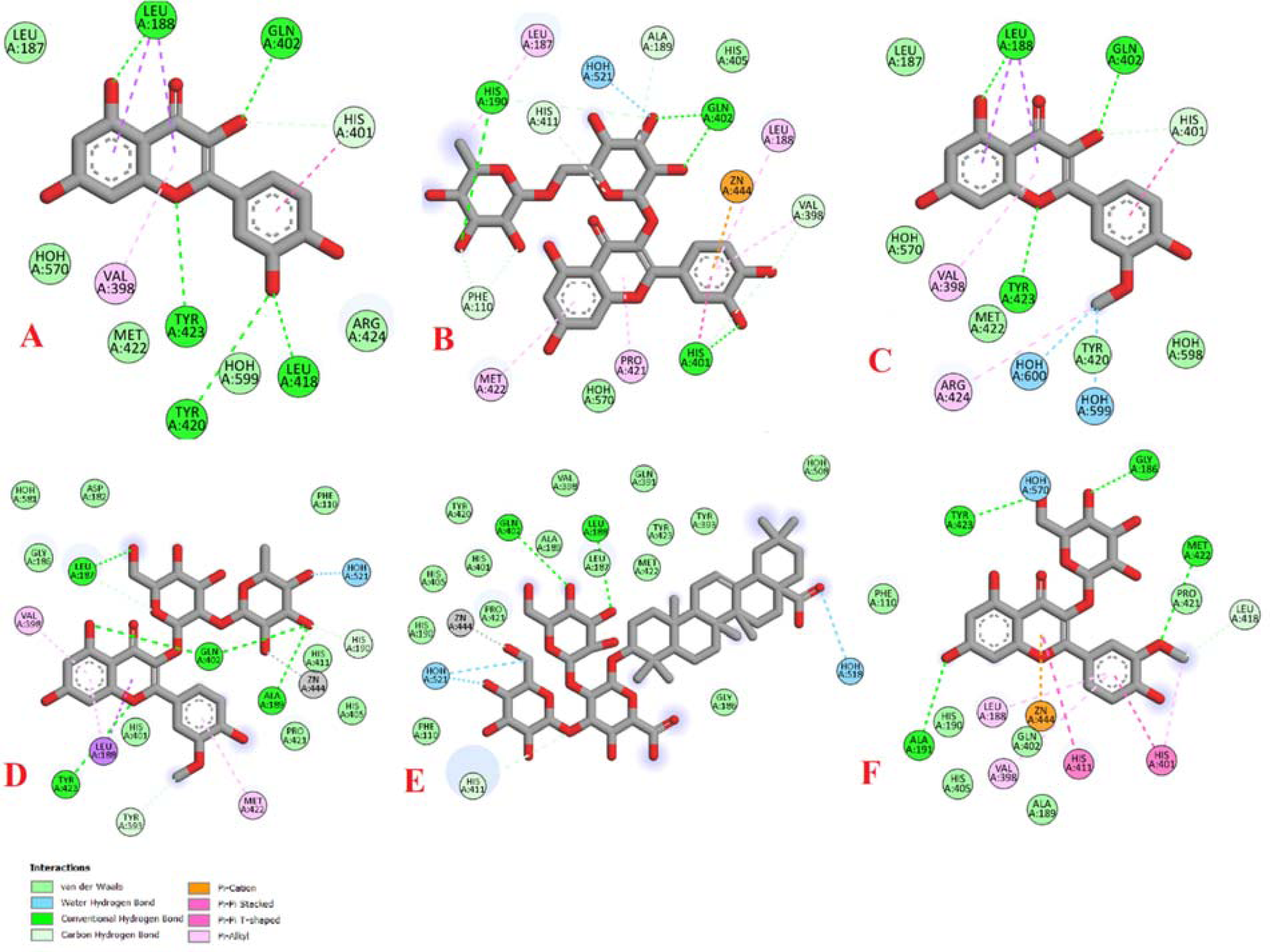
2D interactions of matrixmetalloproteinase-9 (MMP-9) with (A) quercetin, (B) rutin, (C) isorhamnetin, (D) calendoflaviside, (E) Calendula glycoside B, and (F) isorhamnetin 3-O glycoside.

(A) 3D binding pocket of matrixmetalloproteinase-9 (MMP-9) with the histidine triangle in green. (B) 3D binding interactions of quercetin.
In our effort to screen MMP-8 and MMP-9 inhibitors, phylloquinone has proved to be an interesting inhibitor and deserves to be subjected for further understanding. Although it has achieved a low score on MMP-8 and a comparable result on MMP-9 with RND-336, the remarkable difference probed us to study it further. The binding pockets of both proteins had a similar degree of composition with 2 main differences, MMP-9 contained more hydrophobic moieties, as well as its spatial arrangement, is less compact. As a result, the phylloquinone molecule in MMP-9 occurred in a more relaxed form than its counterpart in MMP-8 with less tortional energy (Figure 11). This is also supported by the nearly identical scores of less bulky compounds such as α-cadinol, quercetin, and isoquercetin.

Top: Binding pockets of matrixmetalloproteinase-8 (MMP-8) and matrixmetalloproteinase-9 (MMP-9). Middle and Bottom: 2D and 3D interactions of phylloquinone with MMP-8 and MMP-9.
These promising results identified several dual inhibitors such as quercetin, isoquercetin, isorhamnetin, and isorhamnetin 3-O glycoside that outperformed RND-336. These compounds contained a common benzopyran-4-one moiety and were of suitable size to bind comfortably with the MMP-9 elongated binding site without high steric and torsional stress. Moreover, this benzopyran-4-one is one of the important common features among all potent compounds.
Molecular Dynamics
As many proteins require a high degree of flexibility to conduct their intended function and considering the power of MD simulations in testing the stability of ligand–enzyme complexes,28–30 we employed this technique to evaluate the stability of the ligand-free enzyme against the 2 complex structures obtained from the docking calculations. The 4 structures were subjected to a standard MD protocol with a production phase of 50 ns.
The RMSD values for the 2 free enzyme Cα atoms as well as the 2 complex structures were calculated by comparing the entire trajectories. The retrieved data presented in the RMSD plots show that the unbound enzymes of MMP-8 and MMP-9 had RMSD of 3.27 A° and 3.29 A°, respectively, indicating their high dynamic properties. In addition, the RMSD values for the 2 complexes of MMP-8-quercetin and MMP-9-isorhamnetin-3-O-D glycoside were 1.69 and 1.42 A°, respectively. Those results support the ability of quercetin and isorhamnetin-3-O-D glycoside to inhibit both MMP-8 and MMP-9 (Figure 12).
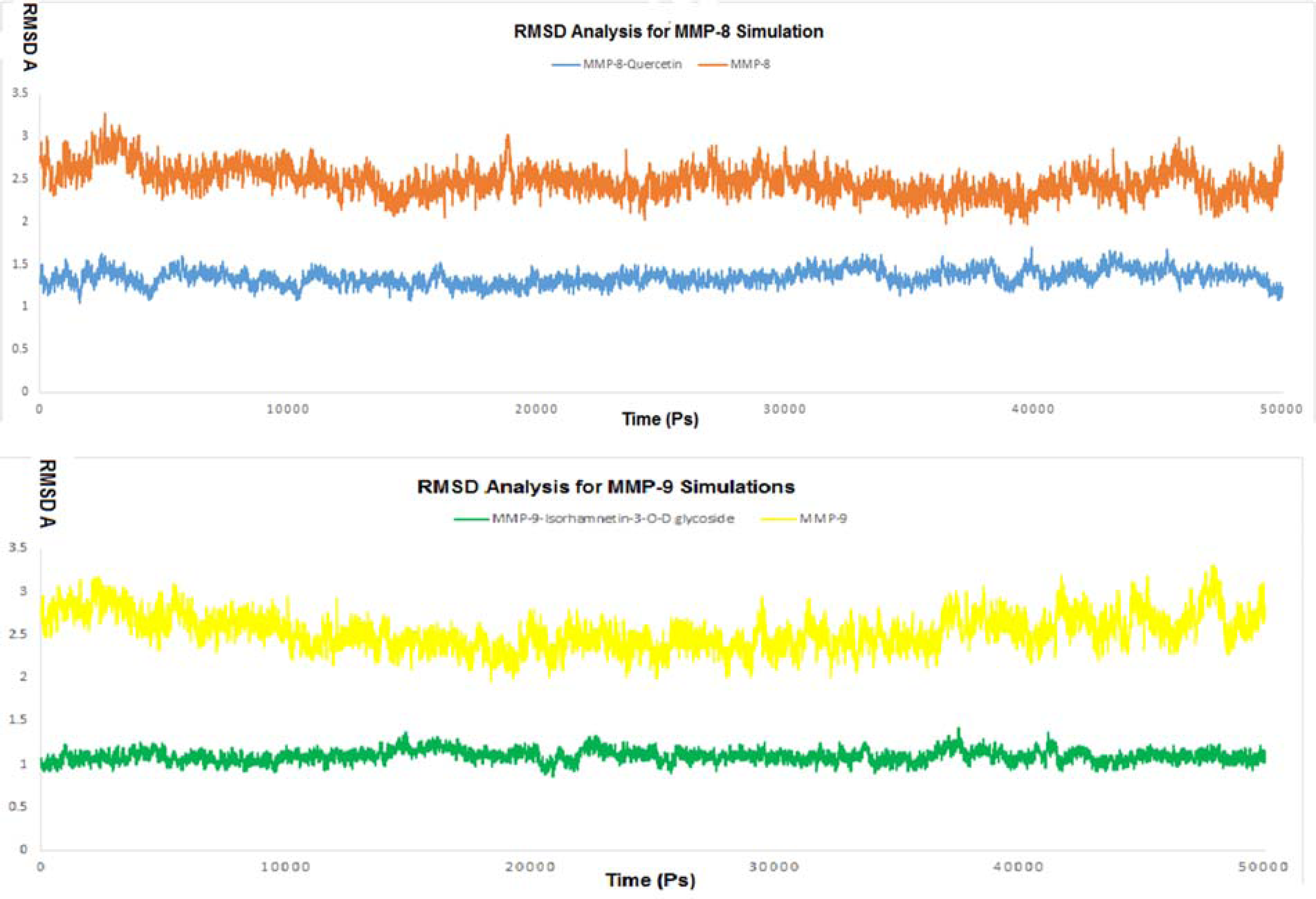
Root-mean-square deviation (RMSD) graph of matrixmetalloproteinase-8 (MMP-8)-quercetin (top) and matrixmetalloproteinase-9 (MMP-9)-isorhamnetin-3-O-D (bottom).
MM-PBSA Binding Free Energy Calculations
The binding energies for quercetin and isorhamnetin-3-O-D glycoside with their potential binding site in MMP-8 and MMP-9 were calculated using the MM-PBSA binding free energy methodology. This method represents a more reliable way to calculate the binding free energy through many MD conformations compared to a single conformation-based score calculated from the docking study. As part of these calculations, the free energy of each component was calculated by the cumulative sum of its molecular mechanics potential energy in a vacuum and the free energy of solvation. The calculated free energy of solvation included the polar solvation energy (electrostatic) and nonpolar solvation energy (non-electrostatic, calculated by the solvent-accessible surface area; SASA model). 31 The resulted interactions, as well as binding free energy values, are summarized in Table 4. The data reveal favorable stability for the 2 complexes as indicated by the high negative energy values across electrostatic, van der Waals, and SASA energies. The only exception was the polar solvation energy, which was in the positive range; however, this can be attributed to the lower solubility of the 2 complexes compared to the ligand-free enzyme. Generally, the average binding free energy of quercetin and isorhamnetin-3-O-D glycoside complexes were −144.66 and −149.75 kJ/mol, respectively, which suggests strong and stable binding for both the compounds. We believe that the results from the MD simulations supported the design and validated the entire virtual screening approach; also, they emphasized and assured the potential inhibitory effect of quercetin and isorhamnetin-3-O-D glycoside.
MM-PBSA Calculations of the Binding Free Energy for the 2 Complexes.
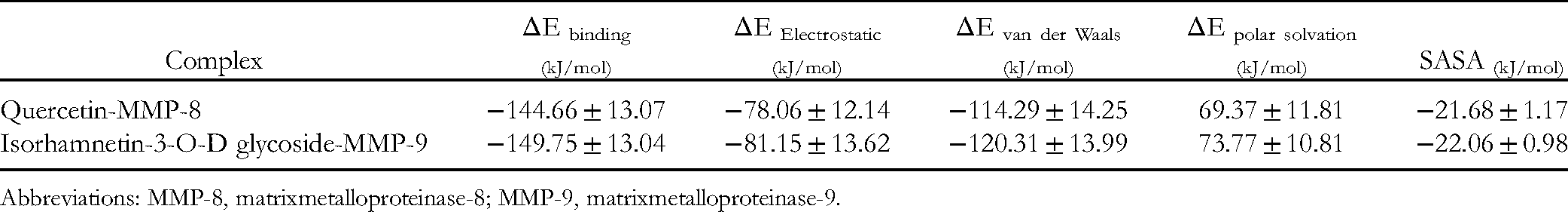

Abbreviations: MMP-8, matrixmetalloproteinase-8; MMP-9, matrixmetalloproteinase-9.
Conclusion
Calendula extract is an established folk medicine for various disorders such as wounds, inflammation, and pain. The rich composition of the extract enables its multifunctional activity. In this research work, we focused on its wound healing properties and its ability to affect MMPs especially types 8 and 9 which are overexpressed in wounds and diabetic foot ulcers. Several chemical constituents are subjugated to various in silico simulations and compared against an established MMP-9 inhibitor (RND-336). Among these compounds, the structures containing benzopyran-4-one moiety are the most potent against MMP-8 and MMP-9 enzymes. Additionally, most compounds outclassed RND-336 while targeting both subtypes simultaneously which points out the impact of screening natural compounds to obtain new successful treatments. Finally, this study suggests the potential effect of Calendula constituents in targeting both MMP-8 and MMP-9, hence their promising treatment effect against wounds and diabetic foot ulcers with overexpression of these target proteins.
Supplemental Material
sj-rar-1-npx-10.1177_1934578X221098848 - Supplemental material for Calendula officinalis Phytochemicals for the Treatment of Wounds Through Matrix Metalloproteinases-8 and 9 (MMP-8 and MMP-9): In Silico Approach
Supplemental material, sj-rar-1-npx-10.1177_1934578X221098848 for
Footnotes
Acknowledgments
Dr Amany Belal would like to thank Taif University, Researchers Supporting Project number (TURSP-2020/35), Taif University, Taif, Saudi Arabia. The authors would like to thank the Deanship of scientific research at Umm Al-Qua University for supporting this work by grant code (22UQU4290559DSR01).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Taif University, Researchers Supporting Project number (TURSP-2020/35), Taif University, Taif, Saudi Arabia. Deanship of scientific research at Umm Al-Qua University grant code (22UQU4290559DSR01).
Ethical Approval
Not applicable, because this article does not contain any studies with human or animal subjects.
Informed Consent
Not applicable, because this article does not contain any studies with human or animal subjects.
Trial Registration
Not applicable, because this article does not contain any clinical trials.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
